Insights into the Bacterial Diversity and Detection of Opportunistic Pathogens in Mexican Chili Powder
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial and Fungal Load
2.2. Salmonella and E. coli Detection
2.3. B. cereus s.l. Detection
2.4. Isolation and Characterization of Ampicillin-Resistant Bacteria
2.5. Bacterial Community Analysis
3. Results
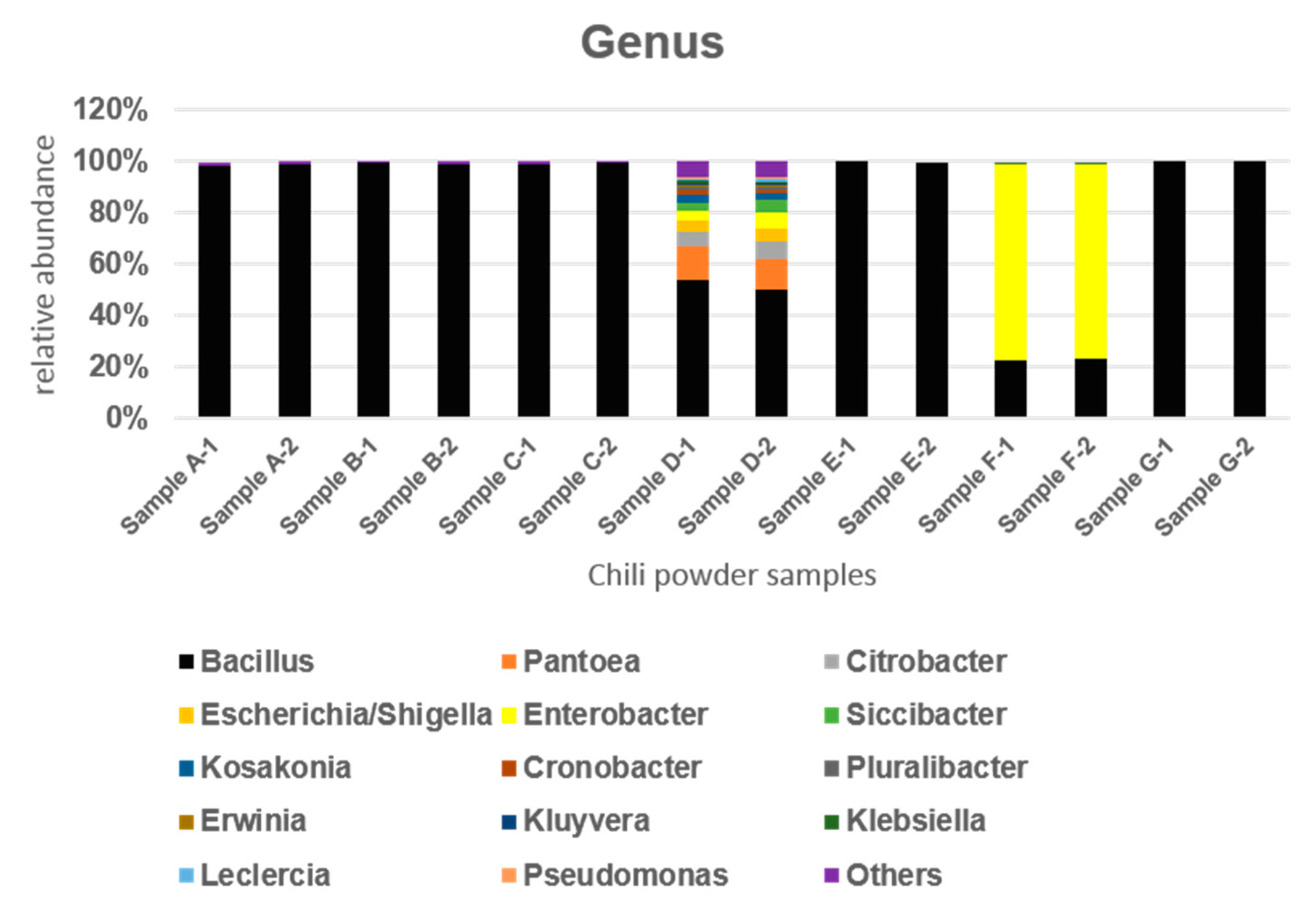
3.1. Bacterial Community Analysis
3.2. Bacterial and Fungal Load
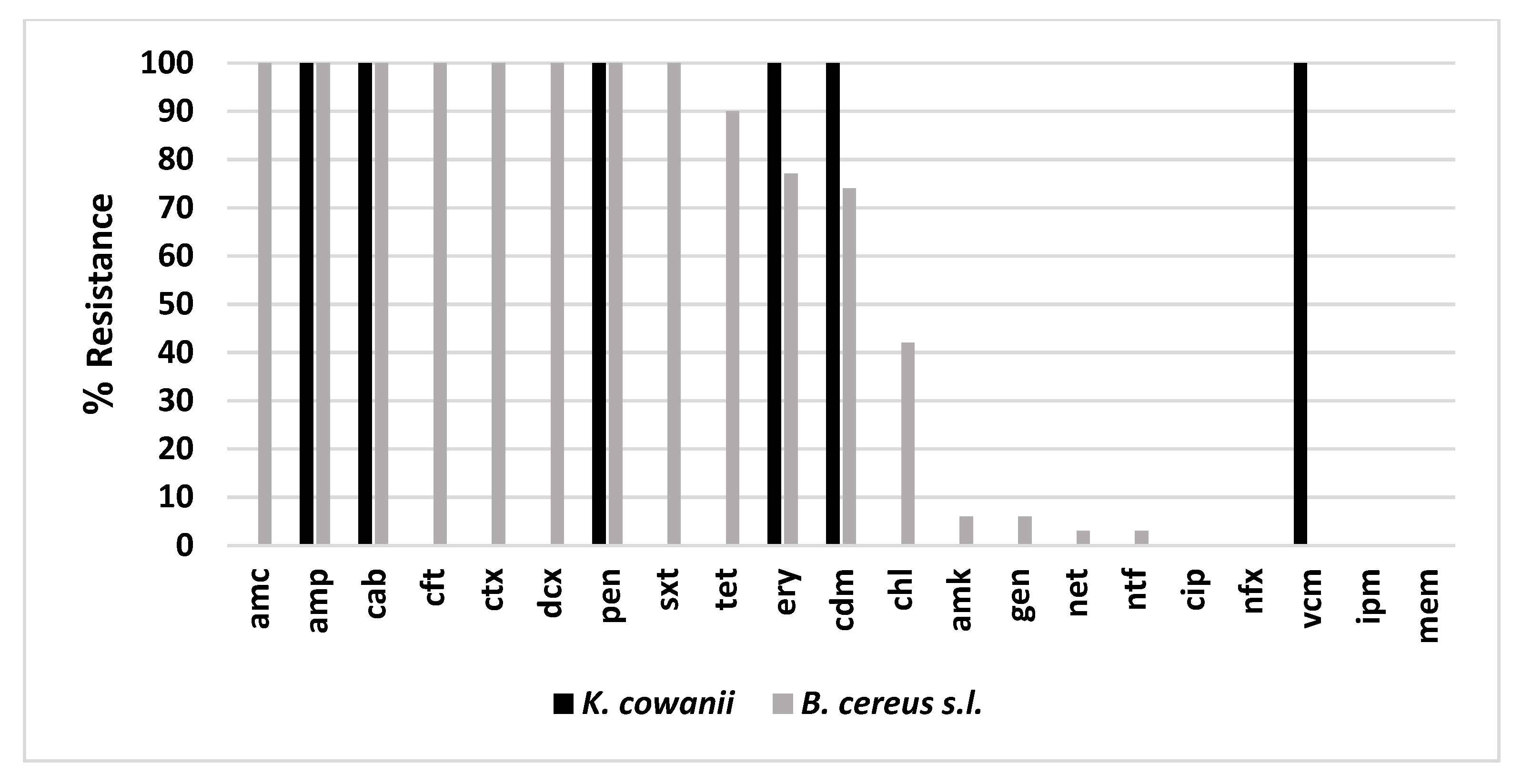
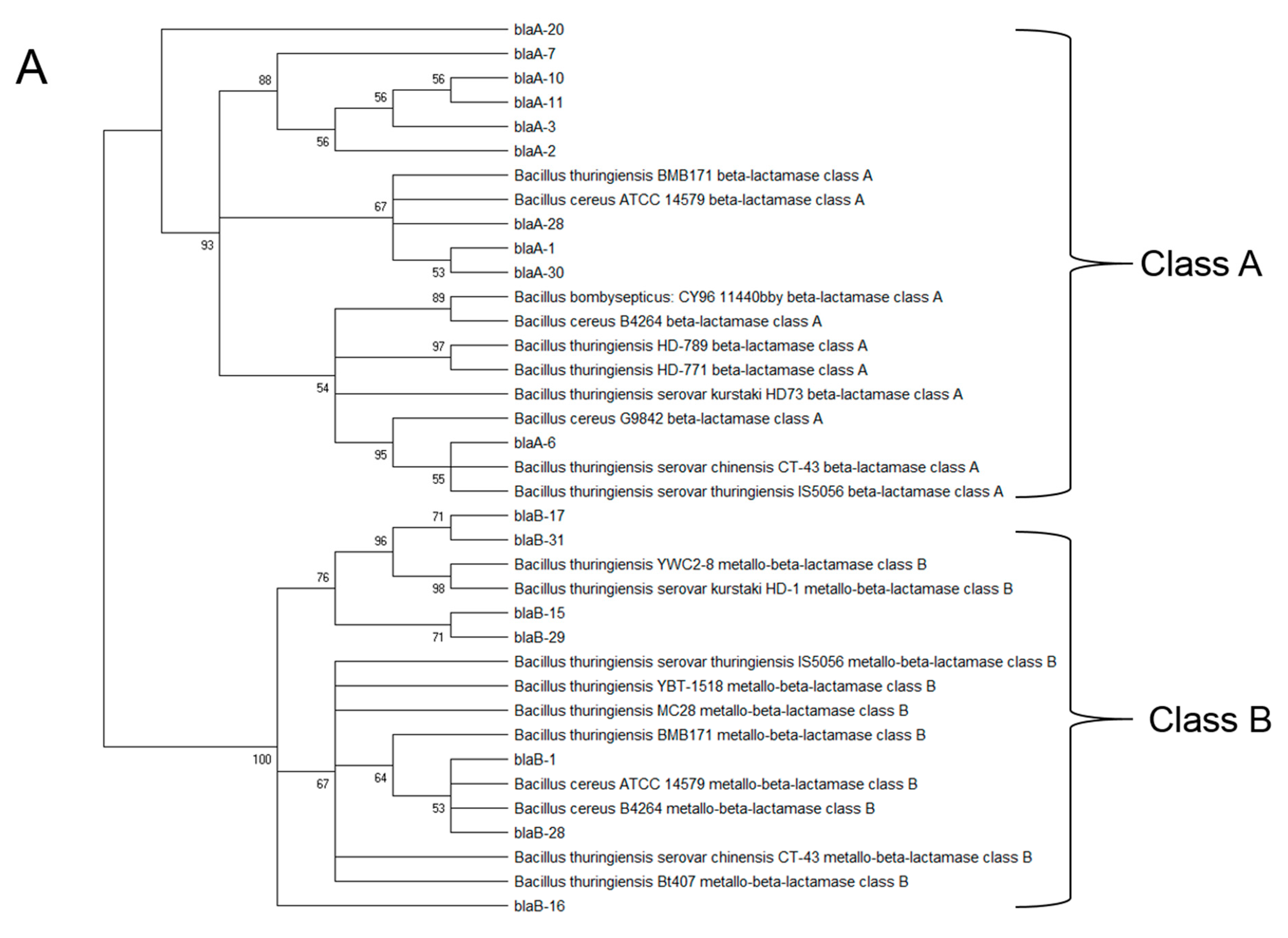
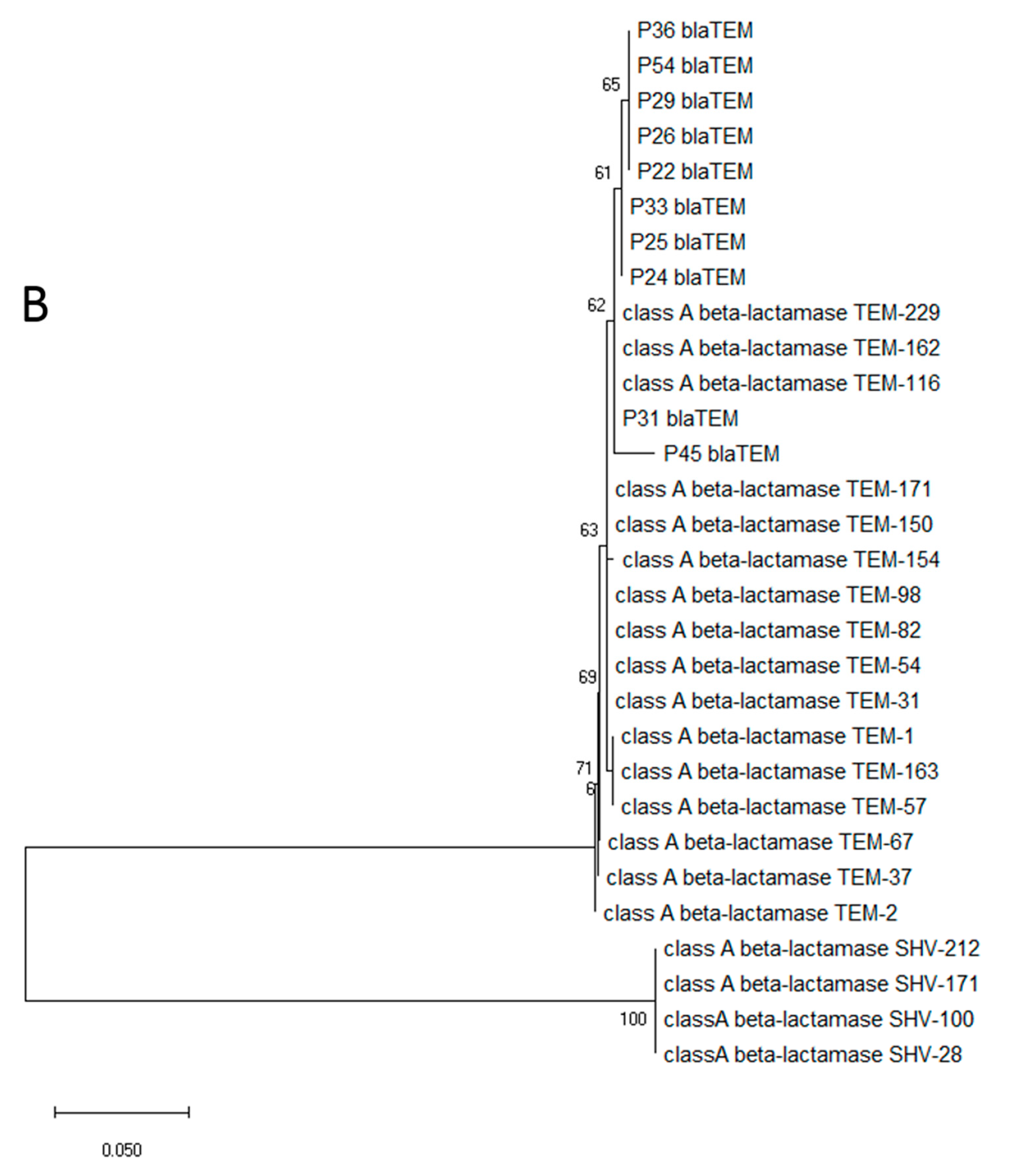
3.3. Isolation and Characterization of Opportunistic Pathogens
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Baenas, N.; Belović, M.; Ilic, N.; Moreno, D.A.; García-Viguera, C. Industrial use of peppr (Capsicum annuem L.) derived products: Technological benefits and biological advantages. Food Chem. 2019, 274, 872–885. [Google Scholar] [CrossRef] [PubMed]
- Rincón, V.H.; Torres, T.C.; López, P.L.; Moreno, L.L.; Meraz, M.R.; Mendoza, H.V.; Castillo, J.A. Los Chiles de México y su Distribución; SINAREFI, COLPOS, INIFAP, ITConkal, UANL, UAN: Montecillo, Texcoco, Estado de México, México, 2010. [Google Scholar]
- Padilha, H.K.; Pereira, E.D.; Munhoz, P.C.; Vizzotto, M.; Valgas, R.A.; Barbieri, R.L. Genetic variability for synthesis of bioactive compounds in peppers (Capsicum annuum) from Brazil. Food Sci. Technol. 2015, 35, 516–523. [Google Scholar] [CrossRef] [Green Version]
