Complete Biodegradation of Diclofenac by New Bacterial Strains: Postulated Pathways and Degrading Enzymes
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection and Isolation of Local Diclofenac-Degrading Bacteria
2.2. Optimization of Bacterial Isolate Growth under Different Physicochemical Conditions
2.3. Biochemical Characterization and Molecular Identification of the Selected Bacterial Isolates
2.4. Quantification of Diclofenac Biodegradation
2.5. Detection of Diclofenac Metabolites Produced by Individual Bacterial Isolates Using Gas Chromatography–Mass Spectrometry (GC–MS) Analysis
2.6. Enzyme Activities
2.7. Statistical Analysis
3. Results
3.1. Screening for Local Diclofenac-Degrading Bacteria
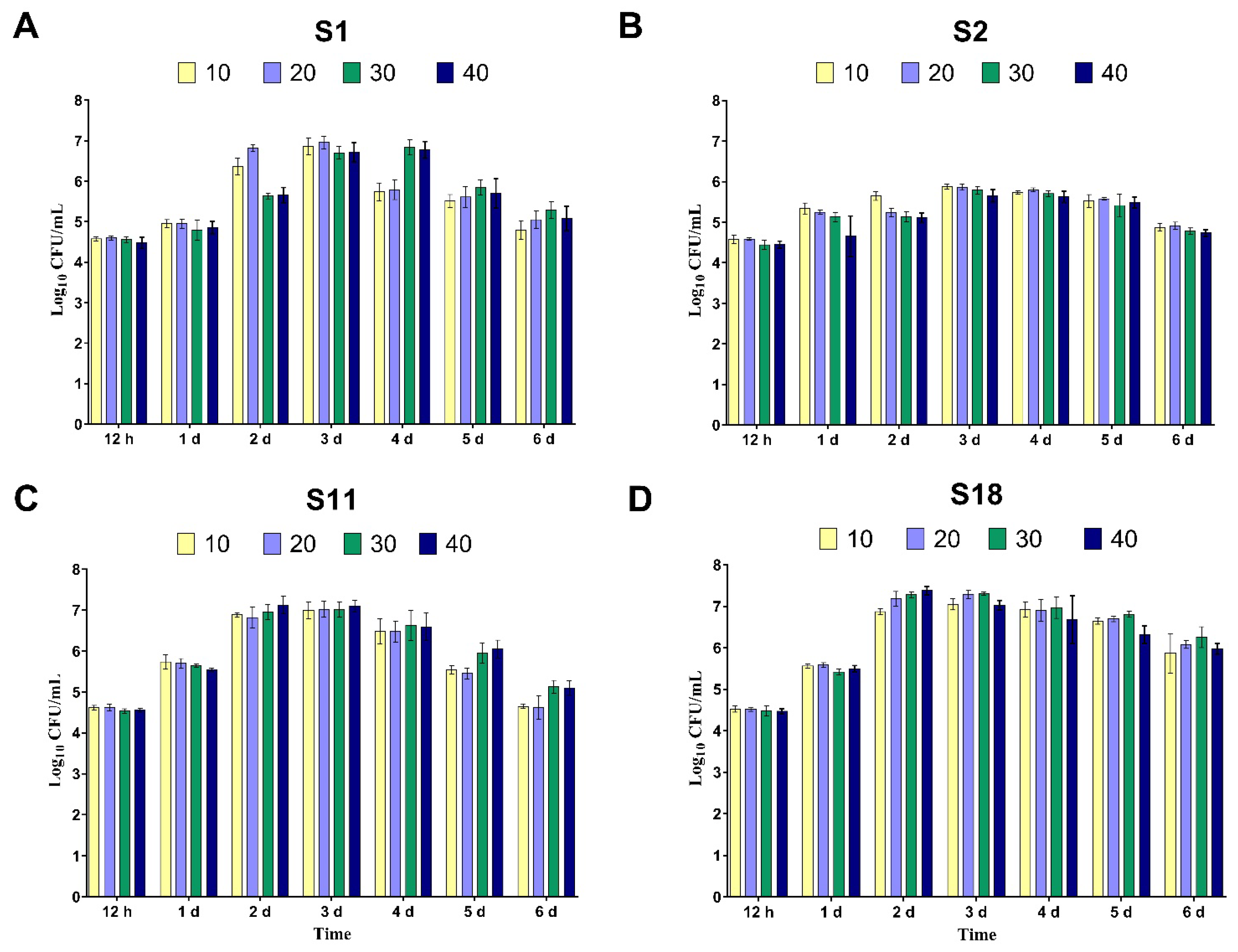
3.2. Ideal Growth Parameters for Diclofenac-Degrading Bacteria
3.3. Removal Efficiency of Isolated Bacterial Strains
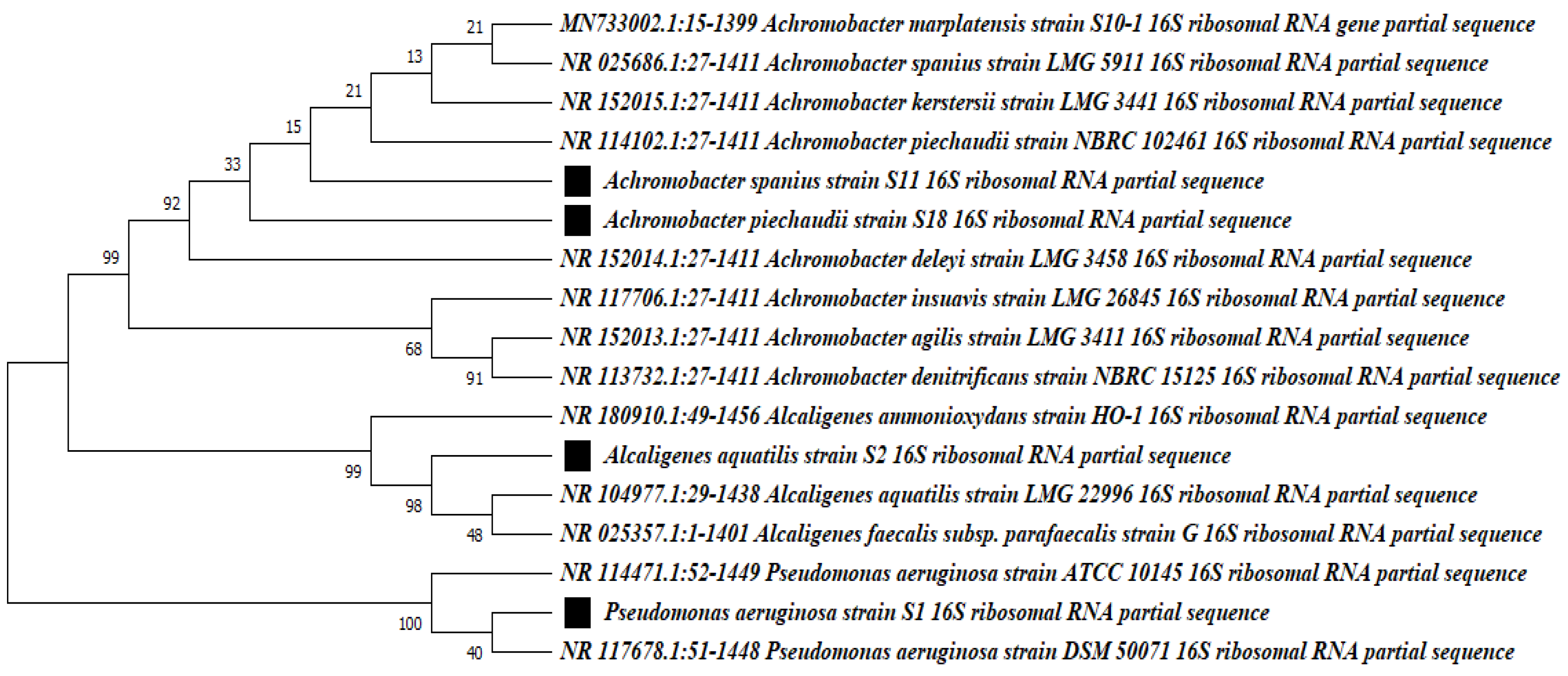
3.4. Identification of DCF-Degrading Bacterial Isolates
3.5. Detection of DCF Biodegradation Intermediate Compounds Produced by Each Bacterial Strain
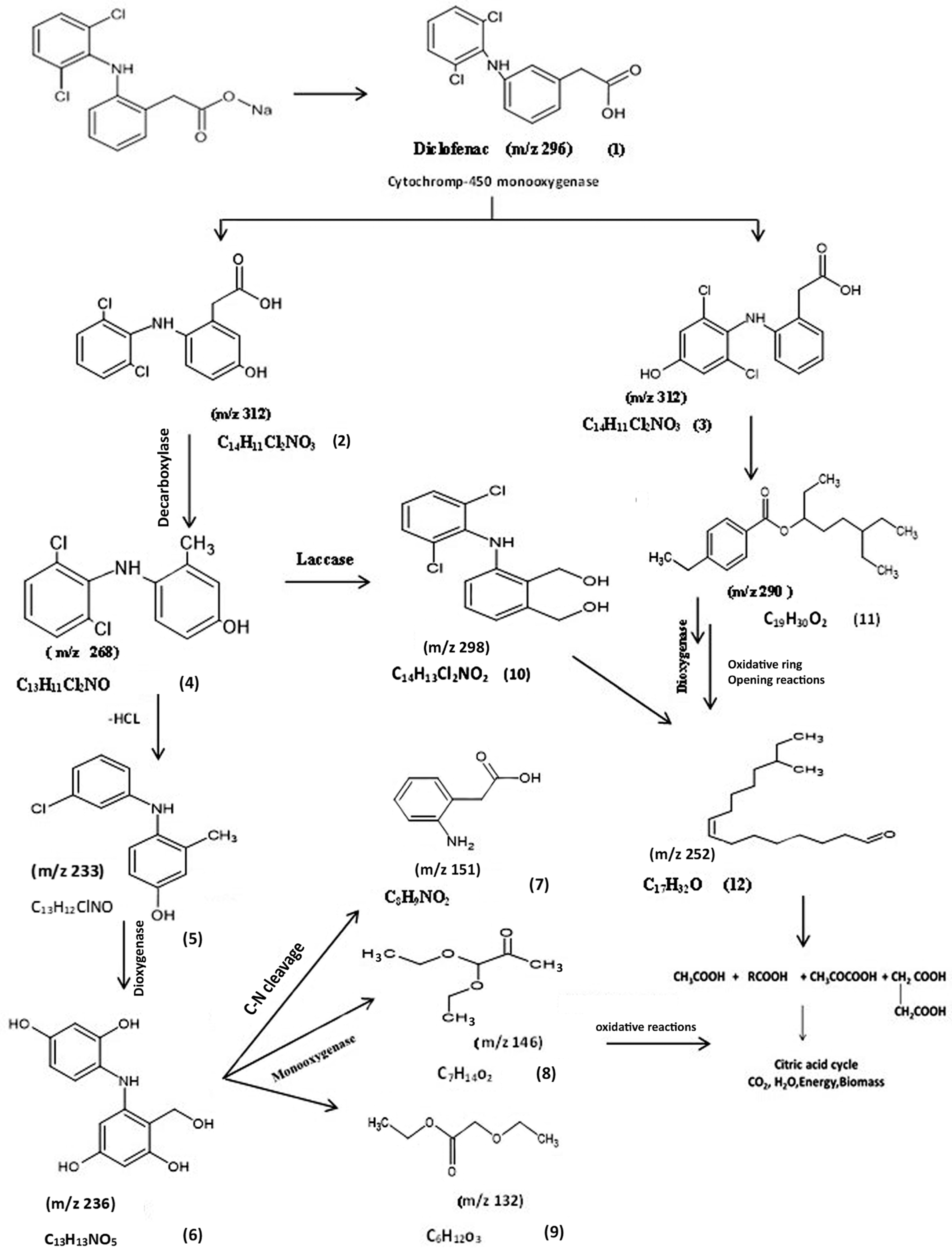
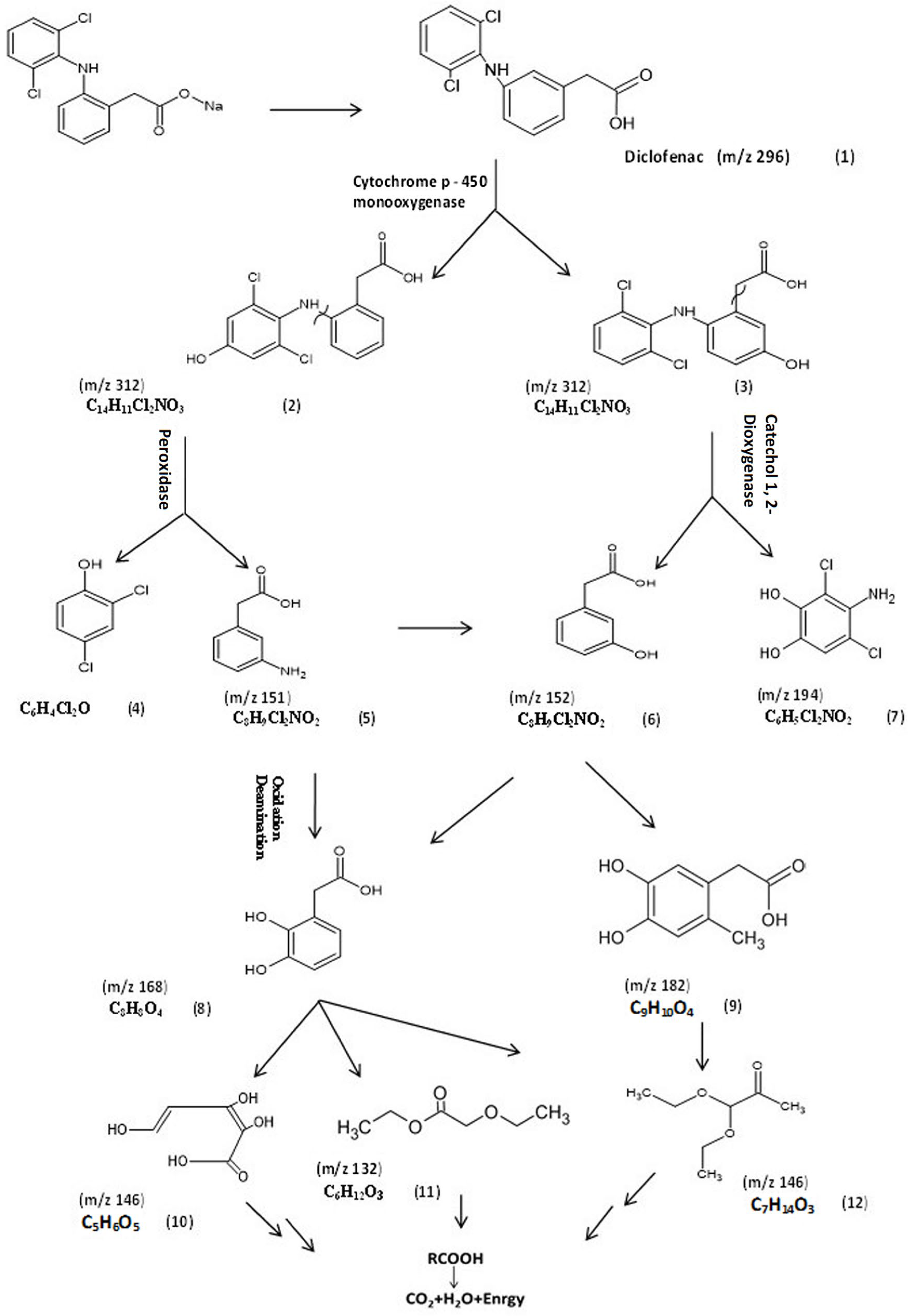
3.5.1. Identification of Intermediate DCF Products of A. spanius S11 and A. piechaudii S18
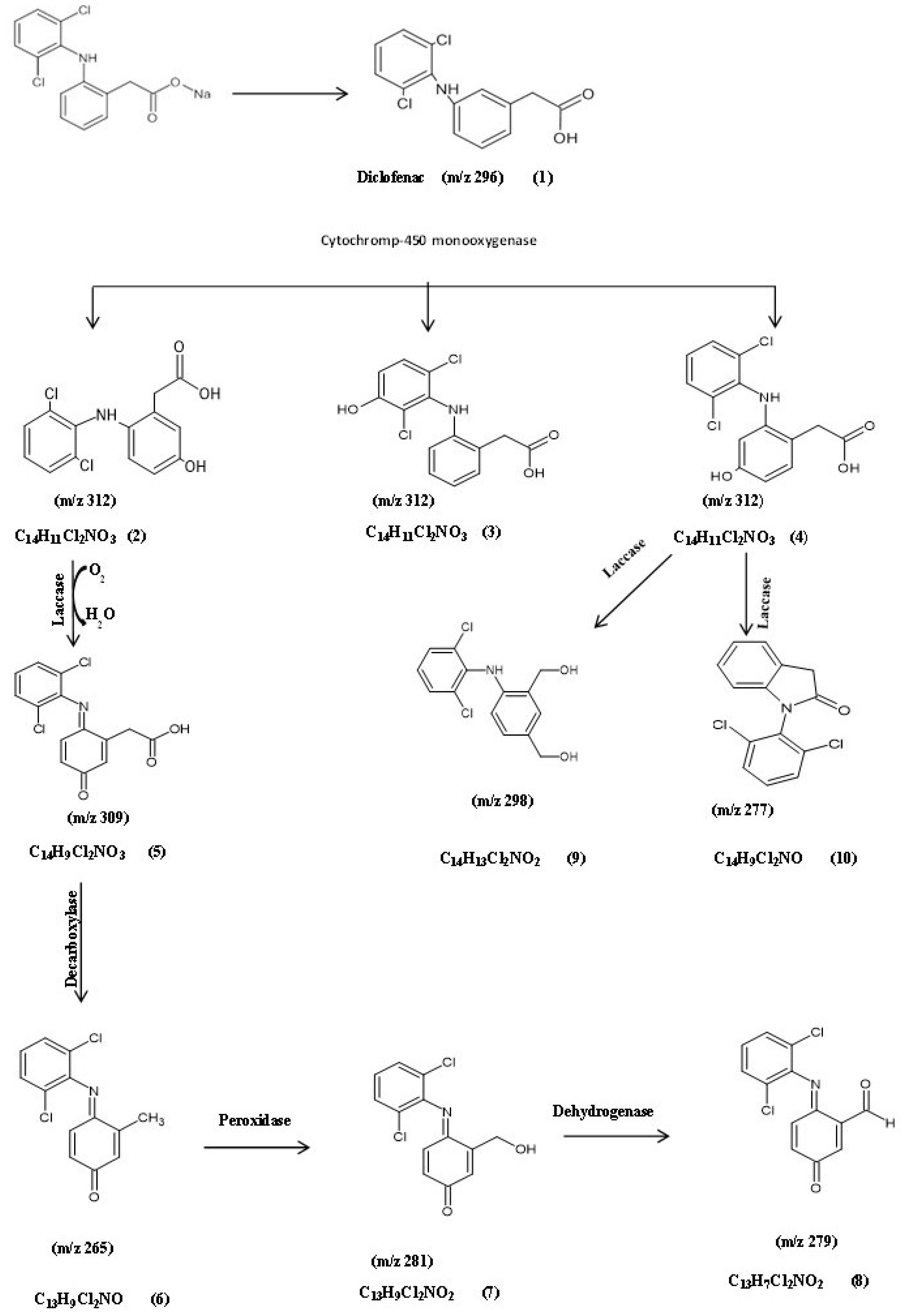
3.5.2. Identification of Intermediate DCF Products of P. aeruginosa S1
3.6. Specific Activities of Enzymes Involved in DCF Degradation
4. Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Luo, Y.; Guo, W.; Ngo, H.H.; Nghiem, L.D.; Hai, F.I.; Zhang, J.; Liang, S.; Wang, X.C. A Review on the Occurrence of Micropollutants in the Aquatic Environment and Their Fate and Removal during Wastewater Treatment. Sci. Total Environ. 2014, 473–474, 619–641. [Google Scholar] [CrossRef] [PubMed]
- Domaradzka, D.; Guzik, U.; Wojcieszyńska, D. Biodegradation and Biotransformation of Polycyclic Non-Steroidal Anti-Inflammatory Drugs. Rev. Environ. Sci. Bio/Technol. 2015, 14, 229–239. [Google Scholar] [CrossRef]
- Parrilla Vázquez, M.M.; Parrilla Vázquez, P.; Martínez Galera, M.; Gil García, M.D.; Uclés, A. Ultrasound-Assisted Ionic Liquid Dispersive Liquid–Liquid Microextraction Coupled with Liquid Chromatography-Quadrupole-Linear Ion Trap-Mass Spectrometry for Simultaneous Analysis of Pharmaceuticals in Wastewaters. J. Chromatogr. A 2013, 1291, 19–26. [Google Scholar] [CrossRef]
