Research of Multicopper Oxidase and Its Degradation of Histamine in Lactiplantibacillus plantarum LPZN19
Abstract
:1. Introduction
2. Materials and Methods
2.1. Primers, Plasmid and Strains
2.2. Homologous Recombination of the MCO/pET-28a
2.3. Heterologous Expression and Purification of Recombinant MCO
2.4. Determination of the Enzymatic Properties of Recombinant MCO
2.4.1. Measurement of Enzyme Activity
2.4.2. The Effect of pH on the Activity and Stability of Recombinant MCO
2.4.3. The Effect of Temperature on the Activity and Stability of Recombinant MCO
2.4.4. Determination of Kinetic Parameters of Recombinant MCO
2.4.5. Effect of Metal Ions on Recombinant MCO Activity
2.4.6. Effect of Sodium Chloride on Recombinant MCO Activity
2.4.7. Investigation of the Substrate Specificity and Optimal Substrate of Recombinant MCO
2.5. Identification and Prediction of Recombinant MCO Structure
2.5.1. Identification of the Secondary Structure
2.5.2. Prediction of the Tertiary Structure
2.6. The Change in Spatial Structure after MCO Acting on Histamine
2.6.1. Preparation of the Histamine–MCO Reaction System
2.6.2. Determination of Ultraviolet Spectrum and Endogenous Fluorescence Spectra
2.6.3. Determination of Secondary Structure Change in MCO
2.6.4. Characterization of the Tertiary Structure Changes of MCO
2.7. Monitoring and Identification of Histamine Degradation Products
2.8. Statistical Analysis
3. Results
3.1. Sequence Analysis of the MCO
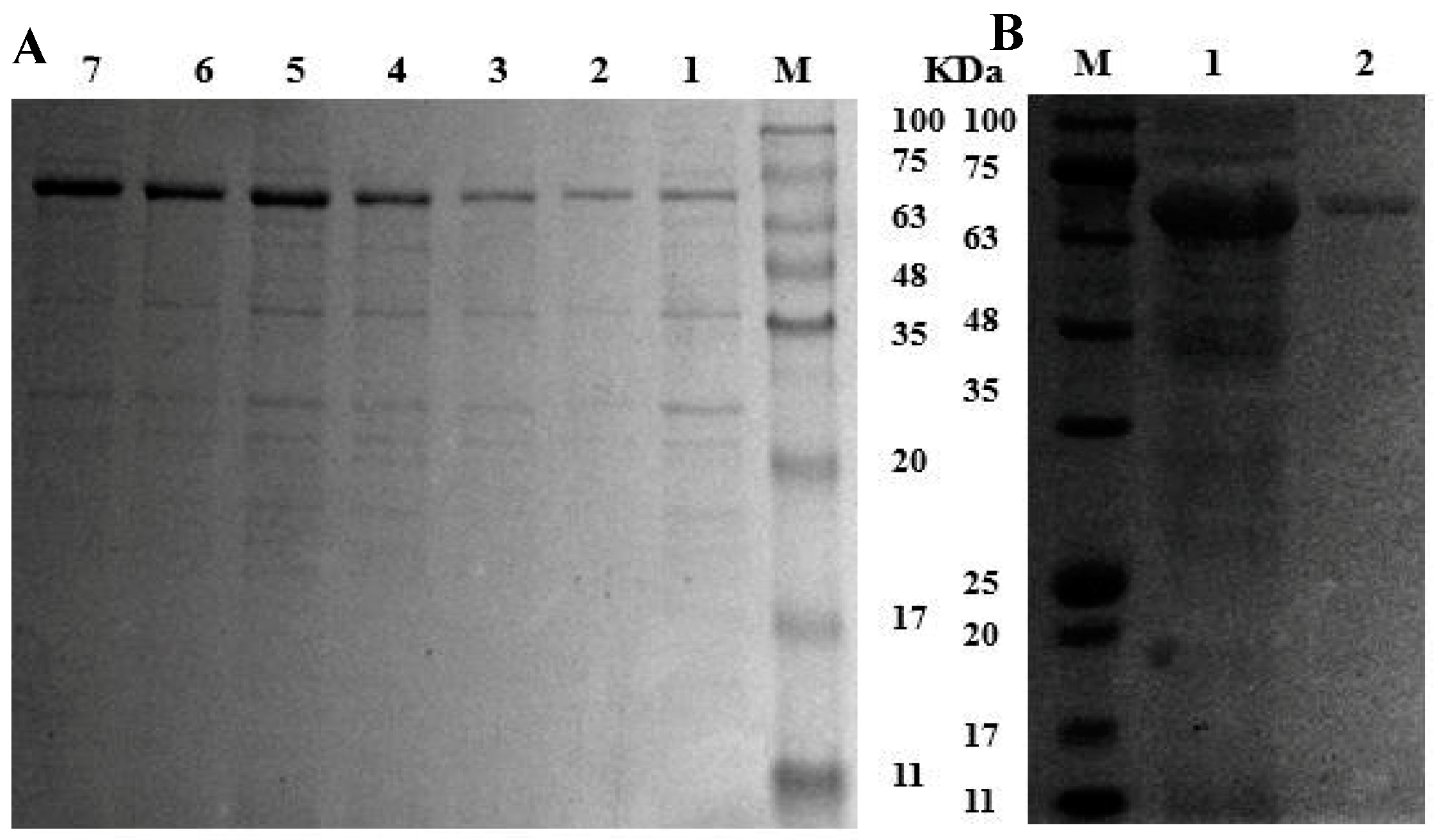
3.2. Heterologous Expression and Purification of Recombinant MCO
3.3. Effects of Physicochemical Factors on Recombinant MCO Activity
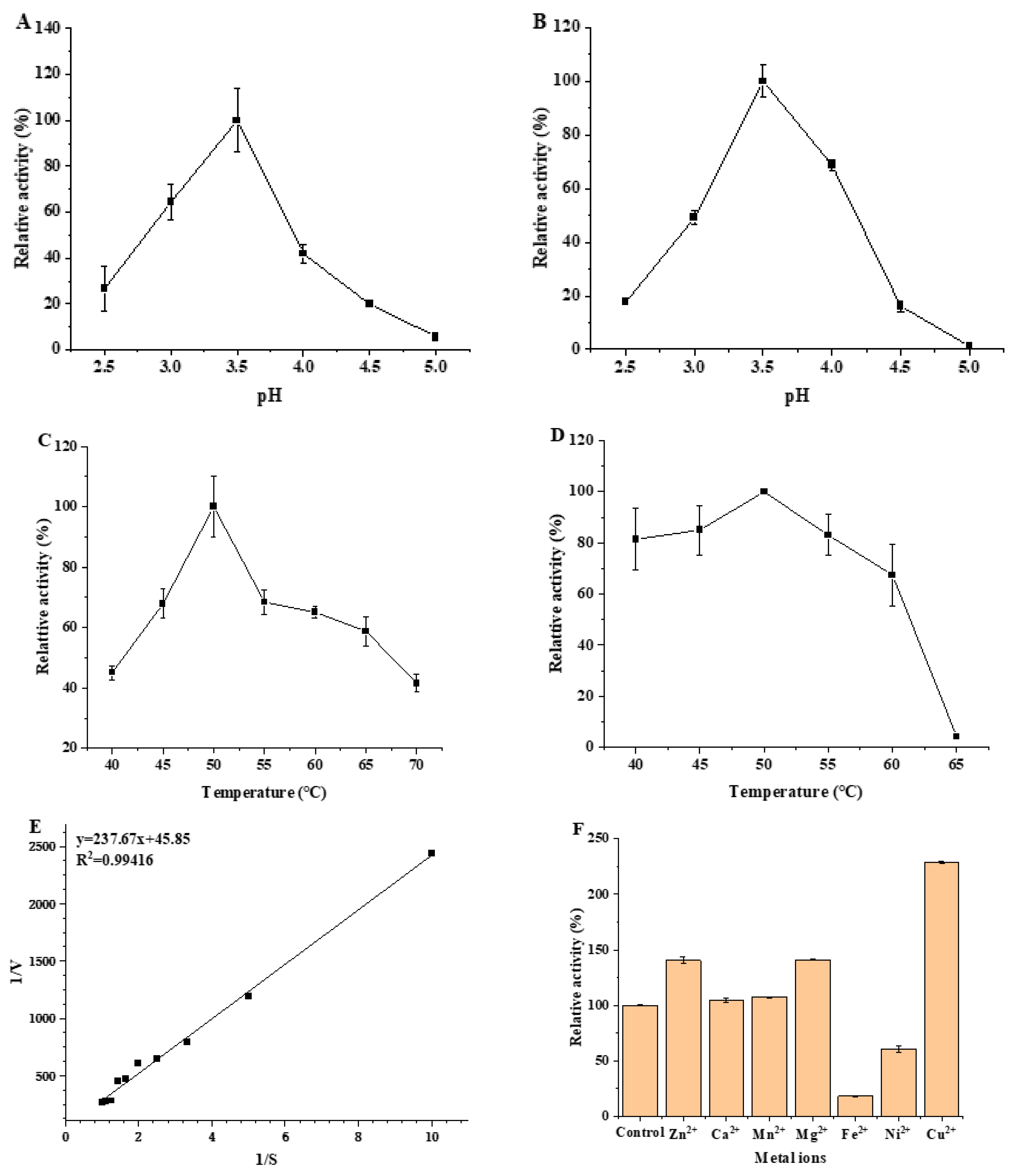
3.3.1. Optimum pH and pH Stability of MCO
3.3.2. Optimum Temperature and Thermal Stability of MCO
3.3.3. Determination of Kinetic Parameters of MCO
3.3.4. Effect of Metal Ions on Recombinant MCO Activity
3.3.5. Effect of Sodium Chloride on Recombinant MCO Activity
3.3.6. Degradation of Biogenic Amine by MCO
3.4. Identification of the Structure of Recombinant MCO
3.4.1. Identification of the Secondary Structure
3.4.2. Prediction of the Tertiary Structure
