Noctiluca scintillans Bloom Reshapes Microbial Community Structure, Interaction Networks, and Metabolism Patterns in Qinhuangdao Coastal Waters, China
Abstract
1. Introduction
2. Materials and Methods
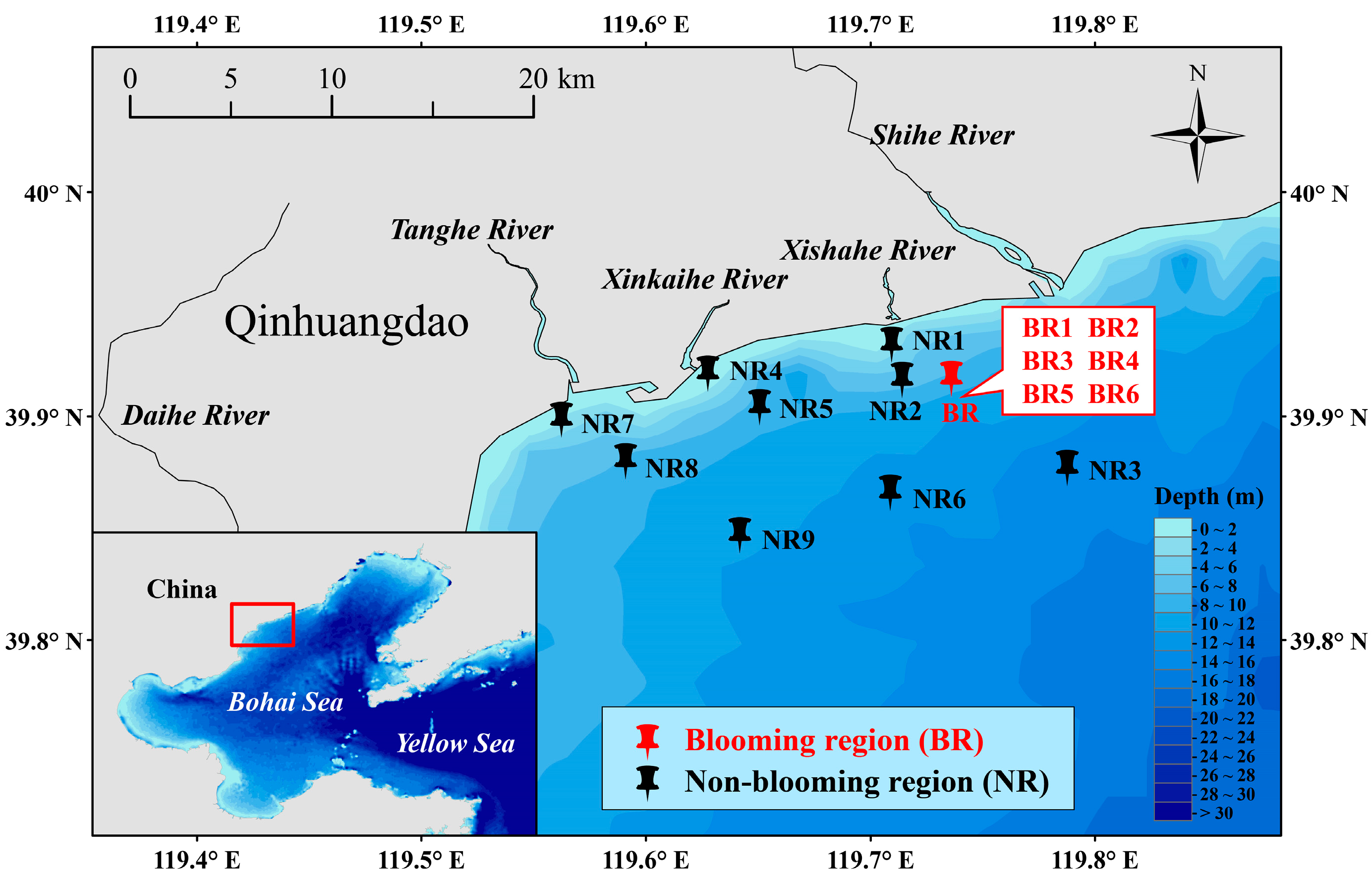
2.1. Study Area and Sampling Sites
2.2. Sampling and Laboratory Analysis
2.3. DNA Extraction, PCR and Sequencing
2.4. Bioinformatics Analysis
2.5. Data Processing and Community Structure Analysis
2.6. Co-Occurrence Network Analysis
2.7. Gene Prediction Analysis
3. Results
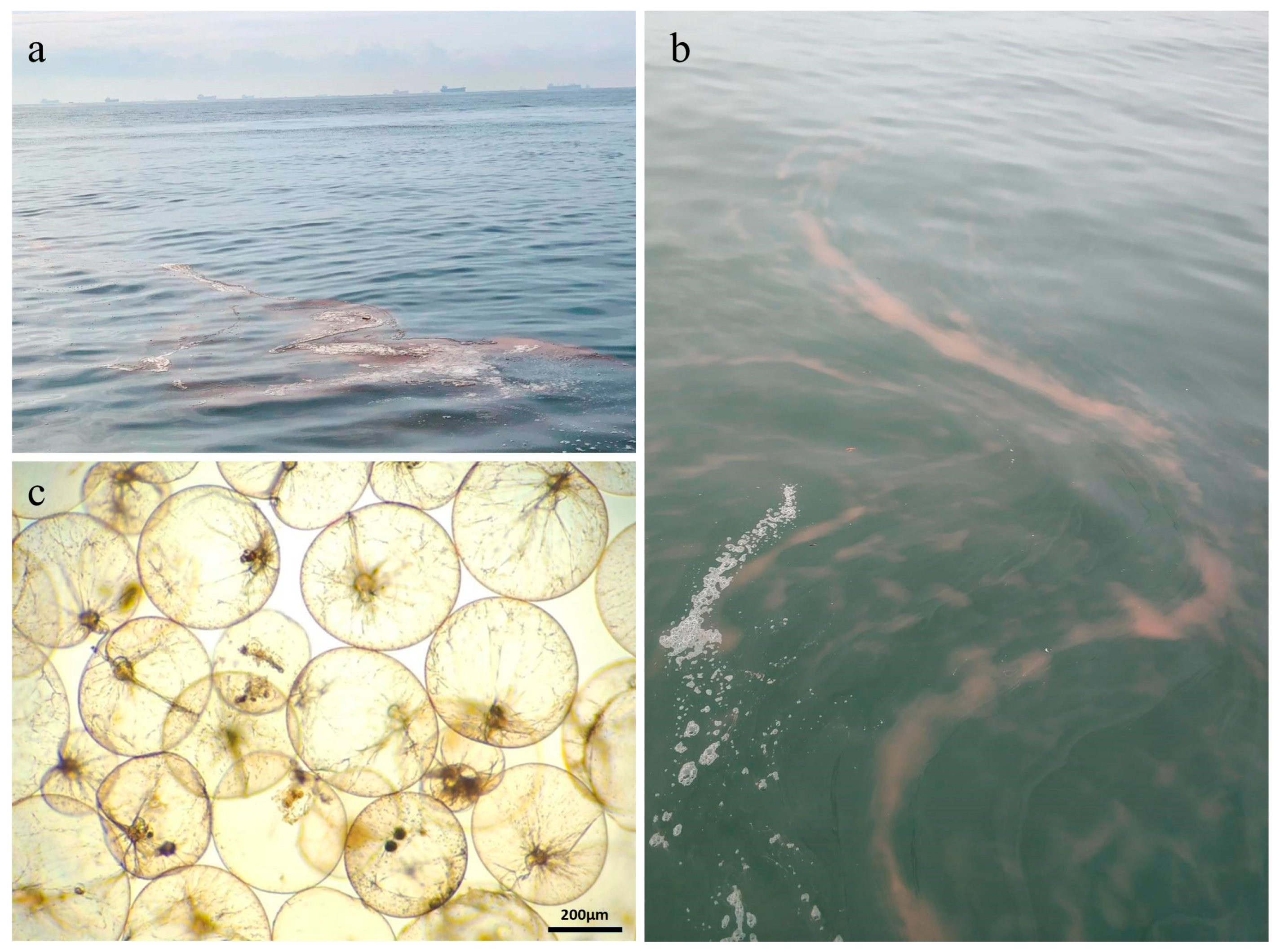
3.1. Noctiluca scintillans Density and Environmental Conditions
3.2. Microbial Community Structure and Diversity
3.2.1. Microbial Community Composition
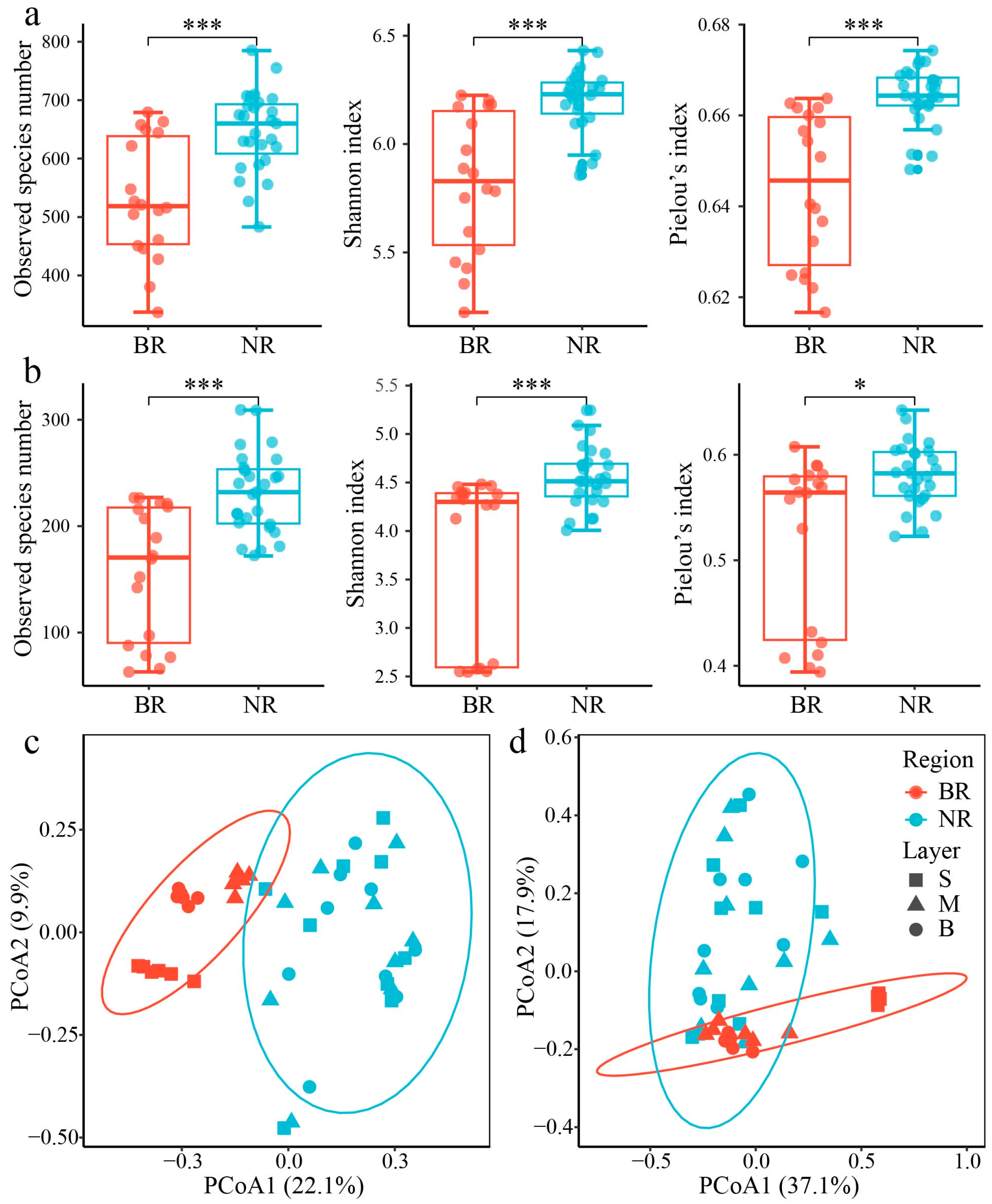
3.2.2. Microbial Diversity
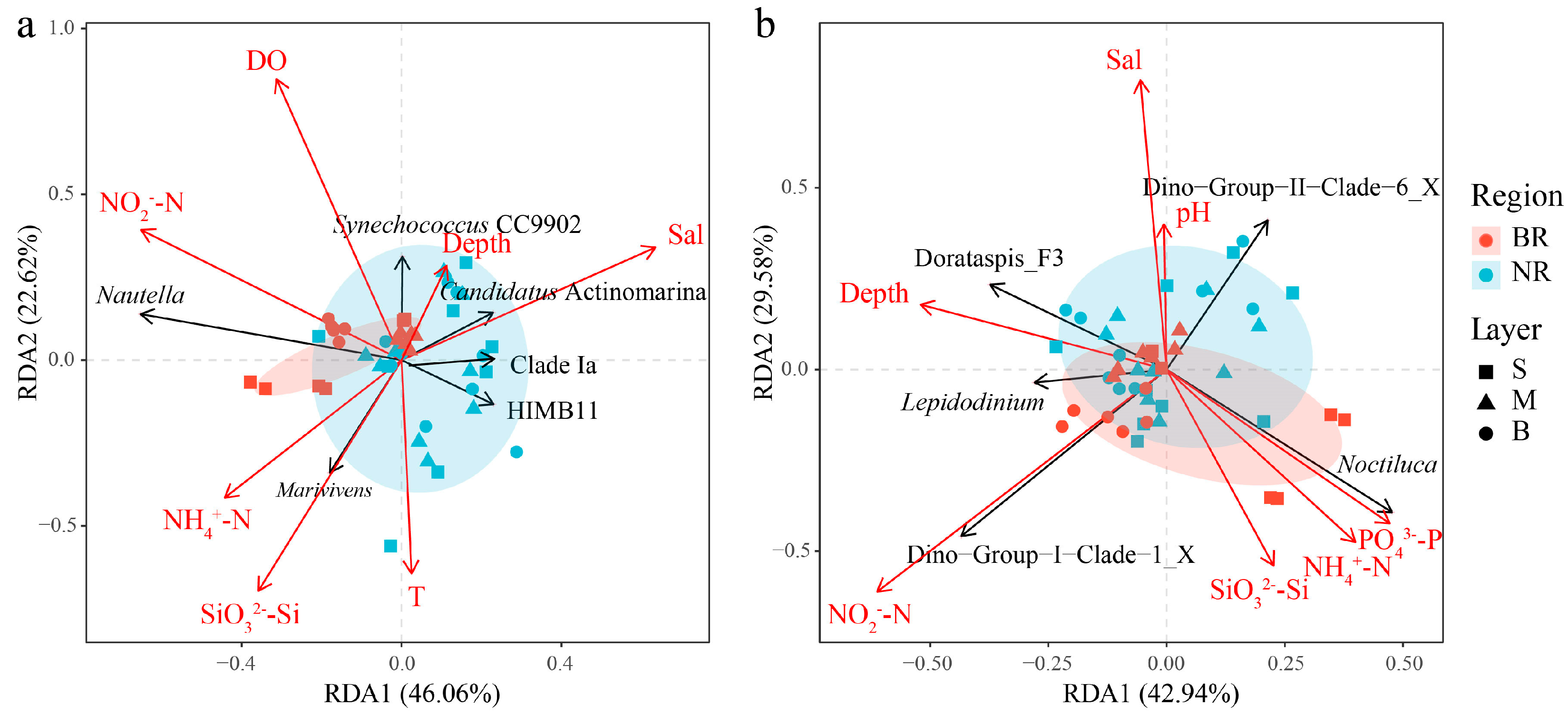
3.3. Relationship Between Microbial Community Composition and Environmental Factors
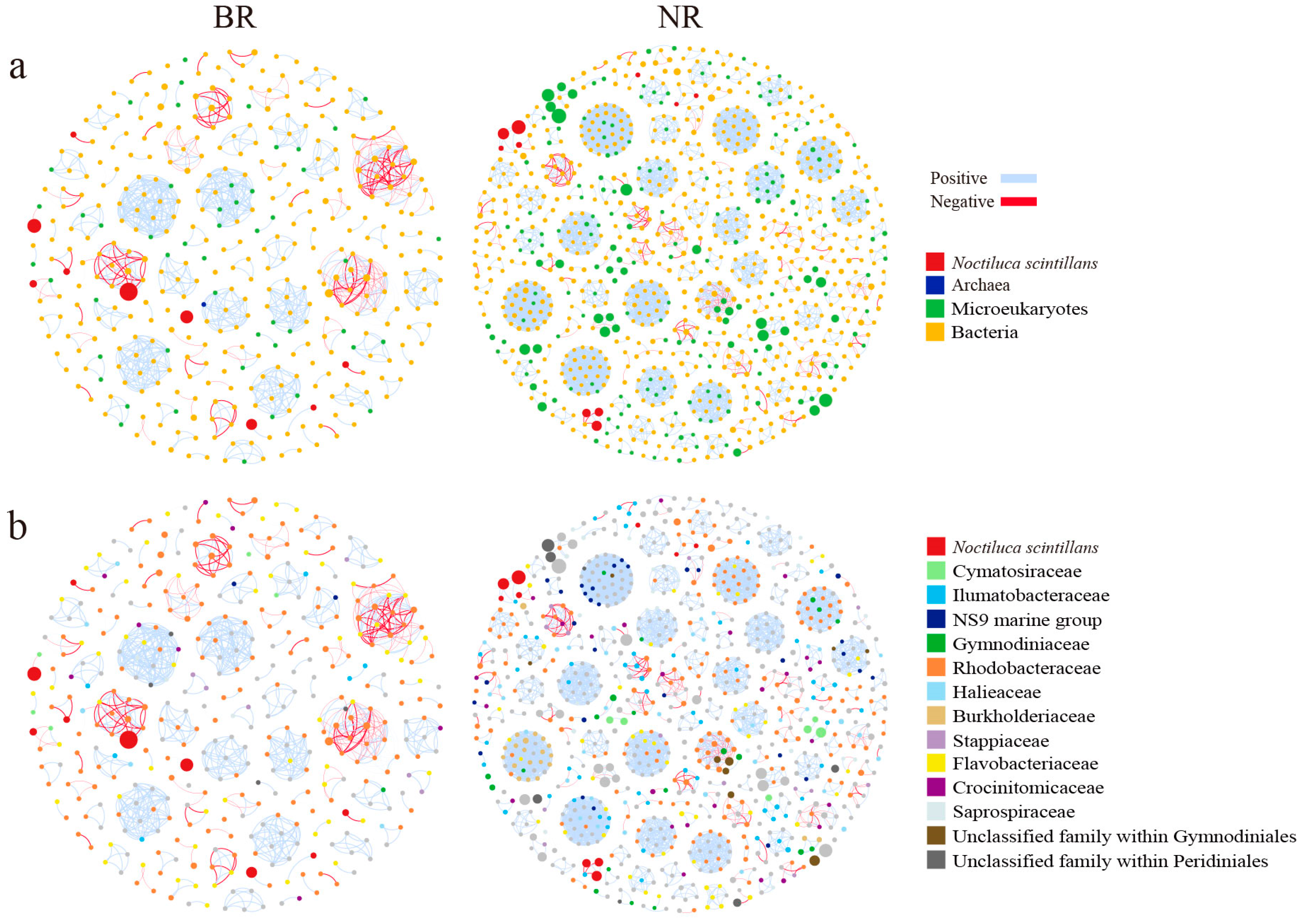
3.4. Microbial Co-Occurrence Network
3.4.1. Network Topological Properties
3.4.2. Network Keystone Taxa
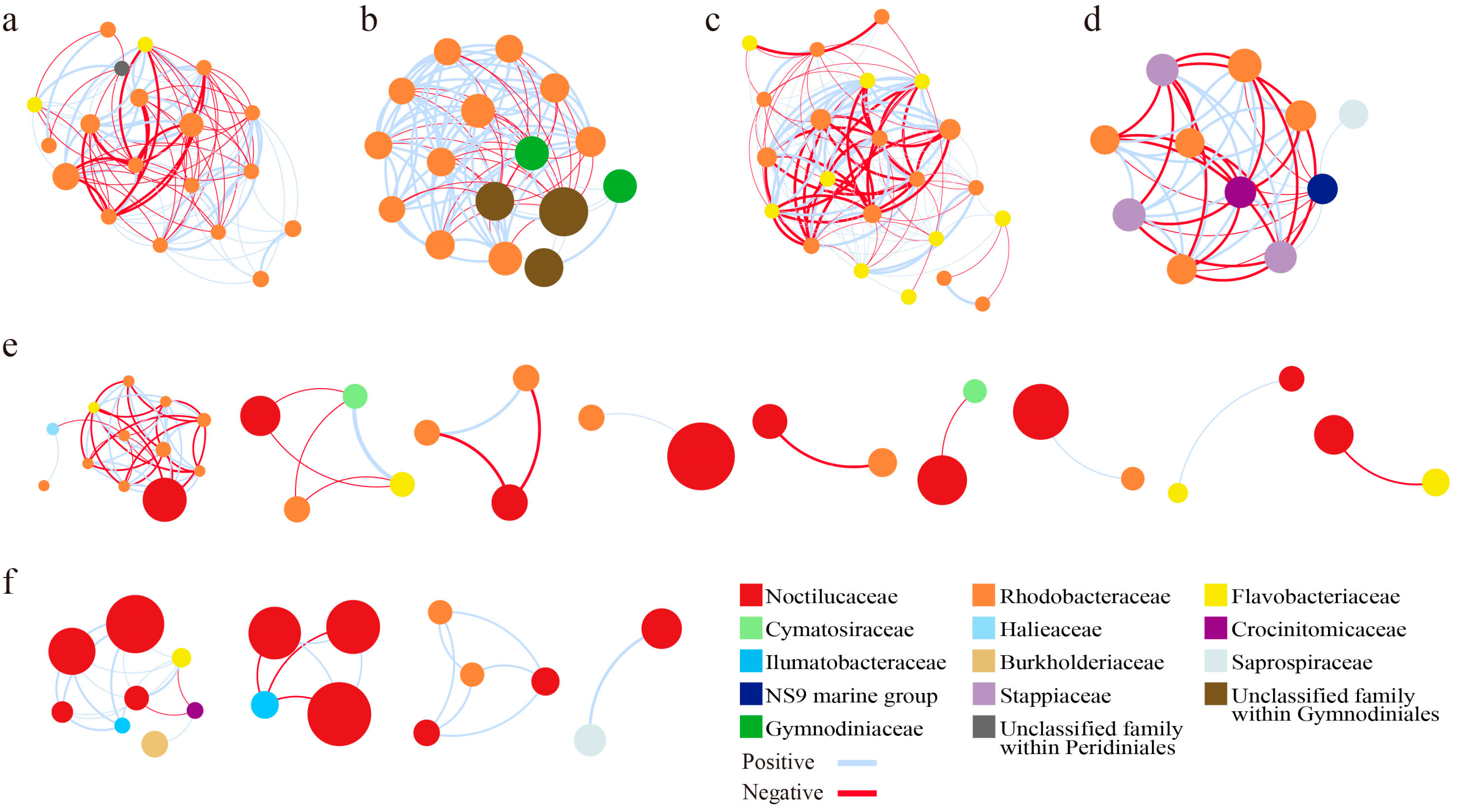
3.4.3. Network Modules
3.5. Microbial Metabolic Functions
3.5.1. Respiratory Chain
3.5.2. Fatty Acid Degradation, Methylmalonyl-CoA Pathway, and TCA Cycle
3.5.3. Pyruvate Metabolism and Gluconeogenesis
3.5.4. Branched-Chain Amino Acid Metabolism
3.5.5. Nitrogen Metabolism
4. Discussion
4.1. Relationship Between the Noctiluca scintillans Bloom and Environmental Conditions
4.2. Impacts of the N. scintillans Bloom on Prokaryotic and Microeukaryotic Community Structure
4.3. Impacts of the N. scintillans Bloom on Microbial Interactions
4.4. Impacts of the N. scintillans Bloom on Microbial Metabolism
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Cai, R.; Han, Z.; Yang, Z. Impacts and risks of changing ocean on marine ecosystems and dependent communities and related responses. Clim. Change Res. 2020, 16, 182–193, (In Chinese with English Abstract). [Google Scholar]
- Hooper, D.U.; Adair, E.C.; Cardinale, B.J.; Byrnes, J.E.; Hungate, B.A.; Matulich, K.L.; Gonzalez, A.; Duffy, J.E.; Gamfeldt, L.; O’Connor, M.I. A global synthesis reveals biodiversity loss as a major driver of ecosystem change. Nature 2012, 486, 105–108. [Google Scholar] [CrossRef] [PubMed]
- Berdalet, E.; Kudela, R.M.; Urban, E.; Enevoldsen, H.O.; Banas, N.S.; Bresnan, E.; Burford, M.A.; Davidson, K.; Gobler, C.J.; Karlson, B.; et al. GlobalHAB: A new program to promote international research, observations, and modeling of harmful algal blooms in aquatic systems. Oceanography 2017, 30, 70–81. [Google Scholar] [CrossRef]
- Berdalet, E.; Fleming, L.E.; Gowen, R.; Davidson, K.; Hess, P.; Backer, L.C.; Moore, S.K.; Hoagland, P.; Enevoldsen, H. Marine harmful algal blooms, human health and wellbeing: Challenges and opportunities in the 21st century. J. Mar. Biol. Assoc. U. K. 2015, 96, 61–91. [Google Scholar] [CrossRef]
