Bacteriocin-Producing Escherichia coli Isolated from the Gastrointestinal Tract of Farm Animals: Prevalence, Molecular Characterization and Potential for Application
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains
2.2. Prevalence of Bacteriocin-Producing Strains
2.3. Molecular Characterization
2.3.1. DNA Extraction
2.3.2. Detection of Genes Encoding Bacteriocins
2.3.3. Detection of Virulence-Associated Genes
2.4. Potential for Appliaction
2.4.1. Preparation of Cell-Free Supernatants (CFS) of Bacteriocin-Producing E. coli
2.4.2. Evaluation of Antagonistic Activity of CFS of Bacteriocin-Producing E. coli
2.4.3. Bacteriophage Induction
2.4.4. Hemolytic Activities
2.5. Statistical Analysis
3. Results
3.1. Prevalence of Bacteriocin-Producing E. coli
3.2. The Effect of CFS of Bacteriocin-Producing E. coli on the Growth of Pathogenic E. coli
3.3. Biological and Molecular-Genetic Properties of Bacteriocin-Producing E. coli Strains
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Vila, J.; Sáez-López, E.; Johnson, J.R.; Römling, U.; Dobrindt, U.; Cantón, R.; Giske, C.G.; Naas, T.; Carattoli, A.; Martínez-Medina, M. Escherichia coli: An old friend with new tidings. FEMS Microbiol. Rev. 2016, 40, 437–463. [Google Scholar] [CrossRef] [PubMed]
- Oporto, B.; Ocejo, M.; Alkorta, M.; Marimón, T.; Montes, M.; Hurtado, A. Zoonotic approach to Shiga toxin-producing Escherichia coli: Integrated analysis of virulence and antimicrobial resistance in ruminants and humans. Epidemiol. Infect. 2019, 147, e164. [Google Scholar] [CrossRef] [PubMed]
- Maturana, V.G.; de Pace, F.; Carlos, C.; Pires, M.M.; de Campos, T.A.; Nakazato, G.; Stheling, E.G.; Logue, C.M.; Nolan, L.K.; da Silveira, W.D. Subpathotypes of avian pathogenic Escherichia coli (APEC) exist as defined by their syndromes and virulence traits. Open Microbiol. J. 2011, 5, 55–64. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Mageiros, L.; Méric, G.; Bayliss, S.C.; Pensar, J.; Pascoe, B.; Mourkas, E.; Calland, J.K.; Yahara, K.; Murray, S.; Wilkinson, T.S.; et al. Genome evolution and the emergence of pathogenicity in avian Escherichia coli. Nat. Commun. 2021, 12, 765. [Google Scholar] [CrossRef] [PubMed]
- Valdes, A.M.; Walter, J.; Segal, E.; Spector, T.D. Role of the gut microbiota in nutrition and health: Discuss strategies for modulating the gut microbiota through diet and probiotics. BMJ 2018, 361, 36–44. [Google Scholar]