3.5. Analysis on the Change in Recombinant MCO Spatial Structure
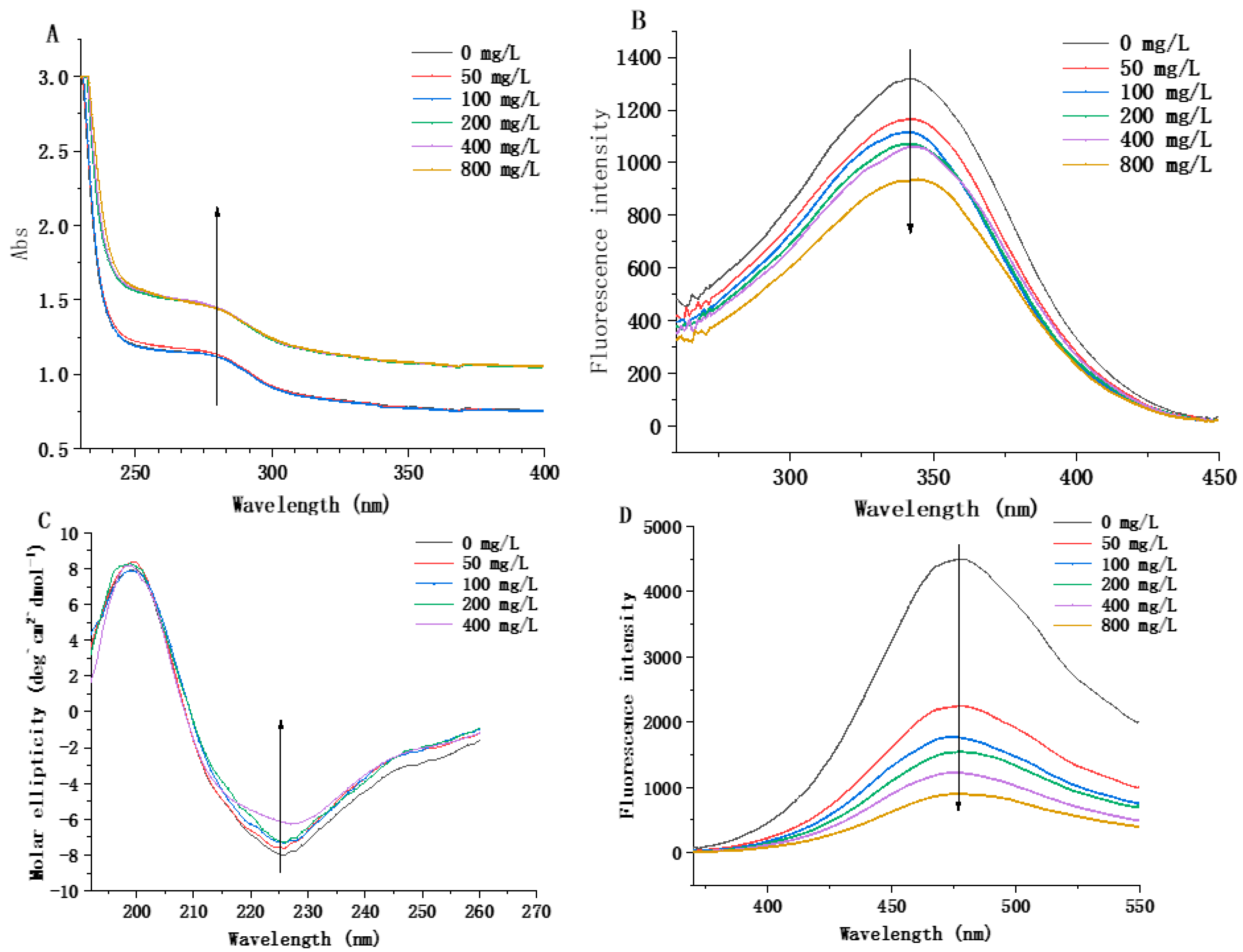
3.5.1. Determination of Structure Induction of MCO by Histamine
3.5.2. Changes in Secondary Structure of Recombinant MCO
3.5.3. Changes in Tertiary Structure of Recombinant MCO
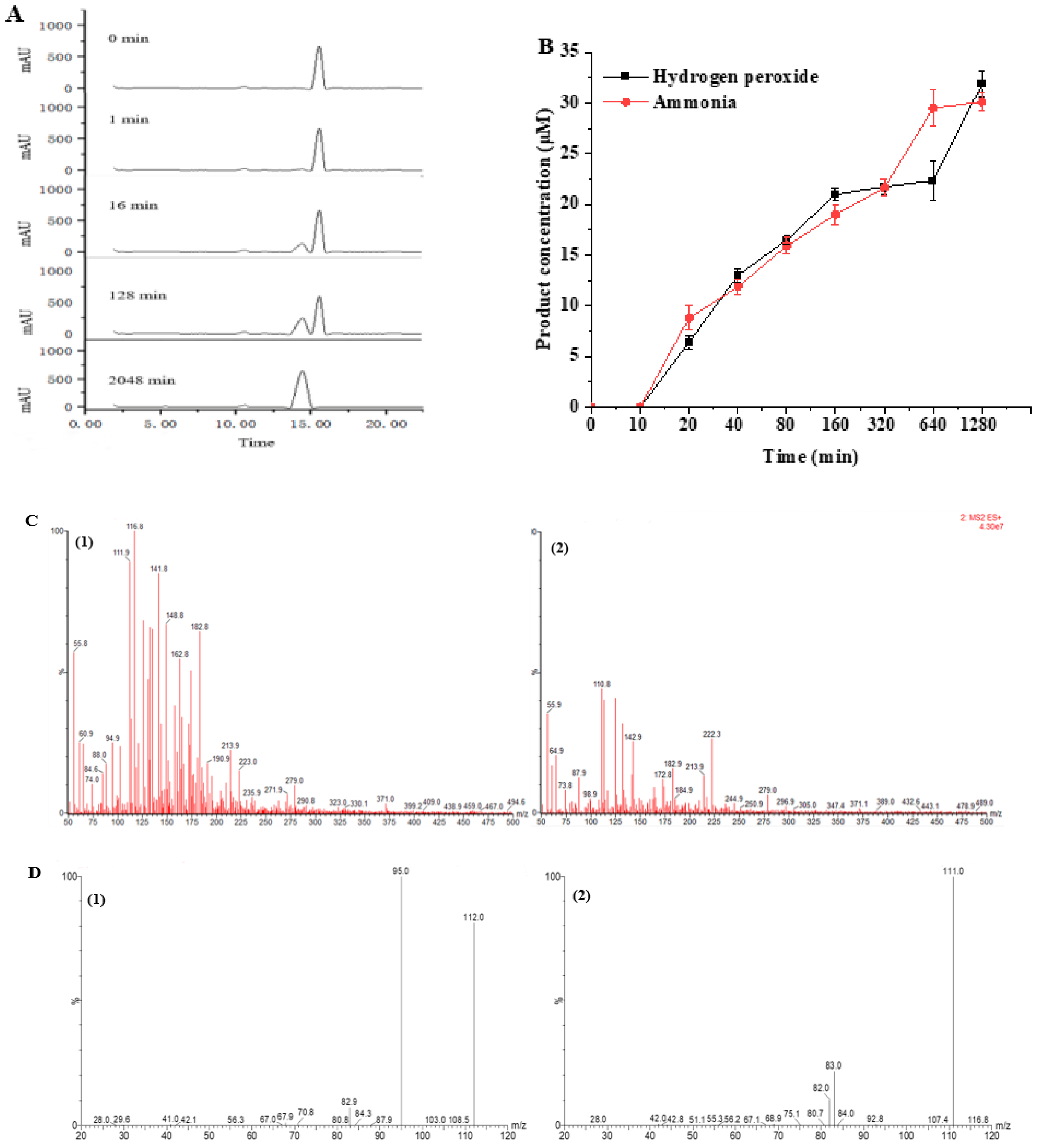
3.6. Monitoring and Identification of Histamine Degradation Products
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Vieira, C.P.; Da Costa, M.P.; Silva, V.L.M.; Da Silva Frasao, B.; De Aquino, L.F.M.C.; De Oliveira Nunes, Y.E.C.; Conte-Junior, C.A. Development and validation of RP-HPLC-DAD method for biogenic amines determination in probiotic yogurts. Arab. J. Chem. 2020, 13, 1582–1597. [Google Scholar] [CrossRef]
- Cueva, C.; García-Ruiz, A.; González-Rompinelli, E.; Bartolome, B.; Martín-Álvarez, J.; Salazar, O.; Vicente, F.; Bills, F.; Moreno-Arribas, V. Degradation of biogenic amines by vineyard ecosystem fungi. Potential use in winemaking. J. Appl. Microbiol. 2012, 112, 672–682. [Google Scholar] [CrossRef] [PubMed]
- Lorenzo, J.M.; Munekata, P.E.S.; Dominguez, R. Role of autochthonous starter cultures in the reduction of biogenic amines in traditional meat products. Curr. Opin. Food Sci. 2017, 14, 61–65. [Google Scholar] [CrossRef]
- Wang, Y.; Pei, H.; Liu, Y.; Huang, X.; Deng, L.; Lan, Q.; Chen, S.; He, L.; Liu, A.; Ao, X. Inhibitory mechanism of cell-free supernatants of Lactobacillus plantarum on Proteus mirabilis and influence of the expression of histamine synthesis-related genes. Food Control 2021, 125, 107982. [Google Scholar] [CrossRef]
- Santos, M.H.S. Biogenic amines: Their importance in foods. Int. J. Food Microbiol. 1996, 29, 213–231. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.H.; Ren, H.Y.; Wang, W.; Bai, T.; Li, J.X. Evaluation of key factors influencing histamine formation and accumulation in fermented sausages. J. Food Saf. 2015, 35, 395–402. [Google Scholar] [CrossRef]
- Naila, A.; Flint, S.; Fletcher, G.; Bremer, P.; Meerdink, G. Control of biogenic amines in food-existing and emerging approaches. J. Food Sci. 2010, 75, R139–R150. [Google Scholar] [CrossRef]