- Brown, A.R.; Lilley, M.K.S.; Shutler, J.D.; Lowe, C.D.; Artioli, Y.; Torres, R.; Berdalet, E.; Tyler, C.R. Assessing risks and mitigating impacts of harmful algal blooms on mariculture and marine fisheries. Rev. Aquac. 2019, 12, 1663–1688. [Google Scholar] [CrossRef]
- Yan, T.; Li, X.-D.; Tan, Z.-J.; Yu, R.-C.; Zou, J.-Z. Toxic effects, mechanisms, and ecological impacts of harmful algal blooms in China. Harmful Algae 2022, 111, 102148. [Google Scholar] [CrossRef]
- Carnicer, O.; Guallar, C.; Andree, K.B.; Diogène, J.; Fernández-Tejedor, M. Ostreopsis cf. ovata dynamics in the NW Mediterranean Sea in relation to biotic and abiotic factors. Environ. Res. 2015, 143, 89–99. [Google Scholar] [PubMed]
- Demuez, M.; González-Fernández, C.; Ballesteros, M. Algicidal microorganisms and secreted algicides: New tools to induce microalgal cell disruption. Biotechnol. Adv. 2015, 33, 1615–1625. [Google Scholar] [CrossRef] [PubMed]
- Gerphagnon, M.; Latour, D.; Colombet, J.; Sime-Ngando, T. Fungal parasitism: Life cycle, dynamics and impact on cyanobacterial blooms. PLoS ONE 2013, 8, e60894. [Google Scholar] [CrossRef]
- Jingzhong, Z.; Liping, D.; Baoping, Q. Preliminary studies on eutrophication and red tide problems in Bohai Bay. Hydrobiologia 1985, 127, 27–30. [Google Scholar] [CrossRef]
- Adachi, M.; Kanno, T.; Okamoto, R.; Itakura, S.; Yamaguchi, M.; Nishijima, T. Population structure of Alexandrium (Dinophyceae) cyst formation-promoting bacteria in Hiroshima Bay, Japan. Appl. Environ. Microbiol. 2003, 69, 6560–6568. [Google Scholar] [CrossRef]
- Zhou, J.; Lyu, Y.; Richlen, M.L.; Anderson, D.M.; Cai, Z.H. Quorum sensing is a language of chemical signals and plays an ecological role in algal-bacterial interactions. Crit. Rev. Plant Sci. 2016, 35, 81–105. [Google Scholar] [CrossRef] [PubMed]
- Faust, K.; Raes, J. Microbial interactions: From networks to models. Nat. Rev. Microbiol. 2012, 10, 538–550. [Google Scholar] [CrossRef] [PubMed]
- De Roy, K.; Marzorati, M.; van den Abbeele, P.; van de Wiele, T.; Boon, N. Synthetic microbial ecosystems: An exciting tool to understand and apply microbial communities. Environ. Microbiol. 2014, 16, 1472–1481. [Google Scholar] [CrossRef] [PubMed]
- Furbino, L.E.; Godinho, V.M.; Santiago, I.F.; Pellizari, F.M.; Alves, T.M.; Zani, C.L.; Junior, P.A.; Romanha, A.J.; Carvalho, A.G.; Gil, L.H.; et al. Diversity patterns, ecology and biological activities of fungal communities associated with the endemic macroalgae across the Antarctic peninsula. Microb. Ecol. 2014, 67, 775–787. [Google Scholar] [CrossRef]
- Seymour, J.R.; Amin, S.A.; Raina, J.B.; Stocker, R. Zooming in on the phycosphere: The ecological interface for phytoplankton-bacteria relationships. Nat. Microbiol. 2017, 2, 17065. [Google Scholar] [CrossRef]
- Matcher, G.; Lemley, D.A.; Adams, J.B. Bacterial community dynamics during a harmful algal bloom of Heterosigma akashiwo. Aquat. Microb. Ecol. 2021, 86, 153–167. [Google Scholar] [CrossRef]
- Zhou, J.; Lao, Y.-m.; Song, J.-t.; Jin, H.; Zhu, J.-m.; Cai, Z.-h. Temporal heterogeneity of microbial communities and metabolic activities during a natural algal bloom. Water Res. 2020, 183, 116020. [Google Scholar] [CrossRef]
