Effects of Lactic Acid Bacteria Inoculants on Fermentation Quality, Bacteria Communities and Antibiotic Resistance Genes in Whole-Crop Corn Silage
Abstract
1. Introduction
2. Materials and Methods
2.1. Silage Preparation
2.2. Fermentation Characteristics and Chemical Compositions Analyses
2.3. DNA Extraction
2.4. Quantification of ARGs and MGEs
2.5. Microbial Diversity Analysis
2.6. Date and Statistical Analysis
3. Results
3.1. Fermentation Characteristics and Chemical Compositions of Whole-Crop Corn Silage
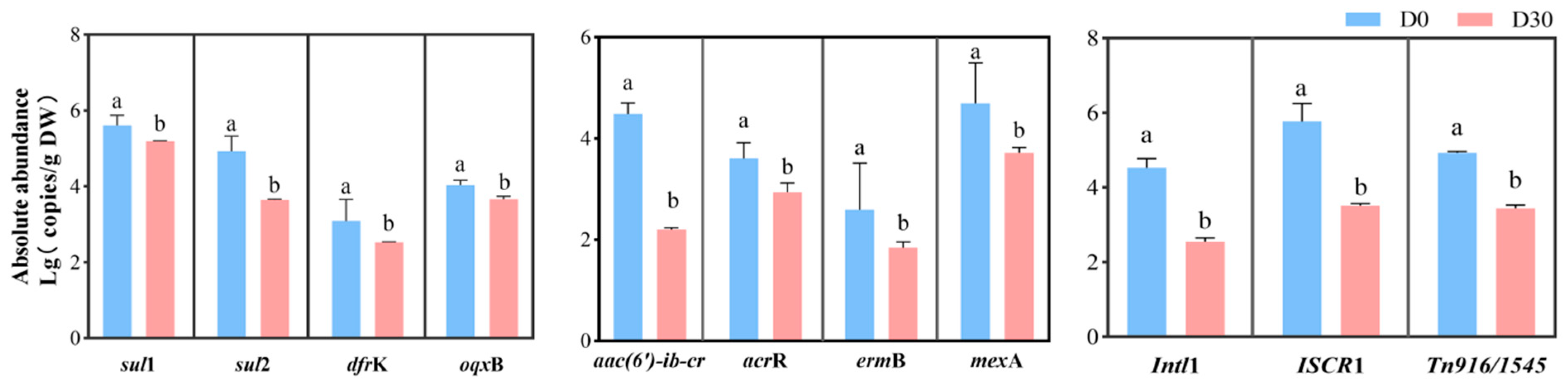
3.2. Abundance Changes in ARGs and MGEs During Silage
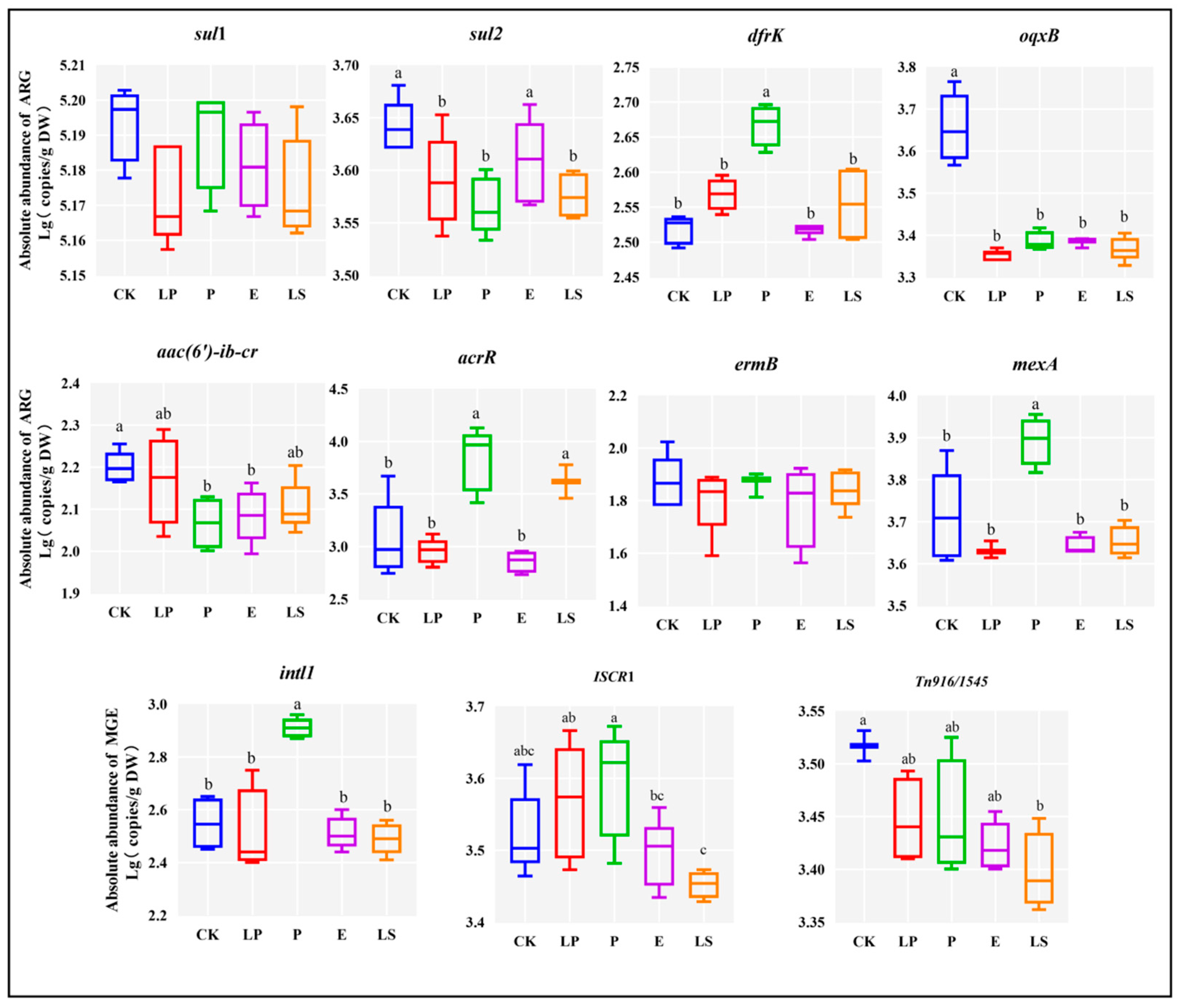
3.3. Effect of LAB on the Removal of ARGs and MGEs in Silage
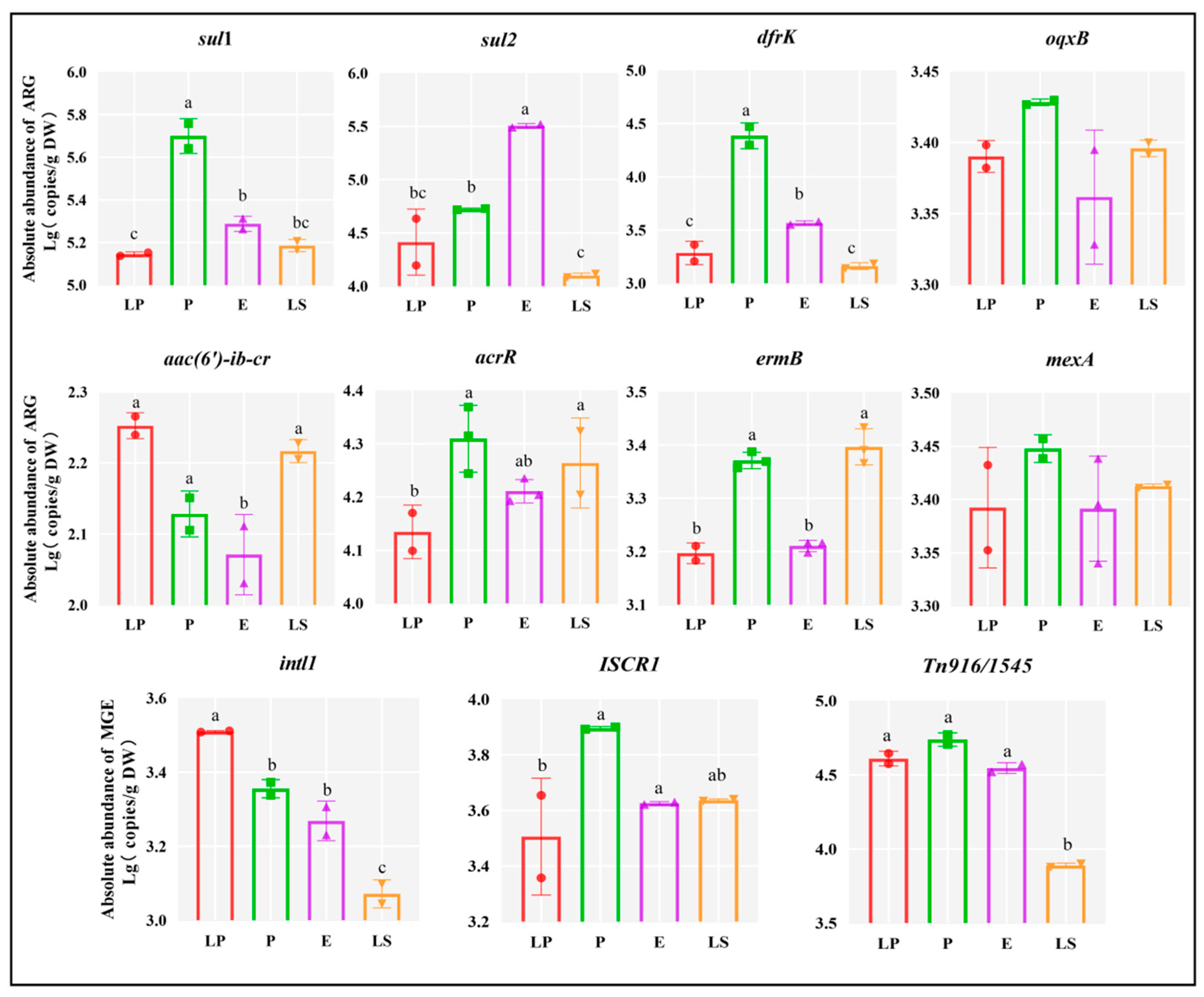
3.4. ARGs and MGEs in Lactic Acid Bacteria Strains
3.5. Changes in the Bacterial Community Structure of Silage After LAB Treatment
3.5.1. Bacterial Alpha Diversity
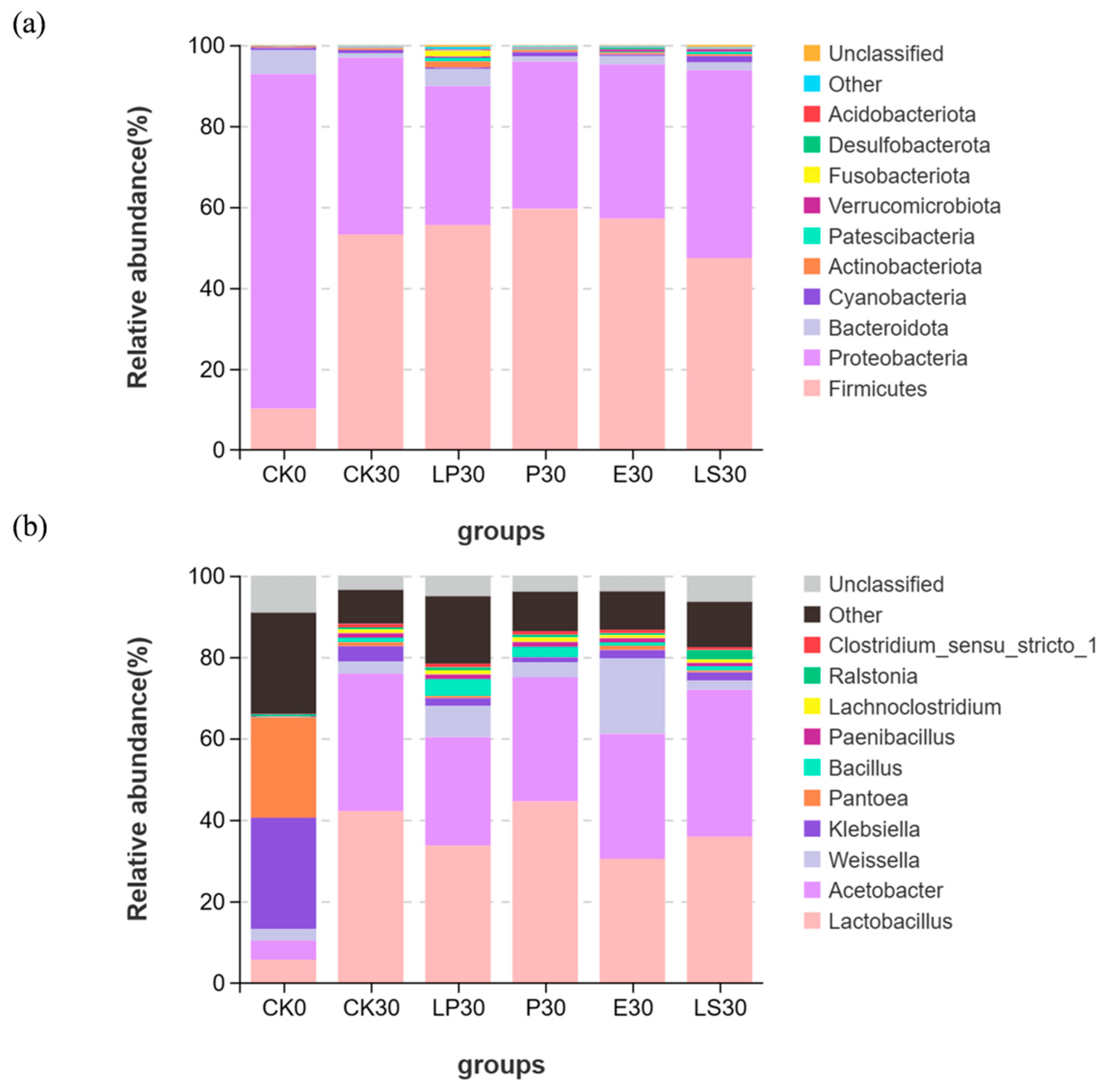
3.5.2. Microbial Community Structure
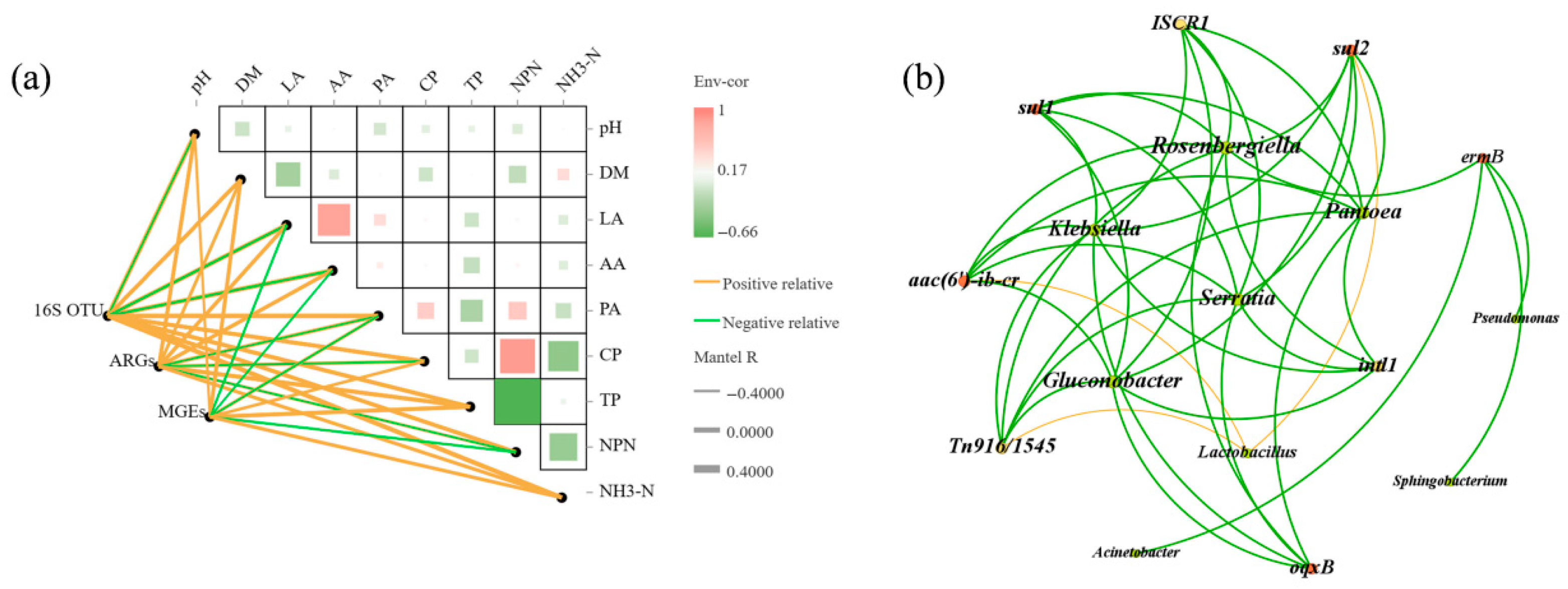
3.6. Relationships Between Silage Quality, ARGs, and the Bacterial Community Structure
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Oliver, J.P.; Gooch, C.A.; Lansing, S.; Schueler, J.; Hurst, J.J.; Sassoubre, L.; Crossette, E.M.; Aga, D.S. Invited review: Fate of antibiotic residues, antibiotic-resistant bacteria, and antibiotic resistance genes in US dairy manure management systems. J. Dairy Sci. 2020, 103, 1051–1071. [Google Scholar] [CrossRef] [PubMed]
- Chaturvedi, P.; Shukla, P.; Giri, B.S.; Chowdhary, P.; Chandra, R.; Gupta, P.; Pandey, A. Prevalence and hazardous impact of pharmaceutical and personal care products and antibiotics in environment: A review on emerging contaminants. Environ. Res. 2021, 194, 110664. [Google Scholar] [CrossRef] [PubMed]
- Deng, F.; Yao, C.; Ke, L.; Chen, M.; Huang, M.; Wen, J.; Chen, Q.; Jiang, J.; Deng, Y. Emerging threat to antibiotic resistance: Impact of mycotoxin deoxynivalenol on gut microbiota and clonal expansion of extensively drug-resistant enterococci. Environ. Int. 2025, 197, 109353. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.L.; An, X.L.; Zheng, B.X.; Ma, Y.B.; Su, J.Q. Long-term organic fertilization increased antibiotic resistome in phyllosphere of maize. Sci. Total Environ. 2018, 645, 1230–1237. [Google Scholar] [CrossRef] [PubMed]
