Sequencing of SARS-CoV-2 RNA Fragments in Wastewater Detects the Spread of New Variants during Major Events
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Extraction of Total Nucleic Acid from Wastewater Samples
2.3. Sequencing
2.4. Variant Mapping
3. Results
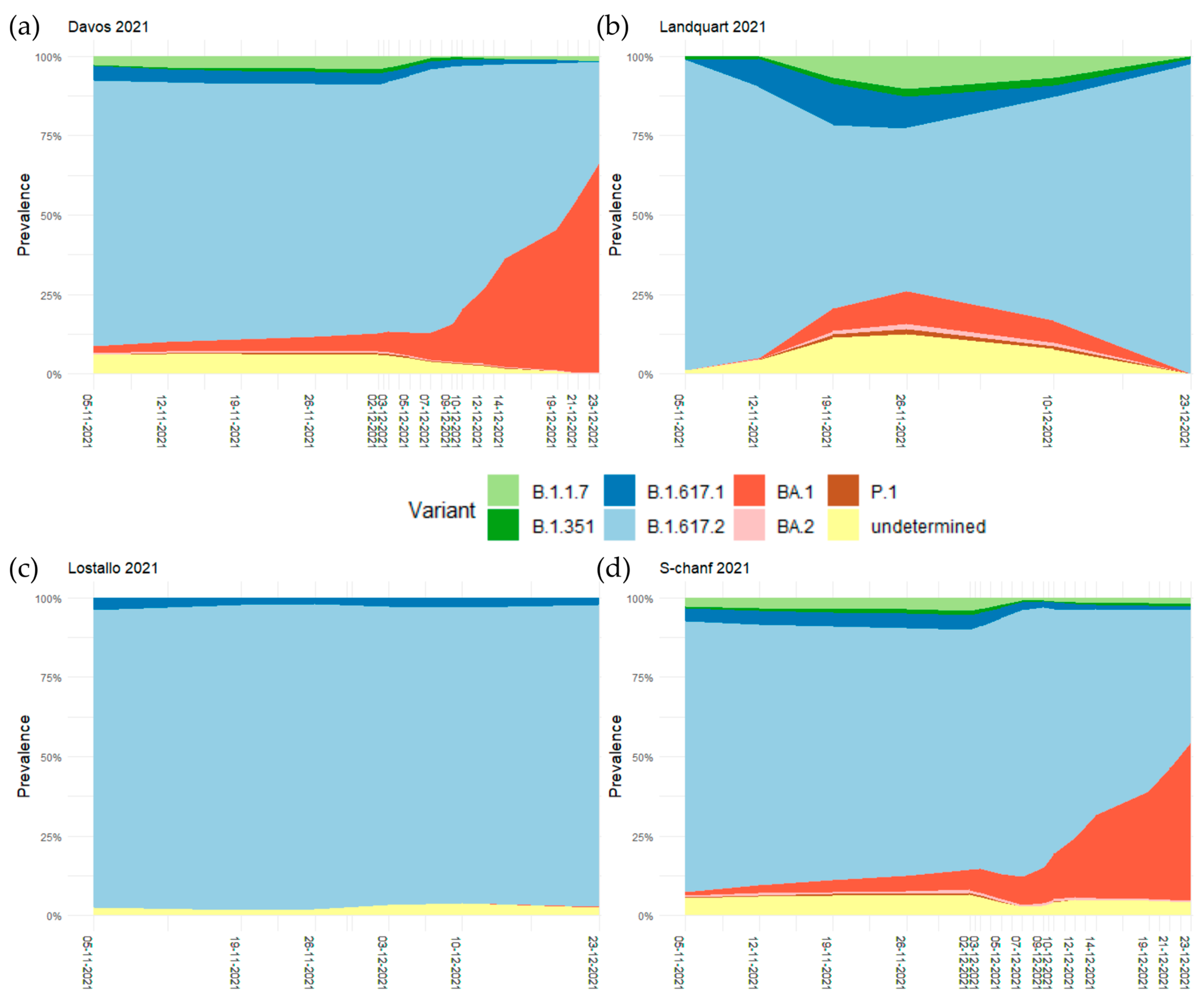
3.1. Spread of Omicron BA.1 during International Sports Competitions in December 2021
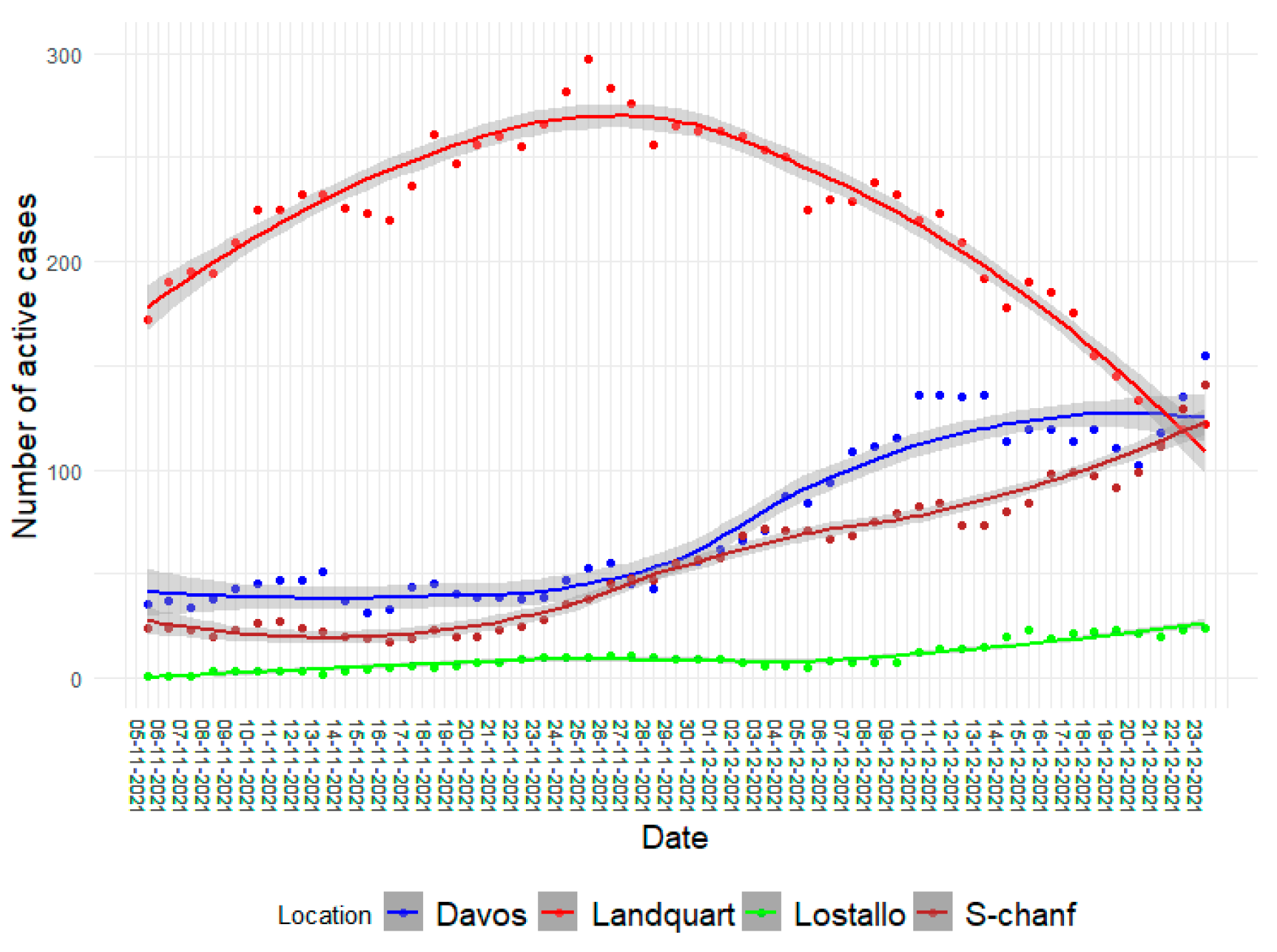
3.2. The Increased Relative Prevalence of Omicron BA.1 Caused Higher Infection Rates
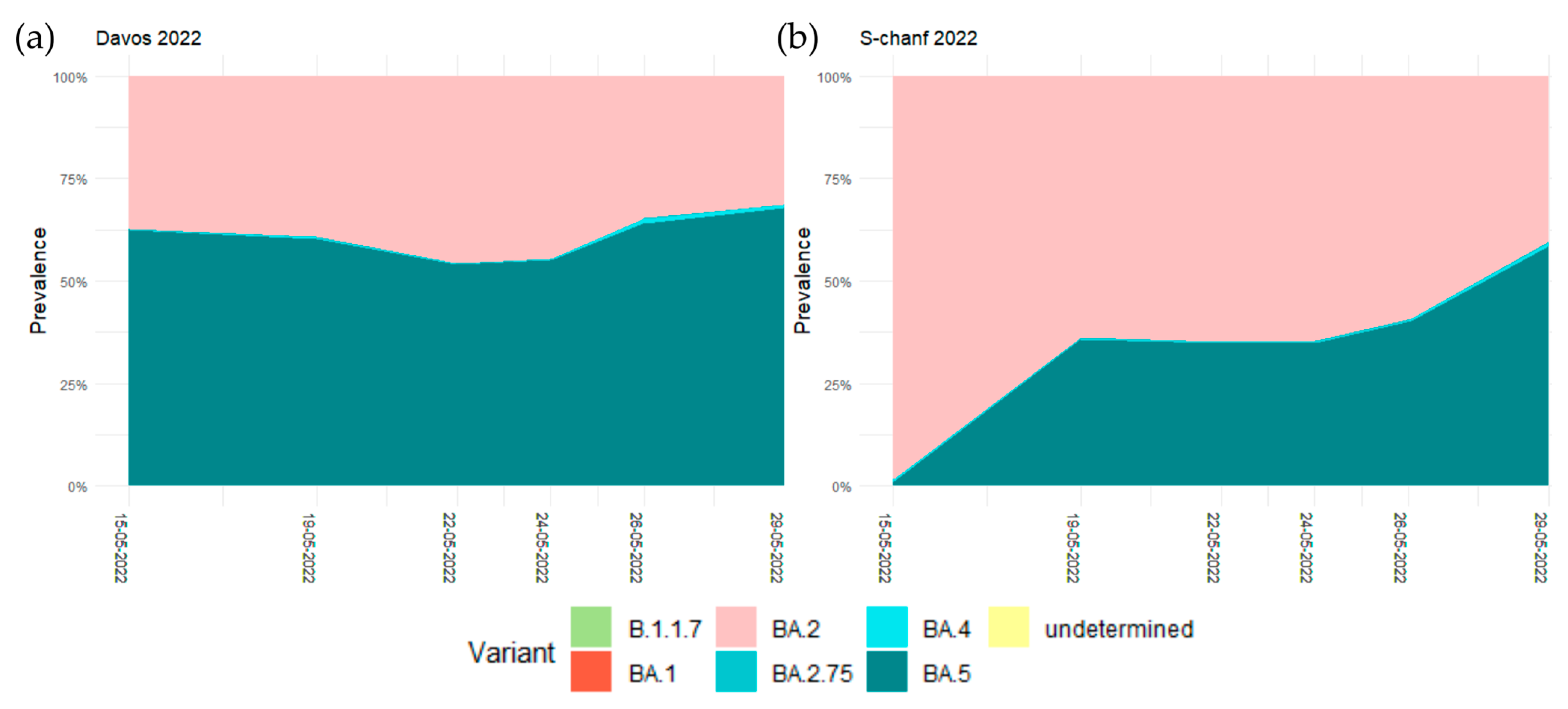
3.3. No Effect of the WEF in May 2022 on the Relative Prevalence of the Dominant Omicron Variants
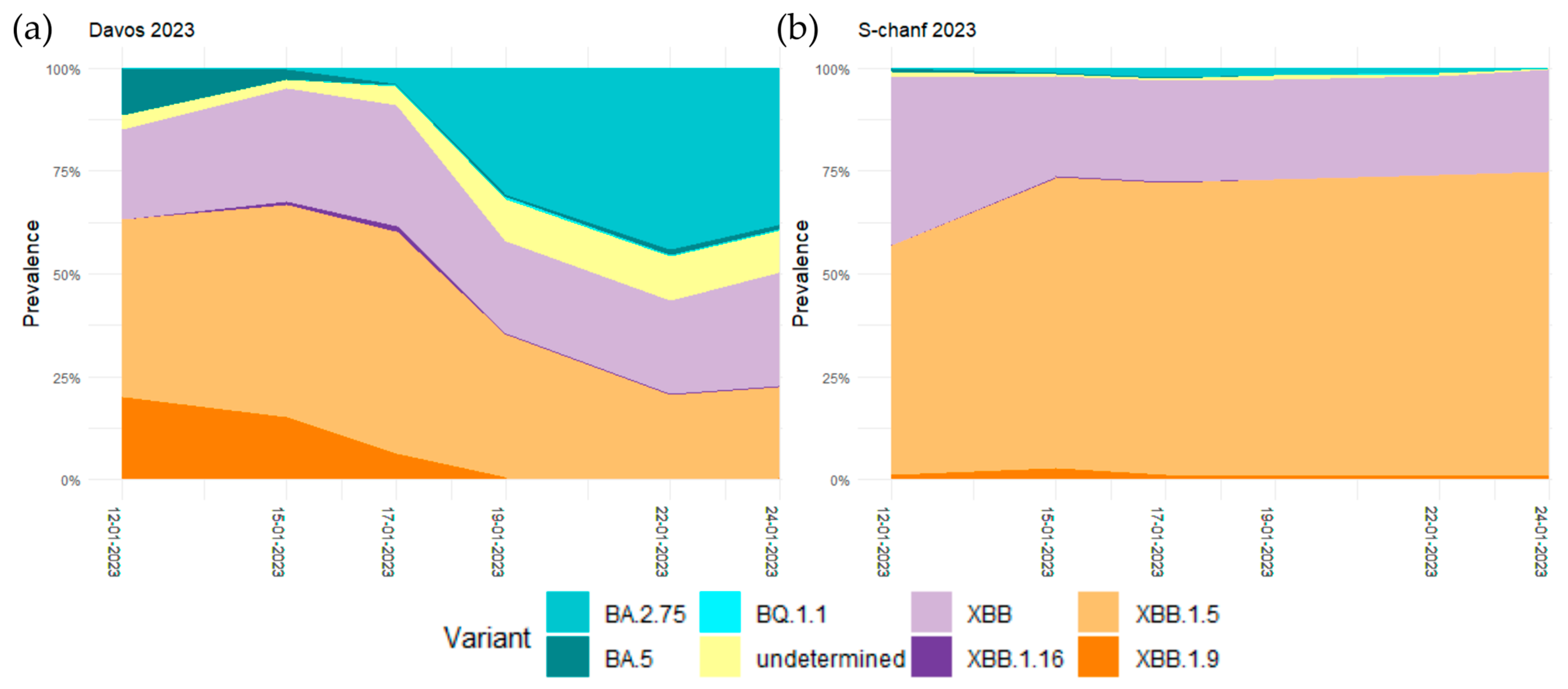
3.4. Arrival and Spread of Omicron BA.2.75 during WEF 2023 in Davos
4. Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gorbalenya, A.E.; Baker, S.C.; Baric, R.S.; de Groot, R.J.; Drosten, C.; Gulyaeva, A.A.; Haagmans, B.L.; Lauber, C.; Leontovich, A.M.; Neuman, B.W.; et al. The Species Severe Acute Respiratory Syndrome-Related Coronavirus: Classifying 2019-NCoV and Naming It SARS-CoV-2. Nat. Microbiol. 2020, 5, 536–544. [Google Scholar] [CrossRef]
- Zhou, P.; Yang, X.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A Pneumonia Outbreak Associated with a New Coronavirus of Probable Bat Origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef]
