Chlamydia trachomatis and Chlamydia pneumoniae Interaction with the Host: Latest Advances and Future Prospective
Abstract
1. Introduction
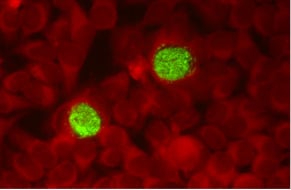
2. Chlamydiae Developmental Cycle
3. Genomic Modification Approaches in Chlamydiae
4. C. trachomatis Interaction with Host Defense Factors
Genital Microbiota Characterization by Metagenomic Analysis
5. C. pneumoniae and Chronic Inflammatory Diseases
5.1. Atherosclerotic Cardiovascular Diseases
5.2. Alzheimer’s Disease
5.3. Reactive Arthritis
6. Conclusions and Future Prospective
Funding
Conflicts of Interest
References
- Mylonas, I. Female genital Chlamydia trachomatis infection: Where are we heading? Arch. Ginecol. Obstet. 2012, 285, 1271–1285. [Google Scholar] [CrossRef] [PubMed]
- Choroszy-Król, I.; Frej-Mądrzak, M.; Hober, M.; Sarowska, J.; Jama-Kmiecik, A. Infections caused by Chlamydophila pneumoniae. Adv. Clin. Exp. Med. 2014, 23, 123–126. [Google Scholar] [CrossRef]
- Wang, Y.; Kahane, S.; Cutcliffe, L.T.; Skilton, R.J.; Lambden, P.R.; Clarke, I.N. Development of a transformation system for Chlamydia trachomatis: Restoration of glycogen biosynthesis by acquisition of a plasmid shuttle vector. PLoS Pathog. 2011, 7, e1002258. [Google Scholar] [CrossRef] [PubMed]
- Van de Wijgert, J.H.; Borgdorff, H.; Verhelst, R.; Crucitti, T.; Francis, S.; Verstraelen, H.; Jespers, V. The vaginal microbiota: What have we learned after a decade of molecular characterization? PLoS ONE 2014, 9, e105998. [Google Scholar] [CrossRef] [PubMed]
- Lewis, F.M.; Bernstein, K.T.; Aral, S.O. Vaginal microbiome and its relationship to behavior, sexual health, and sexually transmitted diseases. Obstet. Gynecol. 2017, 129, 643–654. [Google Scholar] [CrossRef]
- GBD 2016 Disease and Injury Incidence and Prevalence Collaborators. Global, regional, and national incidence, prevalence, and years lived with disability for 328 diseases and injuries for 195 countries, 1990–2016: A systematic analysis for the Global Burden of Disease Study 2016. Lancet 2017, 390, 1211–1259. [Google Scholar] [CrossRef]
- Shaw, K.; Coleman, D.; O’Sullivan, M.; Stephens, N. Public health policies and management strategies for genital Chlamydia trachomatis infection. Risk Manag. Healthc. Policy 2011, 4, 57–65. [Google Scholar] [CrossRef] [PubMed]
- Lanjouw, E.; Ouburg, S.; de Vries, H.J.; Stary, A.; Radcliffe, K.; Unemo, M. 2015 European guideline on the management of Chlamydia trachomatis infections. Int. J. STD AIDS 2016, 27, 333–348. [Google Scholar] [CrossRef] [PubMed]
- Buckner, L.R.; Amedee, A.M.; Albritton, H.L.; Kozlowski, P.A.; Lacour, N.; McGowin, C.L.; Schust, D.J.; Quayle, A.J. Chlamydia trachomatis infection of endocervical epithelial cells enhances early HIV transmission events. PLoS ONE 2016, 11, e0146663. [Google Scholar] [CrossRef]
- Silva, J.; Cerqueira, F.; Medeiros, R. Chlamydia trachomatis infection: Implications for HPV status and cervical cancer. Arch. Gynecol. Obstet. 2014, 289, 715–723. [Google Scholar] [CrossRef]
- Grayston, J.T.; Aldous, M.B.; Easton, A.; Wang, S.P.; Kuo, C.C.; Campbell, L.A.; Altman, J. Evidence that Chlamydia pneumoniae causes pneumonia and bronchitis. J. Infect. Dis. 1993, 168, 1231–1235. [Google Scholar] [CrossRef] [PubMed]
- Moazed, T.C.; Kuo, C.C.; Grayston, J.T.; Campbell, L.A. Evidence of systemic dissemination of Chlamydia pneumoniae via macrophages in the mouse. J. Infect. Dis. 1998, 177, 1322–1325. [Google Scholar] [CrossRef] [PubMed]
- MacIntyre, A.; Abramov, R.; Hammond, C.J.; Hudson, A.P.; Arking, E.J.; Little, C.S.; Appelt, D.M.; Balin, B.J. Chlamydia pneumoniae infection promotes the transmigration of monocytes through human brain endothelial cells. J. Neurosci. Res. 2003, 71, 740–750. [Google Scholar] [CrossRef]
- Sessa, R.; Di Pietro, M.; Schiavoni, G.; Santino, I.; Benedetti-Valentini, F.; Perna, R.; Romano, S.; del Piano, M. Chlamydia pneumoniae DNA in patients with symptomatic carotid atherosclerotic disease. J. Vasc. Surg. 2003, 37, 1027–1031. [Google Scholar] [CrossRef] [PubMed]
- Sessa, R.; di Pietro, M.; Schiavoni, G.; Petrucca, A.; Cipriani, P.; Zagaglia, C.; Nicoletti, M.; Santino, I.; del Piano, M. Measurement of Chlamydia pneumoniae bacterial load in peripheral blood mononuclear cells may be helpful to assess the state of Chlamydial infection in patients with carotid atherosclerotic disease. Atherosclerosis 2007, 195, e224–e230. [Google Scholar] [CrossRef]
- Di Pietro, M.; Schiavoni, G.; Sessa, V.; Pallotta, F.; Costanzo, G.; Sessa, R. Chlamydia pneumoniae and osteoporosis-associated bone loss: A new risk factor? Osteoporos. Int. 2013, 24, 1677–1682. [Google Scholar] [CrossRef] [PubMed]
- Sessa, R.; Nicoletti, M.; Di Pietro, M.; Schiavoni, G.; Santino, I.; Zagaglia, C.; Del Piano, M.; Cipriani, P. Chlamydia pneumoniae and atherosclerosis: Current state and future prospectives. Int. J. Immunopathol. Pharmacol. 2009, 22, 9–14. [Google Scholar] [CrossRef] [PubMed]
- Schiavoni, G.; Di Pietro, M.; Ronco, C.; De Cal, M.; Cazzavillan, S.; Rassu, M.; Nicoletti, M.; Del Piano, M.; Sessa, R. Chlamydia pneumoniae infection as a risk factor for accelerated atherosclerosis in hemodialysis patients. J. Biol. Regul. Homeost. Agents. 2010, 24, 367–375. [Google Scholar]