- Moniente, M.; Garcia-Gonzalo, D.; Ontañón, I.; Pagán, R.; Botello-Morte, L. Histamine accumulation in dairy products: Microbial causes, techniques for the detection of histamine-producing microbiota, and potential solutions. Compr. Rev. Food Sci. Food Saf. 2021, 20, 1481–1523. [Google Scholar] [CrossRef]
- Capozzi, V.; Russo, P.; Ladero, V.; Fernández, M.; Fiocco, D.; Alvarez, M.A.; Grieco, F.; Spano, G. Biogenic amines degradation by Lactobacillus plantarum: Toward a potential application in wine. Front. Microbiol. 2012, 3, 122. [Google Scholar] [CrossRef]
- Lee, J.I.; Kim, Y.W. Characterization of amine oxidases from Arthrobacter aurescens and application for determination of biogenic amines. World J. Microbiol. Biotechnol. 2013, 29, 673–682. [Google Scholar] [CrossRef]
- Yukl, E.T.; Davidson, V.L. Diversity of structures, catalytic mechanisms and processes of cofactor biosynthesis of tryptophylquinone-bearing enzymes. Arch. Biochem. Biophys. 2018, 654, 40–46. [Google Scholar] [CrossRef] [PubMed]
- Guarcello, R.; De Angelis, M.; Settanni, L.; Formiglio, S.; Gaglio, R.; Minervini, F.; Moschetti, G.; Gobbetti, M. Selection of amine-oxidizing dairy lactic acid bacteria and identification of the enzyme and gene involved in the decrease of biogenic amines. Appl. Environ. Microbiol. 2016, 82, 6870–6880. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Pérez, S.; Comas-Basté, O.; Costa-Catala, J.; Iduriaga-Platero, I.; Teresa Veciana-Nogués, M.; Carmen Vidal-Carou, M.; Luz Latorre-Moratalla, M. The rate of histamine degradation by diamine oxidase is compromised by other biogenic amines. Front. Nutr. 2022, 9, 897028. [Google Scholar] [CrossRef] [PubMed]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef]
- Yang, S.; Long, Y.; Yan, H.; Cai, H.; Li, Y.; Wang, X. Gene cloning, identification, and characterization of the multicopper oxidase CumA from Pseudomonas sp. 593. Biotechnol. Appl. Biochem. 2017, 64, 347–355. [Google Scholar] [CrossRef]
