Extracellular Cr(VI) Reduction by the Salt-Tolerant Strain Bacillus safensis BSF-4
Abstract
1. Introduction
2. Materials and Methods
2.1. Isolation, Identification, and Preservation of the BSF-4 Strain
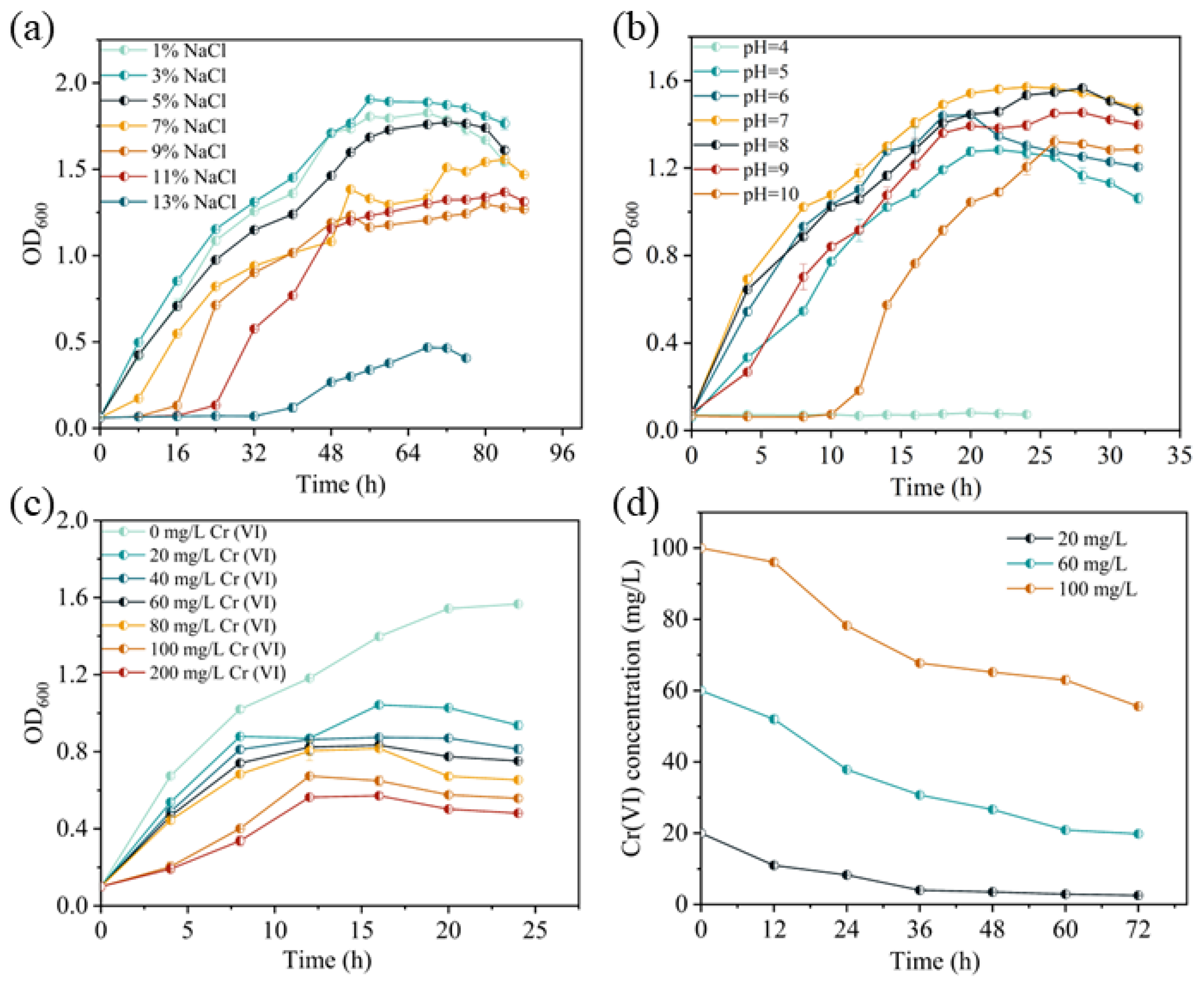
2.2. Culture Conditions of the BSF-4 Strain
2.3. Reduction Mechanism of Cr(VI) by BSF-4 Strain
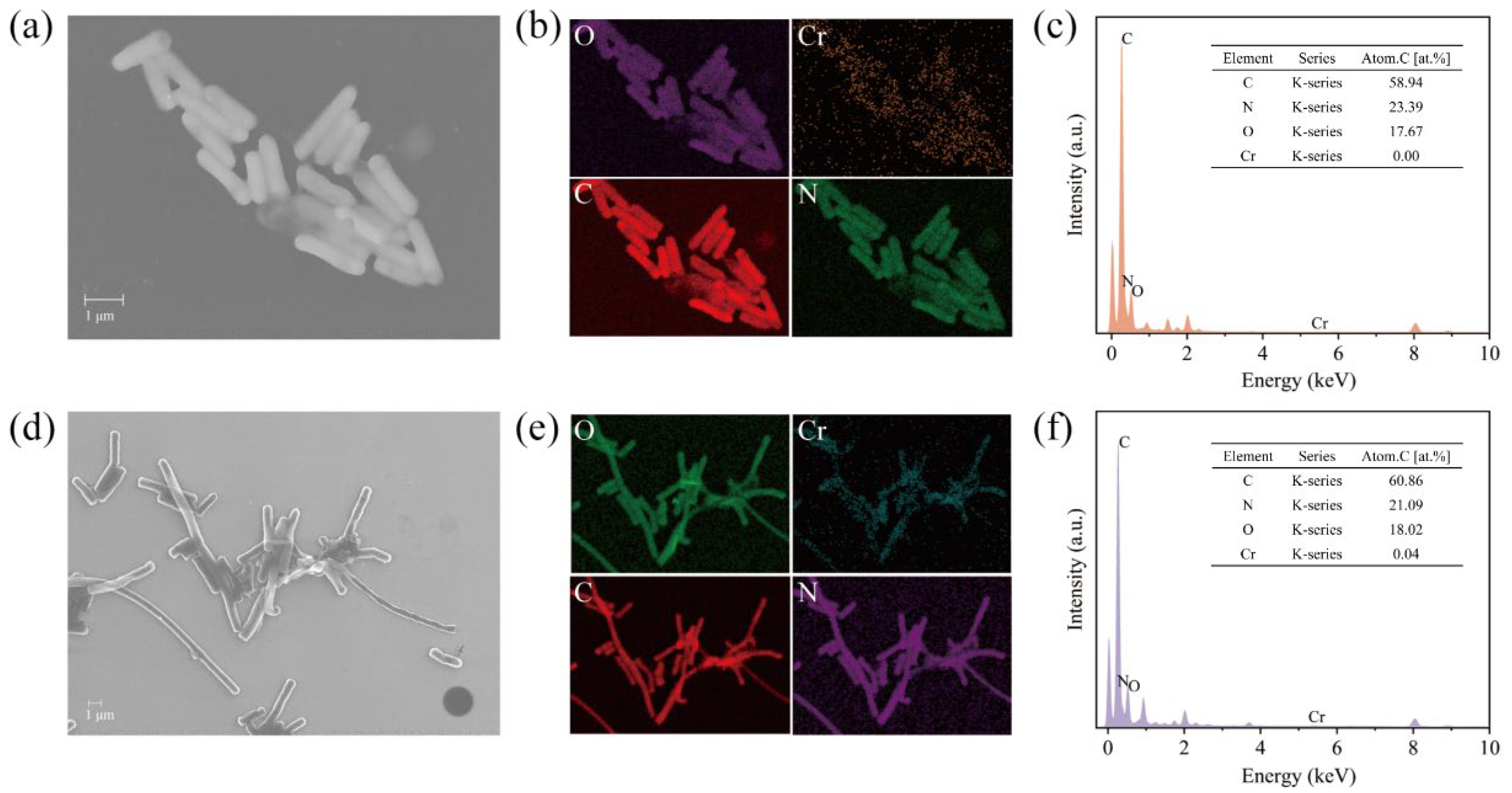
2.3.1. SEM-EDS Analysis [33]
2.3.2. Chromium Speciation Analysis
- (1)
- Extracellular: Transfer the culture medium to be measured into a centrifuge tube, centrifuge at 5000 rpm for 30 min, take the supernatant, and determine Cr(VI) and total Cr concentration.
- (2)
- Intracellular: The bacteria precipitated in the above step were repeatedly cleaned with phosphate buffer (pH = 7) for two times. After re-suspension with deionized water, the bacteria were broken by ultrasound in ice, centrifuged at 5000 rpm for 30 min, then the supernatant was taken to determine Cr(VI) and total Cr concentration.
2.3.3. Cr(VI) Reduction Capacity of Bacterial Cell Components
- (1)
- Extracellular metabolites: The bacteria were inoculated into nutrient medium, incubated in oscillatory culture at 30 °C and 150 rpm until the logarithmic phase, centrifuged at 5000 rpm for 30 min, and the supernatant was taken. Extracellular metabolites were sampled after 0.22 μM.
- (2)
- Intact cells: The cell precipitation in the previous step was repeatedly cleaned with phosphate buffer (pH = 7) for two to three times, divided into three parts, and one part was resuspended with deionized water, that is, to obtain an intact cell sample.
- (3)
- Resting cells: Pelleted biomass underwent two cycles of phosphate-buffered saline (100 mM, pH = 7) washing and reconstitution, yielding metabolically quiescent cellular suspensions.
- (4)
- Intracellular crude enzyme solution: A portion of the bacterial sediment obtained in step (2) was re-suspended with phosphate buffer, placed in an ice bath ultrasonic bacteria removal (100 W, 20 min), centrifuged at 5000 rpm for 30 min, and the supernatant was taken as the crude enzyme extract sample.
