16S-rRNA-Based Metagenomic Profiling of the Bacterial Communities in Traditional Bulgarian Sourdoughs
Abstract
1. Introduction
2. Materials and Methods
2.1. Sourdough Sample Collection
2.2. Physicochemical and Biochemical Characterization of The Sourdoughs
2.3. Enumeration of Lactic Acid Bacteria and Yeasts
2.4. DNA Extraction
2.5. Sequencing and Data Analysis
2.6. Data Availability
2.7. Statistical Analyses
3. Results and Discussion
3.1. Physicochemical and Microbiological Characterization of Bulgarian Sourdoughs
3.2. Sequencing Data Analysis of Sourdough Samples
3.3. Bacterial Community Diversity within the Sourdough Samples (Alpha Diversity)
3.4. Bacterial Communities in the Studied Sourdough Samples
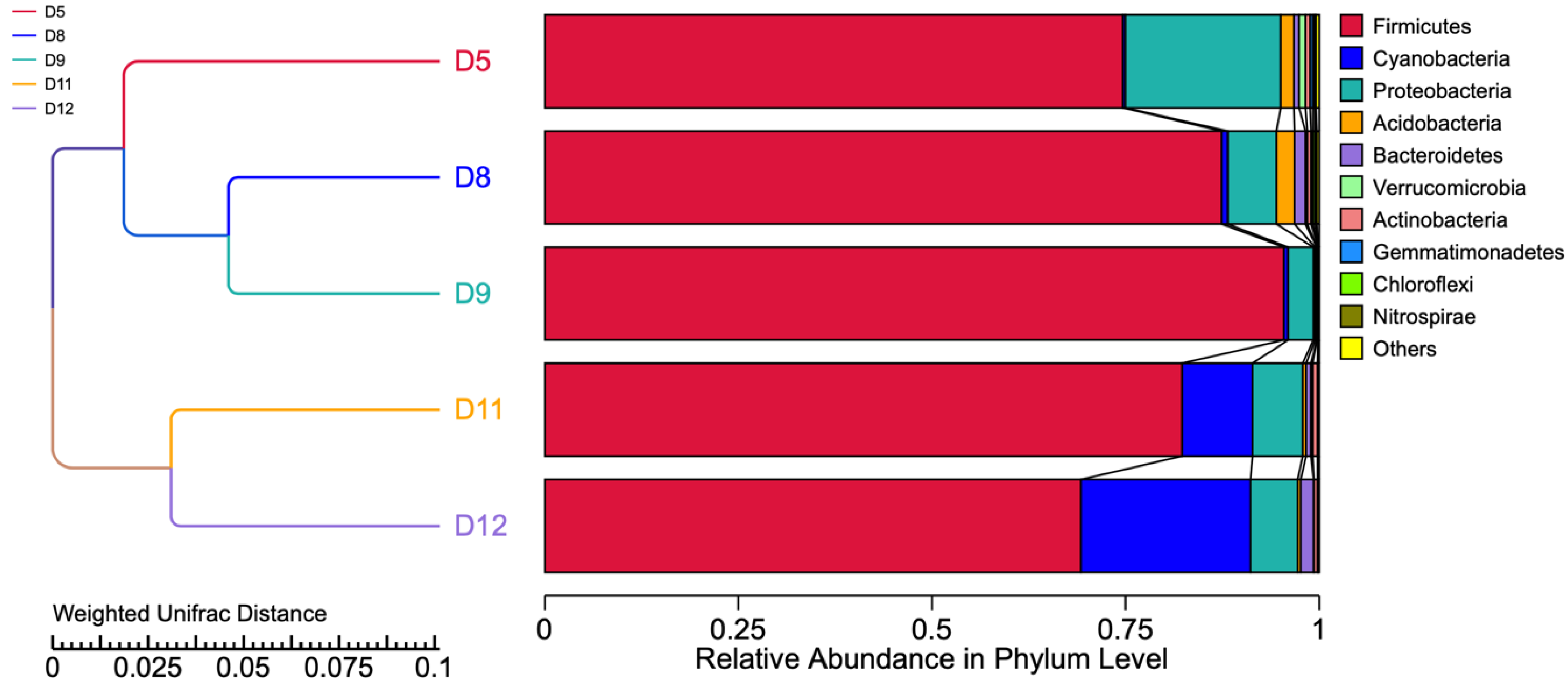
3.5. Similarity among the Different Sourdough Samples
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Butel, M.J. Probiotics, Gut Microbiota and Health. Med. Mal. Infect. 2014, 44, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Petkova, M.; Stefanova, P.; Gotcheva, V.; Angelov, A. Isolation and Characterization of Lactic Acid Bacteria and Yeasts from Typical Bulgarian Sourdoughs. Microorganisms 2021, 9, 1346. [Google Scholar] [CrossRef]
- Petkova, M.; Stefanova, P.; Gotcheva, V.; Kuzmanova, I.; Angelov, A. Microbiological and Physicochemical Characterization of Traditional Bulgarian Sourdoughs and Screening of Lactic Acid Bacteria for Amylolytic Activity. J. Chem. Technol. Metall. 2020, 55, 921–934. [Google Scholar]
- Minervini, F.; di Cagno, R.; Lattanzi, A.; de Angelis, M.; Antonielli, L.; Cardinali, G.; Cappelle, S.; Gobbetti, M. Lactic Acid Bacterium and Yeast Microbiotas of 19 Sourdoughs Used for Traditional/Typical Italian Breads: Interactions between Ingredients and Microbial Species Diversity. Appl. Environ. Microbiol. 2012, 78, 1251–1264. [Google Scholar] [CrossRef]
- Ercolini, D.; Pontonio, E.; de Filippis, F.; Minervini, F.; la Storia, A.; Gobbetti, M.; di Cagno, R. Microbial Ecology Dynamics during Rye and Wheat Sourdough Preparation. Appl. Environ. Microbiol. 2013, 79, 7827–7836. [Google Scholar] [CrossRef]
- Vogel, R.F.; Pavlovic, M.; Ehrmann, M.A.; Wiezer, A.; Liesegang, H.; Offschanka, S.; Voget, S.; Angelov, A.; Böcker, G.; Liebl, W. Genomic Analysis Reveals Lactobacillus Sanfranciscensis as Stable Element in Traditional Sourdoughs. Microb. Cell Fact. 2011, 10, S6. [Google Scholar] [CrossRef]