- Chen, Y.; Wang, L.; Liu, Z.; Su, D.; Wang, Y.; Qi, Y. Biodiversity and interannual variation of harmful algal bloom species in the coastal sea of Qinhuangdao, China. Life 2023, 13, 192. [Google Scholar] [CrossRef]
- Zhang, W.; Ma, X.; Zhang, Y.-F.; Zhang, J.; Zhao, S.; Li, G.-M. An analysis of red tide characteristics in Qinhuangdao coastal seawater. Trans. Oceanol. Limnol. 2020, 5, 48–55, (In Chinese with English Abstract). [Google Scholar]
- Harrison, P.J.; Furuya, K.; Glibert, P.M.; Xu, J.; Liu, H.; Yin, K.; Lee, J.H.W.; Anderson, D.M.; Gowen, R.; Al-Azri, A.R.; et al. Geographical distribution of red and green Noctiluca scintillans. Chin. J. Oceanol. Limnol. 2011, 29, 807–831. [Google Scholar] [CrossRef]
- Morton, B.; Twentyman, P.R. The occurrence and toxicity of a red tide caused by Noctiluca scintillans (Macartney) Ehrenb., in the coastal waters of Hong Kong. Environ. Res. 1971, 4, 544–557. [Google Scholar] [CrossRef]
- Turkoglu, M. Red tides of the dinoflagellate Noctiluca scintillans associated with eutrophication in the Sea of Marmara (the Dardanelles, Turkey). Oceanologia 2013, 55, 709–732. [Google Scholar] [CrossRef]
- Xiang, C.; Tan, Y.; Zhang, H.; Liu, J.; Ke, Z.; Li, G. The key to dinoflagellate (Noctiluca scintillans) blooming and outcompeting diatoms in winter off Pakistan, northern Arabian Sea. Sci. Total Environ. 2019, 694, 133396. [Google Scholar] [CrossRef]
- Wang, K.; Lin, H.; Peng, C.; Sun, L.; Gao, Y.; Chen, B. Long-term changes in Noctiluca scintillans blooms along the Chinese coast from 1933 to 2020. Glob. Change Biol. 2023, 29, 5099–5113. [Google Scholar] [CrossRef]
- Ara, K.; Nakamura, S.; Takahashi, R.; Shiomoto, A.; Hiromi, J. Seasonal variability of the red tide-forming heterotrophic dinoflagellate Noctiluca scintillans in the neritic area of Sagami Bay, Japan: Its role in the nutrient-environment and aquatic ecosystem. Plankton Benthos Res. 2013, 8, 9–30. [Google Scholar] [CrossRef]
- Pithakpol, S.; Tada, K.; Montani, S. Ammonium and phosphate pools of Noctiluca scintillans and their supplies to the water column in Harima Nada, the Seto Inland Sea, Japan. La Mer 2000, 37, 153–162. [Google Scholar]
- Xia, X.; Cheung, S.; Zhang, S.; Lu, Y.; Leung, S.K.; Shi, Z.; Xu, H.; Gu, B.; Tan, Y.; Zeng, H.; et al. Noctiluca scintillans bloom alters the composition and carbohydrate utilization of associated bacterial community and enriches potential pathogenic bacterium Vibrio anguillarum. Water Res. 2024, 249, 120974. [Google Scholar] [CrossRef]
- Xia, X.; Leung, S.K.; Cheung, S.; Zhang, S.; Liu, H. Rare bacteria in seawater are dominant in the bacterial assemblage associated with the bloom-forming dinoflagellate Noctiluca scintillans. Sci. Total Environ. 2020, 711, 135107. [Google Scholar] [CrossRef]
- Parada, A.E.; Needham, D.M.; Fuhrman, J.A. Every base matters: Assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples. Environ. Microbiol. 2016, 18, 1403–1414. [Google Scholar] [CrossRef]