- Yan, R.; Zhang, Y.; Li, Y.; Xia, L.; Guo, Y.; Zhou, Q. Structural Basis for the Recognition of SARS-CoV-2 by Full-Length Human ACE2. Science 2020, 367, 1444–1448. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Zhong, L.; Deng, J.; Peng, J.; Dan, H.; Zeng, X.; Li, T.; Chen, Q. High Expression of ACE2 Receptor of 2019-NCoV on the Epithelial Cells of Oral Mucosa. Int. J. Oral Sci. 2020, 12, 8. [Google Scholar] [CrossRef]
- Radzikowska, U.; Ding, M.; Tan, G.; Zhakparov, D.; Peng, Y.; Wawrzyniak, P.; Wang, M.; Li, S.; Morita, H.; Altunbulakli, C.; et al. Distribution of ACE2, CD147, CD26, and Other SARS-CoV-2 Associated Molecules in Tissues and Immune Cells in Health and in Asthma, COPD, Obesity, Hypertension, and COVID-19 Risk Factors. Allergy Eur. J. Allergy Clin. Immunol. 2020, 75, 2829–2845. [Google Scholar] [CrossRef]
- Wölfel, R.; Corman, V.M.; Guggemos, W.; Seilmaier, M.; Zange, S.; Müller, M.A.; Niemeyer, D.; Jones, T.C.; Vollmar, P.; Rothe, C.; et al. Virological Assessment of Hospitalized Patients with COVID-2019. Nature 2020, 581, 465–469. [Google Scholar] [CrossRef]
- Lescure, F.X.; Bouadma, L.; Nguyen, D.; Parisey, M.; Wicky, P.H.; Behillil, S.; Gaymard, A.; Bouscambert-Duchamp, M.; Donati, F.; Le Hingrat, Q.; et al. Clinical and Virological Data of the First Cases of COVID-19 in Europe: A Case Series. Lancet Infect. Dis. 2020, 20, e148. [Google Scholar] [CrossRef]
- Pan, Y.; Zhang, D.; Yang, P.; Poon, L.L.M.; Wang, Q. Viral Load of SARS-CoV-2 in Clinical Samples. Lancet Infect. Dis. 2020, 20, 411–412. [Google Scholar] [CrossRef]
- Kitajima, M.; Ahmed, W.; Bibby, K.; Carducci, A.; Gerba, C.P.; Hamilton, K.A.; Haramoto, E.; Rose, J.B. SARS-CoV-2 in Wastewater: State of the Knowledge and Research Needs. Sci. Total Environ. 2020, 739, 139076. [Google Scholar] [CrossRef]