- Pickard, J.M.; Núñez, G. Pathogen colonization resistance in the gut and its manipulation for improved health. Am. J. Pathol. 2019, 189, 1300–1310. [Google Scholar] [CrossRef]
- Mazurek-Popczyk, J.; Pisarska, J.; Bok, E.; Baldy-Chudzik, K. Antibacterial activity of bacteriocinogenic commensal Escherichia coli against zoonotic strains resistant and sensitive to antibiotics. Antibiotics 2020, 9, 411. [Google Scholar] [CrossRef]
- Gordon, D.M.; O’Brien, C.L. Bacteriocin diversity and the frequency of multiple bacteriocin production in Escherichia coli. Microbiology 2006, 152, 3239–3244. [Google Scholar] [CrossRef]
- Hammami, R.; Fernandez, B.; Lacroix, C.; Fliss, I. Anti-infective properties of bacteriocins: An update. Cell. Mol. Life Sci. 2013, 70, 2947–2967. [Google Scholar] [CrossRef]
- Rebuffat, S. Bacteriocins from Gram-Negative Bacteria: A Classification? In Prokaryotic Antimicrobial Peptides; Drider, D., Rebuffat, S., Eds.; Springer: New York, NY, USA, 2011; pp. 55–72. [Google Scholar]
- Baquero, F.; Lanza, V.F.; Baquero, M.-R.; del Campo, R.; Bravo-Vázquez, D.A. Microcins in Enterobacteriaceae: Peptide antimicrobials in the eco-active intestinal chemosphere. Front. Microbiol. 2019, 10, 2261. [Google Scholar] [CrossRef]
- Lagha, A.B.; Haas, B.; Gottschalk, M.; Grenier, D. Antimicrobial potential of bacteriocins in poultry and swine production. Vet. Res. 2017, 48, 22. [Google Scholar] [CrossRef] [PubMed]
- Askari, N.; Ghanbarpour, R. Molecular investigation of the colicinogenic strains that are capable of inhibiting E. coli O157:H7 in vitro. BMC Vet. Res. 2019, 15, 14. [Google Scholar] [CrossRef] [PubMed]
- Simons, A.; Alhanout, K.; Duval, R.E. Bacteriocins, antimicrobial peptides from bacterial origin: Overview of their biology and their impact against multidrug-resistant bacteria. Microorganisms 2020, 8, 639. [Google Scholar] [CrossRef] [PubMed]
- Cameron, A.; Zaheer, R.; Adator, E.H.; Barbieri, R.; Reuter, T.; McAllister, T.A. Bacteriocin occurrence and activity in Escherichia coli isolated from bovines and wastewater. Toxins 2019, 11, 475. [Google Scholar] [CrossRef] [PubMed]
- Zaslavskaya, M.I.; Makhrova, T.V.; Aleksandrova, N.A.; Ignatova, N.I.; Belova, I.V.; Tochilina, A.G.; Solovyeva, I.V. Prospects for using bacteriocins of normal microbiota in antibacterial therapy (review). Sovrem. Tehnol. V Med. 2019, 11, 136–145. [Google Scholar] [CrossRef]
- Versalovic, J.; Koeuth, T.; Lupski, J.R. Distribution of repetitive DNA sequences in eubacteria and application to fingerprinting of bacterial genomes. Nucleic Acids Res. 1991, 19, 6823–6831. [Google Scholar] [CrossRef]
