Abstract
Pleosporalean fungi play significant roles as plant pathogens, saprobes, and endophytes in a wide variety of economically important plant hosts. During an investigation of saprobic fungi from Jilin and Hebei, China, five pleosporalean isolates were obtained from the dead stems of Xanthoceras sorbifolium. Morphological evidence and multi-locus sequence analyses using a combined dataset of ITS, LSU, SSU, rpb2, tef1-α, and tub2 indicate that these isolates represent two new species (Alloleptosphaeria xanthoceratis and Lophiostoma multiforme) and a new record of Lophiostoma montanae. Full morphological descriptions and illustrations are provided herein, and phylogenetic relationships of three pleosporalean taxa are also discussed. Detailed morphological descriptions and illustrations are presented, along with phylogenetic affiliations of three pleosporalean taxa.
1. Introduction
Ascomycota, commonly known as sac fungi, constitutes the largest and most ecologically diverse phylum within the fungal kingdom [1]. The phylum exhibits remarkable morphological variability, ranging from unicellular yeasts to multicellular structures producing elaborate fruiting bodies [2,3]. They occupy virtually every terrestrial and aquatic habitat, functioning as decomposers, pathogens, endophytes, and mutualistic symbionts (e.g., lichens and mycorrhizae) [4]. These fungi are economically important as antibiotic producers, industrial agents, model organisms, and sources of bioactive metabolites [5]. Given their ecological dominance and biotechnological relevance, Ascomycota remain a critical focus of mycological research.
Pleosporales, a diverse order in the class Dothideomycetes (Ascomycota), comprises over 10,000 species across more than 90 families [6]. Members of this order are characterized by pseudothecial ascomata, bitunicate and fissitunicate asci, and aseptate or septate ascospores that vary in pigmentation, shape, and gelatinous sheath [7]. Asexual morphs are predominantly coelomycetous or can sometimes be hyphomycetous [8,9,10,11]. Pleosporalean fungi contain saprobic, pathogenic, endophytic, parasitic, hyperparasitic, and lichenized species that have successfully colonized virtually all global habitats, from terrestrial to aquatic environments [7,8,9]. Over the past decade, molecular phylogenetic studies coupled with morphological evidence have significantly refined the taxonomy of Pleosporales, revealing abundant diversity and leading to the establishment of new families and genera [6,12]. Recent investigations in China have revealed numerous novel species associated with woody oil plants [13,14,15].
Xanthoceras sorbifolium (Sapindaceae) is an important tree native to northern China with resistance to drought and could be used for windbreak, sand fixation, and desertification control [16,17]. It also has antitumor and anti-inflammatory activities [18]. Notably, its oil-rich seeds (containing >50% unsaturated fatty acids) have garnered attention as a promising biodiesel feedstock and edible oil source with potential cardiovascular benefits [18]. While extensively studied for its medicinal properties, oil-rich seeds, and stress tolerance, the microfungi associated with this plant remain underexplored despite their potential agricultural and ecological implications [18]. Preliminary investigations have identified several saprobic and pathogenic fungi, including Angularia xanthoceratis [19], Comoclathris xanthoceratis [20], Neocucurbitaria subribicola [21], Fusarium sp., and Verticillium sp. [22], which reflect the abundance of potential microfungal resources in X. sorbifolium.
During a survey of microfungi allied with X. sorbifolium in China, a series of interesting pleosporalean fungi was collected. This study aimed to enrich knowledge of the pleosporalean taxa in X. sorbifolium. Morphological comparisons integrated with multilocus phylogenetic analyses were conducted to determine the classification of these new collections. Two novel species and one newly recorded species are described from Jilin and Hebei Provinces, China.
2. Materials and Methods
2.1. Sample Collection, Isolation, and Morphological Observation
During a series of fieldwork from 2021 to 2022 in Jilin and Hebei Provinces, China, dried woody litter samples were collected. The collected samples, labeled with location, date, host, and collection data in plastic bags, were transported to the lab for morphological study. Pure colonies were obtained through single spore isolation [23] and incubated on potato dextrose agar (PDA) at 25 °C for 2 to 6 weeks. The specimens were preserved at the Herbarium of Mycology, Jilin Agricultural University (HMJAU), Changchun, China, while the pure cultures were stored in the Culture Collection of the International Cooperation Research Center of China for New Germplasm Breeding of Edible Mushrooms (CCMJ). The novel species were documented and assigned accession numbers in MycoBank [24].
Fungal morphological structures were examined and photographed using a Zeiss Stemi 2000C stereo microscope (paired with a Leica DFC450C camera) and a Zeiss AX10 light microscope (fitted with an Axiocam 506 camera) (ZEISS/Leica, Jena/Wetzlar, Germany). Microscopic elements were measured using the ZEN 3.4 (blue edition) program (ZEISS, Jena, Germany), and all images were processed with Adobe Photoshop 2020 (Adobe Systems, San Jose, CA, USA).
2.2. DNA Extraction, PCR Amplification, and Sequencing
Fungal genomic DNA was extracted from mycelium using the NuClean PlantGen DNA Kit (CWBIO, Taizhou, China) following the manufacturer’s protocol. The DNA amplification was conducted by polymerase chain reaction (PCR) using internal transcribed spacers (ITS), large subunit (LSU) rDNA, small subunit (SSU) rDNA, translation elongation factor 1-α (tef1-α), RNA polymerase II second largest subunit (rpb2), and beta-tubulin (tub2). The specific primer pairs for six molecular markers are described in Table 1. Amplification was conducted following Xu et al. [19,21], with PCR products confirmed via 1% agarose gel electrophoresis (stained with 0.5 mL of 10,000× DNA dye; Biotium, Fremont, CA, USA). Successful amplifications were sequenced by Sangon Biotech (Shanghai, China). The newly obtained sequences have been submitted to GenBank [25], with accession numbers provided in Table 2 and Table 3.

Table 1.
PCR primers utilized for amplification and sequencing in the investigation.
2.3. Molecular Phylogeny
The new strains were initially identified by molecular techniques through comparison of individual gene sequences using BLASTn [31], with relevant sequence data acquired from GenBank (Table 2 and Table 3). Alignments were generated using MAFFT v.7 [32] with the L-INS-i algorithm (1PAM/k = 2 scoring matrix, 1.50 gap opening penalty for nucleotide sequences), followed by manual adjustment in BioEdit v7.2.5 [33]. The AliView program was used to convert the alignment data files to PHYLIP and NEXUS formats [34].
Phylogenetic trees were constructed using individual genetic markers and subsequently analyzed in combination, along with a concatenated multi-gene dataset. Maximum likelihood analysis was conducted with RAxML-HPC2 on XSEDE through the CIPRES web portal (http://www.phylo.org/portal2/; accessed 28 March 2025) [35]. The optimal evolutionary models for both individual and concatenated datasets were determined using jModeltest 2.1.10, with model selection based on the Akaike Information Criterion (AIC) for posterior probability analysis [36]. The GTR+GAMMA model of nucleotide evolution was applied to all datasets with 1000 bootstrap repeats. BI analysis was performed using MrBayes v.3.2.6 implemented through the CIPRES Science Gateway portal (https://www.phylo.org; accessed on 29 March 2025) [37]. Simultaneous Markov chains were run for 800 million generations. Trees were sampled every 100 generations, with the first 20% discarded as burn-in. Outgroup taxa selection comprised Didymella exigua (CBS 183.55), D. rumicicola (CBS 683.79), Teichospora rubriostiolata (TR7), and T. trabicola (C134) (Figure 1 and Figure 2). The phylogenetic tree file was downloaded from CIPRES and visualized in FigTree v.1.4.4 [38]. The tree designs were crafted in Adobe Illustrator CS6.

Table 2.
Names and corresponding GenBank accession numbers of Leptosphaeriaceae taxa used in phylogenetic analyses, with new sequences in bold blue and type strains in bold.
Table 2.
Names and corresponding GenBank accession numbers of Leptosphaeriaceae taxa used in phylogenetic analyses, with new sequences in bold blue and type strains in bold.
| Species | Strain/Isolate | GenBank Accession Numbers | |||
|---|---|---|---|---|---|
| ITS | LSU | SSU | tub2 | ||
| Alloleptosphaeria italica | MFLUCC 14-0934 | KT454722 | KT454714 | _ | _ |
| A. clematidis | MFLUCC 17-2071 | MT310604 | MT214557 | MT226674 | _ |
| A. iridicola | CBS 143395 | MH107919 | MH107965 | _ | NA |
| A. shangrilana | HKAS:112210 | MW431059 | MW431315 | MW431058 | NA |
| A. xanthoceratis | CCMJ 13066 | PP151694 | PP153449 | PV569760 | PV670045 |
| Didymella exigua | CBS 183.55 | GU237794 | EU754155 | EU754056 | GU237525 |
| D. rumicicola | CBS 683.79 | KT389503 | KT389721 | _ | KT389800 |
| Heterospora chenopodii | CBS 448.68 | FJ427023 | EU754187 | EU754088 | _ |
| H. chenopodii | CBS 115.96 | JF740227 | EU754188 | EU754089 | _ |
| H. dimorphospora | CBS 165.78 | JF740204 | JF740281 | JF740098 | _ |
| H. dimorphospora | CBS 345.78 | JF740203 | GU238069 | GU238213 | _ |
| Lep. conoidea | CBS 616.75 | JF740201 | JF740279 | _ | KT389804 |
| Lep. doliolum | CBS 155.94 | JF740207 | JF740282 | _ | JF740146 |
| Lep. doliolum | CBS 505.75 | JF740205 | GQ387576 | GQ387515 | JF740144 |
| Lep. ebuli | MFLUCC 14-0828 | KP744446 | KP744488 | KP753954 | _ |
| Lep. irregularis | MFLUCC 15-1118 | KX856056 | KX856055 | _ | _ |
| Lep. slovacica | CBS 389.80 | JF740247 | JF740315 | JF740101 | _ |
| Lep. urticae | MFLU 18-0591 | MK123333 | MK123332 | MK123329 | _ |
| Neoleptosphaeria jonesii | MFLUCC 16-1442 | KY211869 | KY211870 | KY211871 | _ |
| Neol. rubefaciens | CBS 223.77 | JF740243 | JF740312 | _ | _ |
| Neol. rubefaciens | CBS 387.80 | JF740242 | JF740311 | _ | _ |
| Pseudoleptosphaeria etheridgei | CBS 125980 | JF740221 | JF740291 | _ | _ |
| Querciphoma carteri | CBS 105.91 | KF251209 | GQ387594 | GQ387533 | KF252700 |
| Q. carteri | CBS 101633 | KF251210 | GQ387593 | GQ387532 | KF252701 |
| Sclerenchymomyces clematidis | MFLUCC 17-2180 | MT310605 | MT214558 | MT226675 | _ |

Table 3.
Names and corresponding GenBank accession numbers of Lophiostomataceae taxa used in phylogenetic analyses, with new sequences in bold blue and type strains in bold.
Table 3.