- Azevedo-Meleiro, C.; Rodriguez-Amaya, D. Qualitative and quantitative differences in the carotenoid composition of yellow and red peppers determined by HPLC-DAD-MS. J. Sep. Sci. 2009, 32, 3652–3658. [Google Scholar] [CrossRef] [PubMed]
- Eggersdorfer, M.; Wyss, A. Carotenoids in human nutrition and health. Arch. Biochem. Biophys. 2018, 652, 18–26. [Google Scholar] [CrossRef]
- García-Gaytán, V.; Gómez-Merino, F.C.; Trejo-Téllez, L.I.; Baca-Castillo, G.A.; García-Morales, S. The Chilhuacle Chili (Capsicum annuum L.) in Mexico: Description of the Variety, Its Cultivation, and Uses. Int. J. Agron. 2017, 2017, 5641680. [Google Scholar] [CrossRef] [Green Version]
- Molnár, H.; Bata-Vidács, I.; Baka, E.; Cserhalmi, Z.; Ferenczi, S.; Tömösközi-Farkas, R.; Adányi, N.; Székács, A. The effect of different decontamination methods on the microbial load, bioactive components, aroma and colour of spice paprika. Food Control 2018, 83, 131–140. [Google Scholar] [CrossRef]
- Feroz, F.; Shimizu, H.; Nishioka, T.; Mori, M.; Sakagami, Y. Bacterial and Fungal Counts of Dried and Semi-Dried Foods Collected from Dhaka, Bangladesh, and Their Reduction Methods. Biocontrol Sci. 2016, 21, 243–251. [Google Scholar] [CrossRef] [Green Version]
- Doren, J.; Neil, K.; Parish, M.; Gieraltowski, L.; Gould, L.; Gombas, K. Foodborne illness outbreaks from microbial contaminants in spices, 1973–2010. Food Microbiol. 2013, 36, 456–464. [Google Scholar] [CrossRef]
- Melo González, M.G.; Romero, S.M.; Arjona, M.; Larumbe, A.G.; Vaamonde, G. Microbiological quality of Argentinian paprika. Rev. Argent. De Microbiol. 2017, 49, 339–346. [Google Scholar] [CrossRef]
