Occurrence and Molecular Characterization of Potentially Pathogenic Vibrio spp. in Seafood Collected in Sicily
Abstract
1. Introduction
2. Materials and Methods
2.1. Isolation and Identification of Vibrio spp.
2.2. Molecular Analyses
2.3. Antimicrobial Susceptibility
2.4. Data Analysis
3. Results
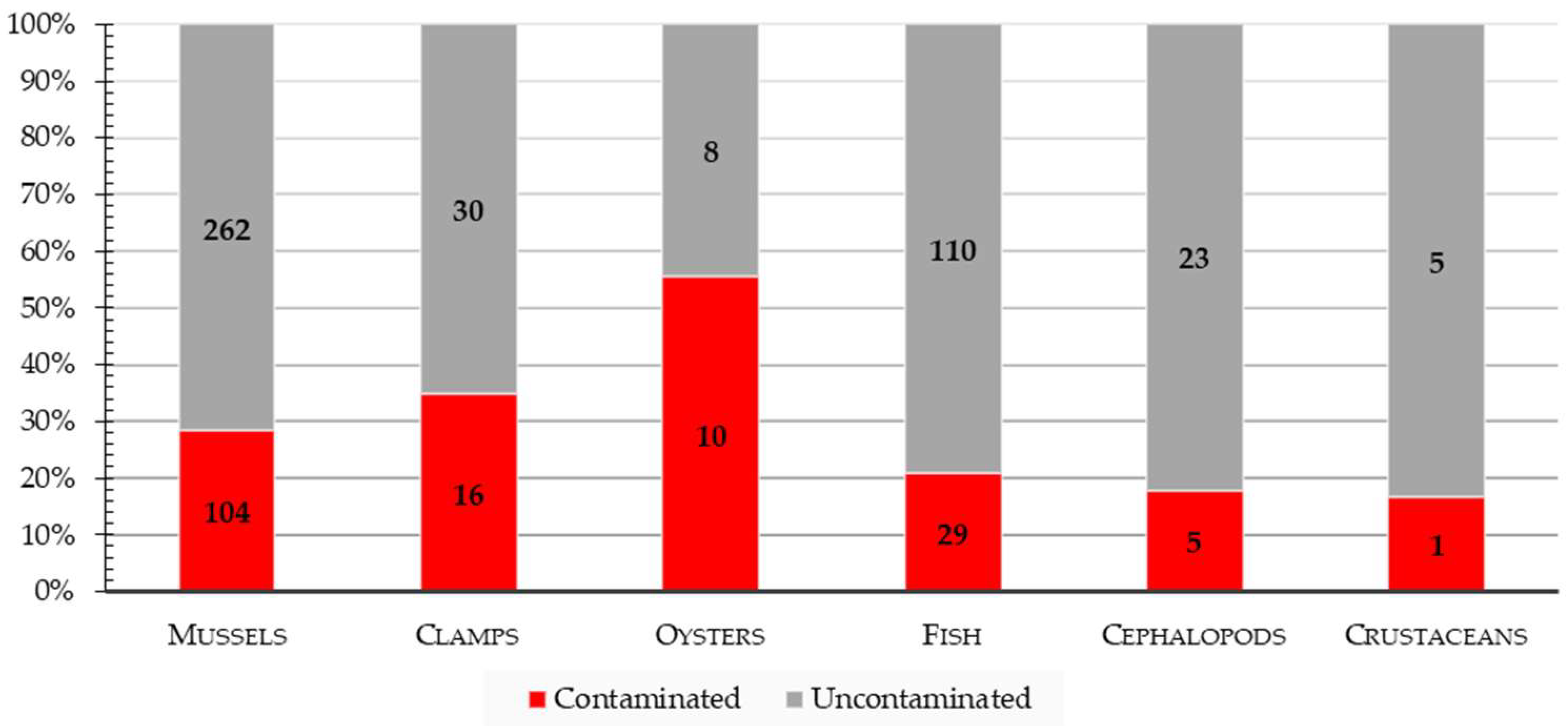
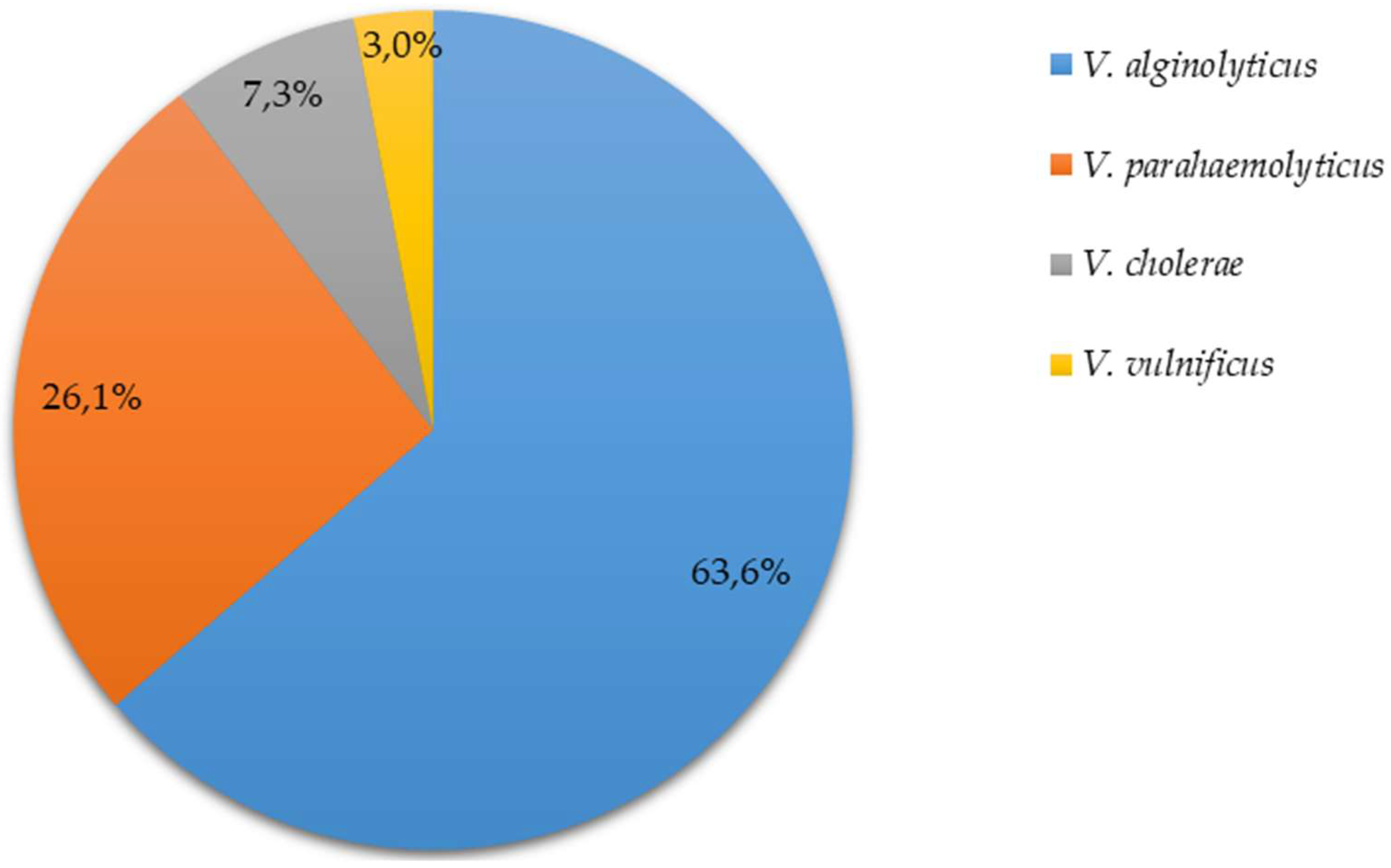
3.1. Isolation and Identification of Vibrio spp.
3.2. Molecular Analyses
3.3. Antimicrobial Susceptibility
- V. parahaemolyticus isolates were resistant to ampicillin, cefazolin, cephalothin, streptomycin, and tetracycline at the rate of 70%, 30.2%, 21%, 19%, and 14%, respectively. Intermediate resistance was revealed to colistin sulphate (21%), ceftazidime and kanamycin (18.6%), and cefotaxime (14%). A pattern of multidrug resistance was exhibited in 2/43 isolates (Table 2). In contrast, all isolates were sensitive to four antimicrobials (ceftriaxone, ciprofloxacin, chloramphenicol, and trimethoprim/sulfamethoxazole), and >90% of the isolates were susceptible to amoxicillin-clavulanic acid and gentamicin.