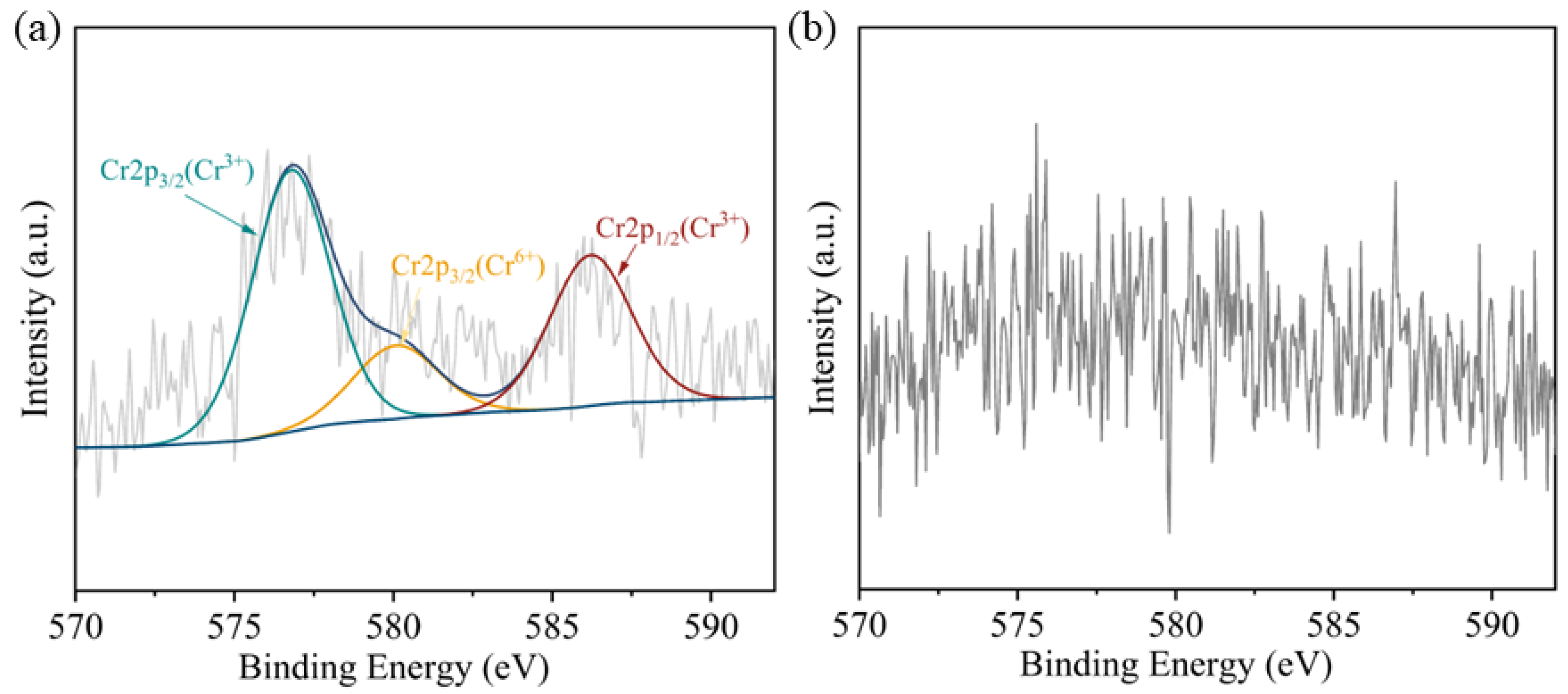
2.3.4. XPS Analysis
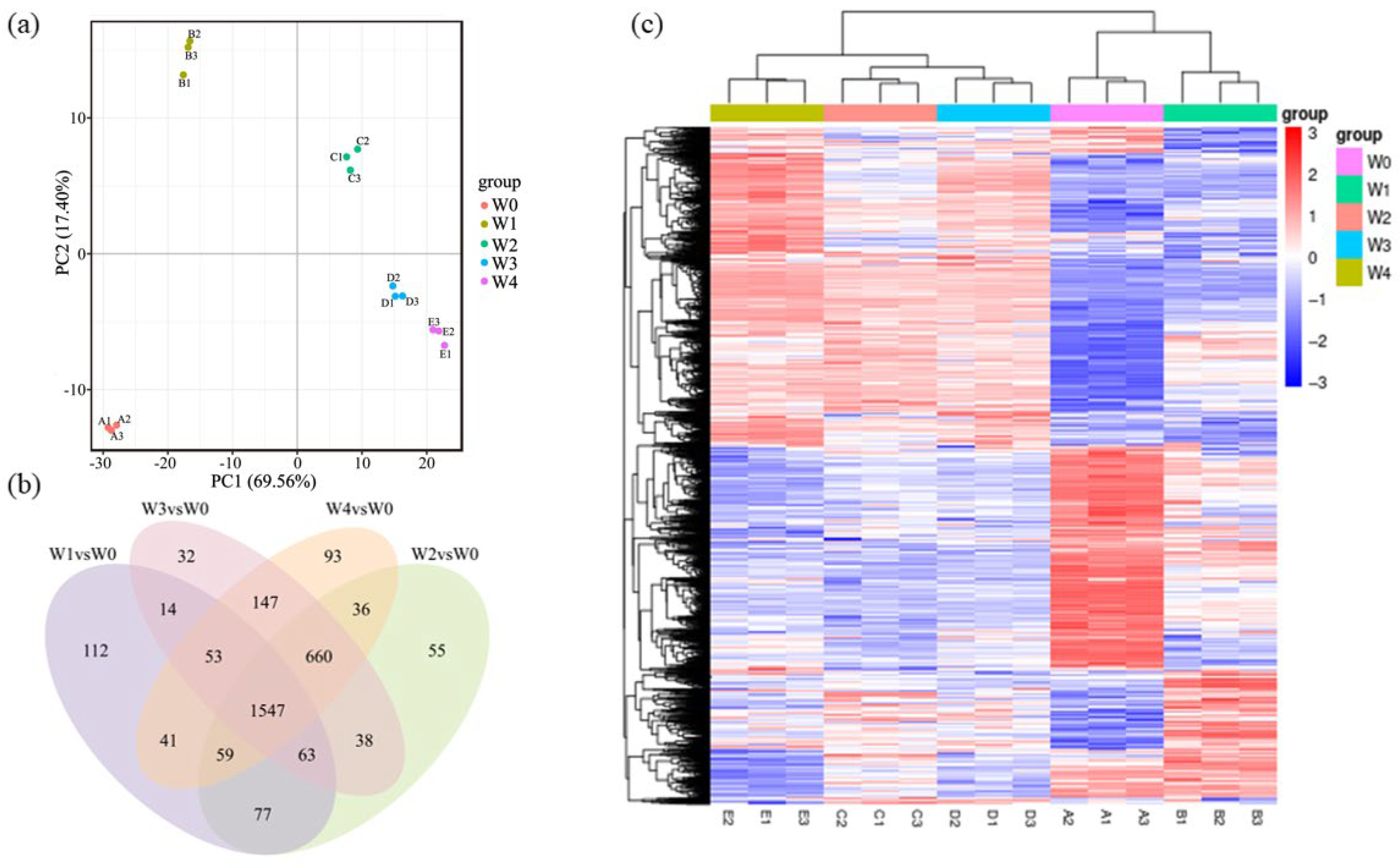
2.4. Transcriptomics Analysis
3. Results and Discussion
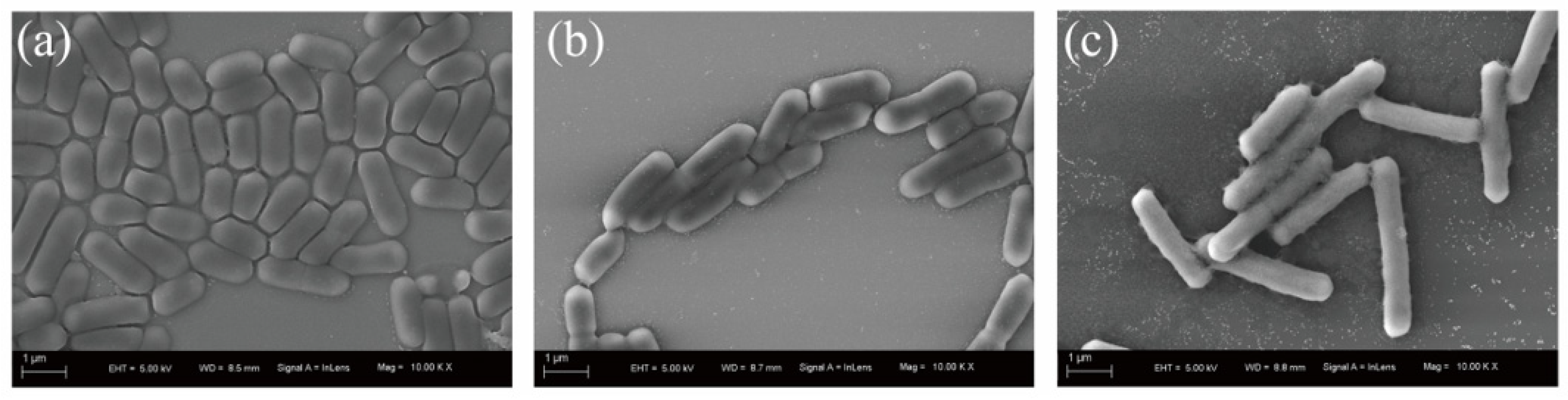
3.1. Identification and Characterization of Strain BSF-4
3.2. Reduction Mechanism of Cr(VI) by Strain BSF-4
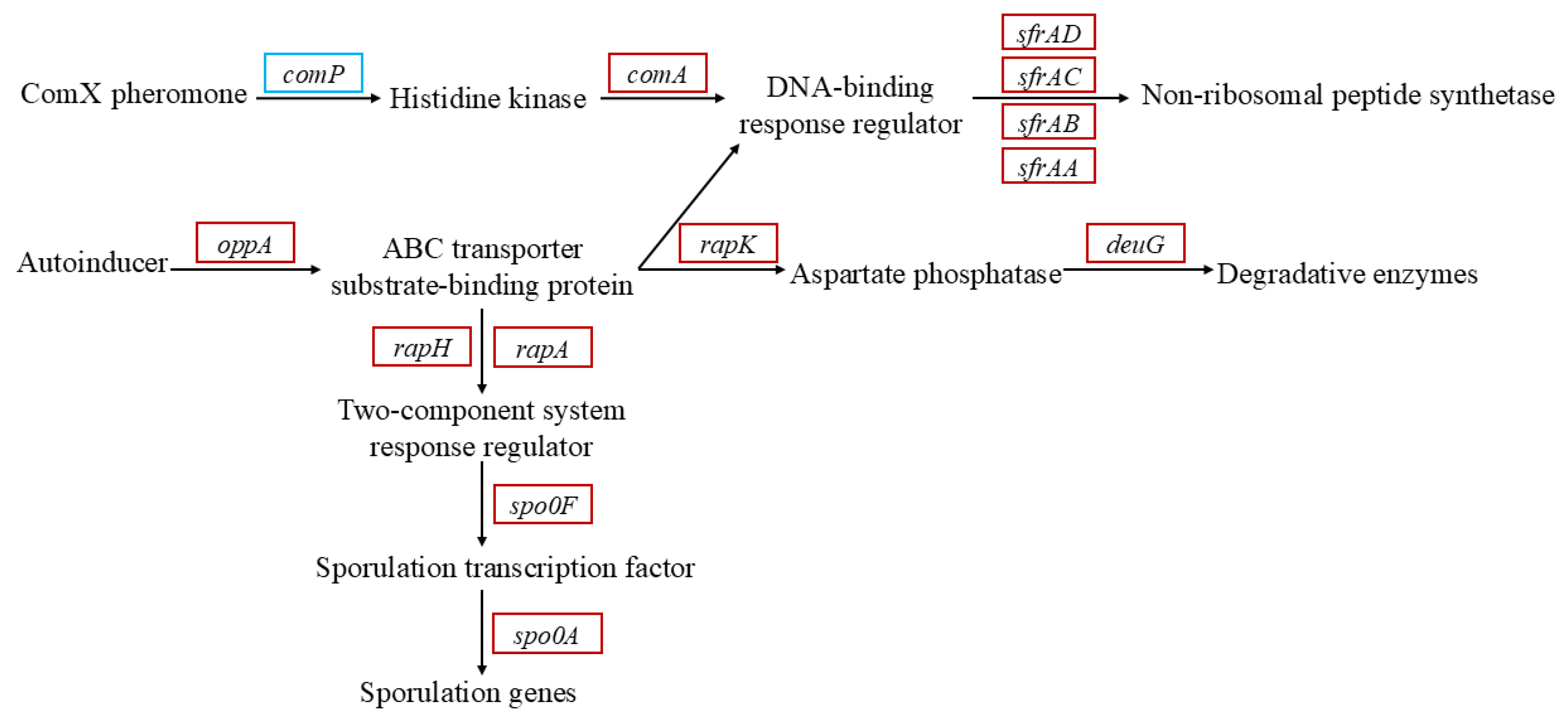
3.3. Identification of Genes Involved in Cr(VI) Reduction
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Dutzi, J.; Boukis, N.; Sauer, J. Supercritical water gasification of heavy metal contaminated plants with focus on separation of heavy metal contaminants. Biomass Bioenergy 2024, 182, 107059. [Google Scholar] [CrossRef]
- Kholisa, B.; Matsena, M.T.; Chirwa, E.M.N. In Situ Bioreduction of Hexavalent Chromium-Contaminated Water Using a Microbial Culture Barrier. Chem. Eng. Technol. 2023, 46, 1307–1311. [Google Scholar] [CrossRef]