- Mamun, A.A.; Aynee, M.A.; Majumder, D.; Ali, M.; Hossen, S.; Maruf, K.M.R. Quality assessment of selected commercial brand of chilli powder in Bangladesh. MOJ Food Processing Technol. 2016, 3, 70–73. [Google Scholar] [CrossRef]
- Bata-Vidács, I.; Baka, E.; Tóth, Á.; Csernus, O.; Luzics, S.; Adányi, N.; Székács, A.; Kukolya, J. Investigation of regional differences of the dominant microflora of spice paprika by molecular methods. Food Control 2018, 83, 109–117. [Google Scholar] [CrossRef]
- György, É.; Eva, L.; Márta, A.; András, C. Antibiotic resistance pattern of the allochthonous bacteria isolated from commercially available spices. Food Sci. Nutr. 2021, 9, 4550–4560. [Google Scholar] [CrossRef] [PubMed]
- Frentzel, H.; Kraushaar, B.; Krause, G.; Bodi, D.; Wichmann-Schauer, H.; Appel, B.; Mader, A. Phylogenetic and toxinogenic characteristics of Bacillus cereus group members isolated from spices and herbs. Food Control 2018, 83, 90–98. [Google Scholar] [CrossRef]
- Hariram, U.; Labbé, R. Spore Prevalence and Toxigenicity of Bacillus cereus and Bacillus thuringiensis Isolates from U.S. Retail Spices. J. Food Prot. 2015, 78, 590–596. [Google Scholar] [CrossRef] [PubMed]
- Lehmacher, A.; Bockemühl, J.; Aleksic, S. Nationwide outbreak of human salmonellosis in Germany due to contaminated paprika and paprika-powdered potato chips. Epidemiol. Infect. 2009, 115, 501–511. [Google Scholar] [CrossRef] [Green Version]