Names and corresponding GenBank accession numbers of Lophiostomataceae taxa used in phylogenetic analyses, with new sequences in bold blue and type strains in bold.
| Species | Strain/Isolate | GenBank Accession Numbers | ||||
|---|---|---|---|---|---|---|
| ITS | LSU | SSU | tef1-α | rpb2 | ||
| Crassiclypeus aquaticus | KH 104 | LC312499 | LC312528 | LC312470 | LC312557 | LC312586 |
| C. aquaticus | KT 970 | LC312501 | LC312530 | LC312472 | LC312559 | LC312588 |
| Dimorphiopsis brachystegiae | CPC 22679 | KF777160 | KF777213 | _ | _ | _ |
| Flabellascoma aquaticum | KUMCC 15-0258 | MN304827 | MN274564 | MN304832 | MN328898 | MN328895 |
| F. cycadicola | KT 2034 | LC312502 | LC312531 | LC312473 | LC312560 | LC312589 |
| F. fusiforme | MFLUCC 18-1584 | MN304830 | MN274567 | _ | MN328902 | _ |
| F. minimum | KT 2013 | LC312503 | LC312532 | LC312474 | LC312561 | LC312590 |
| F. minimum | KT 2040 | LC312504 | LC312533 | LC312475 | LC312562 | LC312591 |
| Lentistoma bipolare | KT 3056 | LC312513 | LC312542 | LC312484 | LC312571 | LC312600 |
| Len. bipolare | CBS 115375 | LC312506 | LC312535 | LC312477 | LC312564 | LC312593 |
| Leptoparies palmarum | KT 1653 | LC312514 | LC312543 | LC312485 | LC312572 | LC312601 |
| Lophiostoma arundinis | KT 606 | JN942964 | AB618998 | AB618679 | LC001737 | JN993482 |
| Lop. arundinis | KT 651 | JN942965 | AB618999 | AB618680 | LC001738 | JN993486 |
| Lop. biappendiculatum | KT 975 | _ | GU205228 | GU205254 | _ | _ |
| Lop. biappendiculatum | KT 1124 | _ | GU205227 | GU205256 | _ | _ |
| Lop. caespitosum | CBS 147391 | MW759252 | MW750387 | _ | MW752404 | MW752383 |
| Lop. caespitosum | MFLUCC 13-0442 | KP899134 | KP888639 | KP899125 | KR075161 | _ |
| Lop. caespitosum | MFLUCC 14-0993 | KP899135 | KP888640 | KP899126 | KR075162 | _ |
| Lop. carabassense | CBS 149324 | MT679671 | OL544969 | OL544968 | OL554876 | _ |
| Lop. carpini | CBS 147279 | MW759258 | MW750386 | _ | MW752405 | MW752384 |
| Lop. caryophyllacearum | MFLUCC 17-0749 | MG828964 | MG829076 | MG829176 | MG829238 | _ |
| Lop. caudatum | KT 530 | LC001723 | AB619000 | AB618681 | LC001739 | _ |
| Lop. caulium | KT 603 | LC001724 | AB619001 | AB618682 | LC001740 | _ |
| Lop. caulium | KT 633 | LC001725 | AB619002 | AB618683 | LC001741 | _ |
| Lop. clavatum | CBS 147278 | MW759259 | MW750385 | _ | MW752406 | MW752385 |
| Lop. clavatum | MFLUCC 18-1316 | _ | MN274566 | MN304835 | MN328901 | _ |
| Lop. clematidicola | MFLUCC 16-0446 | MT310609 | MT214563 | MT226680 | MT394742 | _ |
| Lop. clematidis | MFLUCC 17-2081 | MN393004 | MT214562 | MT226679 | MT394741 | MT394689 |
| Lop. clematidis-subumbellatae | MFLUCC 17-2063 | MT310607 | MT214560 | MT226677 | MT394739 | MT394687 |
| Lop. clematidis-vitalbae | MFLUCC 16-1368 | MT310610 | MT214564 | MT226681 | MT394743 | _ |
| Lop. compressum | CBS 147276 | MW759272 | MW750382 | _ | MW752408 | MW752381 |
| Lop. compressum | IFRD 2014 | _ | FJ795437 | FJ795480 | _ | FJ795457 |
| Lop. cornisporum | KH 322 | LC312515 | LC312544 | LC312486 | LC312573 | LC312602 |
| Lop. coronillae | MFLUCC 14-0941 | KT026120 | KT026112 | KT026116 | _ | _ |
| Lop. crenatum | AFTOL-ID 1581 | _ | DQ678069 | DQ678017 | DQ677912 | DQ677965 |
| Lop. dictyosporum | CBS 147389 | MW759251 | MW750379 | _ | MW752411 | MW752388 |
| Lop. erumpens | CBS 147275 | MW759262 | MW750381 | _ | MW752409 | MW752386 |
| Lop. fusisporum | CBS 147891 | MW759253 | _ | _ | MW752401 | MW752382 |
| Lop. helichrysi | IT-1296 | KT333435 | KT333436 | KT333437 | KT427535 | _ |
| Lop. heterosporum | AFTOL-ID 1036 | GQ203795 | AY016369 | _ | DQ497609 | DQ497615 |
| Lop. japonicum | KT 686-1 | LC001729 | AB619006 | AB618687 | LC001745 | _ |
| Lop. japonicum | KT 573 | LC001728 | AB619005 | AB618686 | LC001744 | _ |
| Lop. jonesii | GAAZ 54-1 | KX687757 | KX687753 | KX687755 | KX687759 | _ |
| Lop. jonesii | GAAZ 54-2 | KX687758 | KX687754 | KX687756 | KX687760 | _ |
| Lop. jotunheimenense | CBS 147522 | MW759261 | MW750394 | _ | MW752392 | _ |
| Lop. junci | MFLUCC 14-0938 | MG828966 | MG829078 | MG829178 | NA | _ |
| Lop. khanzada-kirgizbaeva | TASM 6158 | MZ966265 | OK017520 | OK017525 | MZ997338 | _ |
| Lop. khanzada-kirgizbaeva | TASM 6164 | MZ966266 | OK017521 | OK017526 | MZ997339 | _ |
| Lop. longiappendiculatum | MFLUCC 17-1452 | MT214368 | MT214462 | MT214415 | MT235783 | _ |
| Lop. longiappendiculatum | MFLUCC 17-1457 | MT214369 | MT214463 | MT214416 | MT235784 | MT235821 |
| Lop. macrostomoides | CBS 147523 | MW759256 | MW750389 | _ | _ | _ |
| Lop. macrostomoides | CBS 147277 | MW759257 | MW750384 | _ | MW752407 | MW752380 |
| Lop. macrostomoides | CBS 123097 | _ | FJ795439 | FJ795482 | GU456277 | FJ795458 |
| Lop. macrostomum | KT 508 | JN942961 | AB619010 | _ | LC001751 | JN993491 |
| Lop. macrostomum | KT 709/HHUF 27293 | AB433276 | AB433274 | AB521732 | LC001753 | JN993493 |
| Lop. macrostomum | KT 635/HHUF 27290 | AB433275 | AB433273 | AB521731 | LC001752 | JN993484 |
| Lop. mangiferae | MFLUCC 17-2651 | MG931031 | MG931025 | MG931028 | _ | _ |
| Lop. mangiferae | MFLUCC 17-2653 | MG931032 | MG931026 | MG931029 | _ | _ |
| Lop. medicaginicola | MFLUCC 17-0681 | MG828967 | MG829079 | MG829179 | _ | _ |
| Lop. montanae | MFLUCC 16-0999 | MT310611 | MT214565 | MT226682 | MT394744 | _ |
| Lop. montanae | UESTCC 23.0038 | OR253137 | OR253296 | OR253209 | OR263570 | OR253750 |
| Lop. montanae | UESTCC 23.0039 | OR253138 | OR253297 | OR253210 | OR251148 | OR253751 |
| Lop. montanae | CCMJ 13067 | PV569791 | PV569911 | PV569764 | PV670048 | PV670046 |
| Lop. montanae | CCMJ 13068 | PV569792 | PV569912 | PV569765 | PV670049 | PV670047 |
| Lop. multiforme | CCMJ 13069 | PP151705 | PP153460 | PV569761 | PV670043 | PV670041 |
| Lop. multiforme | CCMJ 13070 | PP151706 | PP153461 | PV569762 | PV670044 | PV670042 |
| Lop. multiseptatum | CBS 623.86 | _ | GU301833 | GU296163 | _ | GU371791 |
| Lop. multiseptatum | KT 604 | LC001726 | AB619003 | AB618684 | LC001742 | _ |
| Lop. neomuriforme | MFLUCC 13-0744 | KY496740 | KY496719 | KY501110 | _ | _ |
| Lop. obtusisporum | KT 3098 | LC312519 | LC312548 | LC312490 | LC312577 | LC312606 |
| Lop. obtusisporum | KT 2838 | LC312518 | LC312547 | LC312489 | LC312576 | LC312605 |