- Wu, X.; He, W.; Yao, L.; Zhang, H.; Liu, Z.; Wang, W.; Ye, Y.; Cao, J. Characterization of binding interactions of (-)-epigallocatechin-3-gallate from green tea and lipase. J. Agric. Food Chem. 2013, 61, 8829–8835. [Google Scholar] [CrossRef]
- Biasini, M.; Bienert, S.; Waterhouse, A.; Arnold, K.; Studer, G.; Schmidt, T.; Kiefer, F.; Cassarino, T.G.; Bertoni, M.; Bordoli, L.; et al. SWISS-MODEL: Modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 2014, 42, W252–W258. [Google Scholar] [CrossRef]
- Zhong, D.; Jiao, Y.; Zhang, Y.; Zhang, W.; Li, N.; Zuo, Q.; Qian, W.; Wei, X.; Liu, Z. Effects of the gene carrier polyethyleneimines on structure and function of blood components. Biomaterials 2013, 34, 294–305. [Google Scholar] [CrossRef]
- Dankowska, A.; Kowalewski, W. Comparison of different classification methods for analyzing fluorescence spectra to characterize type and freshness of olive oils. Eur. Food Res. Technol. 2019, 245, 745–752. [Google Scholar] [CrossRef]
- Si, Y.-X.; Wang, Z.J.; Park, D.; Chung, H.Y.; Wang, S.F.; Yan, L.; Yang, J.M.; Qian, G.Y.; Yin, S.-J.; Park, Y.D. Effect of hesperetin on tyrosinase: Inhibition kinetics integrated computational simulation study. Int. J. Biol. Macromol. 2012, 50, 257–262. [Google Scholar] [CrossRef]
- Mohan, M.; Ramachandran, D.; Sankar, T.; Anandan, R. Influence of pH on the solubility and conformational characteristics of muscle proteins from mullet (Mugil cephalus). Process Biochem. 2007, 42, 1056–1062. [Google Scholar] [CrossRef]
- Li, Y.; Yin, J.; Qu, G.; Lv, L.; Li, Y.; Yang, S.; Wang, X.G. Gene cloning, protein purification, and enzymatic properties of multicopper oxidase, from Klebsiella sp. 601. Can. J. Microbiol. 2008, 54, 725–733. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Zuo, W.; Li, Y.; Wang, X. Cloning of multicopper oxidase gene from Ochrobactrum sp. 531 and characterization of its alkaline laccase activity towards phenolic substrates. Adv. Biol. Chem. 2012, 2, 248–255. [Google Scholar]
- Góralczyk-Bińkowska, A.; Jasińska, A.; Długoński, J. Characteristics and use of multicopper oxidases enzymes. Post. Mikrobiol. 2019, 58, 7–18. [Google Scholar] [CrossRef]