- Muhamad Rizal, N.S.; Neoh, H.M.; Ramli, R.; A/LKPeriyasamy, P.R.; Hanafiah, A.; Abdul Samat, M.N.; Tan, T.L.; Wong, K.K.; Nathan, S.; Chieng, S.; et al. Advantages and Limitations of 16S rRNA Next-Generation Sequencing for Pathogen Identification in the Diagnostic Microbiology Laboratory: Perspectives from a Middle-Income Country. Diagnostics 2020, 10, 816. [Google Scholar] [CrossRef]
- Banerjee, M.; Sarkar, P. Microbiological quality of some retail spices in India. Food Res. Int. 2003, 36, 469–474. [Google Scholar] [CrossRef]
- Celestino Hernández, A.G.; Gómez Ortiz, V.; Arvizu Gómez, J.L.; Ramos López, M.Á.; Rodríguez Morales, J.A.; Flores Macías, A.; Álvarez Hidalgo, E.; Nuñez Ramírez, J.; Flores Gallardo, F.J.; García Gutiérrez, M.C.; et al. Detection of Bacillus cereus sensu lato Isolates Posing Potential Health Risks in Mexican Chili Powder. Microorganisms 2021, 9, 2226. [Google Scholar] [CrossRef]
- Ehling-Schulz, M.; Lereclus, D.; Koehler, T.M.; Fischetti, V.A.; Novick, R.P.; Ferretti, J.J.; Portnoy, D.A.; Braunstein, M.; Rood, J.I. The Bacillus cereus Group: Bacillus Species with Pathogenic Potential. Microbiol. Spectr. 2019, 7. [Google Scholar] [CrossRef]
- Guinebretière, M.H.; Auger, S.; Galleron, N.; Contzen, M.; De Sarrau, B.; De Buyser, M.L.; Lamberet, G.; Fagerlund, A.; Granum, P.E.; Lereclus, D.; et al. Bacillus cytotoxicus sp. nov. is a novel thermotolerant species of the Bacillus cereus Group occasionally associated with food poisoning. Int. J. Syst. Evol. Microbiol. 2013, 63 Pt 1, 31–40. [Google Scholar] [CrossRef]