- Campbell, L.A.; Rosenfeld, M.E. Infection and atherosclerosis development. Arch. Med. Res. 2015, 46, 339–350. [Google Scholar] [CrossRef] [PubMed]
- Porritt, R.A.; Crother, T.R. Chlamydia pneumoniae infection and inflammatory diseases. Forum Immunopathol. Dis. Ther. 2016, 7, 237–254. [Google Scholar] [CrossRef] [PubMed]
- Zeidler, H.; Hudson, A.P. Causality of Chlamydiae in Arthritis and Spondyloarthritis: A Plea for Increased Translational Research. Curr. Rheumatol. Rep. 2016, 18, 9. [Google Scholar] [CrossRef]
- Carter, J.D.; Hudson, A.P. Recent advances and future directions in understanding and treating Chlamydia-induced reactive arthritis. Expert Rev. Clin. Immunol. 2017, 13, 197–206. [Google Scholar] [CrossRef]
- Di Pietro, M.; Filardo, S.; Falasca, F.; Turriziani, O.; Sessa, R. Infectious Agents in Atherosclerotic Cardiovascular Diseases through Oxidative Stress. Int. J. Mol. Sci. 2017, 18, 2459. [Google Scholar] [CrossRef]
- Balin, B.J.; Hammond, C.J.; Little, C.S.; Hingley, S.T.; Al-Atrache, Z.; Appelt, D.M.; Whittum-Hudson, J.A.; Hudson, A.P. Chlamydia pneumoniae: An Etiologic Agent for Late-Onset Dementia. Front. Aging. Neurosci. 2018, 10, 302. [Google Scholar] [CrossRef] [PubMed]
- Abdelrahman, Y.M.; Belland, R.J. The Chlamydial developmental cycle. FEMS Microbiol. Rev. 2005, 29, 949–959. [Google Scholar] [CrossRef] [PubMed]
- Omsland, A.; Sager, J.; Nair, V.; Sturdevant, D.E.; Hackstadt, T. Developmental stage-specific metabolic and transcriptional activity of Chlamydia trachomatis in an axenic medium. Proc. Natl. Acad. Sci. USA 2012, 109, 19781–19785. [Google Scholar] [CrossRef] [PubMed]
- Skipp, P.J.; Hughes, C.; McKenna, T.; Edwards, R.; Langridge, J.; Thomson, N.R.; Clarke, I.N. Quantitative Proteomics of the Infectious and Replicative Forms of Chlamydia trachomatis. PLoS ONE 2016, 11, e0149011. [Google Scholar] [CrossRef] [PubMed]
- Bastidas, R.J.; Elwell, C.A.; Engel, J.N.; Valdivia, R.H. Chlamydial intracellular survival strategies. Cold Spring Harb. Perspect. Med. 2013, 3, a010256. [Google Scholar] [CrossRef] [PubMed]
- Elwell, C.; Mirrashidi, K.; Engel, J. Chlamydia cell biology and pathogenesis. Nat. Rev. Microbiol. 2016, 14, 385–400. [Google Scholar] [CrossRef]
- Dai, W.; Li, Z. Conserved type III secretion system exerts important roles in Chlamydia trachomatis. Int. J. Clin. Exp. Pathol. 2014, 7, 5404–5414. [Google Scholar]
- Nans, A.; Ford, C.; Hayward, R.D. Host-pathogen reorganisation during host cell entry by Chlamydia trachomatis. Microbes Infect. 2015, 17, 727–731. [Google Scholar] [CrossRef] [PubMed]
- Jorgensen, I.; Bednar, M.M.; Amin, V.; Davis, B.K.; Ting, J.P.; McCafferty, D.G.; Valdivia, R.H. The Chlamydia protease CPAF regulates host and bacterial proteins to maintain pathogen vacuole integrity and promote virulence. Cell Host Microbe 2011, 10, 21–32. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Tang, L.; Zhou, Z.; Zhong, G. Neutralizing anti-chlamydial activity of complement by Chlamydia-secreted protease CPAF. Microbes Infect. 2016, 18, 669–674. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Lei, L.; Gong, S.; Chen, D.; Flores, R.; Zhong, G. The chlamydial periplasmic stress response serine protease cHtrA is secreted into host cell cytosol. BMC Microbiol. 2011, 11, 87. [Google Scholar] [CrossRef] [PubMed]
- Hogan, R.J.; Mathews, S.A.; Mukhopadhyay, S.; Summersgill, J.T.; Timms, P. Chlamydial persistence: Beyond the biphasic paradigm. Infect. Immun. 2004, 72, 1843–1855. [Google Scholar] [CrossRef] [PubMed]
- Wyrick, P.B. Chlamydia trachomatis persistence in vitro: An overview. J. Infect. Dis. 2010, 201, S88–S95. [Google Scholar] [CrossRef] [PubMed]
- Raulston, J.E. Response of Chlamydia trachomatis serovar E to iron restriction in vitro and evidence for iron-regulated chlamydial proteins. Infect. Immun. 1997, 65, 4539–4547. [Google Scholar] [PubMed]
- Di Pietro, M.; Tramonti, A.; de Santis, F.; de Biase, D.; Schiavoni, G.; Filardo, S.; Zagaglia, C.; Sessa, R. Analysis of gene expression in penicillin G induced persistence of Chlamydia pneumoniae. J. Biol. Regul. Homeost. Agents. 2012, 26, 277–284. [Google Scholar] [PubMed]
- Kintner, J.; Lajoie, D.; Hall, J.; Whittimore, J.; Schoborg, R.V. Commonly prescribed β-lactam antibiotics induce C. trachomatis persistence/stress in culture at physiologically relevant concentrations. Front. Cell. Infect. Microbiol. 2014, 4, 44. [Google Scholar] [CrossRef] [PubMed]
- Panzetta, M.E.; Valdivia, R.H.; Saka, H.A. Chlamydia Persistence: A Survival Strategy to Evade Antimicrobial Effects in-vitro and in-vivo. Front Microbiol. 2018, 9, 3101. [Google Scholar] [CrossRef]
- Vanover, J.; Kintner, J.; Whittimore, J.; Schoborg, R.V. Interaction of herpes simplex virus type 2 (HSV-2) glycoprotein D with the host cell surface is sufficient to induce Chlamydia trachomatis persistence. Microbiology 2010, 156, 1294–1302. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Romano, J.D.; de Beaumont, C.; Carrasco, J.A.; Ehrenman, K.; Bavoil, P.M.; Coppens, I. A novel co-infection model with Toxoplasma and Chlamydia trachomatis highlights the importance of host cell manipulation for nutrient scavenging. Cell. Microbiol. 2013, 15, 619–646. [Google Scholar] [CrossRef] [PubMed]