- Rimoldi, S.G.; Stefani, F.; Gigantiello, A.; Polesello, S.; Comandatore, F.; Mileto, D.; Maresca, M.; Longobardi, C.; Mancon, A.; Romeri, F.; et al. Presence and Infectivity of SARS-CoV-2 Virus in Wastewaters and Rivers. Sci. Total Environ. 2020, 744, 140911. [Google Scholar] [CrossRef] [PubMed]
- Schenk, H.; Heidinger, P.; Insam, H.; Kreuzinger, N.; Markt, R.; Nägele, F.; Oberacher, H.; Scheffknecht, C.; Steinlechner, M.; Vogl, G.; et al. Prediction of Hospitalisations Based on Wastewater-Based SARS-CoV-2 Epidemiology. Sci. Total Environ. 2023, 873, 162149. [Google Scholar] [CrossRef] [PubMed]
- Mitranescu, A.; Uchaikina, A.; Kau, A.S.; Stange, C.; Ho, J.; Tiehm, A.; Wurzbacher, C.; Drewes, J.E. Wastewater-Based Epidemiology for SARS-CoV-2 Biomarkers: Evaluation of Normalization Methods in Small and Large Communities in Southern Germany. ACS EST Water 2022, 2, 2460–2470. [Google Scholar] [CrossRef] [PubMed]
- Feng, S.; Roguet, A.; McClary-Gutierrez, J.S.; Newton, R.J.; Kloczko, N.; Meiman, J.G.; McLellan, S.L. Evaluation of Sampling, Analysis, and Normalization Methods for SARS-CoV-2 Concentrations in Wastewater to Assess COVID-19 Burdens in Wisconsin Communities. ACS EST Water 2021, 1, 1955–1965. [Google Scholar] [CrossRef]
- Jahn, K.; Dreifuss, D.; Topolsky, I.; Kull, A.; Ganesanandamoorthy, P.; Fernandez-Cassi, X.; Bänziger, C.; Devaux, A.J.; Stachler, E.; Caduff, L.; et al. Early Detection and Surveillance of SARS-CoV-2 Genomic Variants in Wastewater Using COJAC. Nat. Microbiol. 2022, 7, 1151–1160. [Google Scholar] [CrossRef]