- Chang, T.C.; Rosch, J.; Gu, Z.; Hakim, H.; Hewitt, C.; Gaur, A.; Wu, G.; Hayden, R. Whole Genome Characterization of Bacillus cereus Associated with Specific Disease Manifestations. Infect. Immun. 2017, 86, e00574-17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lapidus, A.; Goltsman, E.; Auger, S.; Galleron, N.; Ségurens, B.; Dossat, C.; Land, M.L.; Broussolle, V.; Brillard, J.; Guinebretiere, M.H.; et al. Extending the Bacillus cereus group genomics to putative food-borne pathogens of different toxicity. Chem. Biol. Interact. 2008, 171, 236–249. [Google Scholar] [CrossRef] [PubMed]
- Erickson, B.D.; Elkins, C.A.; Mullis, L.B.; Heinze, T.M.; Wagner, R.D.; Cerniglia, C.E. A metallo-β-lactamase is responsible for the degradation of ceftiofur by the bovine intestinal bacterium Bacillus cereus P41. Vet. Microbiol. 2014, 172, 499–504. [Google Scholar] [CrossRef] [PubMed]
- Luna, V.A.; King, D.S.; Gulledge, J.; Cannons, A.C.; Amuso, P.T.; Cattani, J. Susceptibility of Bacillus anthracis, Bacillus cereus, Bacillus mycoides, Bacillus pseudomycoides and Bacillus thuringiensis to 24 antimicrobials using Sensititre® automated microbroth dilution and Etest® agar gradient diffusion methods. J. Antimicrob. Chemother. 2007, 60, 555–567. [Google Scholar] [CrossRef] [Green Version]
- Ruppé, É.; Woerther, P.L.; Barbier, F. Mechanisms of antimicrobial resistance in Gram-negative bacilli. Ann. Intensive Care 2015, 5, 21. [Google Scholar] [CrossRef] [Green Version]
