Microbiology and Epidemiology of Escherichia albertii—An Emerging Elusive Foodborne Pathogen
Abstract
1. Introduction
2. Biochemical Properties of Escherichia albertii
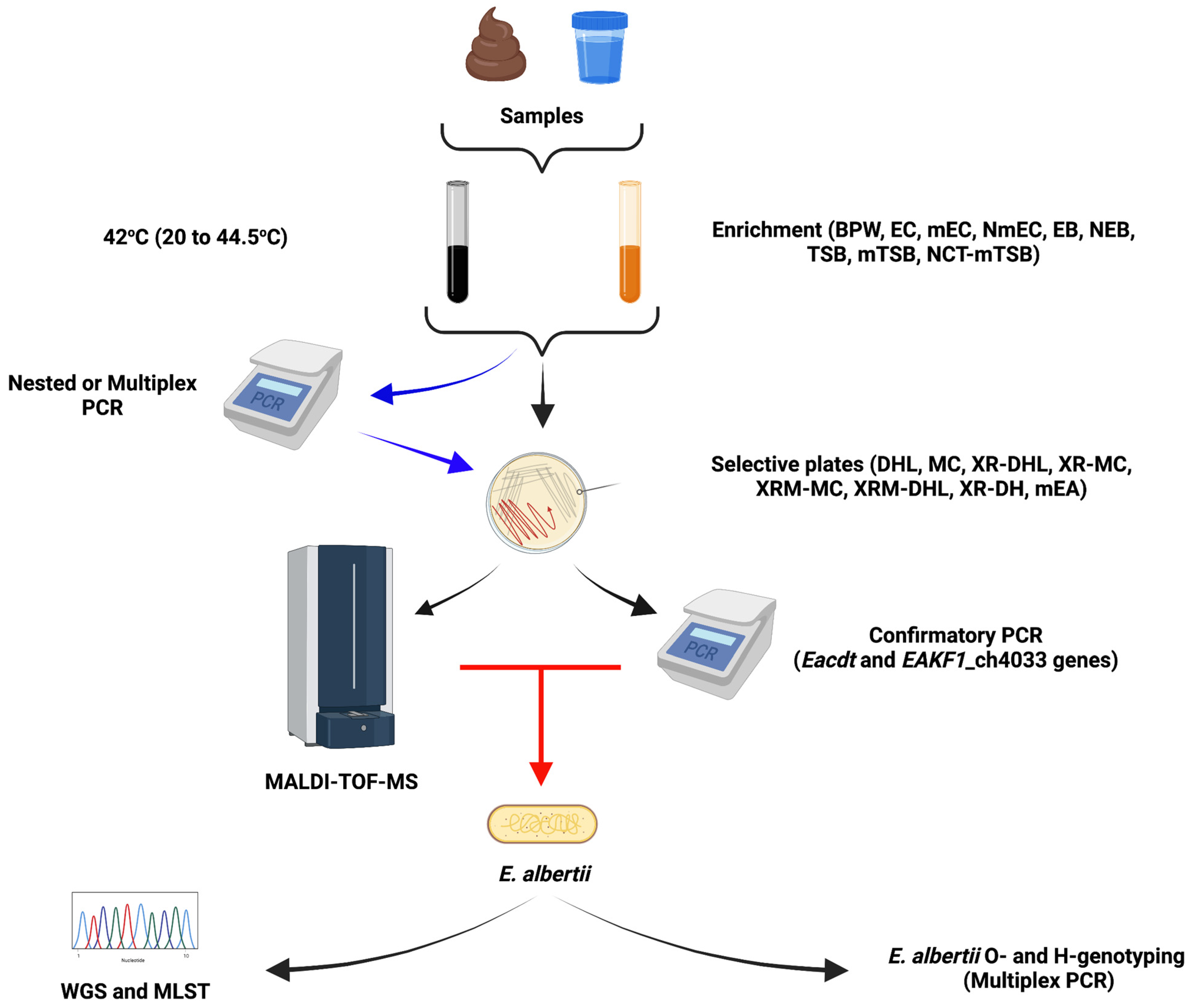
3. Detection and Identification Methods for Escherichia albertii
3.1. Enrichment Broths
3.2. Molecular Approaches
3.3. Selective Agar Plates
Matrix-Assisted Laser Desorption Ionization-Time of Flight Mass Spectrometry
3.4. Characterization by MLST and O-Genotyping
3.5. Escherichia albertii Whole Genome Sequencing
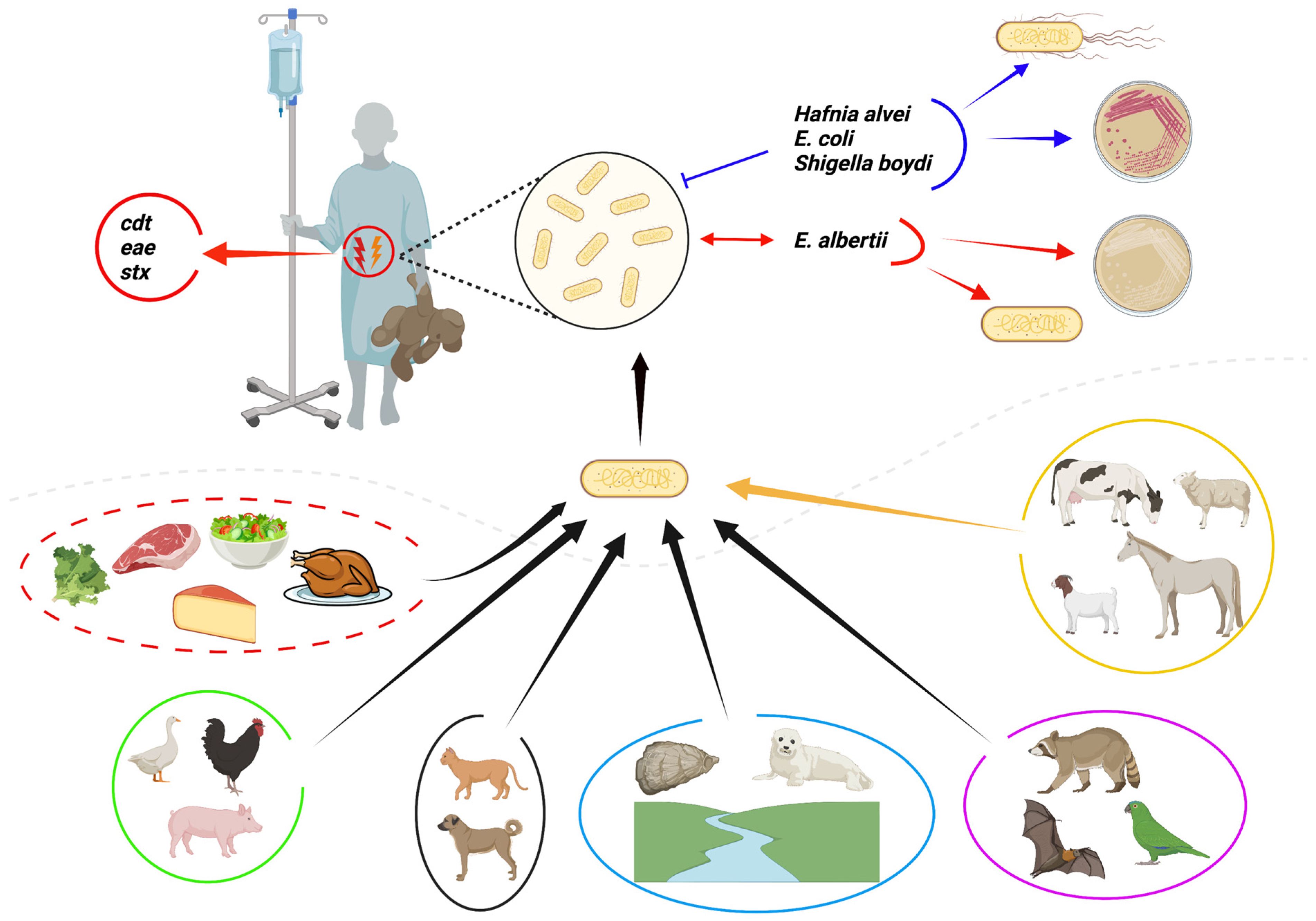
4. Occurrence of Escherichia albertii in Animals
5. Occurrence of Escherichia albertii in Food
6. Pathogenesis and Virulence Factors of Escherichia albertii
7. Clinical Significance of Escherichia albertii
8. Risk Assessment
9. Knowledge Gaps and Future Research Needs
10. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Huys, G.; Cnockaert, M.; Janda, J.M.; Swings, J. Escherichia albertii sp. nov., a diarrhoeagenic species isolated from stool specimens of Bangladeshi children. Int. J. Syst. Evol. Microbiol. 2003, 53 Pt 3, 807–810. [Google Scholar] [CrossRef] [PubMed]
- Oaks, J.L.; Besser, T.E.; Walk, S.T.; Gordon, D.M.; Beckmen, K.B.; Burek, K.A.; Haldorson, G.J.; Bradway, D.S.; Ouellette, L.; Rurangirwa, F.R.; et al. Escherichia albertii in wild and domestic birds. Emerg. Infect. Dis. 2010, 16, 638–646. [Google Scholar] [CrossRef] [PubMed]
- van der Putten, B.C.L.; Matamoros, S.; Mende, D.R.; Scholl, E.R.; Consortium, C.; Schultsz, C. Escherichia ruysiae sp. nov., a novel Gram-stain-negative bacterium, isolated from a faecal sample of an international traveller. Int. J. Syst. Evol. Microbiol. 2021, 71, 004609. [Google Scholar] [CrossRef] [PubMed]
- Ooka, T.; Seto, K.; Ogura, Y.; Nakamura, K.; Iguchi, A.; Gotoh, Y.; Honda, M.; Etoh, Y.; Ikeda, T.; Sugitani, W.; et al. O-antigen biosynthesis gene clusters of Escherichia albertii: Their diversity and similarity to Escherichia coli gene clusters and the development of an O-genotyping method. Microb. Genom. 2019, 5, e000314. [Google Scholar] [CrossRef] [PubMed]
- Gomes, T.A.T.; Ooka, T.; Hernandes, R.T.; Yamamoto, D.; Hayashi, T. Escherichia albertii Pathogenesis. EcoSal Plus 2020, 9. [Google Scholar] [CrossRef]
- Luo, L.; Wang, H.; Payne, M.J.; Liang, C.; Bai, L.; Zheng, H.; Zhang, Z.; Zhang, L.; Zhang, X.; Yan, G.; et al. Comparative genomics of Chinese and international isolates of Escherichia albertii: Population structure and evolution of virulence and antimicrobial resistance. Microb. Genom. 2021, 7, 000710. [Google Scholar] [CrossRef]
- Albert, M.J.; Alam, K.; Islam, M.; Montanaro, J.; Rahaman, A.S.; Haider, K.; Hossain, M.A.; Kibriya, A.K.; Tzipori, S. Hafnia alvei, a probable cause of diarrhea in humans. Infect. Immun. 1991, 59, 1507–1513. [Google Scholar] [CrossRef]