- Ramli, N.N.; Kurniawan, S.B.; Ighalo, J.O.; Mohd Said, N.S.; Marsidi, N.; Buhari, J.; Ramli Shah, R.A.; Zulkifli, M.; Alias, J.; Daud, N.M.; et al. A review of the treatment technologies for hexavalent chromium contaminated water. BioMetals 2023, 36, 1189–1219. [Google Scholar] [CrossRef]
- Azeez, N.A.; Dash, S.S.; Gummadi, S.N.; Deepa, V.S. Nano-remediation of toxic heavy metal contamination: Hexavalent chromium [Cr(VI)]. Chemosphere 2021, 266, 129204. [Google Scholar] [CrossRef]
- Ferro Orozco, A.M.; Contreras, E.M.; Zaritzky, N.E. Biological removal of hexavalent chromium: Evaluation of the metabolic activity of native and Cr(VI)-acclimated activated sludge using a respirometric method. Environ. Technol. 2019, 42, 952–963. [Google Scholar] [CrossRef]
- Zhong, J.; Yin, W.; Li, Y.; Li, P.; Wu, J.; Jiang, G.; Gu, J.; Liang, H. Column study of enhanced Cr(VI) removal and longevity by coupled abiotic and biotic processes using Fe0 and mixed anaerobic culture. Water Res. 2017, 122, 536–544. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.T.A.; Al-Battashi, H.; Al-Hinai, M.; Almdawi, M.; Pracejus, B.; Elshafey, E.-S.I.; Abed, R.M.M. Isolation of Aerobic Heterotrophic Bacteria from a Microbial Mat with the Ability to Grow on and Remove Hexavalent Chromium through Biosorption and Bioreduction. Appl. Biochem. Biotechnol. 2024, 197, 94–112. [Google Scholar] [CrossRef] [PubMed]
- Zhao, F.-J.; Ma, Y.; Zhu, Y.-G.; Tang, Z.; McGrath, S.P. Soil Contamination in China: Current Status and Mitigation Strategies. Environ. Sci. Technol. 2014, 49, 750–759. [Google Scholar] [CrossRef] [PubMed]
- GB15618-2018; Soil Environmental Quality—Risk Control Standard for Soil Contamination of Agricultural Land. National Standard of the People’s Republic of China: Beijing, China, 2018.
- GB 3838-2002; Surface Water Environmental Quality Standard. National Standard of the People’s Republic of China: Beijing, China, 2002.