- Tilahun, M.; Kassa, Y.; Gedefie, A.; Ashagrie, M. Emerging Carbapenem-Resistant Enterobacteriaceae Infection, Its Epidemiology and Novel Treatment Options: A Review. Infect. Drug Resist. 2021, 14, 4363–4374. [Google Scholar] [CrossRef]
- Woodford, N.; Wareham, D.; Guerra, B.; Teale, C. Carbapenemase-producing Enterobacteriaceae and non-Enterobacteriaceae from animals and the environment: An emerging public health risk of our own making? J. Antimicrob. Chemother. 2013, 69, 287–291. [Google Scholar] [CrossRef] [Green Version]
- Center for Food Safety and Applied Nutrition, U.S.F.a.D.A. Bacteriological Analytical Manual Online. 2022. Available online: http://www.cfsan.fda.gov/∼ebam/bam-1.html (accessed on 5 February 2022).
- Hernández Flores, J.L.; Salinas Landaverde, D.; Pacheco Huerta, Y.; Guerra Castillo, V.L.; Barrios Sánchez, M.A.; Arvizu Hernández, I.; Ramos López, M.A.; Álvarez Hidalgo, E.; Jones, G.H.; Campos Guillén, J. Phylogenetic Analysis of Bacillus cereus sensu lato Isolates from Commercial Bee Pollen Using tRNACys-PCR. Microorganisms 2020, 8, 524. [Google Scholar] [CrossRef] [Green Version]
- Resendiz-Nava, C.N.; Silva-Rojas, H.V.; Rebollar-Alviter, A.; Rivera-Pastrana, D.M.; Mercado-Silva, E.M.; Nava, G.M. A Comprehensive Evaluation of Enterobacteriaceae Primer Sets for Analysis of Host-Associated Microbiota. Pathogens 2022, 11, 17. [Google Scholar] [CrossRef]
- M02-A13; Performance standards for antimicrobial disk susceptibility tests; approved standard—12th ed. Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2018.