- Viiard, E.; Mihhalevski, A.; Rühka, T.; Paalme, T.; Sarand, I. Evaluation of the Microbial Community in Industrial Rye Sourdough upon Continuous Back-Slopping Propagation Revealed Lactobacillus Helveticus as the Dominant Species. J. Appl. Microbiol. 2013, 114, 404–412. [Google Scholar] [CrossRef]
- Bessmeltseva, M.; Viiard, E.; Simm, J.; Paalme, T.; Sarand, I. Evolution of Bacterial Consortia in Spontaneously Started Rye Sourdoughs during Two Months of Daily Propagation. PLoS ONE 2014, 9, e95449. [Google Scholar] [CrossRef]
- De Vuyst, L.; van Kerrebroeck, S.; Leroy, F. Microbial Ecology and Process Technology of Sourdough Fermentation. Adv. Appl. Microbiol. 2017, 100, 49–160. [Google Scholar] [CrossRef]
- van Kerrebroeck, S.; Maes, D.; de Vuyst, L. Sourdoughs as a Function of Their Species Diversity and Process Conditions, a Meta-Analysis. Trends Food Sci. Technol. 2017, 68, 152–159. [Google Scholar] [CrossRef]
- Scheirlinck, I.; van der Meulen, R.; van Schoor, A.; Vancanneyt, M.; de Vuyst, L.; Vandamme, P.; Huys, G. Taxonomic Structure and Stability of the Bacterial Community in Belgian Sourdough Ecosystems as Assessed by Culture and Population Fingerprinting. Appl. Environ. Microbiol. 2008, 74, 2414–2423. [Google Scholar] [CrossRef]
- Gobbetti, M.; Minervini, F.; Pontonio, E.; di Cagno, R.; de Angelis, M. Drivers for the Establishment and Composition of the Sourdough Lactic Acid Bacteria Biota. Int. J. Food Microbiol. 2016, 239, 3–18. [Google Scholar] [CrossRef] [PubMed]
- Reese, A.T.; Madden, A.A.; Joossens, M.; Lacaze, G.; Dunn, R.R. Influences of Ingredients and Bakers on the Bacteria and Fungi in Sourdough Starters and Bread. mSphere 2020, 5. [Google Scholar] [CrossRef] [PubMed]
- ben Amor, K.; Vaughan, E.E.; de Vos, W.M. Advanced Molecular Tools for the Identification of Lactic Acid Bacteria. J. Nutr. 2007, 137, 741S–747S. [Google Scholar] [CrossRef]
- Corsetti, A.; Lavermicocca, P.; Morea, M.; Baruzzi, F.; Tosti, N.; Gobbetti, M. Phenotypic and Molecular Identification and Clustering of Lactic Acid Bacteria and Yeasts from Wheat (Species Triticum Durum and Triticum Aestivum) Sourdoughs of Southern Italy. Int. J. Food Microbiol. 2001, 64, 95–104. [Google Scholar] [CrossRef]