| Lop. oleae | CGMCC 3.24426 | OR253081 | OR253233 | OR253172 | OR262141 | OR262130 |
| Lop. oleae | UESTCC 23.0036 | OR253079 | OR253231 | OR253171 | OR262139 | OR262129 |
| Lop. ononidis | MFLUCC 14-0613 | KU243128 | KU243125 | KU243126 | KU243127 | _ |
| Lop. paramacrostomum | MFLUCC 11-0463 | _ | KP888636 | KP899122 | _ | _ |
| Lop. plantaginis | CBS 147527 | MW759250 | MW750378 | _ | _ | MW752375 |
| Lop. pseudodictyosporium | MFLUCC 13-0451 | KR025858 | KR025862 | _ | _ | _ |
| Lop. pseudomacrostomum | CBS 147525 | MW759255 | MW750391 | _ | MW752395 | _ |
| Lop. pseudomacrostomum | CBS 147526 | MW759254 | MW750392 | _ | MW752394 | _ |
| Lop. ravennicum | MFLUCC 14-0005 | KP698413 | KP698414 | KP698415 | _ | _ |
| Lop. rosae-ecae | MFLUCC 17-0807 | MG828924 | MG829033 | MG829139 | MG829217 | _ |
| Lop. rosicola | MFLU 15-1888 | MG828968 | MG829080 | MG829180 | MG829240 | _ |
| Lop. sagittiforme | KT 1934 | AB369268 | AB369267 | AB618693 | LC001756 | _ |
| Lop. scabridisporum | BCC 22835 | _ | GQ925844 | GQ925831 | GU479857 | GU479830 |
| Lop. scabridisporum | BCC 22836 | _ | GQ925845 | GQ925832 | GU479856 | GU479829 |
| Lop. scrophulariicola | MFLUCC 17-0689 | MG828969 | MG829081 | _ | _ | _ |
| Lop. semiliberum | KT 622 | JN942966 | AB619012 | AB618694 | LC001757 | JN993483 |
| Lop. semiliberum | KT 652 | JN942967 | AB619013 | AB618695 | LC001758 | JN993485 |
| Lop. semiliberum | KT 828 | JN942970 | AB619014 | AB618696 | LC001759 | JN993489 |
| Lop. spartii-juncei | MFLUCC 13-0351 | KP899136 | KP888641 | KP899127 | KR075163 | _ |
| Lop. submuriforme | CBS 147274 | MW759260 | MW750380 | _ | MW752410 | MW752387 |
| Lop. terricola | SC-12 | JN662930 | JX985750 | JX985749 | _ | _ |
| Lop. thymi | MFLU 15-2131 | MG828970 | MG829082 | MG829182 | MG829241 | _ |
| Lop. tropicum | KH 352 | LC312521 | LC312550 | LC312492 | LC312579 | LC312608 |
| Lop. tropicum | KT 3134 | LC312522 | LC312551 | LC312493 | LC312580 | LC312609 |
| Lop. vitigenum | HH 26930 | LC001735 | AB619015 | AB618697 | LC001761 | _ |
| Lop. vitigenum | HH 26931 | LC001736 | AB619016 | AB618698 | LC001762 | _ |
| Lop. winteri | KT 740 | JN942969 | AB619017 | AB618699 | LC001763 | JN993487 |
| Lop. winteri | KT 764 | JN942968 | AB619018 | AB618700 | LC001764 | JN993488 |
| Neovaginatispora clematidis | MFLUCC 17-2149 | MT310606 | MT214559 | MT226676 | MT394738 | _ |
| Neov. fuckelii | MFLUCC 17-1334 | MN304828 | MN274565 | MN304833 | MN328899 | MN328896 |
| Neov. fuckelii | KT 634 | LC001732 | AB619009 | AB618690 | LC001750 | _ |
| Parapaucispora pseudoarmatispora | KT 2237 | LC100021 | LC100026 | LC100018 | LC100030 | _ |
| Paucispora kunmingense | MFLUCC 17-0932 | MF173432 | MF173428 | MF173430 | MF173434 | MF173436 |
| Pa. quadrispora | KH 448 | LC001733 | LC001722 | LC001720 | LC001754 | _ |
| Pa. quadrispora | KT 843 | LC001734 | AB619011 | AB618692 | LC001755 | _ |
| Pa. versicolor | KH 110 | AB918731 | AB918732 | LC001721 | LC001760 | _ |
| Pa. xishanensis | HKAS 115905 | MZ966267 | OK017522 | OK017527 | MZ997340 | _ |
| Pa. xishanensis | HKAS 115906 | MZ966268 | OK017523 | OK017528 | MZ997341 | _ |
| Platystomum actinidiae | KT 521 | JN942963 | JN941380 | JN941375 | LC001747 | JN993490 |
| Pl. actinidiae | KT 534 | JN942962 | JN941379 | JN941376 | LC001748 | JN993492 |
| Pl. crataegi | MFLUCC 14-0925 | KT026117 | KT026109 | KT026113 | KT026121 | _ |
| Pl. rosae | MFLUCC 15-0633 | KT026119 | KT026111 | KT026115 | _ | _ |
| Pl. salicicola | MFLUCC 15-0632 | KT026118 | KT026110 | KT026114 | _ | _ |
| Pseudopaucispora brunneospora | KH 227 | LC312523 | LC312552 | LC312494 | LC312581 | LC312610 |
| Vaginatispora amygdali | KT 2248 | LC312524 | LC312553 | LC312495 | LC312582 | LC312611 |
| V. amygdali | MFLUCC 18-1526 | MK085055 | MK085059 | MK085057 | MK087657 | _ |
| V. appendiculata | MFLUCC 16-0314 | KU743217 | KU743218 | KU743219 | KU743220 | _ |
| V. appendiculata | MFLUCC 13-0835 | _ | KY264745 | KY264749 | _ | _ |
| V. aquatica | MFLUCC 11-0083 | KJ591577 | KJ591576 | KJ591575 | _ | _ |
| V. armatispora | MFLUCC 18-0247 | MK085056 | MK085060 | MK085058 | MK087658 | MK087669 |
| V. armatispora | MFLUCC 18-0213 | MN304826 | MN274563 | MN304831 | MN328897 | MN328894 |
| V. microarmatispora | MTCC 12733 | MF142592 | MF142593 | MF142594 | MF142595 | MF142596 |
| V. scabrispora | KT 2443 | LC312525 | LC312554 | LC312496 | LC312583 | LC312612 |
| Teichospora rubriostiolata | TR7 | KU601590 | _ | _ | KU601609 | KU601599 |
| T. trabicola | C134 | KU601591 | _ | _ | KU601601 | KU601600 |
3. Results
3.1. Phylogenetic Analyses
3.1.1. Phylogenetic Analyses of Alloleptosphaeria
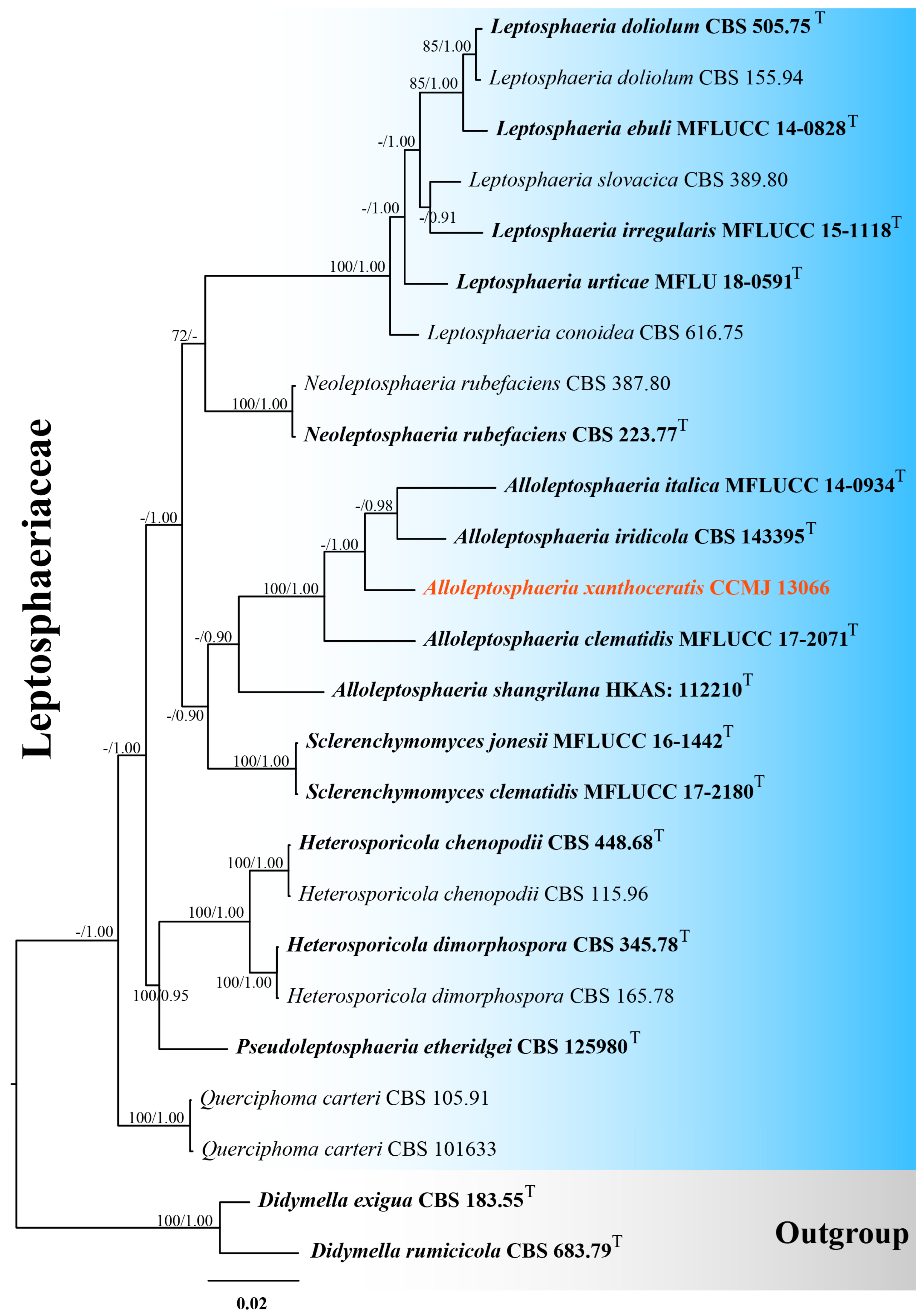
One strain was successfully isolated in the laboratory. Phylogenetic analyses incorporated 25 strains using a 2801-character concatenated alignment (ITS: 556 bp; LSU: 881 bp; SSU: 1023 bp; tub2: 341 bp; gaps included). The optimal RAxML tree (likelihood score: −8462.532314) was derived from an alignment of 504 distinct patterns, including 25.52% undetermined characters/gaps. Estimated base frequencies were as follows: A = 0.244324, C = 0.224019, G = 0.271360, and T = 0.260297; substitution rates, AC = 1.370239, AG = 2.327667, AT = 1.974968, CG = 0.418253, CT = 5.227042, and GT = 1.000000; evolutionary parameters included gamma shape (α = 0.583180) and invariable sites (I = 0.753617). For the Bayesian analysis, a total of 1598 trees were retained after the 20% burn-in with a stop value of 0.008440. Both ML and BI methods yielded consistent topologies (Figure 1 and Figure S1). In our phylogenetic tree, Alloleptosphaeria xanthoceratis (CCMJ 13066) formed a distinct lineage with 42% ML and 1.00 BPP support (Figure 1).
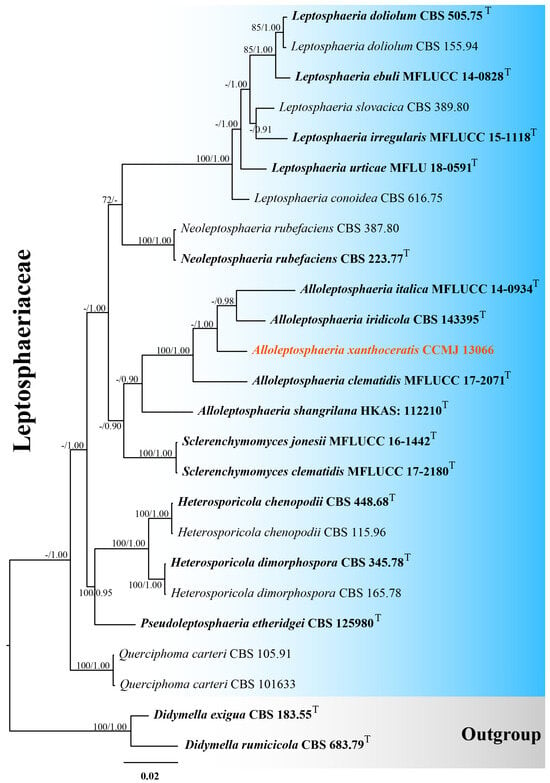
Figure 1.
The Bayesian 50% majority-rule consensus tree based on concatenated ITS, LSU, SSU, and tub2 sequences in Leptosphaeriaceae. The tree is rooted with Didymella exigua (CBS 183.55) and D. rumicicola (CBS 683.79). Maximum likelihood bootstrap support values ≥ 70% (ML) and Bayesian posterior probabilities ≥ 0.9 (BPP) are given at the nodes as ML/BPP. The type strains are in bold and black. The newly generated isolates are shown in bold red and marked with “T”.
Figure 1.