- La Rosa, G.; Mancini, P.; Bonanno Ferraro, G.; Veneri, C.; Iaconelli, M.; Lucentini, L.; Bonadonna, L.; Brusaferro, S.; Brandtner, D.; Fasanella, A.; et al. Rapid Screening for SARS-CoV-2 Variants of Concern in Clinical and Environmental Samples Using Nested RT-PCR Assays Targeting Key Mutations of the Spike Protein. Water Res. 2021, 197, 117104. [Google Scholar] [CrossRef]
- Crits-Christoph, A.; Kantor, R.S.; Olm, M.R.; Whitney, O.N.; Al-Shayeb, B.; Lou, Y.C.; Flamholz, A.; Kennedy, L.C.; Greenwald, H.; Hinkle, A.; et al. Genome Sequencing of Sewage Detects Regionally Prevalent SARS-CoV-2 Variants. mBio 2021, 12, e02703-20. [Google Scholar] [CrossRef]
- Martin, J.; Klapsa, D.; Wilton, T.; Zambon, M.; Bentley, E.; Bujaki, E.; Fritzsche, M.; Mate, R.; Majumdar, M. Tracking SARS-CoV-2 in Sewage: Evidence of Changes in Virus Variant Predominance during COVID-19 Pandemic. Viruses 2020, 12, 1144. [Google Scholar] [CrossRef]
- Rouchka, E.C.; Chariker, J.H.; Saurabh, K.; Waigel, S.; Zacharias, W.; Zhang, M.; Talley, D.; Santisteban, I.; Puccio, M.; Moyer, S.; et al. The Rapid Assessment of Aggregated Wastewater Samples for Genomic Surveillance of SARS-Cov-2 on a City-Wide Scale. Pathogens 2021, 10, 1271. [Google Scholar] [CrossRef]