- Sengupta, B.; Chakraborty, S.; Crawford, M.; Taylor, J.M.; Blackmon, L.E.; Biswas, P.K.; Kramer, W.H. Characterization of diadzein-hemoglobin binding using optical spectroscopy and molecular dynamics simulations. Int. J. Biol. Macromol. 2012, 51, 250–258. [Google Scholar] [CrossRef] [PubMed]
- Haghighi-Poodeh, S.; Kurganov, B.; Navidpour, L.; Yaghmaei, P.; Ebrahim-Habibi, A. Characterization of arginine preventive effect on heat-induced aggregation of insulin. Int. J. Biol. Macromol. 2020, 145, 1039–1048. [Google Scholar] [CrossRef]
- Chen, L.; Chen, J.; Ren, J.; Zhao, M. Effects of ultrasound pretreatment on the enzymatic hydrolysis of soy protein isolates and on the emulsifying properties of hydrolysates. J. Agric. Food Chem. 2011, 59, 2600–2609. [Google Scholar] [CrossRef]
- Perez, A.A.; Sánchez, C.C.; Patino, J.M.R.; Rubiolo, A.C.; Santiago, L.G. Effect of enzymatic hydrolysis and polysaccharide addition on the β-lactoglobulin adsorption at the air-water interface. J. Food Eng. 2012, 109, 712–720. [Google Scholar] [CrossRef]
- Nelis, M.; Decraecker, L.; Boeckxstaens, G.; Augustijns, P.; Cabooter, D. Development of a HILIC-MS/MS method for the quantification of histamine and its main metabolites in human urine samples. Talanta 2020, 220, 121328. [Google Scholar] [CrossRef]
- Xu, Y.; Liu, Y.; Xu, B.; Wang, D.; Jiang, W. Characterisation and application of Halomonas shantousis SWA25, a halotolerant bacterium with multiple biogenic amine degradation activity. Food Addit. Contam. Part A 2016, 33, 674–682. [Google Scholar]
- Hoegger, P.J.; Kilaru, S.; James, T.Y.; Thacker, J.R.; Kües, U. Phylogenetic comparison and classification of laccase and related multicopper oxidase protein sequences. FEBS J. 2006, 273, 2308–2326. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Lu, S. The importance of amine-degrading enzymes on the biogenic amine degradation in fermented foods: A review. Process Biochem. 2020, 99, 331–339. [Google Scholar] [CrossRef]
- Yoon, J.; Solomon, E.I. Electronic structures of exchange coupled trigonal trimeric Cu (II) complexes: Spin frustration, antisymmetric exchange, pseudo-A terms, and their relation to O2 activation in the multicopper oxidases. Coord. Chem. Rev. 2007, 251, 379–400. [Google Scholar] [CrossRef]