- Albert, M.J.; Faruque, S.M.; Ansaruzzaman, M.; Islam, M.M.; Haider, K.; Alam, K.; Kabir, I.; Robins-Browne, R. Sharing of virulence-associated properties at the phenotypic and genetic levels between enteropathogenic Escherichia coli and Hafnia alvei. J. Med. Microbiol. 1992, 37, 310–314. [Google Scholar] [CrossRef]
- Konno, T.; Yatsuyanagi, J.; Takahashi, S.; Kumagai, Y.; Wada, E.; Chiba, M.; Saito, S. Isolation and Identification of Escherichia albertii from a Patient in an Outbreak of Gastroenteritis. Jpn. J. Infect. Dis. 2012, 65, 203–207. [Google Scholar] [CrossRef]
- Ooka, T.; Seto, K.; Kawano, K.; Kobayashi, H.; Etoh, Y.; Ichihara, S.; Kaneko, A.; Isobe, J.; Yamaguchi, K.; Horikawa, K.; et al. Clinical Significance of Escherichia albertii. Emerg. Infect. Dis. 2012, 18, 488–492. [Google Scholar] [CrossRef]
- Ooka, T.; Tokuoka, E.; Furukawa, M.; Nagamura, T.; Ogura, Y.; Arisawa, K.; Harada, S.; Hayashi, T. Human Gastroenteritis Outbreak Associated with Escherichia albertii, Japan. Emerg. Infect. Dis. 2013, 19, 144–146. [Google Scholar] [CrossRef] [PubMed]
- Asoshima, N.; Matsuda, M.; Shigemura, K.; Honda, M.; Yoshida, H.; Hiwaki, H.; Ogata, K.; Oda, T. Identification of Escherichia albertii as a Causative Agent of a Food-Borne Outbreak Occurred in 2003. Jpn. J. Infect. Dis. 2014, 67, 139–140. [Google Scholar] [CrossRef] [PubMed]
- Ooka, T.; Ogura, Y.; Katsura, K.; Seto, K.; Kobayashi, H.; Kawano, K.; Tokuoka, E.; Furukawa, M.; Harada, S.; Yoshino, S.; et al. Defining the Genome Features of Escherichia albertii, an Emerging Enteropathogen Closely Related to Escherichia coli. Genome Biol. Evol. 2015, 7, 3170–3179. [Google Scholar] [CrossRef]
- Masuda, K.; Ooka, T.; Akita, H.; Hiratsuka, T.; Takao, S.; Fukada, M.; Inoue, K.; Honda, M.; Toda, J.; Sugitani, W.; et al. Epidemiological Aspects of Escherichia albertii Outbreaks in Japan and Genetic Characteristics of the Causative Pathogen. Foodborne Pathog. Dis. 2020, 17, 144–150. [Google Scholar] [CrossRef]
- Donnenberg, M.S.; Tzipori, S.; McKee, M.L.; O’Brien, A.D.; Alroy, J.; Kaper, J.B. The role of the eae gene of enterohemorrhagic Escherichia coli in intimate attachment in vitro and in a porcine model. J. Clin. Investig. 1993, 92, 1418–1424. [Google Scholar] [CrossRef] [PubMed]
- Cabal, A.; García-Castillo, M.; Cantón, R.; Gortázar, C.; Domínguez, L.; Álvarez, J. Prevalence of Escherichia coli Virulence Genes in Patients with Diarrhea and a Subpopulation of Healthy Volunteers in Madrid, Spain. Front. Microbiol. 2016, 7, 641. [Google Scholar] [CrossRef]
- Baba, A.; Ebuchi, S.; Uryu, K.; Hiwaki, H.; Ogata, K.; Washimi, E.; Hasegawa, A.; Utiyama, S. An outbreak of water-borne gastroenteritis caused by diarrheagenic Escherichia coli possessing eae gene. Jpn. J. Infect. Dis. 2006, 59, 59–60. [Google Scholar]