- Ehrmann, M.A.; Müller, M.R.A.; Vogel, R.F. Molecular Analysis of Sourdough Reveals Lactobacillus Mindensis Sp. Nov. Int. J. Syst. Evol. Microbiol. 2003, 53, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Rossetti, L.; Giraffa, G. Rapid Identification of Dairy Lactic Acid Bacteria by M13-Generated, RAPD-PCR Fingerprint Databases. J. Microbiol. Methods 2005, 63, 135–144. [Google Scholar] [CrossRef] [PubMed]
- Healy, M.; Huong, J.; Bittner, T.; Lising, M.; Frye, S.; Raza, S.; Schrock, R.; Manry, J.; Renwick, A.; Nieto, R.; et al. Microbial DNA Typing by Automated Repetitive-Sequence-Based PCR. J. Clin. Microbiol. 2005, 43, 199–207. [Google Scholar] [CrossRef]
- Weisburg, W.G.; Barns, S.M.; Pelletier, D.A.; Lane, D.J. 16S Ribosomal DNA Amplification for Phylogenetic Study. J. Bacteriol. 1991, 173, 697–703. [Google Scholar] [CrossRef]
- Fleet, G.H. Yeasts in Foods and Beverages: Impact on Product Quality and Safety. Curr. Opin. Biotechnol. 2007, 18, 170–175. [Google Scholar] [CrossRef]
- Martin, K.J.; Rygiewicz, P.T. Fungal-Specific PCR Primers Developed for Analysis of the ITS Region of Environmental DNA Extracts. BMC Microbiol. 2005, 5, 28. [Google Scholar] [CrossRef]
- Jagadeesan, B.; Gerner-Smidt, P.; Allard, M.W.; Leuillet, S.; Winkler, A.; Xiao, Y.; Chaffron, S.; van der Vossen, J.; Tang, S.; Katase, M.; et al. The Use of next Generation Sequencing for Improving Food Safety: Translation into Practice. Food Microbiol. 2019, 79, 96–115. [Google Scholar] [CrossRef]
- Menezes, L.A.A.; Sardaro, M.L.S.; Duarte, R.T.D.; Mazzon, R.R.; Neviani, E.; Gatti, M.; de Dea Lindner, J. Sourdough Bacterial Dynamics Revealed by Metagenomic Analysis in Brazil. Food Microbiol. 2020, 85, 103302. [Google Scholar] [CrossRef] [PubMed]
- Lhomme, E.; Orain, S.; Courcoux, P.; Onno, B.; Dousset, X. The Predominance of Lactobacillus Sanfranciscensis in French Organic Sourdoughs and Its Impact on Related Bread Characteristics. Int. J. Food Microbiol. 2015, 213, 40–48. [Google Scholar] [CrossRef]
- ISO 15214:1998; Microbiology of Food and Animal Feeding Stuff –Horizontal Method for the Identification of Mesophilic Lactic Acid Bacteria–Colony Count Technique at 30 °C. ISO (International Organization for Standardization): London, UK, 1998.