- Maeng, S.K.; Sharma, S.K.; Lekkerkerker-Teunissen, K.; Amy, G.L. Occurrence and Fate of Bulk Organic Matter and Pharmaceutically Active Compounds in Managed Aquifer Recharge: A Review. Water Res. 2011, 45, 3015–3033. [Google Scholar] [CrossRef]
- Benotti, M.J.; Trenholm, R.A.; Vanderford, B.J.; Holady, J.C.; Stanford, B.D.; Snyder, S.A. Pharmaceuticals and Endocrine Disrupting Compounds in U.S. Drinking Water. Environ. Sci. Technol. 2009, 43, 597–603. [Google Scholar] [CrossRef] [PubMed]
- Jelic, A.; Gros, M.; Ginebreda, A.; Cespedes-Sánchez, R.; Ventura, F.; Petrovic, M.; Barcelo, D. Occurrence, Partition and Removal of Pharmaceuticals in Sewage Water and Sludge during Wastewater Treatment. Water Res. 2011, 45, 1165–1176. [Google Scholar] [CrossRef]
- Żur, J.; Marchlewicz, A.; Piński, A.; Guzik, U.; Wojcieszyńska, D. Degradation of Diclofenac by New Bacterial Strains and Its Influence on the Physiological Status of Cells. J. Hazard. Mater. 2021, 403, 124000. [Google Scholar] [CrossRef]
- Verlicchi, P.; Al Aukidy, M.; Zambello, E. Occurrence of Pharmaceutical Compounds in Urban Wastewater: Removal, Mass Load and Environmental Risk after a Secondary Treatment-A Review. Sci. Total Environ. 2012, 429, 123–155. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Tenorio, R.; González-Juárez, E.; Guzmán-Mar, J.L.; Hinojosa-Reyes, L.; Hernández-Ramírez, A. Review of Occurrence of Pharmaceuticals Worldwide for Estimating Concentration Ranges in Aquatic Environments at the End of the Last Decade. J. Hazard. Mater. Adv. 2022, 8, 100172. [Google Scholar] [CrossRef]
- Gonzalez-Rey, M.; Tapie, N.; Le Menach, K.; Dévier, M.H.; Budzinski, H.; Bebianno, M.J. Occurrence of Pharmaceutical Compounds and Pesticides in Aquatic Systems. Mar. Pollut. Bull. 2015, 96, 384–400. [Google Scholar] [CrossRef]
- Jiang, J.Q.; Zhou, Z.; Sharma, V.K. Occurrence, Transportation, Monitoring and Treatment of Emerging Micro-Pollutants in Waste Water—A Review from Global Views. Microchem. J. 2013, 110, 292–300. [Google Scholar] [CrossRef]
- Munari, M.; Matozzo, V.; Gagné, F.; Chemello, G.; Riedl, V.; Finos, L.; Pastore, P.; Badocco, D.; Marin, M.G. Does Exposure to Reduced PH and Diclofenac Induce Oxidative Stress in Marine Bivalves? A Comparative Study with the Mussel Mytilus galloprovincialis and the Clam Ruditapes philippinarum. Environ. Pollut. 2018, 240, 925–937. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.N.; Nghiem, L.D.; Pramanik, B.K.; Oh, S. Cometabolic Biotransformation and Impacts of the Anti-Inflammatory Drug Diclofenac on Activated Sludge Microbial Communities. Sci. Total Environ. 2019, 657, 739–745. [Google Scholar] [CrossRef]