- Vielee, S.T.; Buchanan, W.J.; Roof, S.H.; Kahloon, R.; Evans, E.; Isibor, J.; Patel, M.; Meaza, I.; Lu, H.; Williams, A.R.; et al. Chromium Selectively Accumulates in the Rat Hippocampus after 90 Days of Exposure to Cr(VI) in Drinking Water and Induces Age- and Sex-Dependent Metal Dyshomeostasis. Toxics 2024, 12, 722. [Google Scholar] [CrossRef]
- Ye, T.; Li, H.; Wang, Z.-X.; Huang, R.; Yu, Y.-J.; Yang, Z.; Gao, C.; Xie, C. Transport and fate of hexavalent chromium in slag–soil system. Environ. Earth Sci. 2019, 78, 239. [Google Scholar] [CrossRef]
- Ng, K.K.; Shi, X.; Ong, S.L.; Lin, C.-F.; Ng, H.Y. An innovative of aerobic bio-entrapped salt marsh sediment membrane reactor for the treatment of high-saline pharmaceutical wastewater. Chem. Eng. J. 2016, 295, 317–325. [Google Scholar] [CrossRef]
- Reddy, G.K.K.; Kavibharathi, K.; Singh, A.; Nancharaiah, Y.V. Growth-dependent cr(VI) reduction by Alteromonas sp. ORB2 under haloalkaline conditions: Toxicity, removal mechanism and effect of heavy metals. World J. Microbiol. Biotechnol. 2024, 40, 165. [Google Scholar] [CrossRef]
- Erabee, I.K.; Ahsan, A.; Imteaz, M.; Alom, M.D.M. Adsorption of hexavalent chromium using activated carbon prepared from garden wastes. Desalination Water Treat. 2019, 164, 293–299. [Google Scholar] [CrossRef]
- Salimin, Z.; Satiyoaji, F.W.; Prasetya, D.A.; Chafidz, A. Chemical Treatment of Liquid Waste Generated from Leather Tannery Industry by Using Alum as Coagulant Material. Mater. Sci. Forum 2020, 991, 178–184. [Google Scholar] [CrossRef]
- Peng, H.; Guo, J. Removal of chromium from wastewater by membrane filtration, chemical precipitation, ion exchange, adsorption electrocoagulation, electrochemical reduction, electrodialysis, electrodeionization, photocatalysis and nanotechnology: A review. Environ. Chem. Lett. 2020, 18, 2055–2068. [Google Scholar] [CrossRef]
- Okeke, B.C.; Frankenberger, W.T. Biodegradation of methyl tertiary butyl ether (MTBE) by a bacterial enrichment consortia and its monoculture isolates. Microbiol. Res. 2003, 158, 99–106. [Google Scholar] [CrossRef]
- Yan, X.; Yan, Z.; Zhu, X.; Zhou, Y.; Ma, G.; Li, S.; Liu, X.; Zhang, M. Comparing Different Strategies for Cr(VI) Bioremediation: Bioaugmentation, Biostimulation, and Bioenhancement. Sustainability 2023, 15, 12522. [Google Scholar] [CrossRef]
- Pradhan, S.K.; Singh, N.R.; Rath, B.P.; Thatoi, H. Bacterial chromate reduction: A review of important genomic, proteomic, and bioinformatic analysis. Crit. Rev. Environ. Sci. Technol. 2016, 46, 1659–1703. [Google Scholar] [CrossRef]
- Zha, S.; Yu, A.; Wang, Z.; Shi, Q.; Cheng, X.; Liu, C.; Deng, C.; Zeng, G.; Luo, S.; Zhao, Z.; et al. Microbial strategies for effective hexavalent chromium removal: A comprehensive review. Chem. Eng. J. 2024, 489, 151457. [Google Scholar] [CrossRef]
- Ziagova, M.G.; Koukkou, A.I.; Liakopoulou-Kyriakides, M. Optimization of cultural conditions of Arthrobacter sp. Sphe3 for growth-associated chromate(VI) reduction in free and immobilized cell systems. Chemosphere 2014, 95, 535–540. [Google Scholar] [CrossRef]
- Murugavelh, S.; Mohanty, K. Isolation, identification and characterization of Cr(VI) reducing Bacillus cereus from chromium contaminated soil. Chem. Eng. J. 2013, 230, 1–9. [Google Scholar] [CrossRef]
- Srivastava, S.; Thakur, I.S. Evaluation of bioremediation and detoxification potentiality of Aspergillus niger for removal of hexavalent chromium in soil microcosm. Soil Biol. Biochem. 2006, 38, 1904–1911. [Google Scholar] [CrossRef]
- He, Y.; Dong, L.; Zhou, S.; Jia, Y.; Gu, R.; Bai, Q.; Gao, J.; Li, Y.; Xiao, H. Chromium resistance characteristics of Cr(VI) resistance genes ChrA and ChrB in Serratia sp. S2. Ecotoxicol. Environ. Saf. 2018, 157, 417–423. [Google Scholar] [CrossRef] [PubMed]
- Kabir, M.M.; Fakhruddin, A.N.M.; Chowdhury, M.A.Z.; Pramanik, M.K.; Fardous, Z. Isolation and characterization of chromium(VI)-reducing bacteria from tannery effluents and solid wastes. World J. Microbiol. Biotechnol. 2018, 34, 126. [Google Scholar] [CrossRef]
- Zhang, B.; Wang, Z.; Shi, J.; Dong, H. Sulfur-based mixotrophic bio-reduction for efficient removal of chromium (VI) in groundwater. Geochim. Cosmochim. Acta 2020, 268, 296–309. [Google Scholar] [CrossRef]
- Viti, C.; Marchi, E.; Decorosi, F.; Giovannetti, L. Molecular mechanisms of Cr(VI) resistance in bacteria and fungi. FEMS Microbiol. Rev. 2014, 38, 633–659. [Google Scholar] [CrossRef]
- Battaglia-Brunet, F.; Foucher, S.; Denamur, A.; Ignatiadis, I.; Michel, C.; Morin, D. Reduction of chromate by fixed films of sulfate-reducing bacteria using hydrogen as an electron source. J. Ind. Microbiol. Biotechnol. 2002, 28, 154–159. [Google Scholar] [CrossRef]
- Thatoi, H.; Das, S.; Mishra, J.; Rath, B.P.; Das, N. Bacterial chromate reductase, a potential enzyme for bioremediation of hexavalent chromium: A review. J. Environ. Manag. 2014, 146, 383–399. [Google Scholar] [CrossRef]
- GB 7467-1987; Water Quality: Determination of Chromium (6). 1.5 Diphenylcarbahydrazide Spectrophotometric Method. National Standard of the People’s Republic of China: Beijing, China, 1987.