- Vassileva, I.; Baev, V.; Yahubyan, G.; Apostolova-Kuzova, E.; Angelov, A.; Koprinarova, M. Identification of Bulgarian Sourdough Microbiota by Metagenomic Approach Using Three Commercially Available DNA Extraction Protocols. Food Technol. Biotechnol. 2023, in press. [Google Scholar] [CrossRef]
- Edgar, R.C. Search and Clustering Orders of Magnitude Faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME Allows Analysis of High-Throughput Community Sequencing Data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA Ribosomal RNA Gene Database Project: Improved Data Processing and Web-Based Tools. Nucleic Acids Res. 2012, 41, D590–D596. [Google Scholar] [CrossRef] [PubMed]
- Andersen, K.S.; Kirkegaard, R.H.; Karst, S.M.; Albertsen, M. Ampvis2: An R Package to Analyse and Visualise 16S RRNA Amplicon Data. bioRxiv 2018. [Google Scholar] [CrossRef]
- Vera, A.; Ly-Chatain, M.H.; Rigobello, V.; Demarigny, Y. Description of a French Natural Wheat Sourdough over 10 Consecutive Days Focussing on the Lactobacilli Present in the Microbiota. Antonie Van Leeuwenhoek Int. J. Gen. Mol. Microbiol. 2012, 101, 369–377. [Google Scholar] [CrossRef]
- Calvert, M.D.; Madden, A.A.; Nichols, L.M.; Haddad, N.M.; Lahne, J.; Dunn, R.R.; McKenney, E.A. A Review of Sourdough Starters: Ecology, Practices, and Sensory Quality with Applications for Baking and Recommendations for Future Research. PeerJ 2021, 9, e11389. [Google Scholar] [CrossRef]
- Rocha, J.M. Microbiological and Lipid Profiles of Broa: Contributions for the Characterization of a Traditional Portuguese Bread. Ph.D Thesis, Instituto Superior de Agronomia, Universidade de Lisboa, Lisbon, Portugal, 2011. [Google Scholar]
- Novotni, D.; Gänzle, M.; Rocha, J.M. Composition and Activity of Microbiota in Sourdough and Their Effect on Bread Quality and Safety. In Trends in Wheat and Bread Making; Elsevier: Amsterdam, The Netherlands, 2021; pp. 129–172. [Google Scholar]
- Gobbetti, M. The Sourdough Microflora: Interactions of Lactic Acid Bacteria and Yeasts. Trends Food Sci. Technol. 1998, 9, 267–274. [Google Scholar] [CrossRef]
- Ottogalli, G.; Galli, A.; Foschino, R. Italian Bakery Products Obtained with Sour Dough: Characterization of the Typical Microflora. Adv. Food Sci. 1996, 18, 131–144. [Google Scholar]
- De Vuyst, L.; Vrancken, G.; Ravyts, F.; Rimaux, T.; Weckx, S. Biodiversity, Ecological Determinants, and Metabolic Exploitation of Sourdough Microbiota. Food Microbiol. 2009, 26, 666–675. [Google Scholar] [CrossRef] [PubMed]
- van Kerrebroeck, S.; Comasio, A.; Harth, H.; de Vuyst, L. Impact of Starter Culture, Ingredients, and Flour Type on Sourdough Bread Volatiles as Monitored by Selected Ion Flow Tube-Mass Spectrometry. Food Res. Int. 2018, 106, 254–262. [Google Scholar] [CrossRef]
- di Cagno, R.; Pontonio, E.; Buchin, S.; de Angelis, M.; Lattanzi, A.; Valerio, F.; Gobbetti, M.; Calasso, M. Diversity of the Lactic Acid Bacterium and Yeast Microbiota in the Switch from Firm- to Liquid-Sourdough Fermentation. Appl. Environ. Microbiol. 2014, 80, 3161–3172. [Google Scholar] [CrossRef] [PubMed]
- Robert, H.; Gabriel, V.; Lefebvre, D.; Rabier, P.; Vayssier, Y.; Fontagné-Faucher, C. Study of the Behaviour of Lactobacillus Plantarum and Leuconostoc Starters during a Complete Wheat Sourdough Breadmaking Process. Lwt 2006, 39, 256–265. [Google Scholar] [CrossRef]