The Bayesian 50% majority-rule consensus tree based on concatenated ITS, LSU, SSU, and tub2 sequences in Leptosphaeriaceae. The tree is rooted with Didymella exigua (CBS 183.55) and D. rumicicola (CBS 683.79). Maximum likelihood bootstrap support values ≥ 70% (ML) and Bayesian posterior probabilities ≥ 0.9 (BPP) are given at the nodes as ML/BPP. The type strains are in bold and black. The newly generated isolates are shown in bold red and marked with “T”.
3.1.2. Phylogenetic Analyses of Lophiostoma
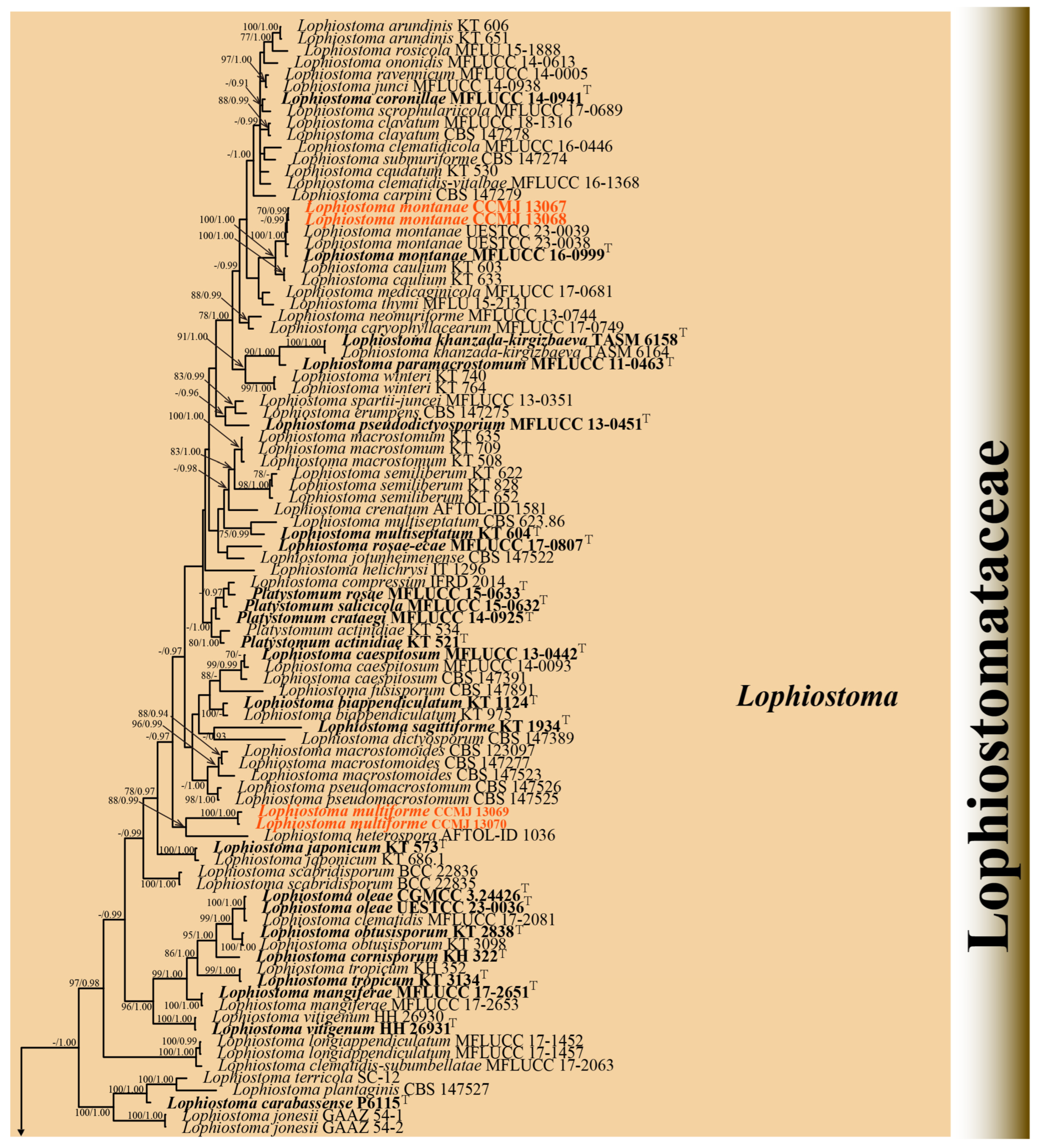
Four strains were successfully isolated in the laboratory. Phylogenetic analyses were performed using a concatenated alignment of ITS (1–603 bp), LSU (604–1521 bp), SSU (1522–2503 bp), tef1-α (2504–3468 bp), and rpb2 (3469–4494 bp) sequences from 125 strains, totaling 4494 characters (including gaps). The RAxML analysis yielded a best-scoring tree (Figure 2) with a final ML optimization likelihood value of −34137.002757. The matrix had 1885 distinct alignment patterns, with 26.95% of undetermined characters or gaps. Estimated base frequencies were as follows: A = 0.249896, C = 0.247227, G = 0.266396, and T = 0.236482; substitution rates AC = 1.541329, AG = 4.043212, AT = 1.269617, CG = 1.398831, CT = 9.131101, and GT = 1.000000; proportion of invariable sites (I) = 0.563654; and gamma distribution shape parameter (α) = 0.661958. Bayesian analysis showed similar topologies with the RAxML analysis (Figure 2 and Figure S2); therefore, only the BI tree is presented herein (Figure 2). Bayesian inference yielded 11,202 post-burn-in trees, with convergence achieved at a stop value of 0.009957.
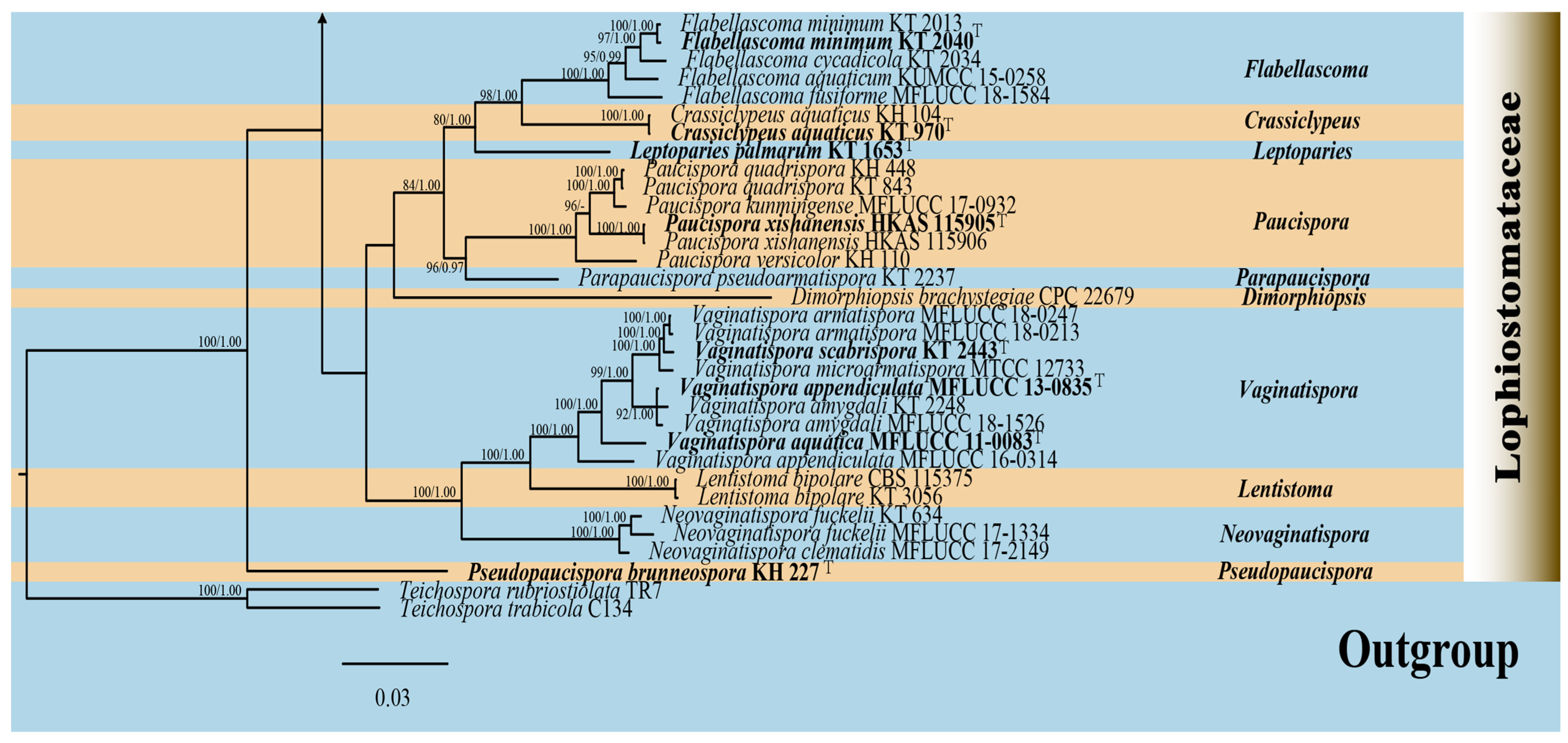
The phylogenetic analyses (ML/BI) strongly supported the monophyly of 11 Lophiostomataceae genera, with high statistical support (100% ML/1.00 BPP). A total of 88 strains of Lophiostoma formed a monophyletic clade with 75% ML support (Figure 2). Lophiostoma multiforme (CCMJ 13069 and CCMJ 13070) formed a well-supported monophyletic lineage with 88% ML and 0.99 BPP values (Figure 2). Lophiostoma montanae (CCMJ 13067 and CCMJ 13068) grouped with L. montanae (MFLUCC 16-0999, UESTCC 23.0038, and UESTCC 23.0039) with 0.99 BPP values.
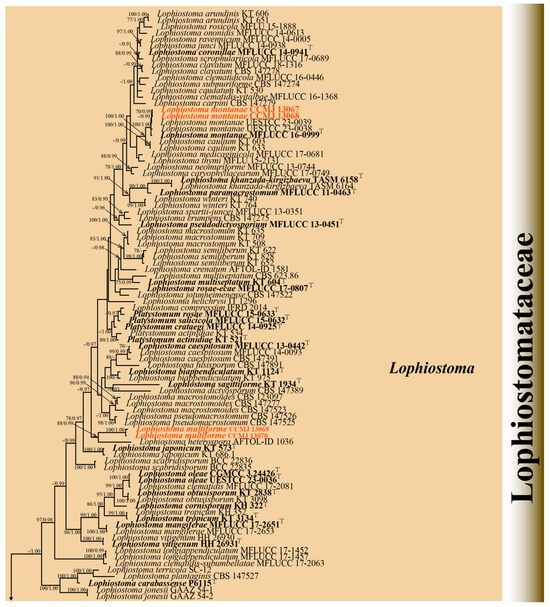
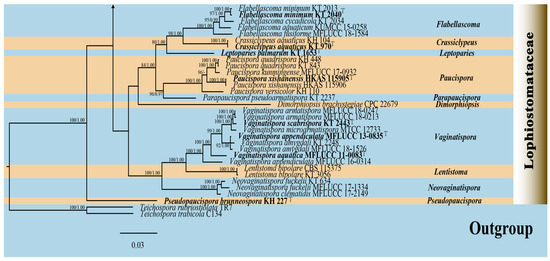
Figure 2.