- Kaur, K.; Sidhu, H.; Capalash, N.; Sharma, P. Multicopper oxidase of Acinetobacter baumannii: Assessing its role in metal homeostasis, stress management and virulence. Microb. Pathog. 2020, 143, 104124. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, X.; Feng, X.; Lin, Y.; Gou, H.; Zhang, Y.; Yang, L. Degradation of polyethylene by Klebsiella pneumoniae Mk-1 isolated from soil. Ecotoxicol. Environ. Saf. 2023, 258, 114965. [Google Scholar] [CrossRef] [PubMed]
- Callejón, S.; Sendra, R.; Ferrer, S.; Pardo, I. Cloning and characterization of a new laccase from Lactobacillus plantarum J16 CECT 8944 catalyzing biogenic amines degradation. Appl. Microbiol. Biotechnol. 2016, 100, 3113–3124. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Kim, K.S.; Lee, J.H.; Lee, Y.C. Cloning, expression in Escherichia coli, and enzymatic properties of laccase from Aeromonas hydrophila WL-11. J. Environ. Sci. 2010, 22, 635–640. [Google Scholar] [CrossRef]
- Brander, S.; Mikkelsen, J.D.; Kepp, K.P. Characterization of an alkali-and halide-resistant laccase expressed in E. coli: CotA from Bacillus clausii. PLoS ONE 2014, 9, e99402. [Google Scholar] [CrossRef]
- Wang, X.; Zhao, Y.; Zhang, S.; Lin, X.; Liang, H.; Chen, Y.; Ji, C. Heterologous expression of the Lactobacillus sakei multiple copper oxidase to degrade histamine and tyramine at different environmental conditions. Foods 2022, 11, 3306. [Google Scholar] [CrossRef]
- Li, B.; Wang, Y.; Xue, L.; Lu, S. Heterologous expression and application of multicopper oxidases from Enterococcus spp. for degradation of biogenic amines. Protein Pept. Lett. 2021, 28, 183–194. [Google Scholar] [CrossRef] [PubMed]
- Jayasimha, A.; Mudambi, R.; Pavan, P.; Lokaksha, B.M.; Bankapur, S.; Patil, N. An effective feature extraction with deep neural network architecture for protein-secondary-structure prediction. Netw. Model. Anal. Health Inform. Bioinform. 2021, 10, 58. [Google Scholar] [CrossRef]