- Zhou, G.; Qiu, X.; Wu, X.; Lu, S. Horizontal gene transfer is a key determinant of antibiotic resistance genes profiles during chicken manure composting with the addition of biochar and zeolite. J. Hazard. Mater. 2021, 408, 124883. [Google Scholar] [CrossRef]
- Wang, X.; Lu, Y.; Yan, Y.; Wang, R.; Wang, Y.; Li, H.; Zhou, L.; Zheng, G.; Zhang, Y. Pivotal role of intracellular oxidation by HOCl in simultaneously removing antibiotic resistance genes and enhancing dewaterability during conditioning of sewage sludge using Fe2+/Ca(ClO)2. Water Res. 2024, 254, 121414. [Google Scholar] [CrossRef]
- Ren, J.; Xu, C.; Zhang, J.; Chen, X.; Yin, D.; Li, C.; Huhe, T.; Niu, D. Anaerobic storage of erythromycin fermentation residue for sustainable management and antibiotic resistance mitigation. ACS Sustain. Chem. Eng. 2025, 13, 3806–3816. [Google Scholar] [CrossRef]
- Zhang, Q.; Zou, X.; Wu, S.; Wu, N.; Chen, X.; Zhou, W. Effects of pyroligneous acid on diversity and dynamics of antibiotic resistance genes in alfalfa silage. Microbiol. Spectr. 2022, 10, e155422. [Google Scholar] [CrossRef]
- Reuben, R.C.; Roy, P.C.; Sarkar, S.L.; Alam, R.; Jahid, I.K. Isolation, characterization, and assessment of lactic acid bacteria toward their selection as poultry probiotics. BMC Microbiol. 2019, 19, 253. [Google Scholar] [CrossRef]
- Qu, T.; Wang, P.; Zhao, X.; Liang, L.; Ge, Y.; Chen, Y. Metagenomics reveals differences in the composition of bacterial antimicrobial resistance and antibiotic resistance genes in pasteurized yogurt and probiotic bacteria yogurt from China. J. Dairy Sci. 2024, 107, 3451–3467. [Google Scholar] [CrossRef]
- Dang, Q.; Kong, L.; Liu, J.; Liu, S.; Wei, X.; Ma, H.; Min, W. Isolation and identification of lactic acid bacteria for fermentation of pickles and analysis of drug resistance. Food Sci. 2019, 40, 166–170. [Google Scholar]