- V. cholerae isolates were resistant to ampicillin (75%), trimethoprim/sulfamethoxazole (42%), ceftriaxone, ceftazidime, and cephalothin (17%). Intermediate resistance to cephalothin was shown in 50% of the isolates, and 17% exhibited intermediate resistance to streptomycin and ceftazidime. All isolates were susceptible to cefotaxime, ciprofloxacin, chloramphenicol, and tetracycline.
- All tested V. vulnificus isolates (n = 5) were susceptible to streptomycin, and a large percentage of strains were sensitive to all the tested antibiotics including drugs recommended by the Center for Disease Control and Prevention (CDC) for the treatment of V. vulnificus infections: tetracycline, ciprofloxacin, and trimethoprim/sulfamethoxazole. One strain, whereas, showed intermediate sensitivity to tetracycline and ciprofloxacin and resistance to cefazolin.
- Among the 105 V. alginolyticus isolates, 88.6% were resistant to ampicillin and 22% to gentamicin. Intermediate resistance to streptomycin and kanamycin were also observed, showing an incidence of 55.2% and 10.5%, respectively. All isolates were sensitive to the remaining antimicrobials.
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Iwamoto, M.; Ayers, T.; Mahon, B.E.; Swerdlow, D.L. Epidemiology of Seafood-Associated Infections in the United States. Clin. Microbiol. Rev. 2010, 23, 399–411. [Google Scholar] [CrossRef] [PubMed]
- Normanno, G.; Parisi, A.; Addante, N.; Quaglia, N.C.; Dambrosio, A.; Montagna, C.; Chiocco, D. Vibrio parahaemolyticus, Vibrio vulnificus and Microorganisms of Fecal Origin in Mussels (Mytilus galloprovincialis) Sold in the Puglia Region (Italy). Int. J. Food Microbiol. 2006, 106, 219–222. [Google Scholar] [CrossRef] [PubMed]
- Passalacqua, P.L.; Zavatta, E.; Bignami, G.; Serraino, A.; Serratore, P. Occurrence of Vibrio Parahaemolyticus, Vibrio Cholerae and Vibrio Vulnificus in the Clam Ruditapes Philippinarum (Adams & Reeve, 1850) from Emilia Romagna and Sardinia, Italy. Ital. J. Food Saf. 2016, 5, 5709. [Google Scholar] [CrossRef] [PubMed]
- Brumfield, K.D.; Usmani, M.; Chen, K.M.; Gangwar, M.; Jutla, A.S.; Huq, A.; Colwell, R.R. Environmental Parameters Associated with Incidence and Transmission of Pathogenic Vibrio spp. Environ. Microbiol. 2021, 23, 7314–7340. [Google Scholar] [CrossRef]
- Ramamurthy, T.; Chowdhury, G.; Pazhani, G.P.; Shinoda, S. Vibrio fluvialis: An Emerging Human Pathogen. Front. Microbiol. 2014, 5, 91. [Google Scholar] [CrossRef] [PubMed]
- Caburlotto, G.; Bianchi, F.; Gennari, M.; Ghidini, V.; Socal, G.; Aubry, F.B.; Bastianini, M.; Tafi, M.; Lleo, M.M. Integrated Evaluation of Environmental Parameters Influencing Vibrio Occurrence in the Coastal Northern Adriatic Sea (Italy) Facing the Venetian Lagoon. Microb. Ecol. 2012, 63, 20–31. [Google Scholar] [CrossRef]
- Jeamsripong, S.; Khant, W.; Chuanchuen, R. Distribution of Phenotypic and Genotypic Antimicrobial Resistance and Virulence Genes in Vibrio Parahaemolyticus Isolated from Cultivated Oysters and Estuarine Water. FEMS Microbiol. Ecol. 2020, 96, fiaa081. [Google Scholar] [CrossRef]
- Ward, L.N.; Bej, A.K. Detection of Vibrio Parahaemolyticus in Shellfish by Use of Multiplexed Real-Time PCR with TaqMan Fluorescent Probes. Appl. Environ. Microbiol. 2006, 72, 2031–2042. [Google Scholar] [CrossRef]