- Tokuoka, E.; Furukawa, K.; Nagamura, T.; Harada, F.; Ekinaga, K.; Tokunaga, H. Food poisoning outbreak due to atypical EPEC OUT:HNM, May 2011–Kumamoto. Infect. Agents Surveill. Rep. 2012, 33, 8–9. [Google Scholar]
- Hyma, K.E.; Lacher, D.W.; Nelson, A.M.; Bumbaugh, A.C.; Janda, J.M.; Strockbine, N.A.; Young, V.B.; Whittam, T.S. Evolutionary Genetics of a New Pathogenic Escherichia Species: Escherichia albertii and Related Shigella boydii Strains. J. Bacteriol. 2005, 187, 619–628. [Google Scholar] [CrossRef]
- Murakami, K.; Etoh, Y.; Tanaka, E.; Ichihara, S.; Horikawa, K.; Kawano, K.; Ooka, T.; Kawamura, Y.; Ito, K. Shiga Toxin 2f-Producing Escherichia albertii from a Symptomatic Human. Jpn. J. Infect. Dis. 2014, 67, 204–208. [Google Scholar] [CrossRef]
- Lindsey, R.L.; Fedorka-Cray, P.J.; Abley, M.; Turpin, J.B.; Meinersmann, R.J. Evaluating the Occurrence of Escherichia albertii in Chicken Carcass Rinses by PCR, Vitek Analysis, and Sequencing of the rpoB Gene. Appl. Environ. Microbiol. 2015, 81, 1727–1734. [Google Scholar] [CrossRef] [PubMed]
- Arai, S.; Yamaya, S.; Ohtsuka, K.; Konishi, N.; Obata, H.; Ooka, T.; Hirose, S.; Kai, A.; Hara-Kudo, Y. Detection of Escherichia albertii in Retail Oysters. J. Food Prot. 2022, 85, 173–179. [Google Scholar] [CrossRef] [PubMed]
- Wakabayashi, Y.; Seto, K.; Kanki, M.; Harada, T.; Kawatsu, K. Proposal of a novel selective enrichment broth, NCT-mTSB, for isolation of Escherichia albertii from poultry samples. J. Appl. Microbiol. 2022, 132, 2121–2130. [Google Scholar] [CrossRef] [PubMed]
- Hinenoya, A.; Ichimura, H.; Yasuda, N.; Harada, S.; Yamada, K.; Suzuki, M.; Iijima, Y.; Nagita, A.; Albert, M.J.; Hatanaka, N.; et al. Development of a specific cytolethal distending toxin (cdt) gene (Eacdt)-based PCR assay for the detection of Escherichia albertii. Diagn. Microbiol. Infect. Dis. 2019, 95, 119–124. [Google Scholar] [CrossRef]
- Maheux, A.F.; Boudreau, D.K.; Bergeron, M.G.; Rodriguez, M.J. Characterization of Escherichia fergusonii and Escherichia albertii isolated from water. J. Appl. Microbiol. 2014, 117, 597–609. [Google Scholar] [CrossRef]
- Hinenoya, A.; Ichimura, H.; Awasthi, S.P.; Yasuda, N.; Yatsuyanagi, J.; Yamasaki, S. Phenotypic and molecular characterization of Escherichia albertii: Further surrogates to avoid potential laboratory misidentification. Int. J. Med. Microbiol. 2019, 309, 108–115. [Google Scholar] [CrossRef]
- Bhatt, S.; Egan, M.; Critelli, B.; Kouse, A.; Kalman, D.; Upreti, C. The Evasive Enemy: Insights into the Virulence and Epidemiology of the Emerging Attaching and Effacing Pathogen Escherichia albertii. Infect. Immun. 2019, 87, e00254-18. [Google Scholar] [CrossRef]
- Takara, T.; Nakama, E.; Kyan, H.; Kakita, T.; Kuba, Y.; Kato, T.; Kudaka, J.; Ameku, C.; Imoto, S.; Nakasone, T.; et al. Escherichia albertii food poisoning due to consumption of Nigana shiraae, seasoned mixed salad of nigana (Ixeris dentata) and tofu, May 2016—Okinawa Prefecture. Infect. Agents Surveill. Rep. 2016, 37, 252–253. [Google Scholar]