- Zhang, X.; Ding, Z.; Usman, S.; Zhang, J.; Chen, M.; Guo, X. Metagenomics insights into the effects of lactic acid bacteria inoculation on the biological reduction of antibiotic resistance genes in alfalfa silage. J. Hazard. Mater. 2023, 443, 130329. [Google Scholar] [CrossRef]
- Xu, D.; Wang, N.; Rinne, M.; Ke, W.; Weinberg, Z.G.; Da, M.; Bai, J.; Zhang, Y.; Li, F.; Guo, X. The bacterial community and metabolome dynamics and their interactions modulate fermentation process of whole crop corn silage prepared with or without inoculants. Microb. Biotechnol. 2021, 14, 561–576. [Google Scholar] [CrossRef]
- Wang, C.; He, L.; Xing, Y.; Zhou, W.; Yang, F.; Chen, X.; Zhang, Q. Fermentation quality and microbial community of alfalfa and stylo silage mixed with Moringa oleifera leaves. Bioresour. Technol. 2019, 284, 240–247. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; He, L.; Xing, Y.; Zheng, Y.; Zhou, W.; Pian, R.; Yang, F.; Chen, X.; Zhang, Q. Dynamics of bacterial community and fermentation quality during ensiling of wilted and unwilted Moringa oleifera leaf silage with or without lactic acid bacterial inoculants. mSphere 2019, 4, e00341-19. [Google Scholar] [CrossRef] [PubMed]
- Sanz, C.; Casadoi, M.; Tadic, D.; Pastor-Lopez, E.J.; Navarro-Martin, L.; Parera, J.; Tugues, J.; Ortiz, C.A.; Bayona, J.M.; Pina, B. Impact of organic soil amendments in antibiotic levels, antibiotic resistance gene loads, and microbiome composition in corn fields and crops. Environ. Res. 2022, 214, 113760. [Google Scholar] [CrossRef]
- Fang, D.; Hua, S.; Chen, H.; Ji, Z.; Wang, D.; Wang, W.; Shao, T.; Dong, Z. Producing high-quality and safe whole-plant quinoa silage through selecting variety and harvest time. BMC Plant Biol. 2025, 25, 333. [Google Scholar] [CrossRef]
- He, L.; Wang, C.; Xing, Y.; Zhou, W.; Pian, R.; Yang, F.; Chen, X.; Zhang, Q. Dynamics of proteolysis, protease activity and bacterial community of Neolamarckia cadamba leaves silage and the effects of formic acid and Lactobacillus farciminis. Bioresour. Technol. 2019, 294, 122127. [Google Scholar] [CrossRef]