- Amaral-Zettler, L.A.; McCliment, E.A.; Ducklow, H.W.; Huse, S.M. A method for studying protistan diversity using massively parallel sequencing of V9 hypervariable regions of small-subunit ribosomal RNA genes. PLoS ONE 2009, 4, e6372, Erratum in PLoS One 2009, 4. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Apprill, A.; McNally, S.P.; Parsons, R.J.; Weber, L. Minor revision to V4 region SSU rRNA 806R gene primer greatly increases detection of SAR11 bacterioplankton. Aquat. Microb. Ecol. 2015, 75, 129–137. [Google Scholar] [CrossRef]
- Oksanen, J. Vegan: Ecological diversity. R Proj. 2013, 368, 1–12. [Google Scholar]
- Liu, J.; Zhu, S.; Liu, X.; Yao, P.; Zhang, X.H. Spatiotemporal dynamics of the archaeal community in coastal sediments: Assembly process and co-occurrence relationship. ISME J. 2020, 14, 1463–1478. [Google Scholar] [CrossRef]
- Kordubel, K.; Martínez-Rincón, R.O.; Baschek, B.; Boersma, M.; Hieronymi, M.; Johns, D.G.; Kirstein, I.V.; Voynova, Y.G.; Möller, K.O. Long-term changes in spatiotemporal distribution of Noctiluca scintillans in the southern North Sea. Harmful Algae 2024, 138, 102699. [Google Scholar] [CrossRef]
- Zhou, C.; Wu, Y.; Zou, J. Nutrient dynamics of Noctiluca scintillans (Macartney). Oceanol. Limnol. Sin. 1994, 25, 152–157, (In Chinese with English Abstract). [Google Scholar]
- Elbrächter, M.; Qi, Z. Aspects of Noctiluca (Dinophyceae) population dynamics. In Proceedings of the NATO Advanced Study Institute, Physiological Ecology of Harmful Algal Blooms, Bermuda, UK, 27 May–6 June 1996; Springer: Berlin/Heidelberg, Germany, 1998; pp. 315–336. [Google Scholar]
- Zheng, X.-R.; Fu, Z.; Xi, Y.-J.; Mu, J.-D.; Li, Y. Grey relationship analysis for the environmental factors affecting the Noctiluca scintillans density in Qinhuangdao coastal area. Prog. Fish. Sci. 2014, 35, 8–15, (In Chinese with English Abstract). [Google Scholar]
- Xu, C.-y.; Huang, M.-z.; Du, Q. Ecological characteristics of important red tide species in Fujian coastal waters. J. Oceanogr. Taiwan Strait 2010, 29, 434–441, (In Chinese with English Abstract). [Google Scholar]
- Miyaguchi, H.; Fujiki, T.; Kikuchi, T.; Kuwahara, V.S.; Toda, T. Relationship between the bloom of Noctiluca scintillans and environmental factors in the coastal waters of Sagami Bay, Japan. J. Plankton Res. 2006, 28, 313–324. [Google Scholar] [CrossRef]
- Li, Y.; Yang, C.; Li, D.; Tian, Y.; Zheng, T. Dynamics of bacterial community during the bloom caused by Skeletonema costatum and Akashiwo sanguinea in Xiamen sea area. Acta Microbiol. Sin. 2012, 52, 1268–1281, (In Chinese with English Abstract). [Google Scholar]
- Qi, S.; Qin, C.; Huang, S.; Zhong, C.; Sun, K. Correlation between the patch distribution of Noctiluca Scintillans bloom and aquatic environmental factors. J. Trop. Biol. 2018, 9, 1–11, (In Chinese with English Abstract). [Google Scholar]