- Pereira, C.S.; Possas, C.d.A.; Viana, C.M.; Rodrigues, D.d.P. Characteristics of Vibrio parahaemolyticus isolated from mussels (Perna perna) commercialized at Niterói, Rio de Janeiro. Rev. Soc. Bras. Med. Trop. 2007, 40, 56–59. [Google Scholar] [CrossRef]
- Silva, I.P.; Carneiro, C.d.S.; Saraiva, M.A.F.; Oliveira, T.A.S.d.; Sousa, O.V.d.; Evangelista-Barreto, N.S. Antimicrobial Resistance and Potential Virulence of Vibrio Parahaemolyticus Isolated from Water and Bivalve Mollusks from Bahia, Brazil. Mar. Pollut. Bull. 2018, 131, 757–762. [Google Scholar] [CrossRef]
- Costa Sobrinho, P.d.S.; Destro, M.T.; Franco, B.D.G.M.; Landgraf, M. Occurrence and Distribution of Vibrio Parahaemolyticus in Retail Oysters in Sao Paulo State, Brazil. Food Microbiol. 2011, 28, 137–140. [Google Scholar] [CrossRef] [PubMed]
- Lamon, S.; Consolati, S.G.; Fois, F.; Cambula, M.G.; Pes, M.; Porcheddu, G.; Agus, V.; Esposito, G.; Mureddu, A.; Meloni, D. Occurrence, Seasonal Distribution, and Molecular Characterization of Vibrio Vulnificus, Vibrio Cholerae, and Vibrio Parahaemolyticus in Shellfish (Mytilus Galloprovincialis and Ruditapes Decussatus) Collected in Sardinia (Italy). J. Food Prot. 2019, 82, 1851–1856. [Google Scholar] [CrossRef] [PubMed]
- Baker-Austin, C.; Trinanes, J.A.; Salmenlinna, S.; Löfdahl, M.; Siitonen, A.; Taylor, N.G.H.; Martinez-Urtaza, J. Heat Wave–Associated Vibriosis, Sweden and Finland, 2014. Emerg. Infect. Dis. 2016, 22, 5. [Google Scholar] [CrossRef]
- Lutz, C.; Erken, M.; Noorian, P.; Sun, S.; McDougald, D. Environmental Reservoirs and Mechanisms of Persistence of Vibrio Cholerae. Front. Microbiol. 2013, 4, 375. [Google Scholar] [CrossRef]
- Clemens, J.D.; Nair, G.B.; Ahmed, T.; Qadri, F.; Holmgren, J. Cholera. Lancet 2017, 390, 1539–1549. [Google Scholar] [CrossRef] [PubMed]
- Pazzani, C.; Scrascia, M.; Dionisi, A.M.; Maimone, F.; Luzzi, I. Molecular Epidemiology and Origin of Cholera Reemergence in Italy and Albania in the 1990s. Res. Microbiol. 2006, 157, 508–512. [Google Scholar] [CrossRef]
- Baker-Austin, C.; Trinanes, J.; Gonzalez-Escalona, N.; Martinez-Urtaza, J. Non-Cholera Vibrios: The Microbial Barometer of Climate Change. Trends Microbiol. 2017, 25, 76–84. [Google Scholar] [CrossRef]
- Ottaviani, D.; Medici, L.; Talevi, G.; Napoleoni, M.; Serratore, P.; Zavatta, E.; Bignami, G.; Masini, L.; Chierichetti, S.; Fisichella, S.; et al. Molecular Characterization and Drug Susceptibility of Non-O1/O139 V. cholerae Strains of Seafood, Environmental and Clinical Origin, Italy. Food Microbiol. 2018, 72, 82–88. [Google Scholar] [CrossRef]
- Roux, F.L.; Wegner, K.M.; Baker-Austin, C.; Vezzulli, L.; Osorio, C.R.; Amaro, C.; Ritchie, J.M.; Defoirdt, T.; Destoumieux-Garzón, D.; Blokesch, M.; et al. The Emergence of Vibrio Pathogens in Europe: Ecology, Evolution, and Pathogenesis (Paris, 11–12th March 2015). Front. Microbiol. 2015, 6, 830. [Google Scholar] [CrossRef]
- Williams, T.C.; Froelich, B.; Oliver, J.D. A New Culture-Based Method for the Improved Identification of Vibrio Vulnificus from Environmental Samples, Reducing the Need for Molecular Confirmation. J. Microbiol. Methods 2013, 93, 277–283. [Google Scholar] [CrossRef]