- Xu, D.; Ding, W.; Ke, W.; Li, F.; Zhang, P.; Guo, X. Modulation of metabolome and bacterial community in whole crop corn silage by inoculating homofermentative Lactobacillus plantarum and heterofermentative Lactobacillus buchneri. Front. Microbiol. 2018, 9, 3299. [Google Scholar] [CrossRef]
- Lu, L.; Zuo, Q.; Cheng, Y.; Huang, Y. The mechanism of microbial structure and flavor characteristics in Qing-Jiang-flavor Jiupei regulated by different fermentation seasons. Food Chem. X 2024, 22, 101392. [Google Scholar] [CrossRef]
- Khan, N.A.; Yu, P.; Ali, M.; Cone, J.W.; Hendriks, W.H. Nutritive value of maize silage in relation to dairy cow performance and milk quality. J. Sci. Food. Agric. 2015, 95, 238–252. [Google Scholar] [CrossRef]
- Chen, D.; Zheng, M.; Zhou, Y.; Gao, L.; Zhou, W.; Wang, M.; Zhu, Y.; Xu, W. Improving the quality of Napier grass silage with pyroligneous acid: Fermentation, aerobic stability, and microbial communities. Front. Microbiol. 2022, 13, 1034198. [Google Scholar] [CrossRef]
- Guan, H.; Li, H.; Gan, L.; Chen, S.; Yan, Y.; Jia, Z.; Liu, W.; Wei, X.; Ma, X.; Zhou, Q. The effects of native lactic acid bacteria on the microbiome, fermentation profile, and nutritive value of Napier grass silage prepared with different legume ratios. Front. Microbiol. 2022, 13, 1112058. [Google Scholar] [CrossRef]
- Peng, Z.; Xiong, T.; Huang, T.; Xu, X.; Fan, P.; Qiao, B.; Xie, M. Factors affecting production and effectiveness, performance improvement and mechanisms of action of bacteriocins as food preservative. Crit. Rev. Food. Sci. Nutr. 2023, 63, 12294–12307. [Google Scholar] [CrossRef]
- He, X.; Cui, Y.; Jia, Q.; Zhuang, Y.; Gu, Y.; Fan, X.; Ding, Y. Response mechanisms of lactic acid bacteria under environmental stress and their application in the food industry. Food Biosci. 2025, 64, 16. [Google Scholar] [CrossRef]
- Zhao, J.; Li, X.; Liu, H.; Jing, Z.; Yin, X.; Li, J.; Dong, Z.; Shao, T. Ensilage using Leuconostoc lactis and Weissella confusa reduces microbial risk and enhances hygienic quality of whole-crop corn. Chem. Biol. Technol. Agric. 2024, 11, 14. [Google Scholar] [CrossRef]