- Fewtrell, L.; Kaufmann, R.B.; Kay, D.; Enanoria, W.; Haller, L.; Colford, J.M., Jr. Water, sanitation, and hygiene interventions to reduce diarrhoea in less developed countries: A systematic review and meta-analysis. Lancet Infect. Dis. 2005, 5, 42–52. [Google Scholar] [CrossRef]
- Karinja, M.; Schlienger, R.; Pillai, G.C.; Esterhuizen, T.; Onyango, E.; Gitau, A.; Ogutu, B. Risk reduction of diarrhea and respiratory infections following a community health education program—A facility-based case-control study in rural parts of Kenya. BMC Public Health 2020, 20, 586. [Google Scholar] [CrossRef]
- Janda, J.M.; Abbott, S.L.; Albert, M.J. Prototypal Diarrheagenic Strains of Hafnia alvei Are Actually Members of the Genus Escherichia. J. Clin. Microbiol. 1999, 37, 2399–2401. [Google Scholar] [CrossRef] [PubMed]
- Abbott, S.L.; O’Connor, J.; Robin, T.; Zimmer, B.L.; Janda, J.M. Biochemical Properties of a Newly Described Escherichia Species, Escherichia albertii. J. Clin. Microbiol. 2003, 41, 4852–4854. [Google Scholar] [CrossRef] [PubMed]
- Nimri, L.F. Escherichia albertii, a newly emerging enteric pathogen with poorly defined properties. Diagn. Microbiol. Infect. Dis. 2013, 77, 91–95. [Google Scholar] [CrossRef]
- Inglis, T.J.; Merritt, A.J.; Bzdyl, N.; Lansley, S.; Urosevic, M.N. First bacteraemic human infection with Escherichia albertii. New Microbes New Infect. 2015, 8, 171–173. [Google Scholar] [CrossRef]
- Maheux, A.F.; Brodeur, S.; Bérubé, È.; Boudreau, D.K.; Abed, J.Y.; Boissinot, M.; Bissonnette, L.; Bergeron, M.G. Method for isolation of both lactose-fermenting and—non-fermenting Escherichia albertii strains from stool samples. J. Microbiol. Methods 2018, 154, 134–140. [Google Scholar] [CrossRef] [PubMed]
- Murakami, K.; Maeda-Mitani, E.; Kimura, H.; Honda, M.; Ikeda, T.; Sugitani, W.; Konno, T.; Kawano, K.; Etoh, Y.; Sera, N.; et al. Non-biogroup 1 or 2 Strains of the Emerging Zoonotic Pathogen Escherichia albertii, Their Proposed Assignment to Biogroup 3, and Their Commonly Detected Characteristics. Front. Microbiol. 2019, 10, 1543. [Google Scholar] [CrossRef] [PubMed]
- Arai, S.; Ohtsuka, K.; Konishi, N.; Ohya, K.; Konno, T.; Tokoi, Y.; Nagaoka, H.; Asano, Y.; Maruyama, H.; Uchiyama, H.; et al. Evaluating Methods for Detecting Escherichia albertii in Chicken Meat. J. Food Prot. 2021, 84, 553–562. [Google Scholar] [CrossRef]
- Stock, I.; Rahman, M.; Sherwood, K.J.; Wiedemann, B. Natural antimicrobial susceptibility patterns and biochemical identification of Escherichia albertii and Hafnia alvei strains. Diagn. Microbiol. Infect. Dis. 2005, 51, 151–163. [Google Scholar] [CrossRef]