- Moreira, I.S.; Bessa, V.S.; Murgolo, S.; Piccirillo, C.; Mascolo, G.; Castro, P.M.L. Biodegradation of Diclofenac by the Bacterial Strain Labrys portucalensis F11. Ecotoxicol. Environ. Saf. 2018, 152, 104–113. [Google Scholar] [CrossRef] [PubMed]
- Murshid, S.; Dhakshinamoorthy, G.P. Biodegradation of Sodium Diclofenac and Mefenamic Acid: Kinetic Studies, Identification of Metabolites and Analysis of Enzyme Activity. Int. Biodeterior. Biodegrad. 2019, 144, 104756. [Google Scholar] [CrossRef]
- Sharma, S.; Setia, H.; Toor, A.P. Assessing the Bioremediation Potential of Indigenously Isolated Klebsiella Sp. WAH1 for Diclofenac Sodium: Optimization, Toxicity and Metabolic Pathway Studies. World J. Microbiol. Biotechnol. 2021, 37, 1–12. [Google Scholar] [CrossRef]
- Yamasato, K.; Akagawa, M.; Oishi, N.; Kuraishi, H. Carbon Substrate Assimilation Profiles and Other Taxonomic Features of Alcaligenes faecalis, Alcaligenes ruhlandii and Achromobacter xylosoxidans. J. Gen. Appl. Microbiol. 1982, 28, 195–213. [Google Scholar] [CrossRef]
- Abdelkader, A.A.; Khalil, M.S.; Mohamed, M.S.M. Simultaneous Biodegradation of λ-Cyhalothrin Pesticide and Vicia faba Growth Promotion under Greenhouse Conditions. AMB Express 2022, 12, 44. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple Sequence Alignment with High Accuracy and High Throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef]
- Ivshina, I.B.; Tyumina, E.A.; Kuzmina, M.V.; Vikhareva, E.V. Features of Diclofenac Biodegradation by Rhodococcus Ruber IEGM 346. Sci. Rep. 2019, 9, 1–13. [Google Scholar] [CrossRef]
- Hatvani, N.; Mécs, I. Production of Laccase and Manganese Peroxidase by Lentinus Edodes on Malt-Containing by-Product of the Brewing Process. Process Biochem. 2001, 37, 491–496. [Google Scholar] [CrossRef]
- Wojcieszyńska, D.; Guzik, U.; Greń, I.; Perkosz, M.; Hupert-Kocurek, K. Induction of Aromatic Ring: Cleavage Dioxygenases in Stenotrophomonas maltophilia Strain KB2 in Cometabolic Systems. World J. Microbiol. Biotechnol. 2011, 27, 805–811. [Google Scholar] [CrossRef]