- Phillips Campbell, R.; Kintner, J.; Whittimore, J.; Schoborg, R.V. Chlamydia muridarum enters a viable but non-infectious state in amoxicillin-treated BALB/c mice. Microbes Infect. 2012, 14, 1177–1185. [Google Scholar] [CrossRef][Green Version]
- Phillips-Campbell, R.; Kintner, J.; Schoborg, R.V. Induction of the Chlamydia muridarum stress/persistence response increases azithromycin treatment failure in a murine model of infection. Antimicrob. Agents. Chemother. 2014, 58, 1782–1784. [Google Scholar] [CrossRef] [PubMed]
- Borel, N.; Summersgill, J.T.; Mukhopadhyay, S.; Miller, R.D.; Ramirez, J.A.; Pospischil, A. Evidence for persistent Chlamydia pneumoniae infection of human coronary atheromas. Atherosclerosis 2008, 199, 154–161. [Google Scholar] [CrossRef] [PubMed]
- Lewis, M.E.; Belland, R.J.; AbdelRahman, Y.M.; Beatty, W.L.; Aiyar, A.A.; Zea, A.H.; Greene, S.J.; Marrero, L.; Buckner, L.R.; Tate, D.J.; et al. Morphologic and molecular evaluation of Chlamydia trachomatis growth in human endocervix reveals distinct growth patterns. Front. Cell. Infect. Microbiol. 2014, 4, 71. [Google Scholar] [CrossRef]
- Wyrick, P.B.; Knight, S.T. Pre-exposure of infected human endometrial epithelial cells to penicillin in vitro renders Chlamydia trachomatis refractory to azithromycin. J. Antimicrob. Chemother. 2004, 54, 79–85. [Google Scholar] [CrossRef] [PubMed]
- Choroszy-Król, I.C.; Frej-Mądrzak, M.; Jama-Kmiecik, A.; Bober, T.; Jolanta Sarowska, J.; Frej-Mądrzak, M.; Jama-Kmiecik, A.; Bober, T.; Jolanta Sarowska, J. Characteristics of the Chlamydia trachomatis species - immunopathology and infections. Adv. Clin. Exp. Med. 2012, 21, 799–808. [Google Scholar]
- Tam, J.E.; Davis, C.H.; Wyrick, P.B. Expression of recombinant DNA introduced into Chlamydia trachomatis by electroporation. Can. J. Microbiol. 1994, 40, 583–591. [Google Scholar] [CrossRef] [PubMed]
- Binet, R.; Maurelli, A.T. The Chlamydial functional homolog of KsgA confers kasugamycin sensitivity to Chlamydia trachomatis and impacts bacterial fitness. BMC Microbiol. 2009, 9, 279. [Google Scholar] [CrossRef]
- Agaisse, H.; Derré, I. A C. trachomatis cloning vector and the generation of C. trachomatis strains expressing fluorescent proteins under the control of a C. trachomatis promoter. PLoS ONE 2013, 8, e57090. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Derré, I. A Co-infection Model System and the Use of Chimeric Proteins to Study Chlamydia Inclusion Proteins Interaction. Front. Cell Infect Microbiol. 2017, 7, 79. [Google Scholar] [CrossRef]
- Wickstrum, J.; Sammons, L.R.; Restivo, K.N.; Hefty, P.S. Conditional gene expression in Chlamydia trachomatis using the tet system. PLoS ONE 2013, 8, e76743. [Google Scholar] [CrossRef]
- Kari, L.; Goheen, M.M.; Randall, L.B.; Taylor, L.D.; Carlson, J.H.; Whitmire, W.M.; Virok, D.; Rajaram, K.; Endresz, V.; McClarty, G.; et al. Generation of targeted Chlamydia trachomatis null mutants. Proc. Natl. Acad. Sci. USA 2011, 108, 7189–7193. [Google Scholar] [CrossRef]
- Nguyen, B.D.; Valdivia, R.H. Virulence determinants in the obligate intracellular pathogen Chlamydia trachomatis revealed by forward genetic approaches. Proc. Natl. Acad. Sci. USA 2012, 109, 1263–1268. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.M.; Fisher, D.J. Site-specific, insertional inactivation of incA in Chlamydia trachomatis using a group II intron. PLoS ONE 2013, 8, e83989. [Google Scholar] [CrossRef] [PubMed]
- Kokes, M.; Dunn, J.D.; Granek, J.A.; Nguyen, B.D.; Barker, J.R.; Valdivia, R.H.; Bastidas, R.J. Integrating chemical mutagenesis and whole-genome sequencing as a platform for forward and reverse genetic analysis of Chlamydia. Cell Host Microbe 2015, 17, 716–725. [Google Scholar] [CrossRef]
- Mueller, K.E.; Wolf, K.; Fields, K.A. Gene Deletion by Fluorescence-Reported Allelic Exchange Mutagenesis in Chlamydia trachomatis. mBio 2016, 7, e01817-15. [Google Scholar] [CrossRef] [PubMed]
- Shima, K.; Wanker, M.; Skilton, R.J.; Cutcliffe, L.T.; Schnee, C.; Kohl, T.A.; Niemann, S.; Geijo, J.; Klinger, M.; Timms, P.; et al. The Genetic Transformation of Chlamydia pneumoniae. mSphere 2018, 3, e00412-18. [Google Scholar] [CrossRef] [PubMed]
- Mendling, W. Vaginal Microbiota. Adv. Exp. Med. Biol. 2016, 902, 83–93. [Google Scholar]
- Srinivasan, S.; Fredricks, D.N. The human vaginal bacterial biota and bacterial vaginosis. Interdiscip. Perspect. Infect. Dis. 2008, 2008, 750479. [Google Scholar] [CrossRef] [PubMed]
- Eckert, L.O. Clinical practice. Acute vulvovaginitis. N. Engl. J. Med. 2006, 355, 1244–1252. [Google Scholar] [CrossRef] [PubMed]
- Machado, A.; Cerca, N. Influence of biofilm formation by Gardnerella vaginalis and other anaerobes on bacterial vaginosis. J. Infect. Dis. 2015, 212, 1856–1861. [Google Scholar] [CrossRef]
- Cauchie, M.; Desmet, S.; Lagrou, K. Candida and its dual lifestyle as a commensal and a pathogen. Res. Microbiol. 2017, 168, 802–810. [Google Scholar] [CrossRef]
- Filardo, S.; Di Pietro, M.; Tranquilli, G.; Sessa, R. Biofilm in Genital Ecosystem: A Potential Risk Factor for Chlamydia trachomatis Infection. Can. J. Infect. Dis. Med. Microbiol. 2019, 2019, 1672109. [Google Scholar] [CrossRef] [PubMed]