- Fujimoto, A.; Ito, K.; Narushima, N.; Miyamoto, T. Identification of Lactic Acid Bacteria and Yeasts, and Characterization of Food Components of Sourdoughs Used in Japanese Bakeries. J. Biosci. Bioeng. 2019, 127, 575–581. [Google Scholar] [CrossRef] [PubMed]
- Buksa, K. Effect of Pentoses, Hexoses, and Hydrolyzed Arabinoxylan on the Most Abundant Sugar, Organic Acid, and Alcohol Contents during Rye Sourdough Bread Production. Cereal Chem. 2020, 97, 642–652. [Google Scholar] [CrossRef]
- Debonne, E.; van Schoors, F.; Maene, P.; van Bockstaele, F.; Vermeir, P.; Verwaeren, J.; Eeckhout, M.; Devlieghere, F. Comparison of the Antifungal Effect of Undissociated Lactic and Acetic Acid in Sourdough Bread and in Chemically Acidified Wheat Bread. Int. J. Food Microbiol. 2020, 321, 108551. [Google Scholar] [CrossRef]
- Komatsuzaki, N.; Izawa, M.; Suzumori, M.; Fujihara, S.; Shima, J. Characteristics of New Sourdough Using Lactic Acid Bacteria and Wild Yeast. J. Food Sci. Nutr. Res. 2019, 2, 001–012. [Google Scholar] [CrossRef]
- Häggman, M.; Salovaara, H. Effect of Fermentation Rate on Endogenous Leavening of Candida Milleri in Sour Rye Dough. Food Res. Int. 2008, 41, 266–273. [Google Scholar] [CrossRef]
- de Vuyst, L.; Harth, H.; van Kerrebroeck, S.; Leroy, F. Yeast Diversity of Sourdoughs and Associated Metabolic Properties and Functionalities. Int. J. Food Microbiol. 2016, 239, 26–34. [Google Scholar] [CrossRef]
- de Vuyst, L.; van Kerrebroeck, S.; Harth, H.; Huys, G.; Daniel, H.M.; Weckx, S. Microbial Ecology of Sourdough Fermentations: Diverse or Uniform? Food Microbiol. 2014, 37, 11–29. [Google Scholar] [CrossRef]
- Narendranath, N.V.; Thomas, K.C.; Ingledew, W.M. Effects of Acetic Acid and Lactic Acid on the Growth of Saccharomyces Cerevisiae in a Minimal Medium. J. Ind. Microbiol. Biotechnol. 2001, 26, 171–177. [Google Scholar] [CrossRef]
- Lattanzi, A.; Minervini, F.; Gobbetti, M. Assessment of Comparative Methods for Storing Type-I Wheat Sourdough. Lwt 2014, 59, 948–955. [Google Scholar] [CrossRef]
- Sterr, Y.; Weiss, A.; Schmidt, H. Evaluation of Lactic Acid Bacteria for Sourdough Fermentation of Amaranth. Int. J. Food Microbiol. 2009, 136, 75–82. [Google Scholar] [CrossRef]
- Carbonetto, B.; Nidelet, T.; Guezenec, S.; Perez, M.; Segond, D.; Sicard, D. Interactions between Kazachstania Humilis Yeast Species and Lactic Acid Bacteria in Sourdough. Microorganisms 2020, 8, 240. [Google Scholar] [CrossRef]
- Wang, F.; Zhao, H.; Xiang, H.; Wu, L.; Men, X.; Qi, C.; Chen, G.; Zhang, H.; Wang, Y.; Xian, M. Species Diversity and Functional Prediction of Surface Bacterial Communities on Aging Flue-Cured Tobaccos. Curr. Microbiol. 2018, 75, 1306–1315. [Google Scholar] [CrossRef]
- Thomas, P.; Sekhar, A.C. Cultivation Versus Molecular Analysis of Banana (Musa Sp.) Shoot-Tip Tissue Reveals Enormous Diversity of Normally Uncultivable Endophytic Bacteria. Microb. Ecol. 2017, 73, 885–899. [Google Scholar] [CrossRef]