- Xu, D.; Zhang, X.; Yuan, X.; Han, H.; Xue, Y.; Guo, X. Hazardous risk of antibiotic resistance genes: Host occurrence, distribution, mobility and vertical transmission from different environments to corn silage. Environ. Pollut. 2023, 338, 11. [Google Scholar] [CrossRef]
- Liu, H.; Ye, X.; Chen, S.; Sun, A.; Duan, X.; Zhang, Y.; Zou, H.; Zhang, Y. Chitosan as additive affects the bacterial community, accelerates the removals of antibiotics and related resistance genes during chicken manure composting. Sci. Total Environ. 2021, 792, 148381. [Google Scholar] [CrossRef]
- Yue, Z.; Zhang, J.; Zhou, Z.; Ding, C.; Wan, L.; Liu, J.; Chen, L.; Wang, X. Pollution characteristics of livestock faeces and the key driver of the spread of antibiotic resistance genes. J. Hazard. Mater. 2021, 409, 124957. [Google Scholar] [CrossRef]
- Liu, R.; Liu, L.; Liu, Y.; Wang, L. Comprehensive evaluation of antibiotic pollution in a typical tributary of the Yellow River, China: Source-specific partitioning and fate analysis. J. Hazard. Mater. 2025, 488, 137294. [Google Scholar] [CrossRef]
- Hameed, P.S.; Kotakonda, H.; Sharma, S.; Nandishaiah, R.; Katagihallimath, N.; Rao, R.; Sadler, C.; Slater, I.; Morton, M.; Chandrasekaran, A.; et al. BWC0977, a broad-spectrum antibacterial clinical candidate to treat multidrug resistant infections. Nat. Commun. 2024, 15, 8202. [Google Scholar] [CrossRef]
- Wu, Z.; Luo, Y.; Bao, J.; Luo, Y.; Yu, Z. Additives affect the distribution of metabolic profile, microbial communities and antibiotic resistance genes in high-moisture sweet corn kernel silage. Bioresour. Technol. 2020, 315, 123821. [Google Scholar] [CrossRef]
- Xu, D.; Zhang, X.; Usman, S.; Bai, J.; Sheoran, N.; Guo, X. Reducing transmission of high-risk antibiotic resistance genes in whole-crop corn silage through lactic acid bacteria inoculation and increasing ensiling temperature. Sci. Total Environ. 2024, 926, 172114. [Google Scholar] [CrossRef]
- Huerta, B.; Marti, E.; Gros, M.; Lopez, P.; Pompeo, M.; Armengol, J.; Barcelo, D.; Balcazar, J.L.; Rodriguez-Mozaz, S.; Marce, R. Exploring the links between antibiotic occurrence, antibiotic resistance, and bacterial communities in water supply reservoirs. Sci. Total Environ. 2013, 456–457, 161–170. [Google Scholar] [CrossRef]