- Schumann, V.-F.; Ricardo De Castro Cuadrat, R.; Wyler, E.; Wurmus, R.; Deter, A.; Quedenau, C.; Dohmen, J.; Faxel, M.; Borodina, T.; Blume, A.; et al. SARS-CoV-2 Infection Dynamics Revealed by Wastewater Sequencing Analysis and Deconvolution. Sci. Total Environ. 2022, 853, 158931. [Google Scholar] [CrossRef]
- Karthikeyan, S.; Levy, J.I.; De Hoff, P.; Humphrey, G.; Birmingham, A.; Jepsen, K.; Farmer, S.; Tubb, H.M.; Valles, T.; Tribelhorn, C.E.; et al. Wastewater Sequencing Reveals Early Cryptic SARS-CoV-2 Variant Transmission. Nature 2022, 609, 101–108. [Google Scholar] [CrossRef]
- Markov, P.V.; Ghafari, M.; Beer, M.; Lythgoe, K.; Simmonds, P.; Stilianakis, N.I.; Katzourakis, A. The Evolution of SARS-CoV-2. Nat. Rev. Microbiol. 2023, 21, 361–379. [Google Scholar] [CrossRef]
- Harvey, W.T.; Carabelli, A.M.; Jackson, B.; Gupta, R.K.; Thomson, E.C.; Harrison, E.M.; Ludden, C.; Reeve, R.; Rambaut, A.; Peacock, S.J.; et al. SARS-CoV-2 Variants, Spike Mutations and Immune Escape. Nat. Rev. Microbiol. 2021, 19, 409–424. [Google Scholar] [CrossRef] [PubMed]
- Martin, D.P.; Weaver, S.; Tegally, H.; San, J.E.; Shank, S.D.; Wilkinson, E.; Lucaci, A.G.; Giandhari, J.; Naidoo, S.; Pillay, Y.; et al. The Emergence and Ongoing Convergent Evolution of the SARS-CoV-2 N501Y Lineages. Cell 2021, 184, 5189–5200.e7. [Google Scholar] [CrossRef] [PubMed]
- Rochman, N.D.; Wolf, Y.I.; Faure, G.; Mutz, P.; Zhang, F.; Koonin, E.V. Ongoing Global and Regional Adaptive Evolution of SARS-CoV-2. Proc. Natl. Acad. Sci. USA 2021, 118, 2104241118. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.; Jian, F.; Wang, J.; Yu, Y.; Song, W.; Yisimayi, A.; Wang, J.; An, R.; Chen, X.; Zhang, N.; et al. Imprinted SARS-CoV-2 Humoral Immunity Induces Convergent Omicron RBD Evolution. Nature 2023, 614, 521–529. [Google Scholar] [CrossRef]