- Krause, N.; Moravek, M.; Dietrich, R.; Wehrle, E.; Slaghuis, J.; Märtlbauer, E. Performance characteristics of the Duopath® Cereus Enterotoxins assay for rapid detection of enterotoxinogenic Bacillus cereus strains. Int. J. Food Microbiol. 2010, 144, 322–326. [Google Scholar] [CrossRef]
- Hansen, B.; Hendriksen, N. Detection of Enterotoxic Bacillus cereus and Bacillus thuringiensis Strains by PCR Analysis. Appl. Environ. Microbiol. 2001, 67, 185–189. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Monstein, H.J.; Östholm-Balkhed, Å.; Nilsson, M.V.; Nilsson, M.; Dornbusch, K.; Nilsson, L.E. Multiplex PCR amplification assay for the detection of blaSHV, blaTEM and blaCTX-M genes in Enterobacteriaceae. APMIS 2007, 115, 1400–1408. [Google Scholar] [CrossRef] [PubMed]
- Naas, T.; Oueslati, S.; Bonnin, R.A.; Dabos, M.L.; Zavala, A.; Dortet, L.; Retailleau, P.; Iorga, B.I. Beta-lactamase database (BLDB)—structure and function. J. Enzym. Inhib. Med. Chem. 2017, 32, 917–919. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [PubMed]
- Felsenstein, J. Confidence Limits on Phylogenies: An Approach Using the Bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Jukes, T.H.; Cantor, C.R. Evolution of Protein Molecules. In Mammalian Protein Metabolism; Munro, H.N., Ed.; Academic Press: Cambridge, MA, USA, 1969; pp. 21–132. [Google Scholar]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [Green Version]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Gonzalez Peña, A.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [Green Version]
- Segata, N.; Izard, J.; Waldron, L.; Gevers, D.; Miropolsky, L.; Garrett, W.S.; Huttenhower, C. Metagenomic biomarker discovery and explanation. Genome Biol. 2011, 12, R60. [Google Scholar] [CrossRef] [Green Version]
- International Association of Microbiological Societies. Microorganisms in Foods. 2. Sampling for Microbiological Analysis: Principles and Specific Applications; Blackwell Scientific: Oxford, UK, 1986; Volume 58, p. 1308. [Google Scholar]
- Armalyte, J.; Skerniškytė, J.; Bakienė, E.; Krasauskas, R.; Šiugždinienė, R.; Kareivienė, V.; Kerzienė, S.; Klimienė, I.; Suziedeliene, E.; Ruzauskas, M. Microbial Diversity and Antimicrobial Resistance Profile in Microbiota From Soils of Conventional and Organic Farming Systems. Front. Microbiol. 2019, 10, 892. [Google Scholar] [CrossRef]
- Cycoń, M.; Mrozik, A.; Piotrowska-Seget, Z. Antibiotics in the Soil Environment—Degradation and Their Impact on Microbial Activity and Diversity. Front. Microbiol. 2019, 10, 338. [Google Scholar] [CrossRef] [PubMed]
- Allende, A.; Bolton, D.; Chemaly, M.; Davies, R.; Fernández Escámez, P.S.; Gironés, R.; Herman, L.; Koutsoumanis, K.; Lindqvist, R.; Nørrung, B.; et al. Risks for public health related to the presence of Bacillus cereus and other Bacillus spp. including Bacillus thuringiensis in foodstuffs. EFSA J. 2016, 14, e04524. [Google Scholar]
- Torkar, K.G.; Bedenić, B. Antimicrobial susceptibility and characterization of metallo-β-lactamases, extended-spectrum β-lactamases, and carbapenemases of Bacillus cereus isolates. Microb. Pathog. 2018, 118, 140–145. [Google Scholar] [CrossRef] [PubMed]
- Berinson, B.; Bellon, E.; Christner, M.; Both, A.; Aepfelbacher, M.; Rohde, H. Identification of Kosakonia cowanii as a rare cause of acute cholecystitis: Case report and review of the literature. BMC Infect. Dis. 2020, 20, 366. [Google Scholar] [CrossRef]