- Solomon, C.M.; Collier, J.L.; Berg, G.M.; Glibert, P. Role of urea in microbial metabolism in aquatic systems: A biochemical and molecular review. Aquat. Microb. Ecol. 2010, 59, 67–88. [Google Scholar] [CrossRef]
- Gómez-Consarnau, L.; Needham, D.M.; Weber, P.K.; Fuhrman, J.A.; Mayali, X. Influence of light on particulate organic matter utilization by attached and free-living marine bacteria. Front. Microbiol. 2019, 10, 1024. [Google Scholar] [CrossRef]
- Zhang, S.; Liu, H.; Xia, X. Community dynamics of free-living and particle-attached bacteria over sequential blooms of heterotrophic dinoflagellate Noctiluca scintillans and mixotrophic ciliate Mesodinium rubrum. Appl. Environ. Microbiol. 2022, 88, e01323-22. [Google Scholar] [CrossRef]
- Teeling, H.; Fuchs, B.M.; Becher, D.; Klockow, C.; Gardebrecht, A.; Bennke, C.M.; Kassabgy, M.; Huang, S.; Mann, A.J.; Waldmann, J.; et al. Substrate-controlled succession of marine bacterioplankton populations induced by a phytoplankton bloom. Science 2012, 336, 608–611. [Google Scholar] [CrossRef]
- Buchan, A.; LeCleir, G.R.; Gulvik, C.A.; González, J.M. Master recyclers: Features and functions of bacteria associated with phytoplankton blooms. Nat. Rev. Microbiol. 2014, 12, 686–698. [Google Scholar] [CrossRef]
- Röttjers, L.; Faust, K. From hairballs to hypotheses-biological insights from microbial networks. FEMS Microbiol. Rev. 2018, 42, 761–780. [Google Scholar] [CrossRef]
- Hernandez, D.J.; David, A.S.; Menges, E.S.; Searcy, C.A.; Afkhami, M.E. Environmental stress destabilizes microbial networks. ISME J. 2021, 15, 1722–1734. [Google Scholar] [CrossRef]
- Zhou, J.; Deng, Y.; Luo, F.; He, Z.; Tu, Q.; Zhi, X. Functional molecular ecological networks. mBio 2010, 1, e00169-10. [Google Scholar] [CrossRef]
- Trombetta, T.; Vidussi, F.; Roques, C.; Scotti, M.; Mostajir, B. Marine microbial food web networks during phytoplankton bloom and non-bloom periods: Warming favors smaller organism interactions and intensifies trophic cascade. Front. Microbiol. 2020, 11, 502336. [Google Scholar] [CrossRef]
- Xu, S. The Variation of Marine Microbial Community Structure and Its Ecological Functions Following a Phaeocystis Bloom, Highlighted by Omics Techniques. Ph.D. Dissertation, University of Chinese Academy of Sciences, Beijing, China, 2021. (In Chinese with English Abstract). [Google Scholar]
- Zhu, J.; Zhou, J.; Wang, H.; Chen, G.; Cai, Z. The ecological network approach to algal-bacterial relationships: Review and prospects. Chin. Sci. Bull. 2021, 66, 4378–4394, (In Chinese with English Abstract). [Google Scholar] [CrossRef]
- Berry, D.; Widder, S. Deciphering microbial interactions and detecting keystone species with co-occurrence networks. Front. Microbiol. 2014, 5, 219. [Google Scholar] [CrossRef]
- Roager, L.; Kempen, P.J.; Bentzon-Tilia, M.; Sonnenschein, E.C.; Gram, L. Impact of host species on assembly, composition, and functional profiles of phycosphere microbiomes. Msystems 2024, 9, e0058324. [Google Scholar] [CrossRef]