- Kuznetsova, M.V.; Gizatullina, J.S.; Nesterova, L.Y.; Starčič Erjavec, M. Escherichia coli isolated from cases of colibacillosis in Russian poultry farms (Perm Krai): Sensitivity to antibiotics and bacteriocins. Microorganisms 2020, 8, 741. [Google Scholar] [CrossRef] [PubMed]
- Mihailovskaya, V.S.; Artamonova, O.A.; Zhdanova, I.N.; Kuznetsova, M.V. Prevalence of pathogenicity determinants among Escherichia coli strains isolated from healthy cows and calves in farms of Perm Krai. Mechanisms of microorganisms adaptation to different habitat condition. In Proceedings of the All-Russian Scientific Conference with International Participation, Irkutsk, Russia, 28 February–6 March 2022; pp. 152–154. [Google Scholar]
- Budič, M.; Rijavec, M.; Petkovšek, Z.; Žgur-Bertok, D. Escherichia coli bacteriocins: Antimicrobial efficacy and prevalence among isolates from patients with bacteraemia. PLoS ONE. 2011, 6, e28769. [Google Scholar] [CrossRef]
- Starčič Erjavec, M.; Petkovšek, Ž.; Kuznetsova, M.; Maslennikova, I.; Žgur-Bertok, D. Strain ŽP—The first bacterial conjugation-based «kill»—«anti-kill» antimicrobial system. Plasmid 2015, 82, 28–34. [Google Scholar] [CrossRef]
- Šmajs, D.; Micenková, L.; Šmarda, J.; Vrba, M.; Sevčíková, A.; Vališová, Z.; Woznicová, V. Bacteriocin synthesis in uropathogenic and commensal Escherichia coli: Colicin E1 is a potential virulence factor. BMC Microbiol. 2010, 10, 288. [Google Scholar] [CrossRef]
- Chapman, T.A.; Wu, X.-Y.; Barchia, I.; Bettelheim, K.A.; Driesen, S.; Trott, D.; Wilson, M.; Chin, J.J.-C. Comparison of virulence gene profiles of Escherichia coli strains isolated from healthy and diarrheic swine. Appl. Environ. Microbiol. 2006, 72, 4782–4795. [Google Scholar] [CrossRef] [PubMed]
- Moulin-Schouleur, M.; Répérant, M.; Laurent, S.; Brée, A.; Mignon-Grasteau, S.; Germon, P.; Rasschaert, D.; Schouler, C. Extraintestinal pathogenic Escherichia coli strains of avian and human origin: Link between phylogenetic relationships and common virulence patterns. J. Clin. Microbiol. 2007, 45, 3366–3376. [Google Scholar] [CrossRef] [PubMed]
- Micenková, L.; Bosák, J.; Štaudová, B.; Kohoutová, D.; Čejková, D.; Woznicová, V.; Vrba, M.; Ševčíková, A.; Bureš, J.; Šmajs, D. Microcin determinants are associated with B2 phylogroup of human fecal Escherichia coli isolates. Microbiologyopen 2016, 5, 490–498. [Google Scholar] [CrossRef] [PubMed]