- Peuthert, A.; Chakrabarti, S.; Pflugmacher, S. Uptake of Microcystins-LR and -LF (Cyanobacterial Toxins) in Seedlings of Several Important Agricultural Plant Species and the Correlation with Cellular Damage (Lipid Peroxidation). Environ. Toxicol. 2007, 22, 436–442. [Google Scholar] [CrossRef] [PubMed]
- Hereman, T.C.; Bittencourt-Oliveira, M.d.C. Bioaccumulation of Microcystins in Lettuce. J. Phycol. 2012, 48, 1535–1537. [Google Scholar] [CrossRef] [PubMed]
- Dietrich, D.; Hoeger, S. Guidance Values for Microcystins in Water and Cyanobacterial Supplement Products (Blue-Green Algal Supplements): A Reasonable or Misguided Approach? Toxicol. Appl. Pharmacol. 2005, 203, 273–289. [Google Scholar] [CrossRef] [PubMed]
- Zanchett, G.; Oliveira-Filho, E. Cyanobacteria and Cyanotoxins: From Impacts on Aquatic Ecosystems and Human Health to Anticarcinogenic Effects. Toxins 2013, 5, 1896–1917. [Google Scholar] [CrossRef]
- Pham, T.-L.; Utsumi, M. An Overview of the Accumulation of Microcystins in Aquatic Ecosystems. J. Environ. Manag. 2018, 213, 520–529. [Google Scholar] [CrossRef]
- Liu, X.; Zhou, M.; Jiaxin, C.; Luo, Y.; Ye, F.; Jiao, S.; Hu, X.; Zhang, J.; Lü, X. Bacterial Diversity in Traditional Sourdough from Different Regions in China. Lwt 2018, 96, 251–259. [Google Scholar] [CrossRef]
- Pontonio, E.; Nionelli, L.; Curiel, J.A.; Sadeghi, A.; di Cagno, R.; Gobbetti, M.; Rizzello, C.G. Iranian Wheat Flours from Rural and Industrial Mills: Exploitation of the Chemical and Technology Features, and Selection of Autochthonous Sourdough Starters for Making Breads. Food Microbiol. 2015, 47, 99–110. [Google Scholar] [CrossRef]
- Fessard, A.; Remize, F. Genetic and Technological Characterization of Lactic Acid Bacteria Isolated from Tropically Grown Fruits and Vegetables. Int. J. Food Microbiol. 2019, 301, 61–72. [Google Scholar] [CrossRef]
- Fusco, V.; Quero, G.M.; Stea, G.; Morea, M.; Visconti, A. Novel PCR-Based Identification of Weissella Confusa Using an AFLP-Derived Marker. Int. J. Food Microbiol. 2011, 145, 437–443. [Google Scholar] [CrossRef]
- Osimani, A.; Garofalo, C.; Aquilanti, L.; Milanović, V.; Clementi, F. Unpasteurised Commercial Boza as a Source of Microbial Diversity. Int. J. Food Microbiol. 2015, 194, 62–70. [Google Scholar] [CrossRef]
- Fusco, V.; Quero, G.M.; Cho, G.-S.; Kabisch, J.; Meske, D.; Neve, H.; Bockelmann, W.; Franz, C.M.A.P. The Genus Weissella: Taxonomy, Ecology and Biotechnological Potential. Front. Microbiol. 2015, 6, 155. [Google Scholar] [CrossRef] [PubMed]
- Giri, A.V.; Anandkumar, N.; Muthukumaran, G.; Pennathur, G. A Novel Medium for the Enhanced Cell Growth and Production of Prodigiosin from Serratia Marcescens Isolated from Soil. BMC Microbiol. 2004, 4, 11. [Google Scholar] [CrossRef] [PubMed]
- Chaves-López, C.; de Angelis, M.; Martuscelli, M.; Serio, A.; Paparella, A.; Suzzi, G. Characterization of the Enterobacteriaceae Isolated from an Artisanal Italian Ewe’s Cheese (Pecorino Abruzzese). J. Appl. Microbiol. 2006, 101, 353–360. [Google Scholar] [CrossRef] [PubMed]
- González-Rodríguez, M.N.; Sanz, J.J.; Santos, J.Á.; Otero, A.; García-López, M.L. Numbers and Types of Microorganisms in Vacuum-Packed Cold-Smoked Freshwater Fish at the Retail Level. Int. J. Food Microbiol. 2002, 77, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Monteiro, R.A.; Balsanelli, E.; Wassem, R.; Marin, A.M.; Brusamarello-Santos, L.C.C.; Schmidt, M.A.; Tadra-Sfeir, M.Z.; Pankievicz, V.C.S.; Cruz, L.M.; Chubatsu, L.S.; et al. Herbaspirillum-Plant Interactions: Microscopical, Histological and Molecular Aspects. Plant Soil 2012, 356, 175–196. [Google Scholar] [CrossRef]