- Brüwer, J.D.; Orellana, L.H.; Sidhu, C.; Klip, H.C.; Meunier, C.L.; Boersma, M.; Wiltshire, K.H.; Amann, R.; Fuchs, B.M. In situ cell division and mortality rates of SAR11, SAR86, Bacteroidetes, and Aurantivirga during phytoplankton blooms reveal differences in population controls. mSystems 2023, 8, e0128722, Erratum in mSystems 2024, 9, e0119623 . [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Liang, J.; Liu, J.; Zhan, Y.; Zhou, S.; Xue, C.-X.; Sun, C.; Lin, Y.; Luo, C.; Wang, X.; Zhang, X.-H. Succession of marine bacteria in response to Ulva prolifera-derived dissolved organic matter. Environ. Int. 2021, 155, 106687. [Google Scholar] [CrossRef]
- Raes, J.; Bork, P. Molecular eco-systems biology: Towards an understanding of community function. Nat. Rev. Microbiol. 2008, 6, 693–699. [Google Scholar] [CrossRef]
- Yang, C.; Wang, Q.; Simon, P.N.; Liu, J.; Liu, L.; Dai, X.; Zhang, X.; Kuang, J.; Igarashi, Y.; Pan, X.; et al. Distinct network interactions in particle-associated and free-living bacterial communities during a Microcystis aeruginosa bloom in a plateau lake. Front. Microbiol. 2017, 8, 1202. [Google Scholar] [CrossRef]
- Tarafdar, L.; Mohapatra, M.; Muduli, P.R.; Kumar, A.; Mishra, D.R.; Rastogi, G. Co-occurrence patterns and environmental factors associated with rapid onset of Microcystis aeruginosa bloom in a tropical coastal lagoon. J. Environ. Manag. 2023, 325, 116580. [Google Scholar] [CrossRef]
- Chen, Y. Ecological Characteristics of Phytoplankton and Physiological Mechanism of Interspecies Relationship Between the Key Phytoplankton Species in a Seallop Culture Sea Area of Bohai Bay. Ph.D. Dissertation, Xiamen University, Xiamen, China, 2017. (In Chinese with English Abstract). [Google Scholar]
- Wang, F.-W.; Li, Y.; Zhen, Y.; Liu, Q. Transcriptome analysis of carbon fixation pathway of Minutocellus polymorphus under different nitrogen source conditions and the comparison with other microalgae. Oceanol. Limnol. Sin. 2020, 51, 1127–1135, (In Chinese with English Abstract). [Google Scholar]
- Chen, J.; Zhong, Y.; Wang, L.; Qiu, D. In situ diets of the bloom-forming dinoflagellate Noctiluca scintillans in Daya Bay. Harmful Algae 2023, 130, 102546. [Google Scholar] [CrossRef]
- de Gier, J.W.L.; Schepper, M.; Reijnders, W.N.; Van Dyck, S.J.; Slotboom, D.J.; Warne, A.; Saraste, M.; Krab, K.; Finel, M.; Stouthamer, A.H.; et al. Structural and functional analysis of aa3-type and cbb3-type cytochrome c oxidases of Paracoccus denitrificans reveals significant differences in proton-pump design. Mol. Microbiol. 1996, 20, 1247–1260. [Google Scholar] [CrossRef]
- Bertero, M.G.; Rothery, R.A.; Palak, M.; Hou, C.; Lim, D.; Blasco, F.; Weiner, J.H.; Strynadka, N.C.J. Insights into the respiratory electron transfer pathway from the structure of nitrate reductase A. Nat. Struct. Mol. Biol. 2003, 10, 681–687. [Google Scholar] [CrossRef]
- Rothery, R.A.; Blasco, F.; Magalon, A.; Weiner, J.H. The diheme cytochrome b subunit (Narl) of Escherichia coli nitrate reductase A (NarGHI): Structure, function, and interaction with quinols. J. Mol. Microbiol. Biotechnol. 2001, 3, 273–283. [Google Scholar]