The Bayesian 50% majority-rule consensus tree based on concatenated ITS, LSU, SSU, tef1-α, and rpb2 sequences in Lophiostomataceae. The tree is rooted with Teichospora rubriostiolata (TR7) and T. trabicola (C134). Maximum likelihood bootstrap support values ≥ 70% (ML) and Bayesian posterior probabilities ≥ 0.9 (BPP) are at the nodes as ML/BPP. The type strains are in bold and black. The newly generated isolates are indicated in bold red and marked with “T”.
Figure 2.
The Bayesian 50% majority-rule consensus tree based on concatenated ITS, LSU, SSU, tef1-α, and rpb2 sequences in Lophiostomataceae. The tree is rooted with Teichospora rubriostiolata (TR7) and T. trabicola (C134). Maximum likelihood bootstrap support values ≥ 70% (ML) and Bayesian posterior probabilities ≥ 0.9 (BPP) are at the nodes as ML/BPP. The type strains are in bold and black. The newly generated isolates are indicated in bold red and marked with “T”.
3.2. Taxonomy
3.2.1. Alloleptosphaeria xanthoceratis R. Xu and Y. Li, sp. nov. (Figure 3)
MycoBank Number: 858873
Etymology: referring to the host plant genus Xanthoceras (Sapindaceae).
Holotype: HMJAU 60190
Description: Saprobic on dead stems of Xanthoceras sorbifolium. Sexual morph: Undetermined. Asexual morph: Conidiomata 131–175 × 110–200 µm ( = 151 × 158 µm, n = 5), pycnidial, solitary to aggregated in small groups, semi-immersed in host substrate, black, elongate, subglobose, without a distinct ostiole. Ostioles absent. Conidiomatal wall multilayered, thick, 12–30 µm wide, comprising 3–4 layers of scleroplectenchymatous tissue, pale brown to brown, arranged in textura angularis. Conidiophores reduced to conidiogenous cells. Conidiogenous cells hyaline, phialidic, discrete, determinate, 9.8–22 × 1.8–3 µm ( = 15 × 2.5 µm, n = 20), arising from inner wall layers. Conidia 3.6–6.4 × 2–6.4 µm ( = 4.8 × 2.6 µm, n = 40), aseptate, hyaline, smooth-walled, guttulate, subcylindrical to narrowly ellipsoid, apex obtuse, base truncate.
Culture characteristics: Colonies reach 2 cm in diameter after 14 days at 25 °C. Colonies are dense, round, milky-white central domes, transitioning to gray toward the periphery, creeping hyphae margins irregular with pale brown; reverse white centrally, becoming dark brown peripherally.
Material examined: CHINA. Jilin Province: Changchun city, on dead stems of Xanthoceras sorbifolium. 16 October 2021, Rong Xu, HMJAU 60190 (holotype); living culture, CCMJ 13066.
GenBank numbers: ITS: PP151694; LSU: PP153449; SSU: PV569760; tub2: PV670045.
Notes: A BLASTn search of the ITS region revealed that our strain shares 96.41% similarity with A. iridicola (CBS 143395, NR_159068). The tub2 sequence is 84.92% similar to Leptosphaeria zhaotongensis (HKAS:124664, OP476695) with 94% query cover. In our phylogenetic study, Alloleptosphaeria xanthoceratis is phylogenetically closely related to three other Alloleptosphaeria species. To date, only A. iridicola has been reported to produce asexual morph within this genus [39]. Crous et al. [39] initially described Subplenodomus iridicola as the causative agent of Iris leaf spots in England, and it was later transferred to Alloleptosphaeria based on molecular evidence [40]. Morphologically, Alloleptosphaeria xanthoceratis is distinguished from A. iridicola by its larger conidiogenous cells (9.8–22 × 1.8–3 µm vs. 4–7 × 4–6 µm). We therefore formally describe it as a new species.
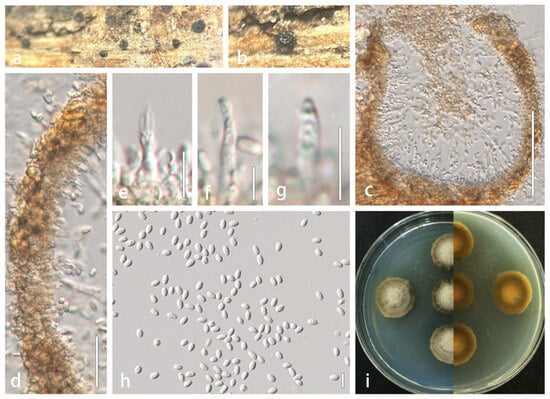
Figure 3.
Alloleptosphaeria xanthoceratis (HMJAU 60190, holotype). (a,b) Conidiomata developing on natural substrate. (c) Longitudinal section through conidioma. (d) Wall of conidioma. (e–g) Conidiogenous cells and conidia. (h) Mature conidia. (i) Colony morphology on PDA. Scale bars: (c) = 100 µm; (d) = 20 µm; (e–g) = 10 µm; (h) = 5 µm.
Figure 3.
Alloleptosphaeria xanthoceratis (HMJAU 60190, holotype). (a,b) Conidiomata developing on natural substrate. (c) Longitudinal section through conidioma. (d) Wall of conidioma. (e–g) Conidiogenous cells and conidia. (h) Mature conidia. (i) Colony morphology on PDA. Scale bars: (c) = 100 µm; (d) = 20 µm; (e–g) = 10 µm; (h) = 5 µm.
3.2.2. Lophiostoma montanae (Phukhams., Sue, and K.D. Hyde) Andreasen, Jaklitsch, and Voglmayr, Persoonia 46: 259 (2021) [41] = Sigarispora montanae Phukhams. Sue et al., Fungal Diversity 102: 55 [42], (Figure 4)
Index Fungorum number: IF557124; Facesofungi number: FoF 07295
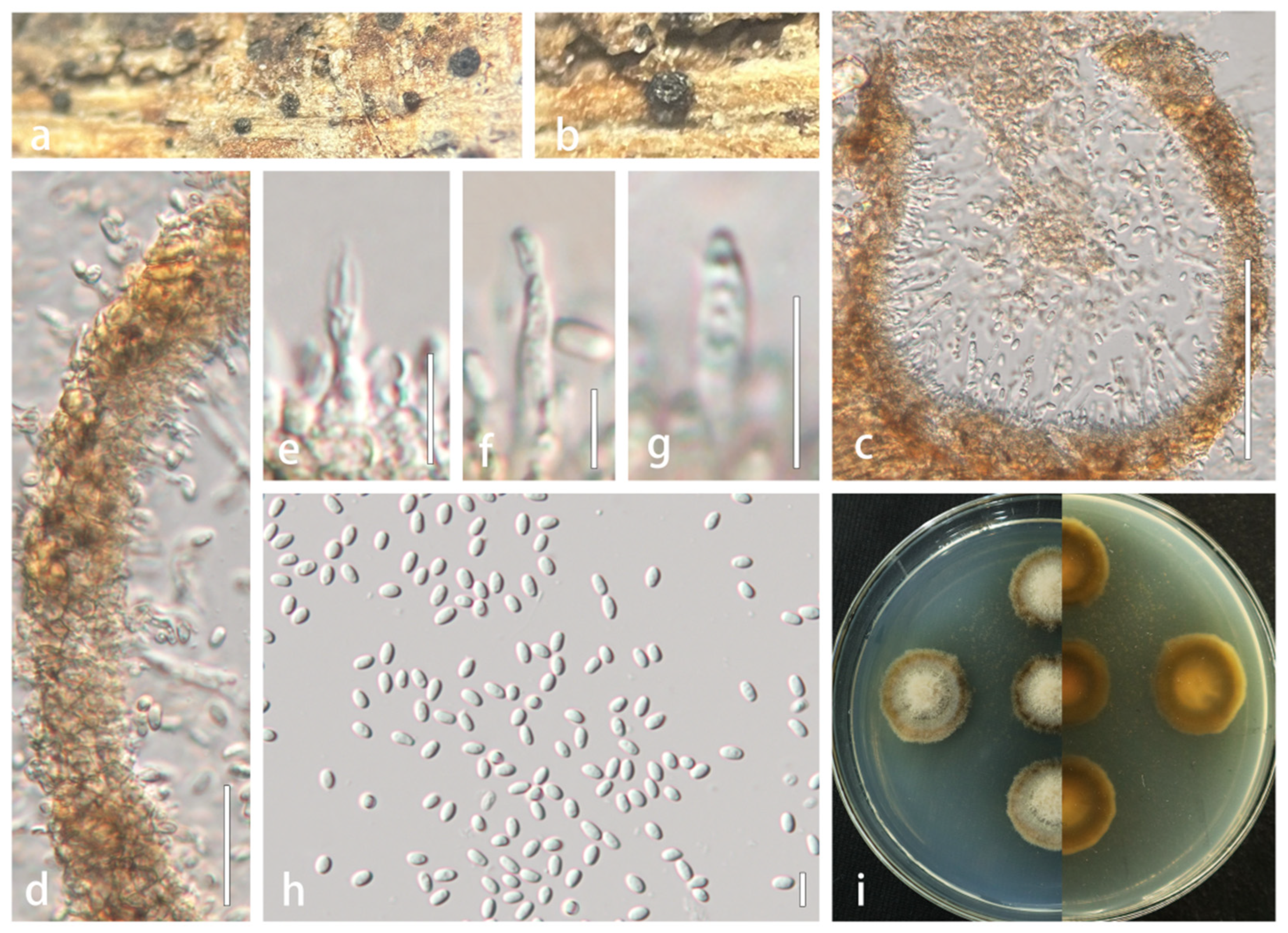
Description: Saprobic on dead stems of Xanthoceras sorbifolium. Sexual morph: Ascomata dark brown to black, solitary, scattered, semi-immersed, globose, 186–350 × 160–278 μm ( = 260 × 216 μm, n = 5), rough-walled, apex partially carbonaceous forming a clypeate structure, ostiolate, coriaceous. Ostioles central, crest-like at apex, 65–93 × 39–57 μm ( = 76.1 × 49.6 μm, n = 5), elongated and laterally compressed, filled with hyaline periphyses, irregular wall. Peridium composed of 6–8 layers of thick-walled, brown to dark brown cells of textura angularis, broader at the apex and gradually tapering toward the base, 22–46 μm wide ( = 33 μm, n = 20). Hamathecium 1–1.9 µm ( = 1.6 μm, n = 20), densely arranged, hyaline, septate, frequently branched, and anastomosing within a gelatinous matrix. Asci bitunicate, fissitunicate, 8-spored, broad cylindrical to clavate, 102–130 × 9–14 µm ( = 118 × 12 µm, n = 20), short pedicellate (pedicel furcate), apically rounded with distinct ocular chamber. Ascospores biseriate and partially overlapping, fusiform with attenuated ends, initially hyaline, becoming yellowish-brown at maturity, 16–29 × 4.5–6.8 µm ( = 21.1 × 5.6 µm, n = 50), 3–5 transversely septa, constricted at the septa, cells above medial septum swollen, wall smooth, guttulate, surrounded by persistent mucilaginous sheath (3–5 μm) extending as polar appendages. Asexual morph: Undetermined.