- Han, F.; Wang, F.; Ge, B. Detecting Potentially Virulent Vibrio Vulnificus Strains in Raw Oysters by Quantitative Loop-Mediated Isothermal Amplification. Appl. Environ. Microbiol. 2011, 77, 2589–2595. [Google Scholar] [CrossRef] [PubMed]
- Jacobs Slifka, K.M.; Newton, A.E.; Mahon, B.E. Vibrio alginolyticus Infections in the USA, 1988–2012. Epidemiol. Infect. 2017, 145, 1491–1499. [Google Scholar] [CrossRef] [PubMed]
- Vu, T.T.T.; Alter, T.; Huehn, S. Prevalence of Vibrio Spp. in Retail Seafood in Berlin, Germany. J. Food Prot. 2018, 81, 593–597. [Google Scholar] [CrossRef] [PubMed]
- Dechet, A.M.; Yu, P.A.; Koram, N.; Painter, J. Nonfoodborne Vibrio Infections: An Important Cause of Morbidity and Mortality in the United States, 1997–2006. Clin. Infect. Dis. 2008, 46, 970–976. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Urtaza, J.; Trinanes, J.; Abanto, M.; Lozano-Leon, A.; Llovo-Taboada, J.; Garcia-Campello, M.; Pousa, A.; Powell, A.; Baker-Austin, C.; Gonzalez-Escalona, N. Epidemic Dynamics of Vibrio parahaemolyticus Illness in a Hotspot of Disease Emergence, Galicia, Spain. Emerg. Infect. Dis. 2018, 24, 852–859. [Google Scholar] [CrossRef]
- Lee, L.-H.; Ab Mutalib, N.-S.; Law, J.W.-F.; Wong, S.H.; Letchumanan, V. Discovery on Antibiotic Resistance Patterns of Vibrio Parahaemolyticus in Selangor Reveals Carbapenemase Producing Vibrio Parahaemolyticus in Marine and Freshwater Fish. Front. Microbiol. 2018, 9, 2513. [Google Scholar] [CrossRef]
- Bier, N.; Schwartz, K.; Guerra, B.; Strauch, E. Survey on Antimicrobial Resistance Patterns in Vibrio Vulnificus and Vibrio Cholerae Non-O1/Non-O139 in Germany Reveals Carbapenemase-Producing Vibrio Cholerae in Coastal Waters. Front. Microbiol. 2015, 6, 1179. [Google Scholar] [CrossRef]
- Canteón, R. Antibiotic Resistance Genes from the Environment: A Perspective through Newly Identified Antibiotic Resistance Mechanisms in the Clinical Setting. Clin. Microbiol. Infect. 2009, 15, 20–25. [Google Scholar] [CrossRef]
- Gillings, M.R.; Paulsen, I.T.; Tetu, S.G. Genomics and the Evolution of Antibiotic Resistance: Genomics and Antibiotic Resistance. Ann. N. Y. Acad. Sci. 2017, 1388, 92–107. [Google Scholar] [CrossRef]
- Christaki, E.; Marcou, M.; Tofarides, A. Antimicrobial Resistance in Bacteria: Mechanisms, Evolution, and Persistence. J. Mol. Evol. 2020, 88, 26–40. [Google Scholar] [CrossRef]
- Mok, J.S.; Ryu, A.; Kwon, J.Y.; Park, K.; Shim, K.B. Abundance, Antimicrobial Resistance, and Virulence of Pathogenic Vibrio Strains from Molluscan Shellfish Farms along the Korean Coast. Mar. Pollut. Bull. 2019, 149, 110559. [Google Scholar] [CrossRef] [PubMed]
- Cabello, F.C. Heavy Use of Prophylactic Antibiotics in Aquaculture: A Growing Problem for Human and Animal Health and for the Environment. Environ. Microbiol. 2006, 8, 1137–1144. [Google Scholar] [CrossRef] [PubMed]
- Shaw, K.S.; Rosenberg Goldstein, R.E.; He, X.; Jacobs, J.M.; Crump, B.C.; Sapkota, A.R. Antimicrobial Susceptibility of Vibrio Vulnificus and Vibrio Parahaemolyticus Recovered from Recreational and Commercial Areas of Chesapeake Bay and Maryland Coastal Bays. PLoS ONE 2014, 9, e89616. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.H.; Mok, J.S.; Jung, Y.J.; Lee, K.J.; Kwon, J.Y.; Park, K.; Moon, S.Y.; Kwon, S.J.; Ryu, A.R.; Lee, T.S. Distribution and Antimicrobial Susceptibility of Vibrio Species Associated with Zooplankton in Coastal Area of Korea. Mar. Pollut. Bull. 2017, 125, 39–44. [Google Scholar] [CrossRef] [PubMed]