- Bloom, J.D.; Neher, R.A. Fitness Effects of Mutations to SARS-CoV-2 Proteins. bioRxiv 2023. [Google Scholar] [CrossRef] [PubMed]
- Planas, D.; Bruel, T.; Staropoli, I.; Guivel-Benhassine, F.; Porrot, F.; Maes, P.; Grzelak, L.; Prot, M.; Mougari, S.; Planchais, C.; et al. Resistance of Omicron Subvariants BA.2.75.2, BA.4.6, and BQ.1.1 to Neutralizing Antibodies. Nat. Commun. 2023, 14, 824. [Google Scholar] [CrossRef]
- Tian, D.; Sun, Y.; Xu, H.; Ye, Q. The Emergence and Epidemic Characteristics of the Highly Mutated SARS-CoV-2 Omicron Variant. J. Med. Virol. 2022, 94, 2376–2383. [Google Scholar] [CrossRef]
- Shu, Y.; McCauley, J.W. Gisaid: Global initiative on sharing all influenza data—From vision to reality. Eurosurveillance 2017, 22. [Google Scholar] [CrossRef]
- Benson, D.A.; Cavanaugh, M.; Clark, K.; Karsch-Mizrachi, I.; Lipman, D.J.; Ostell, J.; Sayers, E.W. GenBank. Nucleic Acids Res. 2013, 41, D36–D42. [Google Scholar] [CrossRef] [PubMed]
- Hadfield, J.; Megill, C.; Bell, S.M.; Huddleston, J.; Potter, B.; Callender, C.; Sagulenko, P.; Bedford, T.; Neher, R.A. NextStrain: Real-Time Tracking of Pathogen Evolution. Bioinformatics 2018, 34, 4121–4123. [Google Scholar] [CrossRef] [PubMed]
- Hodcroft, E. CoVariants: SARS-CoV-2 Mutations and Variants of Interest. Available online: https://covariants.org/ (accessed on 12 October 2023).