Culture characteristics: Colonies on PDA reaching 30 mm in diameter after 35 days at 25 °C. Cultures from above are circular, flat, thick and dense, umbonate, entire edge, grayish white; reverse is cream-colored marginally, gradually darkening to brown at the middle region with a distinct red-brown central disc.
Material examined: CHINA. Jilin Province: Changchun city, Jinyue district, on dead stem of Xanthoceras sorbifolium, 10 June 2022, Rong Xu, XR36.1 (HMJAU 64835), living culture CCMJ 13067; XR36.2 (HMJAU 64836), living culture CCMJ 13068.
GenBank numbers: CCMJ 13067: ITS = PV569791, LSU = PV569911, SSU = PV569764, tef1-α = PV670048, rpb2 = PV670046. CCMJ 13068: ITS = PV569792, LSU = PV569912, SSU = PV569765, tef1-α = PV670049, rpb2 = PV6700467.
Host: Xanthoceras sorbifolium (this study), Clematis montana, Paeonia suffruticosa.
Distribution: Jilin Province (this study), Yunnan Province [42], Sichuan Province [14], China.
Notes: Lophiostoma montanae was originally described by Phukhamsakda et al. [42] from dead leaves of Clematis montana in China, characterized by its distinctive ascomata featuring ostioles with crest-like apices. Our new isolates align morphologically with this species and form a fully supported clade (100% ML/1.00 BPP) with L. montanae in phylogenetic analyses (Figure 2 and Figure 4). Thus, the isolates are identified as L. montanae, representing the first record of this species in X. sorbifolium and expanding its known host range in China.
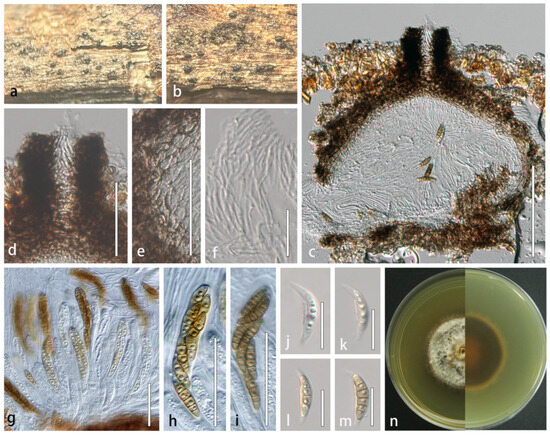
Figure 4.
Lophiostoma montanae (HMJAU 64835). (a,b) Ascomata developing on natural host substrate. (c) Longitudinal section through ascomata. (d) Ostiole. (e) Partial peridium wall. (f) Anastomosing pseudoparaphyses in hamathecium. (g–i) Asci. (j–m) Ascospores. (n) Colony morphology on PDA. Scale bars: (c) = 100 μm; (d,e,g–i) = 50 μm; (f,j–m) = 20 μm.
Figure 4.
Lophiostoma montanae (HMJAU 64835). (a,b) Ascomata developing on natural host substrate. (c) Longitudinal section through ascomata. (d) Ostiole. (e) Partial peridium wall. (f) Anastomosing pseudoparaphyses in hamathecium. (g–i) Asci. (j–m) Ascospores. (n) Colony morphology on PDA. Scale bars: (c) = 100 μm; (d,e,g–i) = 50 μm; (f,j–m) = 20 μm.

3.2.3. Lophiostoma multiforme R. Xu and Y. Li, sp. nov. (Figure 5)
MycoBank number: 858874
Etymology: referring to its multiform of ascospores.
Holotype: HMJAU 64837.
Description: Saprobic on dead stems of Xanthoceras sorbifolium. Sexual morph: Ascomata 223–352 × 187–351 μm ( = 226 × 259 μm, n = 5), black, solitary, superficial, scattered or aggregated in small groups, globose to subglobose, coriaceous. Ostioles 80–100 μm × 20–30 μm ( = 90 × 25.4 μm, n = 5), slit-like, central. Peridium wider at the apex, 31–65 μm ( = 43 μm, n = 20), attenuating toward the base, distinctly layered, comprising 6–8 cell layers; outer layers composed of reddish brown to dark brown, thick-walled cells of textura angularis, gradually transitioning inward to paler, more elongated cells of textura prismatica. Hamathecium composed of filamentous, densely branched, septate pseudoparaphyses, 1–1.7 μm wide, embedded in a gelatinous matrix, extending between and surpassing the asci. Asci 51–102 × 8–12 μm ( = 81.9 × 10 μm, n = 30), bitunicate, fissitunicate, 8-spored, cylindrical to clavate, apex rounded with a distinct ocular chamber, base tapering to a short furcate pedicel. Ascospores 16–23 × 4–7 μm ( = 20.1 × 5.5 μm, n = 50), 1–2-seriate, ellipsoidal to fusiform, gradually tapering toward both ends, (0-)1–2 transversely septa, initially hyaline, becoming dark brown at maturity, guttulate, smooth-walled, surrounded by a distinct mucilaginous sheath. Asexual morph: Undetermined.
Culture characteristics: Colonies on PDA attaining 30 mm diam after 10 days at 25 °C, circular with undulate margins, flat, moderately dense, greenish brown at the center, transitioning gradually to cream-colored at the periphery, with a distinct radiate pattern.
Material examined: CHINA. Hebei Province: Handan city, Qiu county, forest farm in Liangerzhuang Town, on dead stem of Xanthoceras sorbifolium, 9 August 2022, Rong Xu, XR98.1 (HMJAU 64837, holotype), ex-type living culture CCMJ 13069; XR98.2 (HMJAU 648838, isotype), ex-isotype living culture CCMJ 13070.
GenBank numbers: CCMJ 13069: ITS = PP151705, LSU = PP153460, SSU = PV569761, tef1-α = PV670043, rpb2 = PV670041. CCMJ 13070: ITS = PP151706, LSU = PP153461, SSU = PV569762, tef1-α = PV670044, rpb2 = PV670042.
Notes: The new collections fit well with the generic concept of Lophiostoma in having crest-like apices ascomata. In the phylogenetic analyses, two strains of L. multiforme (CCMJ 13069 and CCMJ 13070) are distinct from extant species in Lophiostomataceae and clustered with the strain of L. heterospora (AFTOL-ID 1036) with 88% ML/0.99 BPP support (Figure 2). A BLASTn analysis in GenBank revealed that the ITS sequence of CCMJ 13069 showed the highest similarity (92.77%) to L. compressum (MAL02, MW759267), while its LSU sequence most closely matched Guttulispora crataegi (MFLUCC 13-0442, NG_059563) with 97.87% similarity.
Lophiostoma multiforme differs from L. heterospora in having smaller ascomata (223–352 × 187–351 vs. 275–440 × 220–275 μm), smaller asci (51–102 × 8–12 vs. 70–110 × 15–23 μm), smaller ascospores (16–23 × 4–7 vs. 27–35 × 5–6 μm), and less septate (0–2 transversely septa vs. 8–10 transversely septa) [43]. Additionally, the ascospores of L. heterosporum are hyaline, while those of L. multiforme are initially hyaline and become dark brown at maturity. Based on morphological characteristics and multi-locus phylogenetic analyses, we propose Lophiostoma multiforme as a novel species.
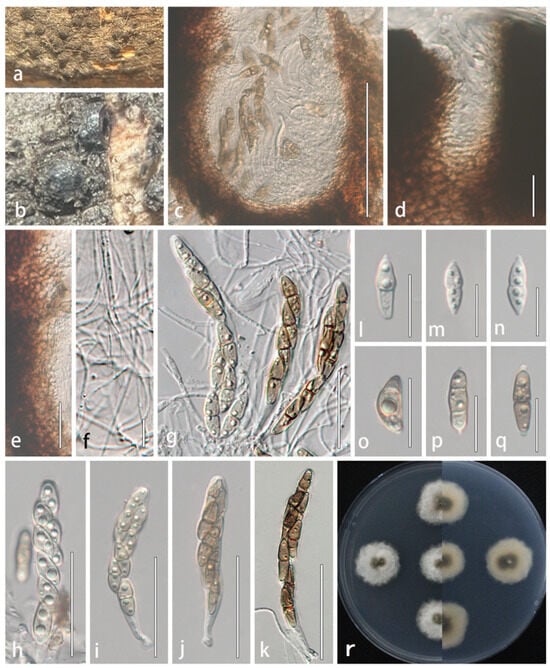
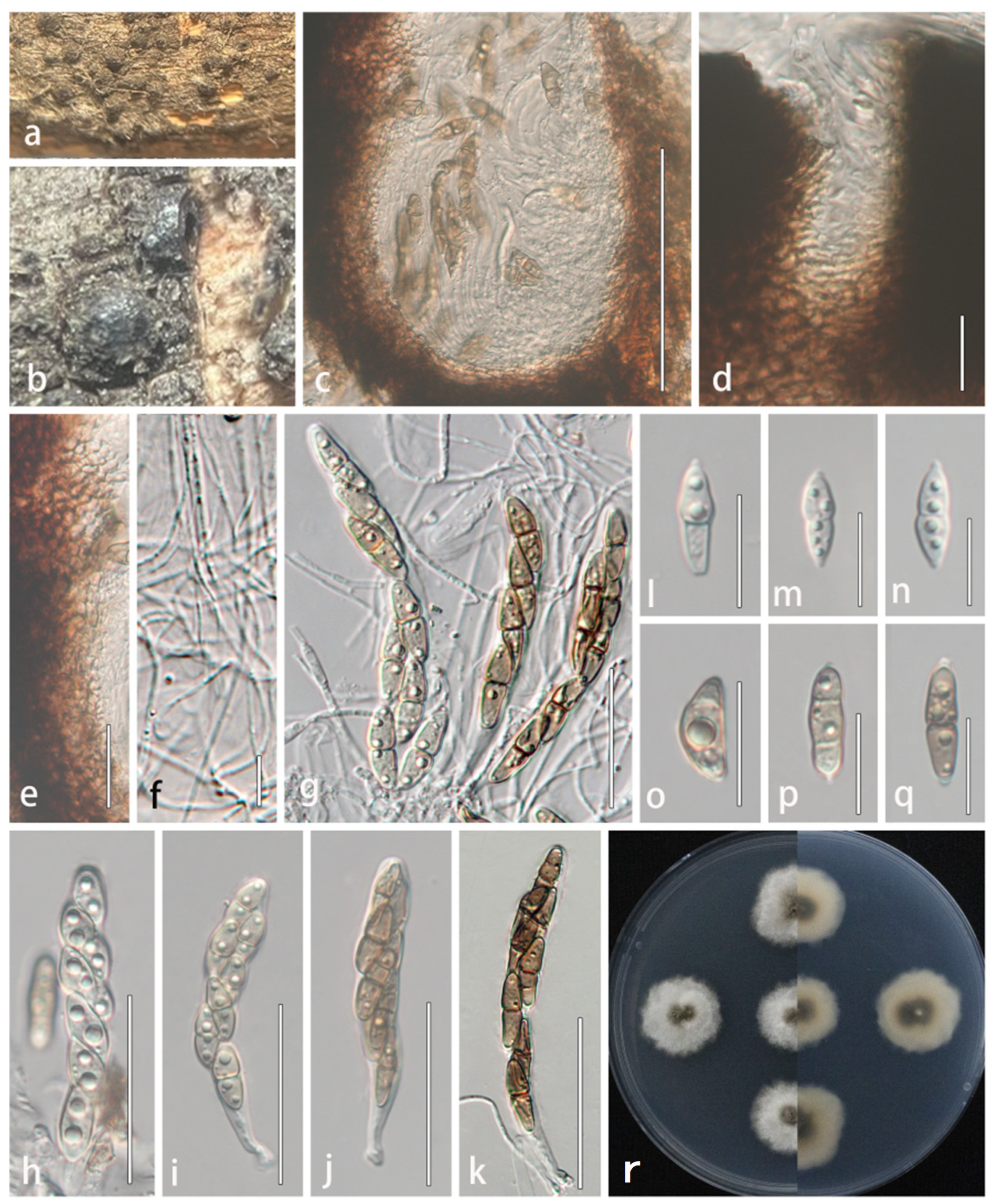
Figure 5.