- Chance, B.; Maehly, A.C. [136] Assay of Catalases and Peroxidases. In Methods in Enzymology; Elsevier: Amsterdam, The Netherlands, 1955; Volume 2, pp. 764–775. [Google Scholar]
- Świacka, K.; Michnowska, A.; Maculewicz, J.; Caban, M.; Smolarz, K. Toxic Effects of NSAIDs in Non-Target Species: A Review from the Perspective of the Aquatic Environment. Environ. Pollut. 2021, 273, 115891. [Google Scholar] [CrossRef] [PubMed]
- Żur, J.; Piński, A.; Wojcieszyńska, D.; Smułek, W.; Guzik, U. Diclofenac Degradation—Enzymes, Genetic Background and Cellular Alterations Triggered in Diclofenac-Metabolizing Strain Pseudomonas Moorei KB4. Int. J. Mol. Sci. 2020, 21, 6786. [Google Scholar] [CrossRef] [PubMed]
- Arisan, E.D.; Akar, R.O.; Rencuzogullari, O.; Obakan Yerlikaya, P.; Coker Gurkan, A.; Akın, B.; Dener, E.; Kayhan, E.; Palavan Unsal, N. The Molecular Targets of Diclofenac Differs from Ibuprofen to Induce Apoptosis and Epithelial Mesenchymal Transition Due to Alternation on Oxidative Stress Management P53 Independently in PC3 Prostate Cancer Cells. Prostate Int. 2019, 7, 156–165. [Google Scholar] [CrossRef]
- Facey, S.J.; Nebel, B.A.; Kontny, L.; Allgaier, M.; Hauer, B. Rapid and Complete Degradation of Diclofenac by Native Soil Microorganisms. Environ. Technol. Innov. 2018, 10, 55–61. [Google Scholar] [CrossRef]
- Stylianou, K.; Hapeshi, E.; Vasquez, M.I.; Fatta-Kassinos, D.; Vyrides, I. Diclofenac Biodegradation by Newly Isolated Klebsiella Sp. KSC: Microbial Intermediates and Ecotoxicological Assessment. J. Environ. Chem. Eng. 2018, 6, 3242–3248. [Google Scholar] [CrossRef]
- Grandclément, C.; Piram, A.; Petit, M.E.; Seyssiecq, I.; Laffont-Schwob, I.; Vanot, G.; Tiliacos, N.; Roche, N.; Doumenq, P. Biological Removal and Fate Assessment of Diclofenac Using Bacillus Subtilis and Brevibacillus laterosporus Strains and Ecotoxicological Effects of Diclofenac and 4′-Hydroxy-Diclofenac. J. Chem. 2020, 2020, 9789420. [Google Scholar] [CrossRef]
- Zhang, X.; Hao, X.; Huo, S.; Lin, W.; Xia, X.; Liu, K.; Duan, B. Isolation and Identification of the Raoultella ornithinolytica-ZK4 Degrading Pyrethroid Pesticides within Soil Sediment from an Abandoned Pesticide Plant. Arch. Microbiol. 2019, 201, 1207–1217. [Google Scholar] [CrossRef]
- Wojcieszyńska, D.; Łagoda, K.; Guzik, U. Diclofenac Biodegradation by Microorganisms and with Immobilised Systems—A Review. Catalysts 2023, 13, 412. [Google Scholar] [CrossRef]