- Mijac, V.D.; Dukić, S.V.; Opavski, N.Z.; Dukić, M.K.; Ranin, L.T.; Dukić, S.V.; Opavski, N.Z.; Dukić, M.K.; Ranin, L.T. Hydrogen peroxide producing lactobacilli in women with vaginal infections. Eur. J. Obstet. Gynecol. Reprod. Biol. 2006, 129, 69–76. [Google Scholar] [CrossRef] [PubMed]
- Vielfort, K.; Sjölinder, H.; Roos, S.; Jonsson, H.; Aro, H. Adherence of clinically isolated lactobacilli to human cervical cells in competition with Neisseria gonorrhoeae. Microbes Infect. 2008, 10, 1325–1334. [Google Scholar] [CrossRef] [PubMed]
- Petrova, M.I.; Lievens, E.; Malik, S.; Imholz, N.; Lebeer, S. Lactobacillus species as biomarkers and agents that can promote various aspects of vaginal health. Front. Physiol. 2015, 6, 81. [Google Scholar] [CrossRef] [PubMed]
- Gong, Z.; Luna, Y.; Yu, P.; Fan, H. Lactobacilli inactivate Chlamydia trachomatis through lactic acid but not H2O2. PLoS ONE 2014, 9, e107758. [Google Scholar] [CrossRef]
- Mastromarino, P.; Di Pietro, M.; Schiavoni, G.; Nardis, C.; Gentile, M.; Sessa, R. Effects of vaginal lactobacilli in Chlamydia trachomatis infection. Int. J. Med. Microbiol. 2014, 304, 654–661. [Google Scholar] [CrossRef]
- Nardini, P.; Ñahui Palomino, R.A.; Parolin, C.; Laghi, L.; Foschi, C.; Cevenini, R.; Vitali, B.; Marangoni, A. Lactobacillus crispatus inhibits the infectivity of Chlamydia trachomatis elementary bodies, in vitro study. Sci. Rep. 2016, 6, 29024. [Google Scholar] [CrossRef] [PubMed]
- Parolin, C.; Frisco, G.; Foschi, C.; Giordani, B.; Salvo, M.; Vitali, B.; Marangoni, A.; Calonghi, N. Lactobacillus crispatus BC5 Interferes with Chlamydia trachomatis Infectivity Through Integrin Modulation in Cervical Cells. Front Microbiol. 2018, 9, 2630. [Google Scholar] [CrossRef] [PubMed]
- Rizzo, A.; Fiorentino, M.; Buommino, E.; Donnarumma, G.; Losacco, A.; Bevilacqua, N. Lactobacillus crispatus mediates anti-inflammatory cytokine interleukin-10 induction in response to Chlamydia trachomatis infection in vitro. Int. J. Med. Microbiol. 2015, 305, 815–827. [Google Scholar] [CrossRef] [PubMed]
- Andersen, J.H.; Jenssen, H.; Gutteberg, T.J. Lactoferrin and lactoferricin inhibit Herpes simplex 1 and 2 infection and exhibit synergy when combined with acyclovir. Antiviral Res. 2003, 58, 209–215. [Google Scholar] [CrossRef]
- Naidu, A.S.; Chen, J.; Martinez, C.; Tulpinski, J.; Pal, B.K.; Fowler, R.S. Activated lactoferrin’s ability to inhibit Candida growth and block yeast adhesion to the vaginal epithelial monolayer. J. Reprod. Med. 2004, 49, 859–866. [Google Scholar] [PubMed]
- Redwan, E.M.; Uversky, V.N.; El-Fakharany, E.M.; Al-Mehdar, H. Potential lactoferrin activity against pathogenic viruses. C. R. Biol. 2014, 337, 581–595. [Google Scholar] [CrossRef] [PubMed]
- Sessa, R.; Di Pietro, M.; Filardo, S.; Bressan, A.; Mastromarino, P.; Biasucci, A.V.; Rosa, L.; Cutone, A.; Berlutti, F.; Paesano, R.; et., al. Lactobacilli-lactoferrin interplay in Chlamydia trachomatis infection. Pathog. Dis. 2017, 75. [Google Scholar] [CrossRef]
- Sessa, R.; di Pietro, M.; Filardo, S.; Bressan, A.; Rosa, L.; Cutone, A.; Frioni, A.; Berlutti, F.; Paesano, R.; Valenti, P. Effect of bovine lactoferrin on Chlamydia trachomatis infection and inflammation. Biochem. Cell. Biol. 2017, 95, 34–40. [Google Scholar] [CrossRef] [PubMed]
- Sawada, M.; Otsuki, K.; Mitsukawa, K.; Yakuwa, K.; Nagatsuka, M.; Okai, T. Cervical inflammatory cytokines and other markers inthe cervical mucus of pregnant womenwith lower genital tract infection. Int. J. Gynaecol. Obstet. 2006, 92, 117–121. [Google Scholar] [CrossRef]
- Spear, G.T.; Kendrick, S.R.; Chen, H.Y.; Thomas, T.T.; Bahk, M.; Balderas, R.; Ghosh, S.; Aaron Weinberg, A.; Landay, A.L. Multiplex immunoassay of lower genital tract mucosal fluid from women attending an urban STD clinic shows broadly increased IL1ß and lactoferrin. PLoS ONE 2011, 6, e19560. [Google Scholar] [CrossRef] [PubMed]
- Filardo, S.; Di Pietro, M.; Tranquilli, G.; Latino, M.A.; Recine, N.; Porpora, M.; Sessa, R. Selected immunological mediators and cervical microbial signatures in women with Chlamydia trachomatis infection. mSystems 2019. In press. [Google Scholar]
- Yasin, B.; Pang, M.; Lehrer, R.I.; Wagar, E.A. Activity of Novispirin G-10, a novel antimicrobial peptide against Chlamydia trachomatis and vaginosis-associated bacteria. Exp. Mol. Pathol. 2003, 74, 190–195. [Google Scholar] [CrossRef]
- Donati, M.; di Leo, K.; Benincasa, M.; Cavrini, F.; Accardo, S.; Moroni, A.; Gennaro, R.; Cevenini, R. Activity of cathelicidin peptides against Chlamydia spp. Antimicrob. Agents Chemother. 2005, 49, 1201–1202. [Google Scholar] [CrossRef] [PubMed]
- Di Francesco, A.; Favaroni, A.; Donati, M. Host defense peptides: General overview and an update on their activity against Chlamydia spp. Expert Rev. Anti. Infect. Ther. 2013, 11, 1215–1224. [Google Scholar] [CrossRef]
- Noda-Nicolau, N.M.; Bastos, L.B.; Bolpetti, A.N.; Pinto, G.V.S.; Marcolino, L.D.; Marconi, C.; Ferreira, C.S.T.; Polettini, J.; Vieira, E.P.; da Silva, M.G. Cervicovaginal Levels of Human β-Defensin 1, 2, 3, and 4 of Reproductive-Aged Women With Chlamydia trachomatis Infection. J. Low Genit. Tract. Dis. 2017, 21, 189–192. [Google Scholar] [CrossRef] [PubMed]