- Ottaviani, D.; Leoni, F.; Rocchegiani, E.; Mioni, R.; Costa, A.; Virgilio, S.; Serracca, L.; Bove, D.; Canonico, C.; Di Cesare, A.; et al. An Extensive Investigation into the Prevalence and the Genetic and Serological Diversity of Toxigenic Vibrio parahaemolyticus in Italian Marine Coastal Waters: Toxigenic V. parahaemolyticus from Italian Coastal Waters. Environ. Microbiol. 2013, 15, 1377–1386. [Google Scholar] [CrossRef]
- Caburlotto, G.; Suffredini, E.; Toson, M.; Fasolato, L.; Antonetti, P.; Zambon, M.; Manfrin, A. Occurrence and Molecular Characterisation of Vibrio Parahaemolyticus in Crustaceans Commercialised in Venice Area, Italy. Int. J. Food Microbiol. 2016, 220, 39–49. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.B.; Okuda, J.; Matsumoto, C.; Takahashi, N.; Hashimoto, S.; Nishibuchi, M. Identification of Vibrio Parahaemolyticus Strains at the Species Level by PCR Targeted to the ToxR Gene. J. Clin. Microbiol. 1999, 37, 1173–1177. [Google Scholar] [CrossRef]
- Bej, A.K.; Patterson, D.P.; Brasher, C.W.; Vickery, M.C.L.; Jones, D.D.; Kaysner, C.A. Detection of Total and Hemolysin-Producing Vibrio Parahaemolyticus in Shellfish Using Multiplex PCR Amplification of Tl, Tdh and Trh. J. Microbiol. Methods 1999, 36, 215–225. [Google Scholar] [CrossRef]
- Hartnell, R.E.; Stockley, L.; Keay, W.; Rosec, J.-P.; Hervio-Heath, D.; Van den Berg, H.; Leoni, F.; Ottaviani, D.; Henigman, U.; Denayer, S.; et al. A Pan-European Ring Trial to Validate an International Standard for Detection of Vibrio Cholerae, Vibrio Parahaemolyticus and Vibrio Vulnificus in Seafoods. Int. J. Food Microbiol. 2019, 288, 58–65. [Google Scholar] [CrossRef]
- Ottaviani, D.; Leoni, F.; Rocchegiani, E.; Santarelli, S.; Masini, L.; Di Trani, V.; Canonico, C.; Pianetti, A.; Tega, L.; Carraturo, A. Prevalence and Virulence Properties of Non-O1 Non-O139 Vibrio Cholerae Strains from Seafood and Clinical Samples Collected in Italy. Int. J. Food Microbiol. 2009, 132, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Performance Standards for Antimicrobial Susceptibility: Supplement M100, 31st ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2021; ISBN 978-1-68440-105-5.
- Hernández-Robles, M.F.; Álvarez-Contreras, A.K.; Juárez-García, P.; Natividad-Bonifacio, I.; Curiel-Quesada, E.; Vázquez-Salinas, C.; Quiñones-Ramírez, E.I. Virulence Factors and Antimicrobial Resistance in Environmental Strains of Vibrio Alginolyticus. Int. Microbiol. 2016, 19, 191–198. [Google Scholar] [CrossRef] [PubMed]
- Hossain, S.; Wickramanayake, M.V.K.S.; Dahanayake, P.S.; Heo, G.-J. Occurrence of Virulence and Extended-Spectrum β-Lactamase Determinants in Vibrio Spp. Isolated from Marketed Hard-Shelled Mussel (Mytilus coruscus). Microb. Drug Resist. 2020, 26, 391–401. [Google Scholar] [CrossRef] [PubMed]