- GB 7466-1987; Water Quality: Determination of Total Chromium. National Standard of the People’s Republic of China: Beijing, China, 1987.
- Yang, X.Y.; Jin, B.-L.; Wang, X.-N.; Bai, L.H. Isolation, identification and reduction characteristics of a salt-tolerant Cr(VI) reducing strain. J. Sichuan Univ. (Nat. Sci. Ed.) 2023, 60, 016004. [Google Scholar] [CrossRef]
- Zhu, Y.; Yan, J.; Xia, L.; Zhang, X.; Luo, L. Mechanisms of Cr(VI) reduction by Bacillus sp. CRB-1, a novel Cr(VI)-reducing bacterium isolated from tannery activated sludge. Ecotoxicol. Environ. Saf. 2019, 186, 109792. [Google Scholar] [CrossRef]
- Bankar, A.; Zinjarde, S.; Telmore, A.; Walke, A.; Ravikumar, A. Morphological response of Yarrowia lipolytica under stress of heavy metals. Can. J. Microbiol. 2018, 64, 511–526. [Google Scholar] [CrossRef]
- Neumann, G.; Veeranagouda, Y.; Karegoudar, T.B.; Sahin, O.; Mäusezahl, I.; Kabelitz, N.; Kappelmeyer, U.; Heipieper, H.J. Cells of Pseudomonas putida and Enterobacter sp. adapt to toxic organic compounds by increasing their size. Extremophiles 2005, 9, 163–168. [Google Scholar] [CrossRef]
- Liu, X.; Chu, G.; Du, Y.; Li, J.; Si, Y. The role of electron shuttle enhances Fe(III)-mediated reduction of Cr(VI) by Shewanella oneidensis MR-1. World J. Microbiol. Biotechnol. 2019, 35, 64. [Google Scholar] [CrossRef]
- Ju, L.; Jiao, Z.; Ge, S.; Zhan, W.; Liu, Y.; Ren, Q.; Liao, Q.; Yang, Z.; Wang, Y. Formation, stability and mobility of soluble Cr(III) during Cr(VI) reduction by Pannonibacter phragmitetus BB. Environ. Technol. Innov. 2022, 27, 102496. [Google Scholar] [CrossRef]
- Puzon, G.J.; Petersen, J.N.; Roberts, A.G.; Kramer, D.M.; Xun, L. A bacterial flavin reductase system reduces chromate to a soluble chromium(III)–NAD+ complex. Biochem. Biophys. Res. Commun. 2002, 294, 76–81. [Google Scholar] [CrossRef] [PubMed]
- Kantar, C.; Demiray, H.; Dogan, N.M. Role of microbial exopolymeric substances (EPS) on chromium sorption and transport in heterogeneous subsurface soils: II. Binding of Cr(III) in EPS/soil system. Chemosphere 2011, 82, 1496–1505. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Pan, D.; Zheng, J.; Cheng, Y.; Ma, X.; Huang, F.; Lin, Z. Microscopic Investigations of the Cr(VI) Uptake Mechanism of Living Ochrobactrum anthropi. Langmuir 2008, 24, 9630–9635. [Google Scholar] [CrossRef]
- Biesinger, M.C.; Payne, B.P.; Grosvenor, A.P.; Lau, L.W.M.; Gerson, A.R.; Smart, R.S.C. Resolving surface chemical states in XPS analysis of first row transition metals, oxides and hydroxides: Cr, Mn, Fe, Co and Ni. Appl. Surf. Sci. 2011, 257, 2717–2730. [Google Scholar] [CrossRef]
- Ye, Y.; Hao, R.; Shan, B.; Zhang, J.; Li, J.; Lu, A. Mechanism of Cr(VI) removal by efficient Cr(VI)-resistant Bacillus mobilis CR3. World J. Microbiol. Biotechnol. 2023, 40, 21. [Google Scholar] [CrossRef]
- Pan, X.; Liu, Z.; Chen, Z.; Cheng, Y.; Pan, D.; Shao, J.; Lin, Z.; Guan, X. Investigation of Cr(VI) reduction and Cr(III) immobilization mechanism by planktonic cells and biofilms of Bacillus subtilis ATCC-6633. Water Res. 2014, 55, 21–29. [Google Scholar] [CrossRef]
- Dou, X.; Su, H.; Xu, D.; Liu, C.; Meng, H.; Li, H.; Zhang, J.; Dang, Y.; Feng, L.; Zhang, L.; et al. Enhancement effects of dissolved organic matter leached from sewage sludge on microbial reduction and immobilization of Cr(VI) by Geobacter sulfurreducens. Sci. Total Environ. 2022, 835, 155301. [Google Scholar] [CrossRef]
- Bencheikh-Latmani, R.; Williams, S.M.; Haucke, L.; Criddle, C.S.; Wu, L.; Zhou, J.; Tebo, B.M. Global Transcriptional Profiling of Shewanella oneidensis- MR-1 during Cr(VI) and Cr(VI) Reduction. Appl. Environ. Microbiol. 2005, 71, 7453–7460. [Google Scholar] [CrossRef]
- Liu, J.Q.; Ma, X.; Liu, D.F.; Yang, C.W.; Li, D.B.; Min, D.; Yu, H.Q. Multiple roles of released c-type cytochromes in tuning electron transport and physiological status of Geobacter sulfurreducens. Biotechnol. Bioeng. 2023, 120, 1346–1356. [Google Scholar] [CrossRef]