- Duployez, C.; Edun-Renard, M.E.; Kipnis, E.; Dessein, R.; Le Guern, R. Bacteremia Due to Kosakonia cowanii in a Preterm Neonate. J. Pediatric. Infect. Dis. 2021, 16, 183–186. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, B.; Li, Q.; Huang, D.; Zhang, Y.; Li, G.; He, H. Isolation and Complete Genome Sequence Analysis of Kosakonia cowanii Pa82, a Novel Pathogen Causing Bacterial Wilt on Patchouli. Front. Microbiol. 2022, 12, 818228. [Google Scholar] [CrossRef]
- Mertschnigg, T.; Patz, S.; Becker, M.; Feierl, G.; Ruppel, S.; Bunk, B.; Spröer, C.; Overmann, J.; Zarfel, G. First Report of Kosakonia radicincitans Bacteraemia from Europe (Austria) - Identification and Whole-Genome Sequencing of Strain DSM 107547. Sci. Rep. 2020, 10, 1948. [Google Scholar] [CrossRef]
- Caulier, S.; Nannan, C.; Gillis, A.; Licciardi, F.; Bragard, C.; Mahillon, J. Overview of the Antimicrobial Compounds Produced by Members of the Bacillus subtilis Group. Front. Microbiol. 2019, 10, 302. [Google Scholar] [CrossRef] [Green Version]
- Jouzani, G.S.; Valijanian, E.; Sharafi, R. Bacillus thuringiensis: A successful insecticide with new environmental features and tidings. Appl. Microbiol. Biotechnol. 2017, 101, 2691–2711. [Google Scholar] [CrossRef]

| Sample | AMB (CFU/g) | SMB (CFU/g) | F/Y (CFU/g) | Enterobacteriaceae (CFU/g) | E. coli | Salmonella |
|---|---|---|---|---|---|---|
| A | 2.0 × 105 | 1.5 × 103 | 2.7 × 102/4.2 × 102 | 1.1 × 103 | ND * | ND * |
| B | 1.6 × 105 | 7.5 × 102 | 1.8 × 103/9.2 × 102 | 4.2 × 102 | ND | ND |
| C | 1.14 × 105 | 2.4 × 103 | 1.7 × 105/4.3 × 104 | 3.7 × 103 | ND | ND |
| D | 1.2 × 106 | 3.5 × 105 | 5.9 × 104/4.0 × 104 | 1.2 × 105 | ND | ND |
| E | 1.0 × 106 | 4.3 × 103 | 4.3 × 105/5.5 × 104 | 4.8 × 103 | ND | ND |
| F | 6.3 × 104 | 1.0 × 102 | 1.1 × 105/3.5 × 104 | 1.4 × 104 | ND | ND |
| G | 1.2 × 105 | 4.5 × 102 | 2.8 × 103/7.5 × 102 | 9.4 × 102 | ND | ND |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hernández Gómez, Y.F.; González Espinosa, J.; Ramos López, M.Á.; Arvizu Gómez, J.L.; Saldaña, C.; Rodríguez Morales, J.A.; García Gutiérrez, M.C.; Pérez Moreno, V.; Álvarez Hidalgo, E.; Nuñez Ramírez, J.; et al. Insights into the Bacterial Diversity and Detection of Opportunistic Pathogens in Mexican Chili Powder. Microorganisms 2022, 10, 1677. https://doi.org/10.3390/microorganisms10081677
Hernández Gómez YF, González Espinosa J, Ramos López MÁ, Arvizu Gómez JL, Saldaña C, Rodríguez Morales JA, García Gutiérrez MC, Pérez Moreno V, Álvarez Hidalgo E, Nuñez Ramírez J, et al. Insights into the Bacterial Diversity and Detection of Opportunistic Pathogens in Mexican Chili Powder. Microorganisms. 2022; 10(8):1677. https://doi.org/10.3390/microorganisms10081677
Chicago/Turabian StyleHernández Gómez, Yoali Fernanda, Jacqueline González Espinosa, Miguel Ángel Ramos López, Jackeline Lizzeta Arvizu Gómez, Carlos Saldaña, José Alberto Rodríguez Morales, María Carlota García Gutiérrez, Victor Pérez Moreno, Erika Álvarez Hidalgo, Jorge Nuñez Ramírez, and et al. 2022. "Insights into the Bacterial Diversity and Detection of Opportunistic Pathogens in Mexican Chili Powder" Microorganisms 10, no. 8: 1677. https://doi.org/10.3390/microorganisms10081677
APA StyleHernández Gómez, Y. F., González Espinosa, J., Ramos López, M. Á., Arvizu Gómez, J. L., Saldaña, C., Rodríguez Morales, J. A., García Gutiérrez, M. C., Pérez Moreno, V., Álvarez Hidalgo, E., Nuñez Ramírez, J., Jones, G. H., Hernández Flores, J. L., & Campos Guillén, J. (2022). Insights into the Bacterial Diversity and Detection of Opportunistic Pathogens in Mexican Chili Powder. Microorganisms, 10(8), 1677. https://doi.org/10.3390/microorganisms10081677