- Konno, T.; Takahashi, S.; Suzuki, S.; Kashio, H.; Ito, Y.; Kumagai, Y. Distribution of the O-Genotypes of Escherichia albertii Isolated from Humans and Environmental Water in Akita Prefecture, Japan. Jpn. J. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Ridell, J.; Siitonen, A.; Paulin, L.; Lindroos, O.; Korkeala, H.; Albert, M.J. Characterization of Hafnia alvei by biochemical tests, random amplified polymorphic DNA PCR, and partial sequencing of 16S rRNA gene. J. Clin. Microbiol. 1995, 33, 2372–2376. [Google Scholar] [CrossRef]
- Hinenoya, A.; Shima, K.; Asakura, M.; Nishimura, K.; Tsukamoto, T.; Ooka, T.; Hayashi, T.; Ramamurthy, T.; Faruque, S.M.; Yamasaki, S. Molecular characterization of cytolethal dis-tending toxin gene-positive Escherichia coli from healthy cattle and swine in Nara, Japan. BMC Microbiol. 2014, 14, 97. [Google Scholar] [CrossRef] [PubMed]
- Nataro, J.P.; Bopp, C.A.; Fields, P.I.; Kaper, J.B.; Strockbine, N.A. Escherichia, shigella, and salmonella. In Manual of Clinical Microbiology, 9th ed.; Murray, P.R., Baron, E.J., Jorgensen, J.H., Landry, M.L., Pfaller, M.A., Eds.; ASM Press: Washington, DC, USA, 2007; pp. 670–687. [Google Scholar]
- Hinenoya, A.; Yasuda, N.; Mukaizawa, N.; Sheikh, S.; Niwa, Y.; Awasthi, S.P.; Asakura, M.; Tsukamoto, T.; Nagita, A.; Albert, M.J.; et al. Association of cytolethal distending toxin-II gene-positive Escherichia coli with Escherichia albertii, an emerging enteropathogen. Int. J. Med. Microbiol. 2017, 307, 564–571. [Google Scholar] [CrossRef] [PubMed]
- Asoshima, N.; Matsuda, M.; Shigemura, K.; Honda, M.; Yoshida, H.; Oda, T.; Hiwaki, H. Isolation of Escherichia albertii from Raw Chicken Liver in Fukuoka City, Japan. Jpn. J. Infect. Dis. 2015, 68, 248–250. [Google Scholar] [CrossRef] [PubMed]
- Brandal, L.T.; Tunsjø, H.S.; Ranheim, T.E.; Løbersli, I.; Lange, H.; Wester, A.L. Shiga Toxin 2a in Escherichia albertii. J. Clin. Microbiol. 2015, 53, 1454–1455. [Google Scholar] [CrossRef]
- Maeda, E.; Murakami, K.; Sera, N.; Ito, K.; Fujimoto, S. Detection of Escherichia albertii from chicken meat and giblets. J. Vet. Med. Sci. 2015, 77, 871–873. [Google Scholar] [CrossRef]
- Wang, H.; Zheng, H.; Li, Q.; Xu, Y.; Wang, J.; Du, P.; Li, X.; Liu, X.; Zhang, L.; Zou, N.; et al. Defining the Genetic Features of O-Antigen Biosynthesis Gene Cluster and Performance of an O-Antigen Serotyping Scheme for Escherichia albertii. Front. Microbiol. 2017, 8, 1857. [Google Scholar] [CrossRef]
- Wang, H.; Li, Q.; Bai, X.; Xu, Y.; Zhao, A.; Sun, H.; Deng, J.; Xiao, B.; Liu, X.; Sun, S.; et al. Prevalence of eae-positive, lactose non-fermenting Escherichia albertii from retail raw meat in China. Epidemiol. Infect. 2016, 144, 45–52. [Google Scholar] [CrossRef]
- Hinenoya, A.; Nagano, K.; Awasthi, S.P.; Hatanaka, N.; Yamasaki, S. Prevalence of Escherichia albertii in Raccoons (Procyon lotor), Japan. Emerg. Infect. Dis. 2020, 26, 1304–1307. [Google Scholar] [CrossRef]