- Liang, D.h.; Hu, Y. Simultaneous Sulfamethoxazole Biodegradation and Nitrogen Conversion by Achromobacter Sp. JL9 Using with Different Carbon and Nitrogen Sources. Bioresour. Technol. 2019, 293, 122061. [Google Scholar] [CrossRef]
- Hong, Y.H.; Ye, C.C.; Zhou, Q.Z.; Wu, X.Y.; Yuan, J.P.; Peng, J.; Deng, H.; Wang, J.H. Genome Sequencing Reveals the Potential of Achromobacter Sp. HZ01 for Bioremediation. Front. Microbiol. 2017, 8, 1507. [Google Scholar] [CrossRef] [PubMed]
- Hahn, V.; Meister, M.; Hussy, S.; Cordes, A.; Enderle, G.; Saningong, A.; Schauer, F. Enhanced Laccase-Mediated Transformation of Diclofenac and Flufenamic Acid in the Presence of Bisphenol A and Testing of an Enzymatic Membrane Reactor. AMB Express 2018, 8, 28. [Google Scholar] [CrossRef] [PubMed]
- Den Braver, M.W.; den Braver-Sewradj, S.P.; Vermeulen, N.P.E.; Commandeur, J.N.M. Characterization of Cytochrome P450 Isoforms Involved in Sequential Two-Step Bioactivation of Diclofenac to Reactive p-Benzoquinone Imines. Toxicol. Lett. 2016, 253, 46–54. [Google Scholar] [CrossRef] [PubMed]
- Marco-Urrea, E.; Pérez-Trujillo, M.; Cruz-Morató, C.; Caminal, G.; Vicent, T. Degradation of the Drug Sodium Diclofenac by Trametes Versicolor Pellets and Identification of Some Intermediates by NMR. J. Hazard. Mater. 2010, 176, 836–842. [Google Scholar] [CrossRef]
- Yusuf, Y. The Utilization of Laccase-Functionalized Graphene Oxide as an Effective Biodegradation of Pharmaceutical Industry Waste: Diclofenac and Ibuprofen. Syst. Rev. Pharm. 2020, 11, 536–544. [Google Scholar] [CrossRef]
- Gao, L.; Zhou, B.; Wang, F.; Yuan, R.; Chen, H.; Han, X. Effect of Dissolved Organic Matters and Inorganic Ions on TiO2 Photocatalysis of Diclofenac: Mechanistic Study and Degradation Pathways. Environ. Sci. Pollut. Res. 2020, 27, 2044–2053. [Google Scholar] [CrossRef]
- Yang, J.; Duan, A.; Wang, J.; Yang, X.; Liu, X.; Xiao, F.; Qin, F.; Yu, Y.; Wang, D. The Fate of Diclofenac in Anaerobic Fermentation of Waste Activated Sludge. J. Environ. Manag. 2022, 322, 116098. [Google Scholar] [CrossRef]
- Lloret, L.; Eibes, G.; Lú-Chau, T.A.; Moreira, M.T.; Feijoo, G.; Lema, J.M. Laccase-Catalyzed Degradation of Anti-Inflammatories and Estrogens. Biochem. Eng. J. 2010, 51, 124–131. [Google Scholar] [CrossRef]