Lophiostoma multiforme (HMJAU 64837, holotype). (a,b) Ascomata developing on natural host substrate. (c) Longitudinal section through ascomata. (d) Ostiole. (e) Partial peridium wall. (f) Anastomosing pseudoparaphyses in hamathecium. (g–k) Asci. (l–q) Ascospores. (r) Colony morphology on PDA. Scale bars: (c) = 100 μm; (d–e,l–q) = 20 μm; (f) = 10 μm; (g–k) = 50 μm.
Figure 5.
Lophiostoma multiforme (HMJAU 64837, holotype). (a,b) Ascomata developing on natural host substrate. (c) Longitudinal section through ascomata. (d) Ostiole. (e) Partial peridium wall. (f) Anastomosing pseudoparaphyses in hamathecium. (g–k) Asci. (l–q) Ascospores. (r) Colony morphology on PDA. Scale bars: (c) = 100 μm; (d–e,l–q) = 20 μm; (f) = 10 μm; (g–k) = 50 μm.
4. Discussion
Pleosporales was formally defined as an order by Luttrell and Barr (1987) [9]. Due to the polyphyly of traditional morphological groupings, the order has undergone extensive reorganization and significant changes in circumscription [9]. Recent taxonomic studies on Pleosporales have significantly refined its classification through integrative approaches combining morphology, multi-locus phylogenetics, and phylogenomic analyses [7,8,44,45,46]. The application of high-throughput sequencing is helpful for the classification and identification of controversial species [45,47]. Despite these advances, the taxonomic delineation of many species remains unresolved. A robust framework integrating evolutionary genomics and functional ecology is imperative to clarify phylogenetic relationships and ecological diversification within this economically and ecologically pivotal fungal clade.
Alloleptosphaeria was established by Ariyawansa et al. [48] to include the single species A. italica, isolated from dead stems of Clematis vitalba in Italy. This genus species is characterized by semi-immersed to erumpent ascomata, papillate ostiole, reddish brown to dark brown pseudoparenchymatous cells present in the thin-walled peridium, septate, cellular pseudoparaphyses, cylindric-clavate asci, and hyaline to brown ascospores that have transverse or longitudinal septa or have transverse and longitudinal septa together [39,40,42,48,49]. The asexual morph has been documented only from A. iridicola [39]. Currently, there are four epithets (Alloleptosphaeria clematidis Phukhams. and K.D. Hyde; A. iridicola (Crous and Denman) Voglmayr; A. italica Wanas., Camporesi, Ariyaw. and K.D. Hyde; and A. shangrilana Thiyagaraja, Tennakoon and K.D. Hyde) listed in Species Fungorum (2022) under this genus. In the present study, Alloleptosphaeria xanthoceratis groups with other Alloleptosphaeria species with strong support (Figure 1) and differs from known asexual morphs in the genus by its larger conidiogenous cells (9.8–22 × 1.8–3 µm vs. 4–7 × 4–6 µm). Both asexual morph species of Alloleptosphaeria were discovered in temperate regions [39], suggesting these areas may be favorable habitats for asexual members of this genus.
The genus Lophiostoma (type genus of Lophiostomataceae) was established by Cesati and De Notaris (1863), with L. macrostomum designated as its type species [8,50]. Species of Lophiostoma are characterized by laterally compressed or crest-like ascomatal apices, typically occurring as saprobes on decaying plant material in both aquatic and terrestrial environments [41,51,52]. Lophiostoma has been subjected to a number of revisions since its introduction [50,51,52,53]. Andreasen et al. [41] proposed the synonymization of 14 genera with Lophiostoma based on multi-gene phylogenetic analysis using ITS, LSU, tef1-α, and rpb2 markers. A new species, Lophiostoma multiforme sp. nov., and a new record of L. montanae are described in this study based on their morphological characteristics and phylogenetic analysis. Lophiostoma montanae has been documented in Yunnan and Sichuan Provinces, China, associated with Clematis montana and Paeonia suffruticosa, respectively [14,42]. Our isolate was isolated from Xanthoceras sorbifolium in Jilin Province. Subtle variations in morphology were observed among these strains, suggesting that host substrates may influence fungal development. The newly identified strain contributes additional molecular data to the genus Lophiostoma.
Current estimates suggest 2.2–3.8 million fungal species exist worldwide, yet approximately 150,000 (3.5–7%) are formally described [1,54,55]. As the largest Ascomycota order, Pleosporales has a global distribution [7,8,9]. However, studies of microfungi in X. sorbifolium are very scattered, and there are a lot of fungal species that remain to be discovered [19,20,21,22]. Our results emphasize that pleosporalean fungi associated with X. sorbifolium are yet to be properly studied. Further research integrating multi-gene phylogenetic analysis, morphological characterization, and genomic study is essential to elucidate the diversity of microfungi associated with X. sorbifolium and their co-evolutionary interactions.
Supplementary Materials
The following are available online at https://www.mdpi.com/article/10.3390/microorganisms13061296/s1, Figure S1: Phylogram of Leptosphaeriaceae generated from maximum likelihood analysis based on combined ITS, LSU, SSU, and tub2 sequence data. Figure S2: Phylogram of Lophiostomataceae generated from maximum likelihood analysis based on combined ITS, LSU, SSU, tef1-α, and rpb2 sequence data.
Author Contributions
Conceptualization, Y.L.; writing—original draft and formal analysis, R.X.; data curation, R.X.; investigation, R.X.; methodology, R.X.; supervision, Y.L.; writing—review and editing, R.X. and Y.L.; funding acquisition, Y.L. and R.X. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Postdoctor Cultivation Project, Yangzhou University, grant number 137070894, and the Program of Creation and Utilization of Germplasm of Mushroom Crop of “111” Project (no. D17014).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
All sequences generated in this study were submitted to GenBank.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Bánki, O.; Roskov, Y.; Döring, M.; Ower, G.; Vandepitte, L.; Hobern, D.; Remsen, D.; Schalk, P.; DeWalt, R.E.; Keping, M.; et al. Catalogue of Life Checklist (Version 2022-01-14); Catalogue of Life: Leiden, NL, USA, 2022. [Google Scholar]
- Nagy, L.G.; Kovács, G.M.; Krizsán, K. Repeated Evolution of Complex Multicellularity in Fungi. Curr. Biol. 2018, 24, 2384–2394. [Google Scholar]
- Schoch, C.L.; Sung, G.H.; López-Giráldez, F.; Townsend, J.P.; Miadlikowska, J.; Hofstetter, V.; Robbertse, B.; Brandon Matheny, P.; Kauff, F.; Wang, Z.; et al. The Ascomycota tree of life: A phylum-wide phylogeny clarifies the origin and evolution of fundamental reproductive and ecological traits. Syst. Biol. 2009, 58, 224–239. [Google Scholar] [CrossRef] [PubMed]
- Esser, K. The Mycota: A Comprehensive Treatise on Fungi as Experimental Systems for Basic and Applied Research; Springer: Berlin/Heidelberg, Germany, 2001. [Google Scholar]
- Hyde, K.D.; Xu, J.; Rapior, S.; Jeewon, R.; Lumyong, S.; Niego, A.G.T.; Abeywickrama, P.D.; Aluthmuhandiram, J.V.; Brahamanage, R.S.; Brooks, S.; et al. The amazing potential of fungi: 50 ways we can exploit fungi industrially. Fungal Divers. 2019, 97, 1–136. [Google Scholar]
- Hyde, K.D.; Noordeloos, M.E.; Savchenko, K.G.; Jeewon, R.; Lumyong, S.; Niego, A.G.T.; Abeywickrama, P.D.; Aluthmuhandiram, J.V.S.; Brahamanage, R.S.; Brooks, S.; et al. The 2024 Outline of Fungi and fungus-like taxa. Mycosphere 2024, 15, 5146–6239. [Google Scholar] [CrossRef]
- Hongsanan, S.; Hyde, K.D.; Phookamsak, R.; Wanasinghe, D.N.; McKenzie, E.H.C.; Sarma, V.V.; Boonmee, S.; Lücking, R.; Pem, D.; Bhat, J.D.; et al. Refned families of Dothideomycetes: Dothideomycetidae and Pleosporomycetidae. Mycosphere 2020, 11, 1553–2107. [Google Scholar] [CrossRef]
- Hyde, K.D.; Jones, E.B.G.; Liu, J.K.; Ariyawansa, H.; Boehm, E.; Boonmee, S.; Braun, U.; Chomnunti, P.; Crous, P.W.; Dai, D.Q.; et al. Families of Dothideomycetes. Fungal Divers. 2013, 63, 1–313. [Google Scholar]
- Zhang, Y.; Crous, P.W.; Schoch, C.L.; Hyde, K.D. Pleosporales. Fungal Divers. 2012, 53, 1–221. [Google Scholar] [CrossRef]
- Ariyawansa, H.A.; Thambugala, K.M.; Manamgoda, D.S.; Jayawardena, R.; Camporesi, E.; Boonmee, S.; Wanasinghe, D.N.; Phookamsak, R.; Hongsanan, S.; Singtripop, C.; et al. Towards a natural classifcation and backbone tree for Pleosporaceae. Fungal Divers. 2015, 71, 85–139. [Google Scholar] [CrossRef]
- Zhang, Y.; Schoch, C.L.; Fournier, J.; Crous, P.W.; de Gruyter, J.; Woudenberg, J.H.; Hirayama, K.; Tanaka, K.; Pointing, S.B.; Spatafora, J.W.; et al. Multi-locus phylogeny of Pleosporales: A taxonomic, ecological and evolutionary re-evaluation. Stud. Mycol. 2009, 64, 85–102. [Google Scholar] [CrossRef]