- Hinenoya, A.; Li, X.; Zeng, X.; Sahin, O.; Moxley, R.A.; Logue, C.M.; Gillespie, B.; Yamasaki, S.; Lin, J. Isolation and characterization of Escherichia albertii in poultry at the pre-harvest level. Zoonoses Public Health 2021, 68, 213–225. [Google Scholar] [CrossRef]
- Lindsey, R.L.; Garcia-Toledo, L.; Fasulo, D.; Gladney, L.M.; Strockbine, N. Multiplex polymerase chain reaction for identification of Escherichia coli, Escherichia albertii and Escherichia fergusonii. J. Microbiol. Methods 2017, 140, 1–4. [Google Scholar] [CrossRef]
- Maeda, E.; Murakami, K.; Okamoto, F.; Etoh, Y.; Sera, N.; Ito, K.; Fujimoto, S. Nonspecificity of Primers for Escherichia albertii Detection. Jpn. J. Infect. Dis. 2014, 67, 503–505. [Google Scholar] [CrossRef] [PubMed]
- Yamasaki, S.; Asakura, M.; Tsukamoto, T.; Faruque, S.M.; Deb, R.; Ramamurthy, T. Cytolethal distending toxin (cdt): Genetic diversity, structure and role in diarrheal disease. Toxin Rev. 2006, 25, 61–88. [Google Scholar] [CrossRef]
- Lukjancenko, O.; Wassenaar, T.M.; Ussery, D.W. Comparison of 61 sequenced Escherichia coli genomes. Microb. Ecol. 2010, 60, 708–720. [Google Scholar] [CrossRef] [PubMed]
- Maheux, A.F.; Bérubé, È.; Boudreau, D.K.; Cantin, P.; Boissinot, M.; Bissonnette, L.; Rodrigue, L.; Bergeron, M.G. Ability of three DNA-based assays to identify presumptive Escherichia coli colonies isolated from water by the culture-based mFC agar method. Water Res. 2011, 45, 2638–2646. [Google Scholar] [CrossRef]
- Hinenoya, A.; Nagano, K.; Okuno, K.; Nagita, A.; Hatanaka, N.; Awasthi, S.P.; Yamasaki, S. Development of XRM-MacConkey agar selective medium for the isolation of Escherichia albertii. Diagn. Microbiol. Infect. Dis. 2020, 97, 115006. [Google Scholar] [CrossRef]
- Bizzini, A.; Durussel, C.; Bille, J.; Greub, G.; Prod’Hom, G. Performance of matrix-assisted laser desorption ionization-time of flight mass spectrometry for identification of bacterial strains routinely isolated in a clinical microbiology laboratory. J. Clin. Microbiol. 2010, 48, 1549–1554. [Google Scholar] [CrossRef]
- Singhal, N.; Kumar, M.; Kanaujia, P.K.; Virdi, J.S. MALDI-TOF mass spectrometry: An emerging technology for microbial identification and diagnosis. Front. Microbiol. 2015, 6, 791. [Google Scholar] [CrossRef]
- Hou, T.-Y.; Chiang-Ni, C.; Teng, S.-H. Current status of MALDI-TOF mass spectrometry in clinical microbiology. J. Food Drug Anal. 2019, 27, 404–414. [Google Scholar] [CrossRef]
- Hatanaka, N.; Awasthi, S.P.; Hinenoya, A.; Ueda, O.; Yamasaki, S. Accurate identification of Escherichia albertii by matrix-assisted laser desorption ionization-time of flight mass spectrometry. J. Microbiol. Methods 2020, 173, 105916. [Google Scholar] [CrossRef]