- Riley, M.A.; Gordon, D.M. A survey of Col plasmids in natural isolates of Escherchia coli and an investigation into the stability of Col plasmid lineages. J. Gen. Microbiol. 1992, 138, 1345–1352. [Google Scholar] [CrossRef] [PubMed]
- Feldgarden, M.; Riley, M.A. The phenotypic and fitness effects of colicin resistance in Escherichia coli K12. Evolution 1999, 53, 1019–1027. [Google Scholar] [PubMed]
- Braun, V.; Patzer, S.I.; Hantke, K. Ton-dependent colicins and microcins: Modular design and evolution. Biochimie 2002, 84, 365–380. [Google Scholar] [CrossRef]
- Murinda, S.E.; Roberts, R.F.; Wilson, R.A. Evaluation of colicins for inhibitory activity against diarrheagenic Escherichia coli strains, including serotype O157:H7. Appl. Environ. Microbiol. 1996, 62, 3196–3202. [Google Scholar] [CrossRef]
- FAO/WHO. Joint F.A.O./W.H.O. (Food and Agriculture Organization/World Health Organization) Working Group Report on Drafting Guidelines for the Evaluation of Probiotics in Food; FAO: Rome, Italy; WHO: Geneva, Switzerland, 2002; pp. 1–11. [Google Scholar]
| Year | Number of Isolates (Strains) Isolated from the Source | |||||
|---|---|---|---|---|---|---|
| Chicken | Turkey | Quail | Pig | Rabbit | Cow | |
| 2019 | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 5 (1) |
| 2020 | 7 (6) | 4 (3) | 12 (7) | 5 (3) | 7 (4) | 18 (12) |
| 2021 | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 39 (36) |
| Strain | Source | Diameter of E. coli DH5 α lysis Zone, mm |
|---|---|---|
| ŽP | Not applicable | 6.0 ± 0.8 |
| Ch1 | chicken | 8.5 ± 1.6 |
| Q5 | quail | 13.0 ± 1.6 * |
| Q8 | quail | 8.0 ± 2.2 |
| Q12 | quail | 6.5 ± 1.3 |
| C18 | cow | 9.3 ± 1.0 |
| C19 | cow | 12.3 ± 2.5 * |
| C23 | cow | 6.0 ± 1.4 |
| C25 | cow | 18.8 ± 1.5 * |
| C32 | cow | 7.5 ± 1.0 |
| C40 | cow | 7.0 ± 0.8 |
| C41 | cow | 7.8 ± 0.5 |
| C45 | cow | 8.0 ± 0.0 |
| C48 | cow | 8.3 ± 1.0 |
| C49 | cow | 7.3 ± 0.5 |
| C51 | cow | 10.0 ± 1.4 * |
| C56 | cow | 13.8 ± 2.5 * |
| C61 | cow | 10.0 ± 0.0 * |
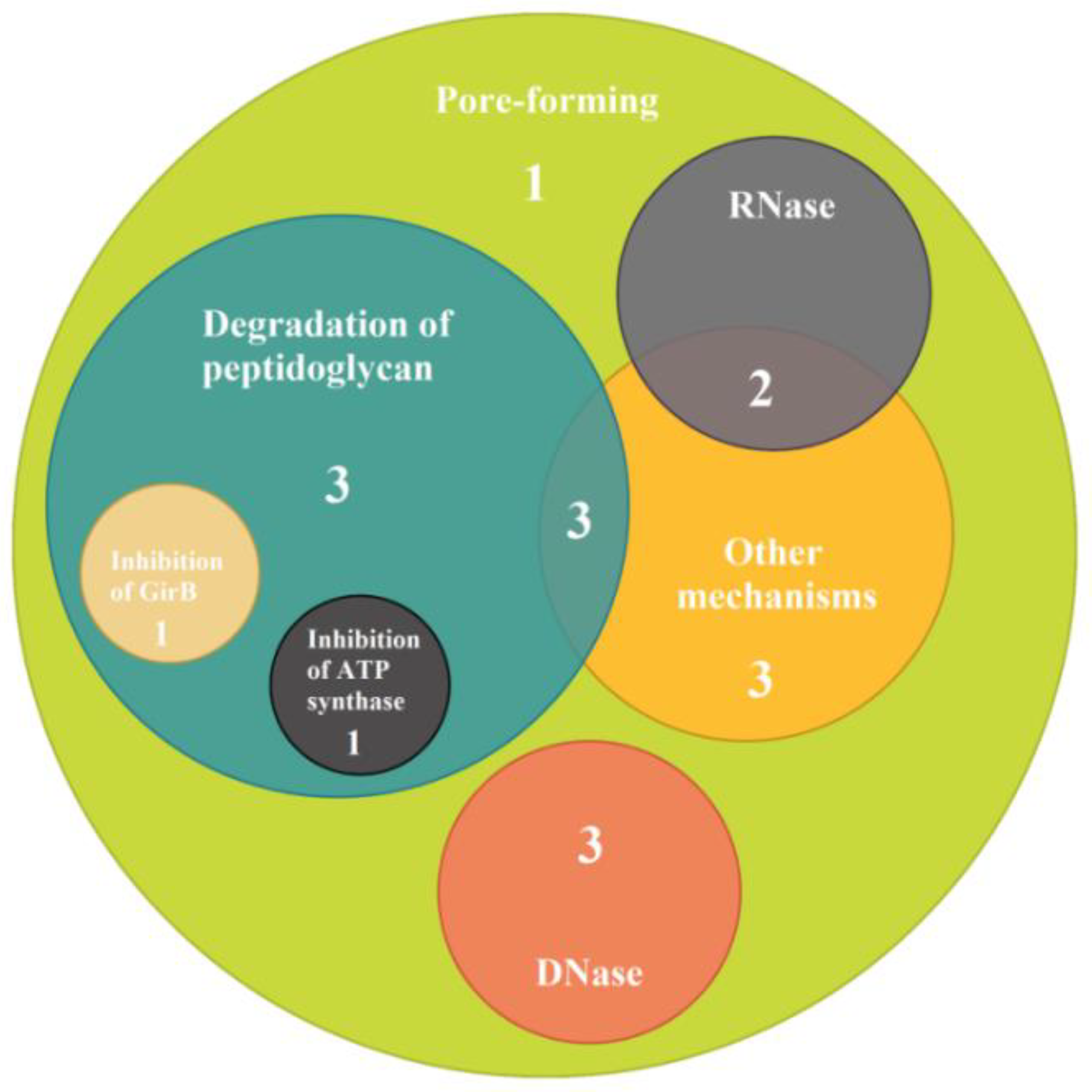
| Mode of Action of Bacteriocins | Bacteriocin Type | Strain | Prevalence, % | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ch1 | Q5 | Q8 | Q12 | C18 | C19 | C23 | C25 | C32 | C40 | C41 | C45 | C48 | C49 | C51 | C56 | C61 | |||
| Pore-forming | mccV | − | − | − | − | − | − | − | − | − | − | − | − | − | − | + | − | − | 5.9 |
| mccL | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | 0.0 | |
| Ia | + | + | − | − | + | − | − | − | − | − | − | − | − | − | + | + | − | 29.4 | |