- Ma, W.; Li, J.; Yang, W.Q.; Zhang, Z.Y.; Yan, C.X.; Huang, P.W.; Sun, X.M. Efficient biosynthesis of odd-chain fatty acids via regulating the supply and consumption of propionyl-CoA in Schizochytrium sp. J. Agric. Food Chem. 2023, 71, 9847–9855. [Google Scholar] [CrossRef]
- Rosenberg, L.E. The inherited methylmalonic acidaemias: A model system for the study of vitamin metabolism and apoenzyme-coenzyme interactions. In Transport and Inherited Disease: Monograph Based upon Proceedings of the Seventeenth Symposium of The Society for the Study of Inborn Errors of Metabolism; Springer: Dordrecht, The Netherlands, 1981; pp. 3–32. [Google Scholar]
- Bratcher, S.C.; Hsu, R.Y. The pyruvate-proton exchange reaction of malic enzyme from pigeon liver. Biochim. Biophys. Acta (BBA)-Protein Struct. Mol. Enzymol. 1982, 702, 54–60. [Google Scholar] [CrossRef]
- Katagiri, T.; Amao, Y. Trivalent metal ions promote the malic enzyme-catalyzed building of carbon–carbon bonds from CO2 and pyruvate. New J. Chem. 2020, 44, 17208–17214. [Google Scholar] [CrossRef]
- Misono, H.; Sugihara, K.; Kuwamoto, Y.; Nagata, S.; Nagasaki, S. Leucine dehydrogenase from Corynebacterium pseudodiphtheriticum: Purification and characterization. Agric. Biol. Chem. 1990, 54, 1491–1498. [Google Scholar] [CrossRef]
- Ohshima, T.; Soda, K. Valine dehydrogenase from a non-spore-forming bacterium Alcaligenes faecalis: Purification and characterization. Biochim. Biophys. Acta (BBA)-Protein Struct. Mol. Enzymol. 1993, 1162, 221–226. [Google Scholar] [CrossRef]
- Wang, H.; Gunsalus, R.P. The nrfA and nirB nitrite reductase operons in Escherichia coli are expressed differently in response to nitrate than to nitrite. J. Bacteriol. 2000, 182, 5813–5822. [Google Scholar] [CrossRef]
- Drepper, T.; Raabe, K.; Giaourakis, D.; Gendrullis, M.; Masepohl, B.; Klipp, W. The Hfq-like protein NrfA of the phototrophic purple bacterium Rhodobacter capsulatus controls nitrogen fixation via regulation of nifA and anfA expression. FEMS Microbiol. Lett. 2002, 215, 221–227. [Google Scholar] [CrossRef][Green Version]
- Pitcher, G.C.; Probyn, T.A. Suffocating phytoplankton, suffocating waters—Red tides and anoxia. Front. Mar. Sci. 2016, 3, 186. [Google Scholar] [CrossRef]
- Durand, S.; Guillier, M. Transcriptional and post-transcriptional control of the nitrate respiration in bacteria. Front. Mol. Biosci. 2021, 8, 667758. [Google Scholar] [CrossRef]



| Layer | T (°C) | Sal | Depth (m) | pH | DO (mg/L) | Chl-a (µg/L) | NO3−-N (mg/L) | NO2−-N (mg/L) | NH4+-N (mg/L) | PO43−-P (mg/L) | SiO32−-Si (mg/L) | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| BR | NR | BR | NR | BR | NR | BR | NR | BR | NR | BR | NR | BR | NR | BR | NR | BR | NR | BR | NR | BR | NR | |
| S | 22.9 * | 23.5 * | 29.1 | 29.1 | 0.2 | 0.1 | 7.7 | 7.6 | 7.5 | 7.1 | 2.7 | 2.1 | 184.4 | 240.8 | 29.7 | 28.9 | 851.8 * | 130.2 * | 72.8 * | 27.2 * | 447.9 | 260.4 |