- Kellogg, E.H.; Leaver-Fay, A.; Baker, D. Role of conformational sampling in computing mutation-induced changes in protein structure and stability. Proteins Struct. Funct. Bioinform. 2011, 79, 830–838. [Google Scholar] [CrossRef]
- Taheri-Kafrani, A.; Choiset, Y.; Faizullin, D.A.; Zuev, Y.F.; Bezuglov, V.V.; Chobert, J.M.; Bordbar, A.K.; Haertlé, T. Interactions of β-lactoglobulin with serotonin and arachidonyl serotonin. Biopolymers 2011, 95, 871–880. [Google Scholar] [CrossRef]
- Komori, H.; Higuchi, Y. Structural insights into the O2 reduction mechanism of multicopper oxidase. J. Biochem. 2015, 158, 293–298. [Google Scholar] [CrossRef]
- Sumner, S.S.; Taylor, S.L. Detection method for histamine-producing, dairy-related bacteria using diamine oxidase and leucocrystal violet. J. Food Prot. 1989, 52, 105–108. [Google Scholar] [CrossRef]
- Sitthisak, S.; Howieson, K.; Amezola, C.; Jayaswal, R.K. Characterization of a multicopper oxidase gene from Staphylococcus aureus. Appl. Environ. Microbiol. 2005, 71, 5650–5653. [Google Scholar] [CrossRef] [PubMed]
- Callejón, S.; Sendra, R.; Ferrer, S.; Pardo, I. Identification of a novel enzymatic activity from lactic acid bacteria able to degrade biogenic amines in wine. Appl. Environ. Microbiol. 2014, 98, 185–198. [Google Scholar] [CrossRef]
- Zofair, S.F.F.; Ahmad, S.; Hashmi, M.A.; Khan, S.H.; Khan, M.A.; Younus, H. Catalytic roles, immobilization and management of recalcitrant environmental pollutants by laccases: Significance in sustainable green chemistry. J. Environ. Manag. 2022, 309, 114676. [Google Scholar] [CrossRef]



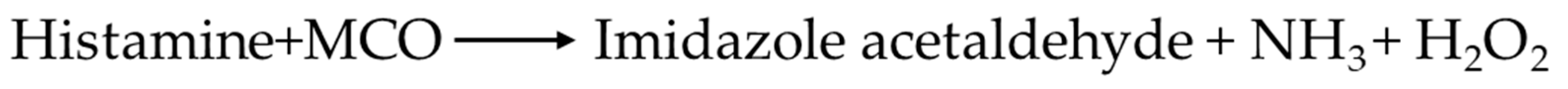
| Protein Name | PI | MW (kDa) | Peptide Sequence |
|---|---|---|---|
| MCO | 4.93 | 57.10 | EFHDDNQFDYR GLATMVIIKDDVEDOLPLPR EFHDDNQFDYR |
| Sample | Volume (mL) | Total Enzyme Activity (U) | Amount of Protein (mg) | Specific Activity (U/mg) | Recovery (%) | Purification Fold |
|---|---|---|---|---|---|---|
| Before purification | 85 | 7.62 | 362.95 | 0.02 | 100 | 1 |
| After purification | 40 | 2.43 | 19.6 | 0.12 | 31.89 | 6.91 |
| Enzyme | pH | Km (mmol/L) | Kcat (s−1) | Kcat/Km (mmol/L·s) |
|---|---|---|---|---|
| MCO | 3.5 | 2.18 ± 0.13 | 3.77 ± 0.09 | 1.73 ± 0.16 |