- Mondal, S.; Feirer, N.; Brockman, M.; Preston, M.A.; Teter, S.J.; Ma, D.; Goueli, S.A.; Moorji, S.; Saul, B.; Cali, J.J. A Direct Capture Method for Purification and Detection of Viral Nucleic Acid Enables Epidemiological Surveillance of SARS-CoV-2. Sci. Total Environ. 2021, 795, 148834. [Google Scholar] [CrossRef] [PubMed]
- Ulhuq, F.R.; Barge, M.; Falconer, K.; Wild, J.; Fernandes, G.; Gallagher, A.; McGinley, S.; Sugadol, A.; Tariq, M.; Maloney, D.; et al. Analysis of the ARTIC V4 and V4.1 SARS-CoV-2 Primers and Their Impact on the Detection of Omicron BA.1 and BA.2 Lineage-Defining Mutations. Microb. Genom. 2023, 9, 000991. [Google Scholar] [CrossRef] [PubMed]
- Posada-Cespedes, S.; Seifert, D.; Topolsky, I.; Jablonski, K.P.; Metzner, K.J.; Beerenwinkel, N. V-Pipe: A Computational Pipeline for Assessing Viral Genetic Diversity from High-Throughput Data. Bioinformatics 2021, 37, 1673–1680. [Google Scholar] [CrossRef]
- Dreifuss, D.; Topolsky, I.; Baykal, P.I.; Beerenwinkel, N. Tracking SARS-CoV-2 Genomic Variants in Wastewater Sequencing Data with LolliPop. medRxiv 2022. [Google Scholar] [CrossRef]
- Gorji, H.; Lunati, I.; Rudolf, F.; Vidondo, B.; Hardt, W.D.; Jenny, P.; Engel, D.; Schneider, J.; Jamnicki, M.; Leuthold, R.; et al. Results from Canton Grisons of Switzerland Suggest Repetitive Testing Reduces SARS-CoV-2 Incidence (February–March 2021). Sci. Rep. 2022, 12, 19538. [Google Scholar] [CrossRef]
- Fontenele, R.S.; Kraberger, S.; Hadfield, J.; Driver, E.M.; Bowes, D.; Holland, L.R.A.; Faleye, T.O.C.; Adhikari, S.; Kumar, R.; Inchausti, R.; et al. High-Throughput Sequencing of SARS-CoV-2 in Wastewater Provides Insights into Circulating Variants. Water Res 2021, 205, 117710. [Google Scholar] [CrossRef]
- Wurtz, N.; Revol, O.; Jardot, P.; Giraud-Gatineau, A.; Houhamdi, L.; Soumagnac, C.; Annessi, A.; Lacoste, A.; Colson, P.; Aherfi, S.; et al. Monitoring the Circulation of SARS-CoV-2 Variants by Genomic Analysis Ofwastewater in Marseille, South-East France. Pathogens 2021, 10, 1042. [Google Scholar] [CrossRef]
- Pérez-Cataluña, A.; Chiner-Oms, Á.; Cuevas-Ferrando, E.; Díaz-Reolid, A.; Falcó, I.; Randazzo, W.; Girón-Guzmán, I.; Allende, A.; Bracho, M.A.; Comas, I.; et al. Spatial and Temporal Distribution of SARS-CoV-2 Diversity Circulating in Wastewater. Water Res. 2022, 211, 118007. [Google Scholar] [CrossRef]
- Agrawal, S.; Orschler, L.; Schubert, S.; Zachmann, K.; Heijnen, L.; Tavazzi, S.; Gawlik, B.M.; de Graaf, M.; Medema, G.; Lackner, S. Prevalence and Circulation Patterns of SARS-CoV-2 Variants in European Sewage Mirror Clinical Data of 54 European Cities. Water Res. 2022, 214, 118162. [Google Scholar] [CrossRef] [PubMed]
- Marcel, S.; Christian, A.L.; Richard, N.; Silvia, S.; Emma, H.; Jacques, F.; Marcel, Z.; Gabriela, S.; Manuel, B.; Annelies, W.S.; et al. COVID-19 Epidemic in Switzerland: On the Importance of Testing, Contact Tracing and Isolation. Swiss Med. Wkly. 2020, 150, w20225. [Google Scholar] [CrossRef]
- Cucinotta, D.; Vanelli, M. WHO Declares COVID-19 a Pandemic. Acta Biomed. 2020, 91, 157–160. [Google Scholar] [PubMed]
| 8 November 2021–22 November 2021 | ||
| Variant | Num seq | Frequency |
| 21J (Delta) | 307 | 0.95 |
| 21I (Delta) | 14 | 0.04 |
| 21A (Delta) (B.1.617.2) | 1 | 0.00 |