- Wijayawardene, N.N.; Hyde, K.D.; Dai, D.Q.; Sánchez-García, M.; Goto, B.T.; Saxena, R.K.; Erdoğdu, M.; Selçuk, F.; Rajeshkumar, K.C.; Aptroot, A.; et al. Outline of Fungi and fungus-like taxa-2021. Mycosphere 2022, 13, 53–453. [Google Scholar] [CrossRef]
- Li, W.L.; Liang, R.R.; Dissanayake, A.J.; Liu, J.K. Botryosphaerialean fungi associated with woody oil plants cultivated in Sichuan Province, China. MycoKeys 2023, 97, 71–116. [Google Scholar] [CrossRef] [PubMed]
- Li, W.L.; Liang, R.R.; Dissanayake, A.J.; Liu, J.K. Mycosphere Notes 413–448: Dothideomycetes associated with woody oil plants in China. Mycosphere 2023, 14, 1436–1529. [Google Scholar] [CrossRef]
- Li, W.L.; Dissanayake, A.J.; Zhang, T.; Maharachchikumbura, S.S.N.; Liu, J.K. Identification and pathogenicity of Pestalotiod fungi associated with woody oil plants in Sichuan Province, China. J. Fungi 2022, 8, 1175. [Google Scholar] [CrossRef]
- Liang, Q.; Li, H.Y.; Li, S.K.; Yuan, F.L.; Sun, J.F.; Duan, Q.C.; Li, Q.Y.; Zhang, R.; Sang, Y.L.; Wang, N.; et al. The genome assembly and annotation of yellowhorn (Xanthoceras sorbifolium Bunge). GigaScience 2019, 8, giz071. [Google Scholar] [CrossRef]
- Zhao, Z.; Liang, C.J.; Zhang, W.; Yang, Y.Y.; Bi, Q.X.; Yu, H.Y.; Wang, L.B. Genome-wide association analysis identifies a candidate gene controlling seed size and yield in Xanthoceras sorbifolium Bunge. Hortic. Res. 2024, 11, 243. [Google Scholar] [CrossRef]
- Zang, E.H.; Qiu, B.; Chen, N.H.; Li, C.F.; Liu, Q.; Zhang, M.; Liu, Y.C.; Li, M.H. Xanthoceras sorbifolium Bunge: A review on botany, phytochemistry, pharmacology, and applications. Front. Pharmacol. 2021, 12, 708549. [Google Scholar] [CrossRef]
- Xu, R.; Su, W.; Tian, S.; Bhunjun, C.S.; Tibpromma, S.; Hyde, K.D.; Li, Y.; Phukhamsakda, C. Synopsis of Leptosphaeriaceae and introduction of three new taxa and one new record from China. J. Fungi 2022, 8, 416. [Google Scholar] [CrossRef]
- Xu, R.; Su, W.; Wang, Y.; Tian, S.; Li, Y.; Phukhamsakda, C. Morphological characteristics and phylogenetic evidence reveal two new species and the first report of Comoclathris (Pleosporaceae, Pleosporales) on dicotyledonous plants from China. MycoKeys 2024, 101, 95–112. [Google Scholar] [CrossRef] [PubMed]
- Xu, R.; Su, W.; Li, Y.; Li, X.; Li, C. Multiple evidence reveals a new species of Neocucurbitaria (Cucurbitariaceae, Pleosporales) from China. Phytotaxa 2024, 650, 249–261. [Google Scholar] [CrossRef]
- Yang, G. Study on Main Diseases and insect pests and control techniques in Xanthoceras sorbifolia Bunge. Hortic. Seed 2022, 42, 37–39. [Google Scholar]
- Senanayake, I.C.; Rathnayaka, A.R.; Marasinghe, D.S.; Calabon, M.S.; Gentekaki, E.; Lee, H.B.; Hurdeal, V.G.; Pem, D.; Dissanayake, L.S.; Wijesinghe, S.N.; et al. Morphological approaches in studying fungi: Collection, examination, isolation, sporulation and preservation. Mycosphere 2020, 11, 2678–2754. [Google Scholar] [CrossRef]
- Crous, P.W.; Gams, W.; Stalpers, J.A.; Robert, V.; Stegehuis, G. MycoBank: An online initiative to launch mycology into the 21st century. Stud. Mycol. 2004, 50, 19–22. [Google Scholar]
- Benson, D.A.; Cavanaugh, M.; Clark, K.; Karsch-Mizrachi, I.; Lipman, D.J.; Ostell, J.; Sayers, E.W. GenBank. Nucleic Acids Res. 2013, 41, D36–D42. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.D.; Lee, S.B.; Taylor, J.W. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protoc. A Guid. Methods Appl. 1990, 18, 315–322. [Google Scholar]
- Vilgalys, R.; Hester, M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J. Bacteriol. 1990, 172, 4238–4246. [Google Scholar] [CrossRef] [PubMed]
- Rehner, S.A.; Buckley, E.A. Beauveria phylogeny inferred from nuclear ITS and EF-1α sequences: Evidence for cryptic diversifcation and links to Cordyceps teleomorphs. Mycologia 2005, 97, 84–98. [Google Scholar]
- Liu, Y.J.; Whelen, S.; Hall, B.D. Phylogenetic relationships among ascomycetes: Evidence from an RNA polymerase II subunit. Mol. Biol. Evol. 1999, 16, 1799–1808. [Google Scholar] [CrossRef]
- Glass, N.L.; Donaldson, G.C. Development of primer sets designed for PCR to amplify conserved genes from filamentous Ascomycetes. Appl. Environ. Microb. 1995, 61, 1323–1330. [Google Scholar] [CrossRef]
- McGinnis, S.; Madden, T.L. BLAST: At the core of a powerful and diverse set of sequence analysis tools. Nucleic Acids Res. 2004, 32, W20–W25. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Larsson, A. AliView: A fast and lightweight alignment viewer and editor for large datasets. Bioinformatics 2014, 30, 3276–3278. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- Nylander, J.A.A. MrModeltest, 2.0; Program Distributed by the Author; Uppsala University: Uppsala, Sweden, 2004. [Google Scholar]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef]
- Rambaut, A. FigTree, v.1.4.4; UnSiversity of Edinburgh: Edinburgh, UK, 2018. [Google Scholar]
- Crous, P.W.; Schumacher, R.K.; Wingfield, M.J.; Akulov, A.; Denman, S.; Roux, J.; Braun, U.; Burgess, T.I.; Carnegie, A.J.; Váczy, K.Z.; et al. New and Interesting Fungi. 1. Fungal Syst. Evol. 2018, 1, 169–216. [Google Scholar] [CrossRef] [PubMed]
- Aiello, D.; Vitale, A.; Polizzi, G.; Voglmayr, H. Ochraceocephala foeniculi gen. et sp. nov., a new pathogen causing crown rot of fennel in Italy. MycoKeys 2020, 66, 1–22. [Google Scholar] [CrossRef]
- Andreasen, M.; Skrede, I.; Jaklitsch, W.M.; Voglmayr, H.; Nordén, B. Multi-locus phylogenetic analysis of lophiostomatoid fungi motivates a broad concept of Lophiostoma and reveals nine new species. Persoonia 2021, 46, 240–271. [Google Scholar] [CrossRef]
- Phukhamsakda, C.; McKenzie, E.H.C.; Phillips, A.J.L.; Jones, E.B.G.; Bhat, D.J.; Stadler, M.; Bhunjun, C.S.; Wanasinghe, D.N.; Thongbai, B.; Camporesi, E.; et al. Microfungi associated with Clematis (Ranunculaceae) with an integrated approach to delimiting species boundaries. Fungal Divers. 2020, 102, 1–203. [Google Scholar] [CrossRef]
- Saccardo, P.A. Sylloge Fungorum. II; Biodiversity Heritage Library: Washington, DC, USA, 1883. [Google Scholar]
- Liu, F.; Ma, Z.Y.; Hou, L.W.; Diao, Y.Z.; Wu, W.P.; Damm, U.; Song, S.; Cai, L. Updating species diveristy of Colletotrichum, with a phylogenomic overview. Stud. Mycol. 2022, 101, 1–56. [Google Scholar] [CrossRef]
- Han, S.L.; Wang, M.M.; Ma, Z.Y.; Raza, M.; Zhao, P.; Liang, J.M.; Gao, M.; Li, Y.J.; Wang, J.W.; Hu, D.M.; et al. Fusarium diversity associated with diseased cereals in China, with an updated phylogenomic assessment of the genus. Stud. Mycol. 2023, 104, 87–148. [Google Scholar] [CrossRef]
- Razaghi, P.; Raza, M.; Han, L.S.; Ma, Z.Y.; Cai, L.; Zhao, P.; Chen, Q.; Phurbu, D.; Liu, F. Sporocadaceae revisited. Stud. Mycol. 2024, 109, 155–272. [Google Scholar] [CrossRef]
- Steenwyk, L.J.; Balamurugan, C.; Raja, A.H.; Gonçalves, C.; Li, N.X.; Martin, F.; Berman, J.; Oberlies, N.H.; Gibbons, J.G.; Goldman, G.H. Phylogenomics reveals extensive misidentification of fungal strains from the genus Aspergillus. Microbiol. Spectr. 2024, 12, e0398023. [Google Scholar] [CrossRef] [PubMed]
- Ariyawansa, H.A.; Phukhamsakda, C.; Thambugala, K.M.; Bulgakov, T.S.; Wanasinghe, D.N.; Perera, R.H.; Mapook, A.; Camporesi, E.; Kang, J.C.; Jones, E.B.G.; et al. Revision and phylogeny of Leptosphaeriaceae. Fungal Divers. 2015, 74, 19–51. [Google Scholar] [CrossRef]
- Thiyagaraja, V.; Wanasinghe, D.N.; Karunarathna, S.C.; Tennakoon, D.S.; Hyde, K.D.; To-anun, C.; Cheewangkoon, R. Alloleptosphaeria shangrilana sp. nov. and first report of the genus (Leptosphaeriaceae, Dothideomycetes) from China. Phytotaxa 2021, 491, 12–22. [Google Scholar] [CrossRef]
- Tanaka, K.; Harada, Y. Pleosporales in Japan (1): The genus Lophiostoma. Mycoscience 2003, 44, 85–96. [Google Scholar] [CrossRef]
- Hirayama, K.; Tanaka, K. Taxonomic revision of Lophiostoma and Lophiotrema based on reevaluation of morphological characters and molecular analyses. Mycoscience 2011, 52, 401–412. [Google Scholar] [CrossRef]
- Thambugala, K.M.; Hyde, K.D.; Tanaka, K.; Tian, Q.; Wanasinghe, D.N.; Ariyawansa, H.A.; Jayasiri, S.C. Towards a natural classification and backbone tree for Lophiostomataceae, Floricolaceae, and Amorosiaceae fam. nov. Fungal Divers. 2015, 74, 199–266. [Google Scholar] [CrossRef]
- Aluthmuhandiram, J.V.; Wanasinghe, D.N.; Thilini Chethana, K.W.; Gafforov, Y.; Saichana, N.; Li, X.; Yan, J.; Mamarakhimov, O.M. Lophiostomataceae (Dothideomycetes): Introducing Lophiostoma khanzada-kirgizbaeva sp. nov. and Paucispora xishanensis sp. nov. Phytotaxa 2022, 559, 247–262. [Google Scholar] [CrossRef]
- Hawksworth, D.L.; Lücking, R. Fungal Diversity Revisited: 2.2 to 3.8 Million Species. Microbiol. Spectr. 2017, 5, 1–17. [Google Scholar] [CrossRef]
- Wanasinghe, D.N.; Mortimer, P.E.; Bezerra, J. Editorial: Fungal Systematics and Biogeography. Front. Microbiol. 2022, 12, 827725. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