- Tang, L.; Chen, J.; Zhou, Z.; Yu, P.; Yang, Z.; Zhong, G. Chlamydia-secreted protease CPAF degrades host antimicrobial peptides. Microbes Infect. 2015, 17, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Lamont, R.F.; Sobel, J.D.; Akins, R.A.; Hassan, S.S.; Chaiworapongsa, T.; Kusanovic, J.P.; Romero, R. The vaginal microbiome: New information about genital tract flora using molecular based techniques. BJOG 2011, 118, 533–549. [Google Scholar] [CrossRef] [PubMed]
- Gajer, P.; Brotman, R.M.; Bai, G.; Sakamoto, J.; Schütte, U.M.; Zhong, X.; Koenig, S.S.; Fu, L.; Ma, Z.S.; Zhou, X.; et al. Temporal dynamics of the human vaginal microbiota. Sci. Transl. Med. 2012, 4, 132. [Google Scholar] [CrossRef]
- Ma, B.; Forney, L.J.; Ravel, J. Vaginal microbiome: Rethinking health and disease. Annu. Rev. Microbiol. 2012, 66, 371–389. [Google Scholar] [CrossRef]
- Petrova, M.I.; Reid, G.; Vaneechoutte, M.; Lebeer, S. Lactobacillus iners: Friend or Foe? Trends Microbiol. 2017, 25, 182–191. [Google Scholar] [CrossRef]
- Ma, B.; Brotman, R.M.; Gajer, P.; Fadrosh, D.; Mahurkar, A.; White, O.; Terplan, M.; Bavoil, P.; Forney, L.J. Association between Chlamydia trachomatis genital infection and the vaginal microbiome. Sex. Transm. Infect. 2013, 89, A1–A428. [Google Scholar] [CrossRef][Green Version]
- Filardo, S.; di Pietro, M.; Porpora, M.G.; Recine, N.; Farcomeni, A.; Latino, M.A.; Sessa, R. Diversity of Cervical Microbiota in Asymptomatic Chlamydia trachomatis Genital Infection: A Pilot Study. Front. Cell. Infect. Microbiol. 2017, 7, 321. [Google Scholar] [CrossRef]
- van der Veer, C.; Bruisten, S.M.; van der Helm, J.J.; de Vries, H.J.; van Houdt, R. The cervicovaginal microbiota in women notified for Chlamydia trachomatis infection: A case-control study at the sexually transmitted infection outpatient clinic in Amsterdam, The Netherlands. Clin. Infect. Dis. 2017, 64, 24–31. [Google Scholar] [CrossRef] [PubMed]
- Balle, C.; Lennard, K.; Dabee, S.; Barnabas, S.L.; Jaumdally, S.Z.; Gasper, M.A.; Maseko, V.; Mbulawa, Z.Z.A.; Williamson, A.L.; Bekker, L.G.; et al. Endocervical and vaginal microbiota in South African adolescents with asymptomatic Chlamydia trachomatis infection. Sci. Rep. 2018, 8, 11109. [Google Scholar] [CrossRef] [PubMed]
- Di Pietro, M.; Filardo, S.; Porpora, M.G.; Recine, N.; Latino, M.A.; Sessa, R. HPV/Chlamydia trachomatis co-infection: Metagenomic analysis of cervical microbiota in asymptomatic women. New Microbiol. 2018, 41, 34–41. [Google Scholar] [PubMed]
- Parolin, C.; Foschi, C.; Laghi, L.; Zhu, C.; Banzola, N.; Gaspari, V.; D’Antuono, A.; Giordani, B.; Severgnini, M.; Consolandi, C.; et al. Insights Into Vaginal Bacterial Communities and Metabolic Profiles of Chlamydia trachomatis Infection: Positioning Between Eubiosis and Dysbiosis. Front. Microbiol. 2018, 9, 600. [Google Scholar] [CrossRef]
- van Houdt, R.; Ma, B.; Bruisten, S.M.; Speksnijder, A.G.C.L.; Ravel, J.; de Vries, H.J.C. Lactobacillus iners-dominated vaginal microbiota is associated with increased susceptibility to Chlamydia trachomatis infection in Dutch women: A case-control study. Sex. Transm. Infect. 2018, 94, 117–123. [Google Scholar] [CrossRef]
- Ziklo, N.; Vidgen, M.E.; Taing, K.; Huston, W.M.; Timms, P. Dysbiosis of the Vaginal Microbiota and Higher Vaginal Kynurenine/Tryptophan Ratio Reveals an Association with Chlamydia trachomatis Genital Infections. Front. Cell. Infect. Microbiol. 2018, 8, 1. [Google Scholar] [CrossRef] [PubMed]
- Tamarelle, J.; Thiébaut, A.C.M.; de Barbeyrac, B.; Bébéar, C.; Ravel, J.; Delarocque-Astagneau, E. The vaginal microbiota and its association with human papillomavirus, Chlamydia trachomatis, Neisseria gonorrhoeae and Mycoplasma genitalium infections: A systematic review and meta-analysis. Clin. Microbiol. Infect. 2019, 25, 35–47. [Google Scholar] [CrossRef]
- Aiyar, A.; Quayle, A.J.; Buckner, L.R.; Sherchand, S.P.; Chang, T.L.; Zea, A.H.; Martin, D.H.; Belland, R.J. Influence of the tryptophan-indole-IFNγ axis on human genital Chlamydia trachomatis infection: Role of vaginal co-infections. Front. Cell. Infect. Microbiol. 2014, 4, 72. [Google Scholar] [CrossRef]
- Ziklo, N.; Huston, W.M.; Hocking, J.S.; Timms, P. Chlamydia trachomatis genital tract infections: When host immune response and the microbiome collide. Trends Microbiol. 2016, 24, 750–765. [Google Scholar] [CrossRef] [PubMed]
- Sessa, R.; di Pietro, M.; Filardo, S.; Turriziani, O. Infectious burden and atherosclerosis: A clinical issue. World J. Clin. Cases. 2014, 2, 240–249. [Google Scholar] [CrossRef] [PubMed]
- Strelić, N.; Bojović, J.; Pavlica, L.; Cikota-Aleksić, B.; Miličić, B.; Magić, Z. Detection of bacteria and analyses of Chlamydia trachomatis viability in patients with postvenereal reactive arthritis. Intern. Med. J. 2014, 44, 1247–1251. [Google Scholar] [CrossRef] [PubMed]
- Bu, X.L.; Yao, X.Q.; Jiao, S.S.; Zeng, F.; Liu, Y.H.; Xiang, Y.; Liang, C.R.; Wang, Q.H.; Wang, X.; Cao, H.Y.; et al. A study on the association between infectious burden and Alzheimer’s disease. Eur. J. Neurol. 2015, 22, 1519–1525. [Google Scholar] [CrossRef]
- Mendis, S.; Puska, P.; Norrving, B.; World Health Organization, World Heart Federation. Global Atlas on Cardiovascular Disease Prevention and Control. Available online: http://whqlibdoc.who.int/publications/2011/9789241564373_eng.pdf (accessed on 15 May 2019).