- Alam, M.; Hassan, T.; Nizami, T.A.; Akter, L.; Rakib, T.M. Draft genome sequences of four lactic acid bacteria from fermented chicken meat unveil biosynthetic gene clusters for antimicrobial compounds. Microbiol. Resour. Announc. 2025, 14, e3825. [Google Scholar] [CrossRef] [PubMed]
- Han, H.; Wang, C.; Huang, Z.; Zhang, Y.; Sun, L.; Xue, Y.; Guo, X. Effects of lactic acid Bacteria-Inoculated corn silage on bacterial communities and metabolites of digestive tract of sheep. Fermentation 2022, 8, 320. [Google Scholar] [CrossRef]
- Pham, H.; Kim, D.; Nguyen, T.L. Wide-genome selection of lactic acid bacteria harboring genes that promote the elimination of antinutritional factors. Front. Plant Sci. 2023, 14, 1145041. [Google Scholar] [CrossRef]
- Bengtsson-Palme, J.; Kristiansson, E.; Larsson, D. Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiol. Rev. 2018, 42, fux053. [Google Scholar] [CrossRef] [PubMed]
- Xie, W.Y.; Shen, Q.; Zhao, F.J. Antibiotics and antibiotic resistance from animal manures to soil: A review. Eur. J. Soil Sci. 2018, 69, 181–195. [Google Scholar] [CrossRef]
- Jacobsen, L.; Wilcks, A.; Hammer, K.; Huys, G.; Gevers, D.; Andersen, S.R. Horizontal transfer of tet(M) and erm(B) resistance plasmids from food strains of Lactobacillus plantarum to Enterococcus faecalis JH2-2 in the gastrointestinal tract of gnotobiotic rats. FEMS Microbiol. Ecol. 2007, 59, 158–166. [Google Scholar] [CrossRef]
- Luo, L.; Wang, Z.; Huang, X.; Gu, J.D.; Yu, C.; Deng, O. The fate of antibiotic resistance genes in wastewater containing microalgae treated by chlorination, ultra-violet, and Fenton reaction. Water Res. 2024, 254, 121392. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Sun, Y.; Huang, K.; Gao, Y.; Lin, Y.; Yuan, B.; Wang, X.; Xu, G.; Nussio, L.G.; Yang, F.; et al. Multi-omics analysis reveals the core microbiome and biomarker for nutrition degradation in alfalfa silage fermentation. mSystems 2024, 9, e68224. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Usman, S.; Bature, I.; Xu, D.; Guo, X. Occurrence and fate of antibiotic-resistance genes and their potential hosts in high-moisture alfalfa silage treated with or without formic acid bactericide. J. Environ. Manag. 2023, 347, 119235. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.; Li, H.; Wu, Z.; Li, S.; Cao, Z.; Ma, B.; Zou, Y.; Zhang, N.; Liu, Z.; Wang, Y.; et al. The addition of nano zero-valent iron during compost maturation effectively removes intracellular and extracellular antibiotic resistance genes by reducing the abundance of potential host bacteria. Bioresour. Technol. 2023, 384, 129350. [Google Scholar] [CrossRef]
| Gene | Primer (5′→3′) | Bp | Tm |
|---|---|---|---|
| oqxB | TTCTCCCCCGGCGGGAAGTAC | 139 | 60 |
| CTCGGCCATTTTGGCGCGTA | |||
| sul1 | CACCGGAAACATCGCTGCA | 158 | 55 |
| AAGTTCCGCCGCAAGGCT | |||
| sul2 | CTCCGATGGAGGCCGGTAT | 190 | 55 |
| GGGAATGCCATCTGCCTTGA | |||
| dfrk | CTGCACCAGCCCCTTGAATA | 211 | 55 |
| CCTGCGAGTACAAACTGGGT | |||
| Tn916/1545 | TCCTACAGCGACAGCCAGTGA | 174 | 54 |
| TGCGTTGCTTTGGTCTGCTGGT | |||
| acrR | TGGCGATCCACTCTCAGTAT | 138 | 55 |
| TGTTGCACAACAGCCATTTC | |||
| ermB | AAAACTTACCCGCCATACCA | 139 | 53 |
| TTTGGCGTGTTTCATTGCTT | |||
| intl1 | CTGGATTTCGATCACGGCACG | 473 | 60 |
| ACATGCGTGTAAATCATCGTCG | |||
| mexA | CTCGAATTCTCCGAGGTTTCCG | 143 | 59 |
| AGGATGGCCTTCTGCTTGAC | |||
| aac(6′)-ib-cr | TTGCGATGCTCTATGAGTGGCTA | 482 | 55 |
| CTCGAATGCCTGGCGTGTTT | |||
| ISCR1 | CGCCCACTCAAACAAACG | 452 | 54 |
| GAGGCTTTGGTGTAACCG |
| Item | Treatment | SEM | p Value | ||||
|---|---|---|---|---|---|---|---|
| CK | LP | P | E | LS | |||
| Dry matter (% FW) | 24.86 | 24.56 | 24.75 | 25.70 | 24.88 | 0.87 | NS |
| pH | 3.69 a | 3.63 b | 3.68 a | 3.67 ab | 3.66 ab | 0.03 | <0.01 |
| Lactic acid bacteria (lg cfu/g FW) | 6.56 | 6.12 | 6.78 | 6.40 | 6.42 | 0.40 | NS |
| Yeasts (lg cfu/g FW) | 5.41 b | 5.45 ab | 5.38 b | 5.53 ab | 5.72 a | 0.21 | <0.05 |
| Mold (lg cfu/g FW) | <2.00 | <2.00 | <2.00 | <2.00 | <2.00 | - | - |
| Coliform bacteria (lg cfu/g FW) | <2.00 | <2.00 | <2.00 | <2.00 | <2.00 | - | - |