- Costa, W.F.; Giambiagi-deMarval, M.; Laport, M.S. Antibiotic and Heavy Metal Susceptibility of Non-Cholera Vibrio Isolated from Marine Sponges and Sea Urchins: Could They Pose a Potential Risk to Public Health? Antibiotics 2021, 10, 1561. [Google Scholar] [CrossRef] [PubMed]
- Vongxay, K.; Wang, S.; Zhang, X.; Wu, B.; Hu, H.; Pan, Z.; Chen, S.; Fang, W. Pathogenetic Characterization of Vibrio Parahaemolyticus Isolates from Clinical and Seafood Sources. Int. J. Food Microbiol. 2008, 126, 71–75. [Google Scholar] [CrossRef] [PubMed]
- Nair, G.B.; Ramamurthy, T.; Bhattacharya, S.K.; Dutta, B.; Takeda, Y.; Sack, D.A. Global Dissemination of Vibrio Parahaemolyticus Serotype O3:K6 and Its Serovariants. Clin. Microbiol. Rev. 2007, 20, 39–48. [Google Scholar] [CrossRef] [PubMed]
- Lozano-Leòn, A.; Torres, J.; Osorio, C.R.; Martínez-Urtaza, J. Identification of Tdh-Positive Vibrio Parahaemolyticus from an Outbreak Associated with Raw Oyster Consumption in Spain. FEMS Microbiol. Lett. 2003, 226, 281–284. [Google Scholar] [CrossRef] [PubMed]
- García, K.; Torres, R.; Uribe, P.; Hernández, C.; Rioseco, M.L.; Romero, J.; Espejo, R.T. Dynamics of Clinical and Environmental Vibrio Parahaemolyticus Strains during Seafood-Related Summer Diarrhea Outbreaks in Southern Chile. Appl. Environ. Microbiol. 2009, 75, 7482–7487. [Google Scholar] [CrossRef]
- Ottaviani, D.; Leoni, F.; Serra, R.; Serracca, L.; Decastelli, L.; Rocchegiani, E.; Masini, L.; Canonico, C.; Talevi, G.; Carraturo, A. Nontoxigenic Vibrio Parahaemolyticus Strains Causing Acute Gastroenteritis. J. Clin. Microbiol. 2012, 50, 4141–4143. [Google Scholar] [CrossRef]
- Ottaviani, D.; Leoni, F.; Talevi, G.; Masini, L.; Santarelli, S.; Rocchegiani, E.; Susini, F.; Montagna, C.; Monno, R.; D’Annibale, L.; et al. Extensive Investigation of Antimicrobial Resistance in Vibrio Parahaemolyticus from Shellfish and Clinical Sources, Italy. Int. J. Antimicrob. Agents 2013, 42, 191–193. [Google Scholar] [CrossRef]
- Kang, C.-H.; Shin, Y.; Jang, S.; Yu, H.; Kim, S.; An, S.; Park, K.; So, J.-S. Characterization of Vibrio Parahaemolyticus Isolated from Oysters in Korea: Resistance to Various Antibiotics and Prevalence of Virulence Genes. Mar. Pollut. Bull. 2017, 118, 261–266. [Google Scholar] [CrossRef]
- Yu, Q.; Niu, M.; Yu, M.; Liu, Y.; Wang, D.; Shi, X. Prevalence and Antimicrobial Susceptibility of Vibrio Parahaemolyticus Isolated from Retail Shellfish in Shanghai. Food Control 2016, 60, 263–268. [Google Scholar] [CrossRef]
- Serratore, P.; Zavatta, E.; Fiocchi, E.; Serafini, E.; Serraino, A.; Giacometti, F.; Bignami, G. Preliminary Study on the Antimicrobial Susceptibility Pattern Related to the Genotype of Vibrio Vulnificus Strains Isolated in the North-Western Adriatic Sea Coastal Area. Ital. J. Food Saf. 2017, 6, 6843. [Google Scholar] [CrossRef] [PubMed]
- Elmahdi, S.; DaSilva, L.V.; Parveen, S. Antibiotic Resistance of Vibrio Parahaemolyticus and Vibrio Vulnificus in Various Countries: A Review. Food Microbiol. 2016, 57, 128–134. [Google Scholar] [CrossRef] [PubMed]
- Han, F.; Walker, R.D.; Janes, M.E.; Prinyawiwatkul, W.; Ge, B. Antimicrobial Susceptibilities of Vibrio parahaemolyticus and Vibrio vulnificus Isolates from Louisiana Gulf and Retail Raw Oysters. Appl. Environ. Microbiol. 2007, 73, 7096–7098. [Google Scholar] [CrossRef]