- Saikku, P.; Leinonen, M.; Mattila, K.; Ekman, M.R.; Nieminen, M.S.; Mäkelä, P.H.; Huttunen, J.K.; Valtonen, V. Serological evidence of an association of a novel Chlamydia, TWAR, with chronic coronary heart disease and acute myocardial infarction. Lancet 1988, 2, 983–986. [Google Scholar] [CrossRef]
- Ramirez, J.A. Isolation of Chlamydia pneumoniae from the coronary artery of a patient with coronary atherosclerosis. The Chlamydia pneumoniae/Atherosclerosis Study Group. Ann. Intern. Med. 1996, 125, 979–982. [Google Scholar] [CrossRef] [PubMed]
- Campbell, L.A.; Kuo, C.C.; Rodriguez, D.I.; Lee, A.; Grayston, J.T. Isolation of Chlamydia pneumoniae from a carotid endarterectomy specimen. J. Infect. Dis. 1997, 176, 292–295. [Google Scholar]
- Apfalter, P.; Loidl, M.; Nadrchal, R.; Makristathis, A.; Rotter, M.; Bergmann, M.; Polterauer, P.; Hirschl, A.M. Isolation and continuous growth of Chlamydia pneumoniae from arterectomy specimens. Eur. J. Clin. Microbiol. Infect. Dis. 2000, 19, 305–308. [Google Scholar] [CrossRef]
- Romano, S.; Penco, M.; Fratini, S.; Di Pietro, M.; Sessa, R.; del Piano, M.; Fedele, F.; Dagianti, A. Chlamydia pneumoniae infection is associated with coronary artery disease but not implicated in inducing plaque instability. Int. J. Cardiol. 2004, 95, 95–99. [Google Scholar] [CrossRef] [PubMed]
- Sessa, R.; Di Pietro, M.; Schiavoni, G.; Galdiero, M.; Cipriani, P.; Romano, S.; Zagaglia, C.; Santino, I.; Faccilongo, S.; Del Piano, M. Chlamydia pneumoniae in asymptomatic carotid atherosclerosis. Int. J. Immunopathol. Pharmacol. 2006, 19, 111–118. [Google Scholar] [CrossRef] [PubMed]
- Watson, C.; Alp, N.J. Role of Chlamydia pneumoniae in atherosclerosis. Clin. Sci. 2008, 114, 509–531. [Google Scholar] [CrossRef] [PubMed]
- Joshi, R.; Khandelwal, B.; Joshi, D.; Gupta, O.P. Chlamydophila pneumoniae infection and cardiovascular disease. N. Am. J. Med. Sci. 2013, 5, 169–181. [Google Scholar] [CrossRef] [PubMed]
- Liuba, P.; Pesonen, E.; Paakkari, I.; Batra, S.; Forslid, A.; Kovanen, P.; Pentikäinen, M.; Persson, K.; Sandström, S. Acute Chlamydia pneumoniae infection causes coronary endothelial dysfunction in pigs. Atherosclerosis 2000, 167, 215–222. [Google Scholar] [CrossRef]
- De Kruif, M.D.; van Gorp, E.C.; Keller, T.T.; Ossewaarde, J.M.; ten Cate, H. Chlamydia pneumoniae infections in mouse models: Relevance for atherosclerosis research. Cardiovasc. Res. 2005, 65, 317–327. [Google Scholar] [CrossRef]
- Di Pietro, M.; de Santis, F.; Schiavoni, G.; Filardo, S.; Sessa, R. Resveratrol in Chlamydia pneumoniae-induced foam cell formation and interleukin-17A synthesis. J. Biol. Regul. Homeost. Agents. 2013, 27, 509–518. [Google Scholar] [PubMed]
- Di Pietro, M.; Filardo, S.; de Santis, F.; Mastromarino, P.; Sessa, R. Chlamydia pneumoniae and oxidative stress in cardiovascular disease: State of the art and prevention strategies. Int. J. Mol. Sci. 2014, 16, 724–735. [Google Scholar] [CrossRef]
- Mueller, K.E.; Wolf, K.C. pneumoniae disrupts eNOS trafficking and impairs NO production in human aortic endothelial cells. Cell. Microbiol. 2015, 17, 119–130. [Google Scholar] [CrossRef] [PubMed]
- Gaydos, C.A. Growth in vascular cells and cytokine production by Chlamydia pneumoniae. J. Infect. Dis. 2000, 181, S473–S478. [Google Scholar] [CrossRef]
- Di Pietro, M.; Filardo, S.; de Santis, F.; Sessa, R. Chlamydia pneumoniae infection in atherosclerotic lesion development through oxidative stress: A brief overview. Int. J. Mol. Sci. 2013, 14, 15105–15120. [Google Scholar] [CrossRef]
- Tumurkhuu, G.; Dagvadorj, J.; Porritt, R.A.; Crother, T.R.; Shimada, K.; Tarling, E.J.; Erbay, E.; Arditi, M.; Chen, S. Chlamydia pneumoniae hijacks a host autoregulatory IL-1β loop to drive foam cell formation and accelerate atherosclerosis. Cell Metab. 2018, 28, 432–448.e4. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Shimada, K.; Crother, T.R.; Erbay, E.; Shah, P.K.; Arditi, M. Chlamydia and lipids engage a common sigOfnaling pathway that promotes atherogenesis. J. Am. Coll. Cardiol. 2018, 71, 1553–1570. [Google Scholar] [CrossRef] [PubMed]
- Filardo, S.; Di Pietro, M.; Farcomeni, A.; Schiavoni, G.; Sessa, R. Chlamydia pneumoniae-mediated inflammation in atherosclerosis: A metaanalysis. Mediators Inflamm. 2015, 2015, 378658. [Google Scholar] [CrossRef] [PubMed]
- Campbell, L.A.; Rosenfeld, M.E. Persistent C. pneumoniae infection in atherosclerotic lesions: Rethinking the clinical trials. Front. Cell. Infect. Microbiol. 2014, 4, 34. [Google Scholar] [CrossRef] [PubMed]
- Sotuneh, N.; Hosseini, S.R.; Shokri-Shirvani, J.; Bijani, A.; Ghadimi, R. Helicobacter pylori infection and metabolic parameters: Is there an association in elderly population? Int. J. Prev. Med. 2014, 5, 1537–1542. [Google Scholar] [PubMed]