- van den Beld, M.J.C.; Rossen, J.W.A.; Evers, N.; Kooistra-Smid, M.A.M.D.; Reubsaet, F.A.G. MALDI-TOF MS Using a Custom-Made Database, Biomarker Assignment, or Mathematical Classifiers Does Not Differentiate Shigella spp. and Escherichia coli. Microorganisms 2022, 10, 435. [Google Scholar] [CrossRef]
- Paauw, A.; Jonker, D.; Roeselers, G.; Heng, J.M.; Mars-Groenendijk, R.H.; Trip, H.; Molhoek, E.M.; Jansen, H.-J.; van der Plas, J.; de Jong, A.L.; et al. Rapid and reliable discrimination between Shigella species and Escherichia coli using MALDI-TOF mass spectrometry. Int. J. Med. Microbiol. 2015, 305, 446–452. [Google Scholar] [CrossRef] [PubMed]
- Naumenko, O.I.; Zheng, H.; Senchenkova, S.N.; Wang, H.; Li, Q.; Shashkov, A.S.; Wang, J.; Knirel, Y.A.; Xiong, Y. Structures and gene clusters of the O-antigens of Escherichia albertii O3, O4, O6, and O7. Carbohydr. Res. 2017, 449, 17–22. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Shashkov, A.S.; Xiong, Y.; Naumenko, O.I.; Wang, H.; Senchenkova, S.N.; Wang, J.; Knirel, Y.A. Structure and gene cluster of the O-antigen of Escherichia albertii O1 resembling the O-antigen of Pseudomonas aeruginosa O5. Carbohydr. Res. 2017, 446–447, 28–31. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Naumenko, O.I.; Wang, H.; Xiong, Y.; Wang, J.; Shashkov, A.S.; Li, Q.; Knirel, Y.A. Colitose-containing O-polysaccharide structure and O-antigen gene cluster of Escherichia albertii HK18069 related to those of Escherichia coli O55 and E. coli O128. Carbohydr. Res. 2019, 480, 73–79. [Google Scholar] [CrossRef] [PubMed]
- Nakae, K.; Ooka, T.; Murakami, K.; Hara-Kudo, Y.; Imuta, N.; Gotoh, Y.; Ogura, Y.; Hayashi, T.; Okamoto, Y.; Nishi, J. Diversification of Escherichia albertii H-Antigens and Development of H-Genotyping PCR. Front. Microbiol. 2021, 12, 737979. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Zhang, L.; Cao, L.; Zeng, X.; Gillespie, B.; Lin, J. Isolation and characterization of Escherichia albertii originated from the broiler farms in Mississippi and Alabama. Vet. Microbiol. 2022, 267, 109379. [Google Scholar] [CrossRef]
- Foster, G.; Ross, H.M.; Pennycott, T.W.; Hopkins, G.F.; McLaren, I.M. Isolation of Escherichia coli O86:K61 producing cyto-lethal distending toxin from wild birds of the finch family. Lett. Appl. Microbiol. 1998, 26, 395–398. [Google Scholar] [CrossRef]
- Pennycott, T.W.; Ross, H.M.; McLaren, I.M.; Park, A.; Hopkins, G.F.; Foster, G. Causes of death of wild birds of the family Fringillidae in Britain. Vet. Rec. 1998, 143, 155–158. [Google Scholar] [CrossRef]
- Gordon, D.M. Reservoirs of Infection: The Epidemiological Characteristics of an Emerging Pathogen, Escherichia albertii; Research School of Biology, Australian National University: Canberra, Australia, 2011; Available online: https://pdfs.semanticscholar.org/08fe/1278d6d23e907affbaa25f3b71c413758f3f.pdf (accessed on 4 February 2022).