- DeWaal, C.S.; Vaughn Grooters, S. Antibiotic Resistance in Foodborne Pathogens; Center for Science in the Public Interest: Washington, DC, USA, 2013; pp. 1–22. [Google Scholar]
- Letchumanan, V.; Pusparajah, P.; Tan, L.T.-H.; Yin, W.-F.; Lee, L.-H.; Chan, K.-G. Occurrence and Antibiotic Resistance of Vibrio Parahaemolyticus from Shellfish in Selangor, Malaysia. Front. Microbiol. 2015, 6, 1417. [Google Scholar] [CrossRef]
| Target Species | Genetic Marker | Primer Sequences (5′-3′) | Product Size | References |
|---|---|---|---|---|
| Vibrio parahaemolyticus | toxR | F: GTCTTCTGACGCAATCGTTG R: ATACGAGTGGTTGCTGTCATG | 368 bp | [37] |
| tdh | F: GTAAAGGTCTCTGACTTTTGGAC R: TGGAATAGAACCTTCATCTTCAC | 269 bp | [38] | |
| trh | F: TTGGCTTCGATATTTTCAGTATCT R: CATAACAAACATATGCCCATTTCCG | 500 bp | [38] | |
| Vibrio vulnificus | VVH | F: CCGGCGGTACAGCTTGGCGC R: CGCCACCCACTTTCGGGCC | 519 bp | [39] |
| Vibrio cholerae | prVC | F: TTAAGCSTTTTCRCTGAGAATG R: AGTCACTTAACCATACAACCCG | 295–310 bp | [39] |
| ctxA | F: CGGGCAGATTCTAGACCTCCTG R: CGATGATCTTGGAGCATTCCCAC | 564 bp | [40] | |
| stn/sto | F: TCGCATTTAGCCAAACAGTAGAAA R: GCTGGATTGCAACATATTTCGC | 172 bp | [40] | |
| tcpA | F: CACGATAAGAAAACCGGTCAAGAG R: TTACCAAATGCAACGCCGAATG | 620 bp | [40] | |
| hlyAET | F: GGCAAACAGCGAAACAAATACC R: CTCAGCGGGCTAATACGGTTTA | 481 bp | [40] |
| Drugs | Strains | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| V. parahaemolyticus | V. vulnificus | V. cholerae NCV | V. alginolyticus | |||||||||
| n = 43 | n = 5 | n = 12 | n = 105 | |||||||||
| R a | I b | MDR c | R | I | MDR | R | I | MDR | R | I | MDR | |
| Amp | 30 | 2 d | 9 | 93 | ||||||||
| Ctx | 6 | |||||||||||
| Cro | 2 | |||||||||||
| Caz | 8 | 2 | 2 | 12 | 13 | |||||||
| Kf | 9 | 9 | 2 | 6 | ||||||||
| Cip | 1 | |||||||||||
| C | ||||||||||||
| Cn | 2 | 23 | 12 | |||||||||
| K | 2 | 8 | 11 | |||||||||
| S | 8 | 9 | 5 | 2 | 58 | |||||||
| Stx | 5 | |||||||||||
| Te | 6 | 4 | 1 | 12 | ||||||||
| Ct | 2 | 9 | ||||||||||
| Kz | 13 | 1 | ||||||||||
| Amc | 2 | 2 | ||||||||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Castello, A.; Alio, V.; Sciortino, S.; Oliveri, G.; Cardamone, C.; Butera, G.; Costa, A. Occurrence and Molecular Characterization of Potentially Pathogenic Vibrio spp. in Seafood Collected in Sicily. Microorganisms 2023, 11, 53. https://doi.org/10.3390/microorganisms11010053
Castello A, Alio V, Sciortino S, Oliveri G, Cardamone C, Butera G, Costa A. Occurrence and Molecular Characterization of Potentially Pathogenic Vibrio spp. in Seafood Collected in Sicily. Microorganisms. 2023; 11(1):53. https://doi.org/10.3390/microorganisms11010053
Chicago/Turabian StyleCastello, Annamaria, Vincenzina Alio, Sonia Sciortino, Giuseppa Oliveri, Cinzia Cardamone, Gaspare Butera, and Antonella Costa. 2023. "Occurrence and Molecular Characterization of Potentially Pathogenic Vibrio spp. in Seafood Collected in Sicily" Microorganisms 11, no. 1: 53. https://doi.org/10.3390/microorganisms11010053
APA StyleCastello, A., Alio, V., Sciortino, S., Oliveri, G., Cardamone, C., Butera, G., & Costa, A. (2023). Occurrence and Molecular Characterization of Potentially Pathogenic Vibrio spp. in Seafood Collected in Sicily. Microorganisms, 11(1), 53. https://doi.org/10.3390/microorganisms11010053