- Zuhair, M.; Smit, G.S.A.; Wallis, G.; Jabbar, F.; Smith, C.; Devleesschauwer, B.; Griffiths, P. Estimation of the worldwide seroprevalence of cytomegalovirus: A systematic review and meta-analysis. Rev. Med. Virol. 2019, 31, e2034. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Quyyumi, A.A.; Norman, J.E.; Csako, G.; Waclawiw, M.A.; Shearer, G.M.; Epstein, S.E. Effects of total pathogen burden on coronary artery disease risk and C-reactive protein levels. Am. J. Cardiol. 2000, 85, 140–146. [Google Scholar] [CrossRef]
- Budzyński, J.; Wiśniewska, J.; Marek Ciecierski, M.; Anna Kędzia, A. Association between bacterial infection and peripheral vascular disease: A review. Int. J. Angiol. 2016, 25, 3–13. [Google Scholar]
- Filardo, S.; di Pietro, M.; Schiavoni, G.; Minniti, G.; Ortolani, E.; Romano, S.; Sessa, R. Chlamydia pneumoniae Clinical Isolate from Gingival Crevicular Fluid: A Potential Atherogenic Strain. Front. Cell. Infect. Microbiol. 2015, 5, 86. [Google Scholar] [CrossRef] [PubMed]
- Mitra, S.; Drautz-Moses, D.I.; Alhede, M.; Maw, M.T.; Liu, Y.; Purbojati, R.W.; Yap, Z.H.; Kushwaha, K.K.; Gheorghe, A.G.; Bjarnsholt, T.; et al. In silico analyses of metagenomes from human atherosclerotic plaque samples. Microbiome 2015, 3, 38. [Google Scholar] [CrossRef]
- Lindskog Jonsson, A.; Hållenius, F.F.; Akrami, R.; Johansson, E.; Wester, P.; Arnerlöv, C.; Bäckhed, F.; Bergström, G. Bacterial profile in human atherosclerotic plaques. Atherosclerosis 2017, 263, 177–183. [Google Scholar] [CrossRef]
- Ascher, S.; Reinhardt, C. The gut microbiota: An emerging risk factor for cardiovascular and cerebrovascular disease. Eur. J. Immunol. 2018, 48, 564–575. [Google Scholar] [CrossRef]
- Wisniewski, T.; Goñi, F. Immunotherapeutic approaches for Alzheimer’s disease. Neuron 2015, 85, 1162–1176. [Google Scholar] [CrossRef]
- Balin, B.J.; Gérard, H.C.; Arking, E.J.; Appelt, D.M.; Branigan, P.J.; Abrams, J.T.; Whittum-Hudson, J.A.; Hudson, A.P. Identification and localization of Chlamydia pneumoniae in the Alzheimer’s brain. Med. Microbiol. Immunol. 1998, 187, 23–42. [Google Scholar] [CrossRef] [PubMed]
- Dreses-Werringloer, U.; Gérard, H.C.; Whittum-Hudson, J.; Hudson, A.P. Chlamydophila (Chlamydia) pneumoniae infection of human astrocytes and microglia in culture displays an active, rather than a persistent, phenotype. Am. J. Med. Sci. 2006, 332, 168–174. [Google Scholar] [CrossRef] [PubMed]
- Sessa, R.; Schiavoni, G.; Borriello, G.; Zagaglia, C.; Marinelli, F.; del Piano, M.; Pozzilli, C. Real time PCR for detection of Chlamydophila pneumoniae in peripheral blood mononuclear cells of patients with multiple sclerosis. J. Neurol. 2007, 254, 1293–1295. [Google Scholar] [CrossRef] [PubMed]
- Hammond, C.J.; Hallock, L.R.; Howanski, R.J.; Appelt, D.M.; Little, C.S.; Balin, B.J. Immunohistological detection of Chlamydia pneumoniae in the Alzheimer’s disease brain. BMC Neurosci. 2010, 11, 121. [Google Scholar] [CrossRef] [PubMed]
- Maheshwari, P.; Eslick, G.D. Bacterial infection and Alzheimer’s disease: A meta-analysis. J. Alzheimers Dis. 2015, 43, 957–966. [Google Scholar] [CrossRef]
- Little, C.S.; Hammond, C.J.; MacIntyre, A.; Balin, B.J.; Appelt, D.M. Chlamydia pneumoniae induces alzheimer-like amyloid plaques in brains of BALB/C mice. Neurobiol. Aging 2004, 25, 419–429. [Google Scholar] [CrossRef]
- Little, C.S.; Joyce, T.A.; Hammond, C.J.; Matta, H.; Cahn, D.; Appelt, D.M.; Balin, B.J. Detection of bacterial antigens and Alzheimer’s disease-like pathology in the central nervous system of BALB/C mice following intranasal infection with a laboratory isolate of Chlamydia pneumoniae. Front. Aging Neurosci. 2014, 6, 304. [Google Scholar] [CrossRef]
- Di Pietro, M.; Filardo, S.; Cazzavillan, S.; Segala, C.; Bevilacqua, P.; Bonoldi, E.; D’Amore, E.S.; Rassu, M.; Sessa, R. Could past Chlamydial vascular infection promote the dissemination of Chlamydia pneumoniae to the brain? J. Biol. Regul. Homeost. Agents. 2013, 27, 155–164. [Google Scholar]
- Appelt, D.M.; Roupas, M.; Hammond, C.J.; Little, C.S.; Balin, B.J. Apoptotic effects of Chlamydophila pneumoniae following infection of neuronal cells. BMC Neurosci. 2008, 9, 13–23. [Google Scholar] [CrossRef]
- Carter, C. Alzheimer’s disease: APP, gamma secretase, APOE, CLU, CR1, PICALM, ABCA7, BIN1, CD2AP, CD33, EPHA1, and MS4A2, and their relationships with herpes simplex, C. pneumoniae, other suspect pathogens, and the immune system. Int. J. Alzheimers Dis. 2011, 2011, 501862. [Google Scholar] [CrossRef] [PubMed]
- Loeb, M.B.; Molloy, D.; Smieja, M.; Standish, T.; Goldsmith, C.H.; Mahony, J.; Smith, S.; Borrie, M.; Decoteau, E.; Davidson, W. A randomized, controlled trial of doxycycline and rifampin for patients with Alzheimer’s disease. J. Am. Geriatr. Soc. 2004, 52, 381–387. [Google Scholar] [CrossRef] [PubMed]
- Beaudreuil, J.; Hayem, G.; Meyer, O.; Kahn, M.F. Reactive arthritis ascribed to Chlamydia pneumoniae. Report of a case. Rev. Rhum. Engl. Ed. 1995, 62, 224. [Google Scholar] [PubMed]
- Moling, O.; Pegoretti, S.; Rielli, M.; Rimenti, G.; Vedovelli, C.; Pristerá, R.; Mian, P. Chlamydia pneumoniae--reactive arthritis and persistent infection. Br. J. Rheumatol. 1996, 35, 1189–1190. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Hannu, T.; Puolakkainen, M.; Leirisalo-Repo, M. Chlamydia pneumoniae as a triggering infection in reactive arthritis. Rheumatology 1999, 38, 411–414. [Google Scholar] [CrossRef] [PubMed]