| Ib | − | + | + | + | − | − | − | − | − | − | − | − | − | − | − | − | + | 23.5 | |
| E1 | + | − | − | − | − | − | − | + | + | + | + | + | + | + | + | − | − | 52.9 | |
| B | + | − | − | − | − | − | + | + | − | − | − | − | − | − | − | − | + | 23.5 | |
| K | − | − | − | − | + | − | − | − | − | − | − | − | − | − | − | − | − | 5.9 | |
| A | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | 0.0 | |
| N | + | − | − | − | − | − | − | + | − | − | − | − | − | − | − | − | − | 11.8 | |
| U | − | + | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | 5.9 | |
| Y | − | + | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | 5.9 | |
| S4 | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | 0.0 | |
| 5 | − | − | − | − | + | + | − | − | − | − | − | − | − | − | − | − | − | 11.8 | |
| 10 | − | − | − | − | + | + | − | − | − | − | − | − | − | − | − | − | − | 11.8 | |
| DNase | E2 | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | 0.0 |
| E7 | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | 0.0 | |
| E8 | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | 0.0 | |
| E9 | − | − | − | − | − | − | − | − | + | + | + | − | − | − | − | − | − | 17.7 | |
| RNase | E3 | − | + | − | − | − | − | − | − | − | − | − | − | − | − | + | − | − | 11.8 |
| E4 | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | 0.0 | |
| E5 | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | 0.0 | |
| E6 | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | 0.0 | |
| D | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | 0.0 | |
| Inhibition of DNA gyrase | mccB17 | + | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | 5.9 |
| Inhibition of RNA polymerase | mccJ25 | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | 0.0 |
| Degradation of peptidoglycan | M | + | − | + | + | − | − | + | + | − | − | − | − | − | − | − | − | + | 35.3 |
| Inhibition of Asp-tRNA synthesis | mccC7 | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | − | 0.0 |
| Inhibition of ATP synthase | mccH47 | − | − | − | − | − | − | − | + | − | − | − | − | − | − | − | + | − | 11.8 |
| Other mechanisms | mccM | + | + | − | − | − | + | + | + | − | − | − | + | + | + | + | + | − | 58.8 |
| Js | − | − | − | − | − | − | − | − | -− | -− | − | − | − | − | − | − | − | 0.0 | |
| Strain | Inhibition Index, % | |||||
|---|---|---|---|---|---|---|
| APEC | IPEC | |||||
| BR4 | BR35 | BR37 | CA29 | CA43 | CA46 | |
| ŽP | 9.2 ± 6.2 | 5.6 ± 9.6 | 6.1 ± 7.9 | 1.7 ± 1.7 | 8.1 ± 7.6 | 0.8 ± 1.4 |
| Ch1 | 31.5 ± 15.8 | 46.1 ± 9.0 * | 37.2 ± 6.0 * | 17.2 ± 0.4 * | 19.8 ± 6.6 * | 20.8 ± 6.7 * |