- Monga, A.; Fulke, A.B.; Sonker, S.; Dasgupta, D. Unveiling the chromate stress response in the marine bacterium Bacillus enclensis AGM_Cr8: A multifaceted investigation. World J. Microbiol. Biotechnol. 2024, 40, 394. [Google Scholar] [CrossRef]
- Li, J.; Xie, W.; Qi, H.; Sun, S.; Deng, T.; Tang, Y.; Qiu, R. Hexavalent chromium uptake in rice (Oryza sativa L.) mediated by sulfate and phosphate transporters OsSultr1;2 and OsPht1;1. J. Hazard. Mater. 2024, 478, 135559. [Google Scholar] [CrossRef]
- Zhang, X.; Wu, W.; Virgo, N.; Zou, L.; Liu, P.; Li, X. Global transcriptome analysis of hexavalent chromium stress responses in Staphylococcus aureus LZ-01. Ecotoxicology 2014, 23, 1534–1545. [Google Scholar] [CrossRef] [PubMed]
- Marín, R.; Bose, R.; Dabrowski, B.; Kolesnik, S. Magnetic Characterization of Chromium Intermediates in the Reduction of Chromium (VI) by Glutathione in Acidic Solutions. Magnetochemistry 2018, 4, 23. [Google Scholar] [CrossRef]
- Fathima, A.; Rao, J.R. Is Cr(III) toxic to bacteria: Toxicity studies using Bacillus subtilis and Escherichia coli as model organism. Arch. Microbiol. 2017, 200, 453–462. [Google Scholar] [CrossRef] [PubMed]
- Abdullah; Wani, K.I.; Naeem, M.; Jha, P.K.; Jha, U.C.; Aftab, T.; Prasad, P.V.V. Systems biology of chromium-plant interaction: Insights from omics approaches. Front. Plant Sci. 2024, 14, 1305179. [CrossRef] [PubMed]
- Ahemad, M. Bacterial mechanisms for Cr(VI) resistance and reduction: An overview and recent advances. Folia Microbiol. 2014, 59, 321–332. [Google Scholar] [CrossRef]
- Bonde, N.J.; Henry, C.; Wood, E.A.; Cox, M.M.; Keck, J.L. Interaction with the carboxy-terminal tip of SSB is critical for RecG function in E. coli. Nucleic Acids Res. 2023, 51, 3735–3753. [Google Scholar] [CrossRef]
- Romero, Z.J.; Chen, S.H.; Armstrong, T.; Wood, E.A.; van Oijen, A.; Robinson, A.; Cox, M.M. Resolving Toxic DNA repair intermediates in every E. coli replication cycle: Critical roles for RecG, Uup and RadD. Nucleic Acids Res. 2020, 48, 8445–8460. [Google Scholar] [CrossRef]
- Chai, L.; Ding, C.; Li, J.; Yang, Z.; Shi, Y. Multi-omics response of Pannonibacter phragmitetus BB to hexavalent chromium. Environ. Pollut. 2019, 249, 63–73. [Google Scholar] [CrossRef]
- Moreno-Gámez, S.; Hochberg, M.E.; van Doorn, G.S. Quorum sensing as a mechanism to harness the wisdom of the crowds. Nat. Commun. 2023, 14, 3415. [Google Scholar] [CrossRef]
- Rodrigues, M.V.; Kis, P.; Xavier, K.B.; Ventura, M.R. Synthesis and Potential of Autoinducer-2 and Analogs to Manipulate Inter-Species Quorum Sensing. Isr. J. Chem. 2023, 63, e202200091. [Google Scholar] [CrossRef]

| Location | Initial Cr(VI) (mg/L) | Residual Cr(VI) (mg/L) | Total Cr (mg/L) |
|---|---|---|---|
| extracellular | 19.25 | 7.93 | 13.58 |
| intracellular | 0.00 | 0.31 | 1.15 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Y.; Yu, W.; Nie, T.; Wang, L.; Niu, Y. Extracellular Cr(VI) Reduction by the Salt-Tolerant Strain Bacillus safensis BSF-4. Microorganisms 2025, 13, 1961. https://doi.org/10.3390/microorganisms13081961
Liu Y, Yu W, Nie T, Wang L, Niu Y. Extracellular Cr(VI) Reduction by the Salt-Tolerant Strain Bacillus safensis BSF-4. Microorganisms. 2025; 13(8):1961. https://doi.org/10.3390/microorganisms13081961
Chicago/Turabian StyleLiu, Yilan, Weiping Yu, Tianying Nie, Lu Wang, and Yusheng Niu. 2025. "Extracellular Cr(VI) Reduction by the Salt-Tolerant Strain Bacillus safensis BSF-4" Microorganisms 13, no. 8: 1961. https://doi.org/10.3390/microorganisms13081961
APA StyleLiu, Y., Yu, W., Nie, T., Wang, L., & Niu, Y. (2025). Extracellular Cr(VI) Reduction by the Salt-Tolerant Strain Bacillus safensis BSF-4. Microorganisms, 13(8), 1961. https://doi.org/10.3390/microorganisms13081961