- Melby, K.K.; Kvien, T.K.; Glennås, A.; Anestad, G. Chlamydia pneumoniae as a trigger of reactive arthritis. Scand. J. Infect. Dis. 1999, 31, 327–328. [Google Scholar] [PubMed]
- Schumacher, H.R., Jr.; Gérard, H.C.; Arayssi, T.K.; Pando, J.A.; Branigan, P.J.; Saaibi, D.L.; Hudson, A.P. Lower prevalence of Chlamydia pneumoniae DNA compared with Chlamydia trachomatis DNA in synovial tissue of arthritis patients. Arthritis Rheum. 1999, 42, 1889–1893. [Google Scholar] [CrossRef]
- Gérard, H.C.; Schumacher, H.R.; El-Gabalawy, H.; Goldbach-Mansky, R.; Hudson, A.P. Chlamydia pneumoniae present in the human synovium are viable and metabolically active. Microb. Pathog. 2000, 29, 17–24. [Google Scholar]
- Cascina, A.; Marone Bianco, A.; Mangiarotti, P.; Montecucco, C.M.; Meloni, F. Cutaneous vasculitis and reactive arthritis following respiratory infection due to Chlamydia pneumoniae: Report of a case. Clin. Exp. Rheumatol. 2002, 20, 845–847. [Google Scholar]
- Carter, J.D.; Gérard, H.C.; Espinoza, L.R.; Ricca, L.R.; Valeriano, J.; Snelgrove, J.; Oszust, C.; Vasey, F.B.; Hudson, A.P. Chlamydiae as etiologic agents in chronic undifferentiated spondylarthritis. Arthritis Rheum. 2009, 60, 1311–1316. [Google Scholar] [CrossRef] [PubMed]
- Contini, C.; Grilli, A.; Badia, L.; Guardigni, V.; Govoni, M.; Seraceni, S. Detection of Chlamydophila pneumoniae in patients with arthritis: Significance and diagnostic value. Rheumatol Int. 2011, 31, 1307–1313. [Google Scholar] [CrossRef] [PubMed]
- Rizzo, A.; Domenico, M.D.; Carratelli, C.R.; Paolillo, R. The role of Chlamydia and Chlamydophila infections in reactive arthritis. Intern Med. 2012, 51, 113–117. [Google Scholar] [CrossRef] [PubMed]
- Kvien, T.K.; Glennås, A.; Melby, K.; Granfors, K.; Andrup, O.; Karstensen, B.; Thoen, J.E. Reactive arthritis: Incidence, triggering agents and clinical presentation. J. Rheumatol. 1994, 21, 115–122. [Google Scholar]
- Telyatnikova, N.; Hill Gaston, J.S. Prior exposure to infection with Chlamydia pneumoniae can influence the T-cell-mediated response to Chlamydia trachomatis. FEMS Immunol. Med. Microbiol. 2006, 47, 190–198. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bojović, J.; Strelić, N.; Pavlica, L. Reiter’s syndrome—Disease of young men—Analysis of 312 patients. Med. Pregl. 2014, 67, 222–230. [Google Scholar] [CrossRef]
- Pal, Z.; Urbán, E.; Dósa, E.; Pál, A.; Nagy, E. Biofilm formation on intrauterine devices in relation to duration of use. J. Med. Microbiol. 2005, 54, 1199–1203. [Google Scholar] [CrossRef] [PubMed]
- Auler, M.E.; Morreira, D.; Rodrigues, F.F.; Abr Ao, M.S.; Margarido, P.F.; Matsumoto, F.E.; Silva, E.G.; Silva, B.C.; Schneider, R.P.; Paula, C.R. Biofilm formation on intrauterine devices in patients with recurrent vulvovaginal candidiasis. Med. Mycol. 2010, 48, 211–216. [Google Scholar] [CrossRef]
- Dean, D.; Suchland, R.J.; Stamm, W.E. Evidence for long-term cervical persistence of Chlamydia trachomatis by omp1 genotyping. J. Infect. Dis. 2000, 182, 909–916. [Google Scholar] [CrossRef] [PubMed]
- Suchland, R.J.; Dimond, Z.E.; Putman, T.E.; Rockey, D.D. Demonstration of persistent infections and genome stability by whole-genome sequencing of repeat-positive, same-serovar Chlamydia trachomatis collected from the female genital tract. J. Infect. Dis. 2017, 215, 1657–1665. [Google Scholar] [CrossRef] [PubMed]
- Di Pietro, M.; Filardo, S.; De Santis, F.; Sessa, R. New insights into Chlamydiae persistence: An energy metabolism strategy? Int. J. Immunopathol. Pharmacol. 2013, 26, 525–528. [Google Scholar] [CrossRef] [PubMed]
- Frohlich, K.M.; Hua, Z.; Quayle, A.J.; Wang, J.; Lewis, M.E.; Chou, C.W.; Luo, M.; Buckner, L.R.; Shen, L. Membrane vesicle production by Chlamydia trachomatis as an adaptive response. Front. Cell. Infect. Microbiol. 2014, 4, 73. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Di Pietro, M.; Filardo, S.; Romano, S.; Sessa, R. Chlamydia trachomatis and Chlamydia pneumoniae Interaction with the Host: Latest Advances and Future Prospective. Microorganisms 2019, 7, 140. https://doi.org/10.3390/microorganisms7050140
Di Pietro M, Filardo S, Romano S, Sessa R. Chlamydia trachomatis and Chlamydia pneumoniae Interaction with the Host: Latest Advances and Future Prospective. Microorganisms. 2019; 7(5):140. https://doi.org/10.3390/microorganisms7050140
Chicago/Turabian StyleDi Pietro, Marisa, Simone Filardo, Silvio Romano, and Rosa Sessa. 2019. "Chlamydia trachomatis and Chlamydia pneumoniae Interaction with the Host: Latest Advances and Future Prospective" Microorganisms 7, no. 5: 140. https://doi.org/10.3390/microorganisms7050140
APA StyleDi Pietro, M., Filardo, S., Romano, S., & Sessa, R. (2019). Chlamydia trachomatis and Chlamydia pneumoniae Interaction with the Host: Latest Advances and Future Prospective. Microorganisms, 7(5), 140. https://doi.org/10.3390/microorganisms7050140