- Salter, S.J.; Cox, M.J.; Turek, E.M.; Calus, S.T.; Cookson, W.O.; Moffatt, M.F.; Turner, P.; Parkhill, J.; Loman, N.J.; Walker, A.W. Reagent and Laboratory Contamination Can Critically Impact Sequence-Based Microbiome Analyses. BMC Biol. 2014, 12, 87. [Google Scholar] [CrossRef]
- Barton, H.A.; Taylor, N.M.; Lubbers, B.R.; Pemberton, A.C. DNA Extraction from Low-Biomass Carbonate Rock: An Improved Method with Reduced Contamination and the Low-Biomass Contaminant Database. J. Microbiol. Methods 2006, 66, 21–31. [Google Scholar] [CrossRef]
- Scheirlinck, I.; van der Meulen, R.; van Schoor, A.; Vancanneyt, M.; de Vuyst, L.; Vandamme, P.; Huys, G. Influence of Geographical Origin and Flour Type on Diversity of Lactic Acid Bacteria in Traditional Belgian Sourdoughs. Appl. Environ. Microbiol. 2007, 73, 6262–6269. [Google Scholar] [CrossRef] [PubMed]
- Landis, E.A.; Oliverio, A.M.; McKenney, E.A.; Nichols, L.M.; Kfoury, N.; Biango-Daniels, M.; Shell, L.K.; Madden, A.A.; Shapiro, L.; Sakunala, S.; et al. The Diversity and Function of Sourdough Starter Microbiomes. Elife 2021, 10, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Gobbetti, M.; Corsetti, A.; Rossi, J.; Rosa, F. la Identification and Clustering of Lactic Acid Bacteria and Yeasts from Wheat Sourdoughs of Central Italy. Ital. J. Food Sci. (Italy) 1994, 6, 85–94. [Google Scholar]
- Stanzer, D.; Kazazić, S.; Ivanuša, I.; Mrvčić, J.; Hanousek Čiča, K. Diversity of Lactic Acid Bacteria on Organic Flours and Application of Isolates in Sourdough Fermentation. Croat. J. Food Technol. Biotechnol. Nutr. 2017, 12, 44–51. [Google Scholar]
- Golshan Tafti, A.; Peighambardoust, S.H.; Hejazi, M.A.; Moosavy, M.H. Diversity of Lactobacillus Strains in Iranian Traditional Wheat Sourdough. J. Food Qual. Hazards Control 2014, 1, 41–45. [Google Scholar]
- Shi, S.; Cheng, B.; Gu, B.; Sheng, T.; Tu, J.; Shao, Y.; Qi, K.; Zhou, D. Evaluation of the Probiotic and Functional Potential of Lactobacillus Agilis 32 Isolated from Pig Manure. Lett. Appl. Microbiol. 2021, 73, 9–19. [Google Scholar] [CrossRef] [PubMed]
- Falasconi, I.; Fontana, A.; Patrone, V.; Rebecchi, A.; Garrido, G.D.; Principato, L.; Callegari, M.L.; Spigno, G.; Morelli, L. Genome-Assisted Characterization of Lactobacillus Fermentum, Weissella Cibaria, and Weissella Confusa Strains Isolated from Sorghum as Starters for Sourdough Fermentation. Microorganisms 2020, 8, 1388. [Google Scholar] [CrossRef] [PubMed]
- Bavaro, A.R.; di Biase, M.; Conte, A.; Lonigro, S.L.; Caputo, L.; Cedola, A.; del Nobile, M.A.; Logrieco, A.F.; Lavermicocca, P.; Valerio, F. Weissella Cibaria Short-Fermented Liquid Sourdoughs Based on Quinoa or Amaranth Flours as Fat Replacer in Focaccia Bread Formulation. Int. J. Food Sci. Technol. 2021, 56, 3197–3208. [Google Scholar] [CrossRef]
- Comasio, A.; Verce, M.; van Kerrebroeck, S.; de Vuyst, L. Diverse Microbial Composition of Sourdoughs From Different Origins. Front. Microbiol. 2020, 11, 1212. [Google Scholar] [CrossRef]