- Tran, N.H.; Urase, T.; Kusakabe, O. Biodegradation Characteristics of Pharmaceutical Substances by Whole Fungal Culture Trametes Versicolor and Its Laccase. J. Water Environ. Technol. 2010, 8, 125–140. [Google Scholar] [CrossRef]
- Jewell, K.S.; Falås, P.; Wick, A.; Joss, A.; Ternes, T.A. Transformation of Diclofenac in Hybrid Biofilm–Activated Sludge Processes. Water Res. 2016, 105, 559–567. [Google Scholar] [CrossRef] [PubMed]
- Fetyan, N.A.H.; Azeiz, A.Z.A.; Ismail, I.M.; Shaban, S.A. Oxidative Decolorization of Direct Blue 71 Azo Dye by Saccharomyces cerevisiae Catalyzed by Nano Zero-Valent Iron. Annu. Res. Rev. Biol. 2016, 11, 1–12. [Google Scholar] [CrossRef]
- Yu, H.; Nie, E.; Xu, J.; Yan, S.; Cooper, W.J.; Song, W. Degradation of Diclofenac by Advanced Oxidation and Reduction Processes: Kinetic Studies, Degradation Pathways and Toxicity Assessments. Water Res. 2013, 47, 1909–1918. [Google Scholar] [CrossRef] [PubMed]
- Fetyan, N.; Azeiz, A.; Ismail, I.; Salem, T. Biodegradation of Cibacron Redazo Dye and Industrial Textile Effluent by Pseudomonas aeruginosa Immobilized on Chitosan-Fe2O3 Composite. J. Adv. Biol. Biotechnol. 2017, 12, 1–15. [Google Scholar] [CrossRef]
- Vijayalakshmi, V.; Senthilkumar, P.; Mophin-Kani, K.; Sivamani, S.; Sivarajasekar, N.; Vasantharaj, S. Bio-Degradation of Bisphenol A by Pseudomonas Aeruginosa PAb1 Isolated from Effluent of Thermal Paper Industry: Kinetic Modeling and Process Optimization. J. Radiat. Res. Appl. Sci. 2018, 11, 56–65. [Google Scholar] [CrossRef]
- Van scnte, P.M.; Young, L.Y. Biodegradation of Phenol: Mechanisms and Applications. Bioremediat. J. 2000, 4, 1–18. [Google Scholar] [CrossRef]
- Ma, Y.L.; Lu, W.; Wan, L.L.; Luo, N. Elucidation of Fluoranthene Degradative Characteristics in a Newly Isolated Achromobacter xylosoxidans DN002. Appl. Biochem. Biotechnol. 2015, 175, 1294–1305. [Google Scholar] [CrossRef]

| Isolates | Degradation of Diclofenac (%) * | |
|---|---|---|
| 3 Days | 6 Days | |
| S1 | 37.76 ± 0.93 c | 75.20 ± 0.71 c |
| S2 | 12.85 ± 1.58 b | 38.99 ± 0.90 b |
| S11 | 86.60 ± 1.19 e | 97.79 ± 0.84 e |
| S18 | 53.15 ±1.55 d | 88.72 ± 0.89 d |
| Control | 2. 29 ± 1.32 a | 3.51 ± 0.87 a |
| No. | Compound Name | RT (min) | Fragments (m/z) | MW | Formula |
|---|---|---|---|---|---|
| 1 | Diclofenac | 12.77 | 249, 277, 295 | 296 | C14H11Cl2N2O2 |