| Histamine Concentration (mg/L) | |||||
|---|---|---|---|---|---|
| 0 | 50 | 100 | 200 | 400 | |
| Alpha-helix (%) | 2.9 ± 0.02 | 3.0 ± 0.01 | 2.9 ± 0.02 | 3.1 ± 0.03 | 3.2 ± 0.01 |
| Beta-folding (%) | 39.7 ± 0.15 | 39.9 ± 0.24 | 40.1 ± 0.17 | 40.4 ± 0.26 | 41.8 ± 0.22 |
| Beta-turn (%) | 21.2 ± 0.12 | 21.3 ± 0.08 | 21.2 ± 0.15 | 21.2 ± 0.04 | 21.3 ± 0.09 |
| Random coil (%) | 36.1 ± 0.14 | 35.7 ± 0.31 | 35.7 ± 0.27 | 35.2 ± 0.22 | 33.6 ± 0.19 |
| Compound Name | Molecular Formula | Precursor Ion (m/z) | Production (m/z) | Mass Transition | CE (KV) | Cone Voltage (V) |
|---|---|---|---|---|---|---|
| Histamine | C5H9N3 | 112 | 94.4 | 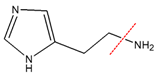 | 11 | 30 |
| Imidazole acetaldehyde | C5H6N2O | 111 | 82.9 | 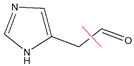 | 15 | 30 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pei, H.; Wang, Y.; He, W.; Deng, L.; Lan, Q.; Zhang, Y.; Yang, L.; Hu, K.; Li, J.; Liu, A.; et al. Research of Multicopper Oxidase and Its Degradation of Histamine in Lactiplantibacillus plantarum LPZN19. Microorganisms 2023, 11, 2724. https://doi.org/10.3390/microorganisms11112724
Pei H, Wang Y, He W, Deng L, Lan Q, Zhang Y, Yang L, Hu K, Li J, Liu A, et al. Research of Multicopper Oxidase and Its Degradation of Histamine in Lactiplantibacillus plantarum LPZN19. Microorganisms. 2023; 11(11):2724. https://doi.org/10.3390/microorganisms11112724
Chicago/Turabian StylePei, Huijie, Yilun Wang, Wei He, Lin Deng, Qinjie Lan, Yue Zhang, Lamei Yang, Kaidi Hu, Jianlong Li, Aiping Liu, and et al. 2023. "Research of Multicopper Oxidase and Its Degradation of Histamine in Lactiplantibacillus plantarum LPZN19" Microorganisms 11, no. 11: 2724. https://doi.org/10.3390/microorganisms11112724
APA StylePei, H., Wang, Y., He, W., Deng, L., Lan, Q., Zhang, Y., Yang, L., Hu, K., Li, J., Liu, A., Ao, X., Teng, H., Liu, S., Zou, L., Li, R., & Yang, Y. (2023). Research of Multicopper Oxidase and Its Degradation of Histamine in Lactiplantibacillus plantarum LPZN19. Microorganisms, 11(11), 2724. https://doi.org/10.3390/microorganisms11112724