- Juega, M.; Costantini, A.; Bonello, F.; Cravero, M.C.; Martinez-Rodriguez, A.J.; Carrascosa, A.V.; Garcia-Moruno, E. Effect of Malolactic Fermentation by Pediococcus Damnosus on the Composition and Sensory Profile of Albariño and Caiño White Wines. J. Appl. Microbiol. 2014, 116, 586–595. [Google Scholar] [CrossRef]
- Immerstrand, T.; Paul, C.J.; Rosenquist, A.; Deraz, S.; Mårtensson, O.B.; Ljungh, Å.; Blücher, A.; Öste, R.; Holst, O.; Karlsson, E.N. Characterization of the Properties of Pediococcus Parvulus for Probiotic or Protective Culture Use. J. Food Prot. 2010, 73, 960–966. [Google Scholar] [CrossRef]



| Sample Code | Depository | Flour Origin | Dry Matter, % | pH | TTA, ml 0.1 NaOH | LA, mM | AA, mM | FQ | LAB, Log cfu/g | Yeast, Log cfu/g |
|---|---|---|---|---|---|---|---|---|---|---|
| D 5 | Bakery “8”, Smolyan (Rhodope mountains, South Bulgaria) | Triticum aestivum, white | 56.7 ± 0.96 a | 4.28 ± 0.06 a | 12.2 ± 0.12 d | 79.7 ± 1.1 b | 37.8 ± 0.9 a | 2.11 ± 0.82 b | 11.68 ± 0.58 a | 8.79 ± 0.44 ab |
| D 8 | Bakery Samun, Bansko (Southwest Bulgaria) | Triticum aestivum, wholegrain | 41.95 ± 0.47 d | 3.56 ± 0.04 c | 15.6 ± 0.18 a | 97.2 ± 1.5 a | 15.4 ± 1.1 c | 6.31 ± 0.72 a | 11.41 ± 0.88 a | 7.99 ± 0.84 ab |
| D9 | Bakery Samun, Bansko (Southwest Bulgaria) | Triticum monococcum, wholegrain | 46.67 ± 0.62 c | 4.04 ± 0.08 b | 13.8 ± 0.16 b | 83.5 ± 2.3 b | 16.4 ± 0.8 c | 5.09 ± 0,91 a | 11.36 ± 0.46 a | 9.59 ± 0.68 a |
| D 11 | Bakery Kusi 1, Ruse (North Bulgaria) | Triticum aestivum, white | 48.78 ± 0.54 b | 4.02 ± 0.02 b | 13.6 ± 0.08 b | 74.5 ± 2.5 c | 15.1 ± 1.0 c | 4.93 ± 0.88 a | 9.92 ± 0.96 a | 7.72 ± 0.34 b |
| D 12 | Bakery Kusi 2, Ruse (North Bulgaria) | Triticum aestivum, white | 56.35 ± 0.56 a | 4.12 ± 0.02 b | 12.8 ± 0.12 c | 70.9 ± 1.8 c | 29.8 ± 0.8 b | 2.38 ± 0.84 b | 10.81 ± 0.32 a | 8.04 ± 0.86 ab |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Baev, V.; Apostolova, E.; Gotcheva, V.; Koprinarova, M.; Papageorgiou, M.; Rocha, J.M.; Yahubyan, G.; Angelov, A. 16S-rRNA-Based Metagenomic Profiling of the Bacterial Communities in Traditional Bulgarian Sourdoughs. Microorganisms 2023, 11, 803. https://doi.org/10.3390/microorganisms11030803
Baev V, Apostolova E, Gotcheva V, Koprinarova M, Papageorgiou M, Rocha JM, Yahubyan G, Angelov A. 16S-rRNA-Based Metagenomic Profiling of the Bacterial Communities in Traditional Bulgarian Sourdoughs. Microorganisms. 2023; 11(3):803. https://doi.org/10.3390/microorganisms11030803
Chicago/Turabian StyleBaev, Vesselin, Elena Apostolova, Velitchka Gotcheva, Miglena Koprinarova, Maria Papageorgiou, João Miguel Rocha, Galina Yahubyan, and Angel Angelov. 2023. "16S-rRNA-Based Metagenomic Profiling of the Bacterial Communities in Traditional Bulgarian Sourdoughs" Microorganisms 11, no. 3: 803. https://doi.org/10.3390/microorganisms11030803
APA StyleBaev, V., Apostolova, E., Gotcheva, V., Koprinarova, M., Papageorgiou, M., Rocha, J. M., Yahubyan, G., & Angelov, A. (2023). 16S-rRNA-Based Metagenomic Profiling of the Bacterial Communities in Traditional Bulgarian Sourdoughs. Microorganisms, 11(3), 803. https://doi.org/10.3390/microorganisms11030803