| 2 | 5-Hydroxydiclofenac | 10.93 | 214, 231, 249, 312 | 312 | C14H11Cl2NO3 |
| 3 | 4/-Hydroxydiclofenac | ||||
| 4 | 3-Hydroxydiclofenac | ||||
| 5 | DF-2,5-benzoquinone imine | 13.72 | 242, 279, 309 | 309 | C14H9Cl2NO3 |
| 6 | (6E)-6-[(2,5-dichlorophenyl) imino]-3-oxocyclohexa-1,4-diene-1-carbaldehyde | 12.88 | 111, 222, 265 | 265 | C13H9Cl2NO |
| 7 | (4E)-4-[(2,5-dichlorophenyl) imino]-3-(hydroxymethyl) cyclohexa-2,5-dien-1-one | -------- | ----- | 281 | C13H9Cl2NO2 |
| 8 | (6E)-6-[(2,5-dichlorophenyl) amino]-3-oxocyclohexa-1,4-diene-1-carbaldehyde | 7.66 | 242, 277, 278 | 279 | C13H7Cl2NO2 |
| 9 | 4-(2,6-dichlorophenylamino)-1,3-benzenedimethanol | 13.07 | 179, 214, 242, 279 | 298 | C14H13Cl2NO2 |
| 10 | 1-(2,6-Dichlorophenyl)indolin-2-one diclofenac amide | 13.72 | 151, 214, 242, 277 | 277 | C14H9Cl2NO |
| No. | Compound Name | RT (min) | Fragments (m/z) | MW | Formula |
|---|---|---|---|---|---|
| 1 | Diclofenac | 12.77 | 249, 277, 295 | 296 | C14H11Cl2N2O2 |
| 2 | 5-Hydroxydiclofenac | 6.32 | 214, 231, 249, 312 | 312 | C14H11Cl2NO3 |
| 3 | 4/-Hydroxydiclofenac | 6.54 | 214, 231, 249, 312 | 312 | C14H11Cl2NO3 |
| 4 | (2, 4-Dichloro-3-(2-methylanilino) phenol | 7.15 | 249, 268, 176 | 268 | C13H11Cl2NO |
| 5 | 3-(3-Chloroanilino)-2-methylphenol | 15.73 | 133, 203, 233, 107 | 233 | C13H12ClNO |
| 6 | 5-(2,4-Dihydroxyanilino)-4-(hydroxymethyl) benzene-1,3-diol | 4.6 | 133, 151, 203, 263 | 263 | C13H13No5 |
| 7 | (2-Aminophenyl) acetic acid | 5.04 | 77, 133, 151 | 151 | C8H9NO2 |
| 8 | Pyruvaldehyde, 1-(diethyl acetal) | 47, 101, 144, 146 | 146 | C7H14o2 | |
| 9 | 2-Ethoxyethyl acetate | 4.6 | 59, 88, 103, 132 | 132 | C6H12o3 |
| 10 | 4-Dichlorophenylamino-1,3benzen dimethanol | 14.67 | 214, 242, 277, 298 | 298 | C14H13Cl2NO2 |
| 11 | 4-Ethylbenzoic acid, 6-ethyl-3-octyl ester | 55, 70, 135, 290 | 290 | C19H30O2 | |
| 12 | 4-Methyl-8-hexadecenal, Z | 7.17 | 55, 70, 135, 252 | 252 | C17H32O |
| No. | Compound Name | RT (min) | Fragments (m/z) | MW | Formula |
|---|---|---|---|---|---|
| 1 | Diclofenac | 33.66 | 249, 277, 295 | 296 | C14H11Cl2N2O2 |
| 2 | 4/-Hydroxydiclofenac | 6.26 | 214, 231, 249, 312 | 312 | C14H11Cl2NO3 |
| 3 | 5-Hydroxydiclofenac | 6.67 | 214, 231, 249, 312 | 312 | C14H11Cl2NO3 |
| 4 | 2,4-Dichlorophenol | ------- | ------- | 163 | C6H4Cl2O |
| 5 | (3-Aminophenyl) acetic acid | 5 | 51, 77, 133, 151 | 151 | C8H9NO2 |