| 22 November 2021–6 December 2021 | ||
| Variant | Num seq | Frequency |
| 21J (Delta) | 241 | 0.96 |
| 21I (Delta) | 8 | 0.03 |
| 21K (Omicron) (BA.1) | 3 | 0.01 |
| 6 December 2021–20 December 2021 | ||
| Variant | Num seq | Frequency |
| 21J (Delta) | 163 | 0.67 |
| 21K (Omicron) (BA.1) | 76 | 0.31 |
| 21I (Delta) | 3 | 0.01 |
| 20 December 2021–3 January 2022 | ||
| Variant | Num seq | Frequency |
| 21K (Omicron) (BA.1) | 410 | 0.86 |
| 21J (Delta) | 66 | 0.14 |
| 21I (Delta) | 1 | 0.00 |
| Asia | Eastern Asia | South Korea | 0.53 |
| Mauritius | 0.45 | ||
| South-eastern Asia | Viet Nam | 0.92 | |
| Thailand | 0.84 | ||
| Cambodia | 0.44 | ||
| Singapore | 0.19 | ||
| Malaysia | 0.14 | ||
| Western Asia | Georgia | 0.65 | |
| Qatar | 0.28 | ||
| Europe | Eastern Europe | Romania | 0.20 |
| Bulgaria | 0.19 | ||
| Czech Republic | 0.17 | ||
| Slovakia | 0.15 | ||
| Northern Europe | Lithuania | 0.22 | |
| Iceland | 0.18 | ||
| Denmark | 0.17 | ||
| Southern Europe | Greece | 0.15 | |
| Croatia | 0.12 | ||
| Oceania | Australia/ New Zealand | Australia | 0.33 |
| New Zealand | 0.17 |
| 19 December 2022–2 January 2023 | ||
| Variant | Num seq | Frequency |
| 22E (Omicron) (BQ.1) | 32 | 0.71 |
| 22B (Omicron) (BA.5) | 11 | 0.24 |
| others | 1 | 0.02 |
| 23C (Omicron) (CH.1.1) | 1 | 0.02 |
| 2 January 2023–16 January 2023 | ||
| Variant | Num seq | Frequency |
| 22E (Omicron) (BQ.1) | 26 | 0.72 |
| 22B (Omicron) (BA.5) | 4 | 0.11 |
| 22F (Omicron) (XBB) | 2 | 0.06 |
| 22D (Omicron) (BA.2.75) | 2 | 0.06 |
| 23C (Omicron) (CH.1.1) | 1 | 0.03 |
| 21L (Omicron) (BA.2) | 1 | 0.03 |
| Africa | 139 | North Africa | 20 |
| Eastern Africa | 38 | ||
| Middle Africa | 6 | ||
| Southern Africa | 52 | ||
| Western Africa | 23 | ||
| Americas | 881 | Caribbean | 4 |
| Central America | 28 | ||
| South America | 88 | ||
| Northern America | 761 | ||
| Asia | 622 | Central Asia | 2 |
| Eastern Asia | 184 | ||
| South-eastern Asia | 106 | ||
| Southern Asia | 117 | ||
| Western Asia | 213 | ||
| Europe | 992 | Eastern Europe | 32 |
| Northern Europe | 338 | ||
| Southern Europe | 71 | ||
| Western Europe | 551 | ||
| Oceania | 19 | Australia/New Zealand | 18 |
| Melanesia | 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhakparov, D.; Quirin, Y.; Xiao, Y.; Battaglia, N.; Holzer, M.; Bühler, M.; Kistler, W.; Engel, D.; Zumthor, J.P.; Caduff, A.; et al. Sequencing of SARS-CoV-2 RNA Fragments in Wastewater Detects the Spread of New Variants during Major Events. Microorganisms 2023, 11, 2660. https://doi.org/10.3390/microorganisms11112660
Zhakparov D, Quirin Y, Xiao Y, Battaglia N, Holzer M, Bühler M, Kistler W, Engel D, Zumthor JP, Caduff A, et al. Sequencing of SARS-CoV-2 RNA Fragments in Wastewater Detects the Spread of New Variants during Major Events. Microorganisms. 2023; 11(11):2660. https://doi.org/10.3390/microorganisms11112660
Chicago/Turabian StyleZhakparov, Damir, Yves Quirin, Yi Xiao, Nicole Battaglia, Michael Holzer, Martin Bühler, Walter Kistler, Doortje Engel, Jon Paulin Zumthor, Alexa Caduff, and et al. 2023. "Sequencing of SARS-CoV-2 RNA Fragments in Wastewater Detects the Spread of New Variants during Major Events" Microorganisms 11, no. 11: 2660. https://doi.org/10.3390/microorganisms11112660