| Q5 | 38.3 ± 11.0 * | 49.9 ± 7.3 * | 41.3 ± 8.3 * | 22.8 ± 4.5 * | 29.9 ± 3.2 | 31.0 ± 3.7 * |
| Q8 | 47.1 ± 8.8 * | 55.0 ± 7.8 * | 47.6 ± 3.6 * | 20.0 ± 5.9 * | 23.4 ± 4.7 | 24.1 ± 4.2 * |
| Q12 | 28.4 ± 6.3 * | 38.9 ± 3.8 * | 42.7 ± 5.1 * | 24.8 ± 3.2 * | 17.0 ± 6.0 | 17.0 ± 5.3 * |
| C18 | 41.3 ± 11.8 * | 44.8 ± 3.1 * | 35.5 ± 9.3 | 22.7 ± 1.5 * | 11.4 ± 2.1 | 12.7 ± 1.4 * |
| C19 | 30.6 ± 7.5 * | 44.2 ± 3.9 * | 31.5 ± 5.7 * | 24.1 ± 3.1 * | 14.8 ± 3.0 | 15.7 ± 1.6 * |
| C23 | 58.6 ± 11.5 * | 51.7 ± 11.2 * | 43.8 ± 3.9 * | 10.4 ± 1.7 * | 2.7 ± 2.7 | 3.6 ± 2.3 |
| C25 | 34.4 ± 7.5 | 35.6 ± 4.1 | 36.9 ± 5.6 * | 7.8 ± 2.9 * | 6.3 ± 2.6 | 7.1 ± 1.2 * |
| C32 | 42.8 ± 10.4 * | 51.6 ± 1.9 * | 42.1 ± 4.6 * | 20.2 ± 2.6 * | 33.9 ± 7.8 | 32.5 ± 8.6 * |
| C40 | 45.4 ± 5.7 * | 51.8 ± 3.5 * | 40.0 ± 4.2 * | 19.8 ± 0.7 * | 11.8 ± 4.6 | 12.3 ± 4.7 |
| C41 | 46.9 ± 17.6 * | 58.6 ± 6.9 * | 43.2 ± 6.9 * | 24.3 ± 5.9 * | 38.5 ± 5.7 | 36.8 ± 5.9 * |
| C45 | 27.6 ± 3.7 | 34.6 ± 2.9 * | 38.2 ± 2.8 * | 15.4 ± 1.7 * | 3.6 ± 3.1 | 4.5 ± 3.4 |
| C48 | 14.8 ± 16.0 | 31.0 ± 6.0 | 42.3 ± 5.3 * | 17.4 ± 2.6 * | 1.4 ± 2.4 | 3.0 ± 2.6 |
| C49 | 40.9 ± 12.2 * | 47.4 ± 3.6 * | 34.3 ± 6.0 * | 16.6 ± 2.4 * | 0 | 0 |
| C51 | 33.7 ± 10.6 | 41.1 ± 3.8 * | 34.8 ± 5.1 | 18.5 ± 3.1 * | 7.4 ± 6.6 | 7.8 ± 6.1 |
| C56 | 42.4 ± 4.9 * | 48.8 ± 4.1 * | 38.1 ± 2.7 * | 25.7 ± 4.0 * | 21.2 ± 3.5 | 21.4 ± 3.2 * |
| C61 | 19.1 ± 2.6 | 38.3 ± 3.2 * | 31.9 ± 6.9 | 36.6 ± 7.6 * | 42.0 ± 3.6 * | 42.1 ± 6.4 * |
| Test-Strain ID | Pathotype | Insensitivity Profile | The Number of E. coli Strains Whose CFS Inhibited the Growth of Test-Strains after 22 h of Cultivation, % | |||
|---|---|---|---|---|---|---|
| To Antibiotics | To Bacteriocins | <20% | 20–40% | >40% | ||
| BR4 | APEC | Tet | A+B+D+E1-E7+Ia+Ib+K+N+M+S4+B17+C7+V | 2 | 7 | 8 |
| BR35 | APEC | Amp+Cpf+Lfc+Gen+Tet | A+B+D+Ia+Ib+N+M+S4+B17+C7+V | 0 | 5 | 12 |
| BR37 | APEC | Amp+Cpf+Cfo+Cft+Amk+Gen+Tet | A+B+D+E4+E5+Ia+Ib+K+N+M+C7+V | 0 | 9 | 8 |
| CA29 | STEC/ETEC | - | not defined | 8 | 9 | 0 |
| CA43 | STEC | Amp+Cfp+Cfp+Cfr+Azt | not defined | 11 | 6 | 0 |
| CA46 | ETEC | Amp+Cfp+Cfr+Azt+Tet | not defined | 10 | 7 | 0 |
| Strain | Virulence-Associated Genes | Bacteriophage | Hemolytic Activity | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| east1 | ehxA | estI | estII | eltA | stx1 | stx2 | hlyA | hlyF | cnf1 | fimH | afa/draBC | papC | |||
| Ch1 | + | + | − | − | + | − | − | − | + | − | + | + | − | − | − |