| 6 | (3-Hydroxyphenyl) acetic acid | 5.21 | 77, 133, 151 | 152 | C8H8O3 |
| 7 | 4-Amino-3,5-dichlorobenzene | 5 | 151, 179, 194 | 194 | C6H5Cl2NO2 |
| -1,2-diol | |||||
| 8 | (2,3-Dihydroxyphenyl) acetic acid | 7.19 | 55, 124, 168 | 168 | C8H8O4 |
| 9 | (4,5-Dihydroxy-2-methylphenyl) acetic acid | 7.19 | 151, 179, 182 | 182 | C9H10O4 |
| 10 | (2Z,4E)-2,3,5-Trihydroxypenta-2,4-dienoic acid | 9.18 | 47, 101, 144 | 146 | C5H6O5 |
| 11 | Ethyl ethoxyacetate | 4.57 | 31, 59, 88, 103, 132 | 132 | C6H12O3 |
| 12 | Pyruvaldehyde,1-(diethyl acetal) | 9.14 | 47, 101, 144 | 146 | C7H14O3 |
| Enzyme | Specific Enzyme Activity (mU/mg Protein) | |||||
|---|---|---|---|---|---|---|
| A. spanius S11 | A. piechaudii S18 | P. aeruginosa S1 | ||||
| Control * | MS + DCF ** | Control | MS + DCF | Control | MS + DCF | |
| Catechol 1,2-dioxygenase | 0.777 ± 0.064 | 0.928 ± 0.026 | 1.34 ± 0.256 | 2.714 ± 0.147 | 1.137 ± 0.293 | 1.832 ± 0.20 |
| Catechol 2,3-dioxygenase | 0.0021 ± 0.008 | 0.005 ± 0.002 | 0.334 ± 0.097 | 0.131 ± 0.083 | 0.059 ± 0.016 | 0. 386 ± 0.032 |
| Laccase | 0.192 ± 0.037 | 0.807 ± 0.127 | 0.390 ± 0.036 | 0.672 ± 0.096 | 0.004 ± 0.002 | 0.002 ± 0.001 |
| Peroxidase | 0.0457 ± 0.04 | 0.107 ± 0.076 | 0.654 ± 0.041 | 0.747 ± 0.054 | 0. 328 ± 0.071 | 0. 880 ± 0.127 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mohamed, M.S.M.; Asair, A.A.; Fetyan, N.A.H.; Elnagdy, S.M. Complete Biodegradation of Diclofenac by New Bacterial Strains: Postulated Pathways and Degrading Enzymes. Microorganisms 2023, 11, 1445. https://doi.org/10.3390/microorganisms11061445
Mohamed MSM, Asair AA, Fetyan NAH, Elnagdy SM. Complete Biodegradation of Diclofenac by New Bacterial Strains: Postulated Pathways and Degrading Enzymes. Microorganisms. 2023; 11(6):1445. https://doi.org/10.3390/microorganisms11061445
Chicago/Turabian StyleMohamed, Mahmoud S. M., Ayan A. Asair, Nashwa A. H. Fetyan, and Sherif M. Elnagdy. 2023. "Complete Biodegradation of Diclofenac by New Bacterial Strains: Postulated Pathways and Degrading Enzymes" Microorganisms 11, no. 6: 1445. https://doi.org/10.3390/microorganisms11061445
APA StyleMohamed, M. S. M., Asair, A. A., Fetyan, N. A. H., & Elnagdy, S. M. (2023). Complete Biodegradation of Diclofenac by New Bacterial Strains: Postulated Pathways and Degrading Enzymes. Microorganisms, 11(6), 1445. https://doi.org/10.3390/microorganisms11061445