| Lactic acid (g/kg DM) | 71.22 ab | 60.76 b | 91.35 a | 69.43 ab | 65.20 b | 17.79 | <0.05 |
| Acetic acid (g/kg DM) | 30.90 ab | 24.83 b | 37.49 a | 27.74 ab | 26.48 b | 7.68 | <0.05 |
| Propionic acid (g/kg DM) | 2.91 ab | 1.35 b | 7.60 a | 4.96 ab | 5.34 ab | 3.89 | <0.05 |
| Crude protein (g/kg DM) | 83.95 | 85.54 | 85.92 | 85.44 | 85.80 | 3.91 | NS |
| True protein (g/kg DM) | 43.92 a | 40.91 b | 38.68 b | 39.84 b | 44.26 a | 2.96 | <0.01 |
| NH3-N (g/kg DM) | 1.35 a | 0.89 b | 0.89 b | 1.19 ab | 0.89 b | 0.32 | <0.05 |
| Treatment | Sobs | Shannon | Simpson | Chao | Ace | Goods_Coverage |
|---|---|---|---|---|---|---|
| CK0 | 644.00 | 3.69 | 0.83 | 790.78 | 837.57 | 0.99 |
| CK30 | 625.00 | 4.04 | 0.87 | 733.92 | 756.81 | 0.99 |
| LP30 | 667.40 | 4.69 | 0.90 | 768.32 | 797.93 | 0.99 |
| P30 | 592.40 | 3.99 | 0.84 | 703.86 | 723.74 | 0.99 |
| E30 | 539.40 | 3.99 | 0.84 | 631.55 | 640.88 | 0.99 |
| LS30 | 525.60 | 4.14 | 0.87 | 650.42 | 677.36 | 0.99 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, D.; Yang, D.; Guo, T.; Zhang, Q. Effects of Lactic Acid Bacteria Inoculants on Fermentation Quality, Bacteria Communities and Antibiotic Resistance Genes in Whole-Crop Corn Silage. Microorganisms 2025, 13, 1977. https://doi.org/10.3390/microorganisms13091977
Chen D, Yang D, Guo T, Zhang Q. Effects of Lactic Acid Bacteria Inoculants on Fermentation Quality, Bacteria Communities and Antibiotic Resistance Genes in Whole-Crop Corn Silage. Microorganisms. 2025; 13(9):1977. https://doi.org/10.3390/microorganisms13091977
Chicago/Turabian StyleChen, Dandan, Dan Yang, Tianxin Guo, and Qing Zhang. 2025. "Effects of Lactic Acid Bacteria Inoculants on Fermentation Quality, Bacteria Communities and Antibiotic Resistance Genes in Whole-Crop Corn Silage" Microorganisms 13, no. 9: 1977. https://doi.org/10.3390/microorganisms13091977
APA StyleChen, D., Yang, D., Guo, T., & Zhang, Q. (2025). Effects of Lactic Acid Bacteria Inoculants on Fermentation Quality, Bacteria Communities and Antibiotic Resistance Genes in Whole-Crop Corn Silage. Microorganisms, 13(9), 1977. https://doi.org/10.3390/microorganisms13091977