| Q5 | − | − | − | − | − | − | − | − | − | − | + | − | − | − | − |
| Q8 | + | + | − | − | − | − | + | − | + | − | + | + | − | + | − |
| Q12 | + | − | − | − | − | − | − | − | + | − | + | + | − | + | − |
| C18 | − | − | − | − | − | − | − | − | − | − | + | − | − | + | − |
| C19 | − | − | − | − | − | − | − | − | − | − | + | + | + | + | − |
| C23 | − | − | − | − | − | − | − | − | − | − | + | − | + | − | − |
| C25 | − | − | − | − | − | − | − | − | − | − | + | + | − | + | − |
| C32 | − | − | − | − | − | − | − | − | − | − | + | + | − | + | − |
| C40 | + | + | + | − | − | − | + | − | − | − | + | + | − | − | − |
| C41 | − | − | − | − | − | − | − | − | − | − | + | − | − | − | − |
| C45 | − | − | − | − | − | − | − | − | + | − | + | − | − | − | + |
| C48 | − | − | − | − | − | − | − | − | + | − | + | + | − | − | − |
| C49 | − | − | − | − | − | − | − | − | + | − | + | + | − | − | − |
| C51 | − | − | − | − | − | − | − | − | + | − | + | + | − | + | − |
| C56 | − | − | − | − | − | − | − | + | − | − | + | − | − | + | − |
| C61 | − | − | − | − | − | − | − | + | − | − | + | + | + | − | − |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kuznetsova, M.V.; Mihailovskaya, V.S.; Remezovskaya, N.B.; Starčič Erjavec, M. Bacteriocin-Producing Escherichia coli Isolated from the Gastrointestinal Tract of Farm Animals: Prevalence, Molecular Characterization and Potential for Application. Microorganisms 2022, 10, 1558. https://doi.org/10.3390/microorganisms10081558
Kuznetsova MV, Mihailovskaya VS, Remezovskaya NB, Starčič Erjavec M. Bacteriocin-Producing Escherichia coli Isolated from the Gastrointestinal Tract of Farm Animals: Prevalence, Molecular Characterization and Potential for Application. Microorganisms. 2022; 10(8):1558. https://doi.org/10.3390/microorganisms10081558
Chicago/Turabian StyleKuznetsova, Marina V., Veronika S. Mihailovskaya, Natalia B. Remezovskaya, and Marjanca Starčič Erjavec. 2022. "Bacteriocin-Producing Escherichia coli Isolated from the Gastrointestinal Tract of Farm Animals: Prevalence, Molecular Characterization and Potential for Application" Microorganisms 10, no. 8: 1558. https://doi.org/10.3390/microorganisms10081558
APA StyleKuznetsova, M. V., Mihailovskaya, V. S., Remezovskaya, N. B., & Starčič Erjavec, M. (2022). Bacteriocin-Producing Escherichia coli Isolated from the Gastrointestinal Tract of Farm Animals: Prevalence, Molecular Characterization and Potential for Application. Microorganisms, 10(8), 1558. https://doi.org/10.3390/microorganisms10081558