| M | 22.8 | 22.9 | 29.6 | 29.5 | 5.6 | 5.1 | 7.7 | 7.6 | 7.4 | 7.1 | 2.5 | 2.1 | 96.5 | 97.3 | 24.5 | 21.0 | 291.5 | 316.1 | 13.7 | 10.8 | 395.0 | 574.4 |
| B | 22.7 | 22.9 | 28.8 ** | 29.6 ** | 11.1 | 10.0 | 7.7 | 7.6 | 7.6 | 7.1 | 3.0 | 3.1 | 147.3 * | 90.9 * | 37.3 ** | 21.4 ** | 222.0 * | 311.1 | 51.6 | 12.4 | 516.7 | 502.2 |
| Network Properties | BR | NR |
|---|---|---|
| Number of nodes | 401 | 903 |
| Number of prokaryotic nodes | 342 | 697 |
| Number of microeukaryotic nodes | 59 | 206 |
| Number of edges | 1040 | 4300 |
| Proportion of positive correlations | 79.1% | 94.1% |
| Average degree | 5.187 | 9.524 |
| Average path length | 1.303 | 1.123 |
| Network diameter | 6 | 6 |
| Clustering coefficient | 0.953 | 0.995 |
| Betweenness centrality | 0.0005 | 0.0001 |
| Number of modules | 95 | 159 |
| Modularity | 0.943 | 0.946 |
| BR | NR | ||||
|---|---|---|---|---|---|
| Genus | Identifiable Taxonomic Level Above Genus | Betweenness Centrality | Genus | Identifiable Taxonomic Level Above Genus | Betweenness Centrality |
| Aurantivirga | Flavobacteriaceae | 38 | Unclassified | Rhodobacteraceae | 42 |
| HIMB11 | Rhodobacteraceae | 25.5 | Unclassified | Cryptomonadales | 30 |
| HIMB11 | Rhodobacteraceae | 25.5 | Unclassified | Verrucomicrobiales | 24 |
| Donghicola | Rhodobacteraceae | 18 | Gymnodinium | Gymnodiniaceae | 24 |
| Aurantivirga | Flavobacteriaceae | 16 | Protaspa | Cryomonadida | 21 |
| Donghicola | Rhodobacteraceae | 16 | Marivivens | Rhodobacteraceae | 20 |
| Aurantivirga | Flavobacteriaceae | 16 | Unclassified | Chlorarachniophyceae | 20 |
| Unclassified | Rhodobacteraceae | 16 | Unclassified | Ilumatobacteraceae | 18 |
| Sva0996 marine group | Microtrichaceae | 15 | Roseibacillus | Rubritaleaceae | 18 |
| HIMB11 | Rhodobacteraceae | 15 | Ca. Actinomarina | Actinomarinaceae | 17.5 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Y.; Zhou, M.; Yue, X.; Chen, Y.; Su, D.; Liu, Z. Noctiluca scintillans Bloom Reshapes Microbial Community Structure, Interaction Networks, and Metabolism Patterns in Qinhuangdao Coastal Waters, China. Microorganisms 2025, 13, 1959. https://doi.org/10.3390/microorganisms13081959
Wang Y, Zhou M, Yue X, Chen Y, Su D, Liu Z. Noctiluca scintillans Bloom Reshapes Microbial Community Structure, Interaction Networks, and Metabolism Patterns in Qinhuangdao Coastal Waters, China. Microorganisms. 2025; 13(8):1959. https://doi.org/10.3390/microorganisms13081959
Chicago/Turabian StyleWang, Yibo, Min Zhou, Xinru Yue, Yang Chen, Du Su, and Zhiliang Liu. 2025. "Noctiluca scintillans Bloom Reshapes Microbial Community Structure, Interaction Networks, and Metabolism Patterns in Qinhuangdao Coastal Waters, China" Microorganisms 13, no. 8: 1959. https://doi.org/10.3390/microorganisms13081959
APA StyleWang, Y., Zhou, M., Yue, X., Chen, Y., Su, D., & Liu, Z. (2025). Noctiluca scintillans Bloom Reshapes Microbial Community Structure, Interaction Networks, and Metabolism Patterns in Qinhuangdao Coastal Waters, China. Microorganisms, 13(8), 1959. https://doi.org/10.3390/microorganisms13081959