- Oh, J.-Y.; Kang, M.-S.; Hwang, H.-T.; An, B.-K.; Kwon, J.-H.; Kwon, Y.-K. Epidemiological investigation of eaeA-positive Escherichia coli and Escherichia albertii strains isolated from healthy wild birds. J. Microbiol. 2011, 49, 747–752. [Google Scholar] [CrossRef]
- Grillová, L.; Sedláček, I.; Páchníková, G.; Staňková, E.; Švec, P.; Holochová, P.; Micenková, L.; Bosák, J.; Slaninova, I.; Šmajs, D. Characterization of four Escherichia albertii isolates collected from animals living in Antarctica and Patagonia. J. Vet. Med. Sci. 2018, 80, 138–146. [Google Scholar] [CrossRef]
- Gordon, D.M.; Cowling, A. The distribution and genetic structure of Escherichia coli in Australian vertebrates: Host and geographic effects. Microbiology 2003, 149, 3575–3586. [Google Scholar] [CrossRef] [PubMed]
- Lindberg, A.-M.; Ljungh, Å.; Ahrné, S.; Löfdahl, S.; Molin, G. Enterobacteriaceae found in high numbers in fish, minced meat and pasteurised milk or cream and the presence of toxin encoding genes. Int. J. Food Microbiol. 1998, 39, 11–17. [Google Scholar] [CrossRef]
- Felfö Ldi, T.; Heéger, Z.; Vargha, M.; Márialigeti, K. Detection of potentially pathogenic bacteria in the drinking water distribution system of a hospital in Hungary. Clin. Microbiol. Infect. 2010, 16, 89–92. [Google Scholar] [CrossRef]
- Sharma, M.; Kniel, K.E.; Derevianko, A.; Ling, J.; Bhagwat, A.A. Sensitivity of Escherichia albertii, a potential food-borne pathogen, to food preservation treatments. Appl. Environ. Microbiol. 2007, 73, 4351–4353. [Google Scholar] [CrossRef]
- Lamark, T.; Røkenes, T.P.; McDougall, J.; Strøm, A.R. The complex bet promoters of Escherichia coli: Regulation by oxygen (ArcA), choline (BetI), and osmotic stress. J. Bacteriol. 1996, 178, 1655–1662. [Google Scholar] [CrossRef]
- Hernandes, R.T.; Silva, R.M.; Carneiro, S.M.; Salvador, F.A.; Fernandes, M.C.D.C.; Padovan, A.C.B.; Yamamoto, D.; Mortara, R.A.; Elias, W.P.; Briones, M.R.D.S.; et al. The localized adherence pattern of an atypical enteropathogenic Escherichia coli is mediated by intimin omicron and unexpectedly promotes HeLa cell invasion. Cell. Microbiol. 2008, 10, 415–425. [Google Scholar] [CrossRef]
- Yamamoto, D.; Hernandes, R.T.; Liberatore, A.M.; Abe, C.M.; De Souza, R.B.; Romão, F.T.; Sperandio, V.; Koh, I.; Gomes, T.A.T. Escherichia albertii, a novel human enteropathogen, colonizes rat enterocytes and translocates to extra-intestinal sites. PLoS ONE 2017, 12, e0171385. [Google Scholar] [CrossRef]
- DeVinney, R.; Stein, M.; Reinscheid, D.; Abe, A.; Ruschkowski, S.; Finlay, B.B. Enterohemorrhagic Escherichia coli O157:H7 produces Tir, which is translocated to the host cell membrane but is not tyrosine phosphorylated. Infect. Immun. 1999, 67, 2389–2398. [Google Scholar] [CrossRef]
- Hartland, E.L.; Batchelor, M.; Delahay, R.M.; Hale, C.; Matthews, S.; Dougan, G.; Knutton, S.; Connerton, I.; Frankel, G. Binding of intimin from enteropathogenic Escherichia coli to Tir and to host cells. Mol. Microbiol. 1999, 32, 151–158. [Google Scholar] [CrossRef]
- Liu, H.; Magoun, L.; Luperchio, S.; Schauer, D.B.; Leong, J.M. The Tir-binding region of enterohaemorrhagic Escherichia coli intimin is sufficient to trigger actin condensation after bacterial-induced host cell signalling. Mol. Microbiol. 1999, 34, 67–81. [Google Scholar] [CrossRef]
- Goosney, D.L.; DeVinney, R.; Pfuetzner, R.; Frey, E.A.; Strynadka, N.C.; Finlay, B.B. Enteropathogenic E. coli translocated intimin receptor, Tir, interacts directly with alpha-actinin. Curr. Biol. 2000, 10, 735–738. [Google Scholar] [CrossRef]
- Shaw, R.K.; Daniell, S.; Frankel, G.; Knutton, S. Enteropathogenic Escherichia coli translocate Tir and form an intimin–Tir intimate attachment to red blood cell membranes. Microbiology 2002, 148, 1355–1365. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Donato, K.A.; Zareie, M.; Jassem, A.N.; Jandu, N.; Alingary, N.; Carusone, S.C.; Johnson-Henry, K.; Sherman, P. Escherichia albertii and Hafnia alvei are candidate enteric pathogens with divergent effects on intercellular tight junctions. Microb. Pathog. 2008, 45, 377–385. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, D.; Hernandes, R.T.; Blanco, M.; Greune, L.; Schmidt, M.A.; Carneiro, S.M.; Dahbi, G.; Blanco, J.E.; Mora, A.; Blanco, J.; et al. Invasiveness as a putative additional virulence mechanism of some atypical Enteropathogenic Escherichia coli strains with different uncommon intimin types. BMC Microbiol. 2009, 9, 146. [Google Scholar] [CrossRef] [PubMed]
- Pacheco, V.C.; Yamamoto, D.; Abe, C.M.; Hernandes, R.T.; Mora, A.; Blanco, J.; Gomes, T.A. Invasion of differentiated intestinal Caco-2 cells is a sporadic property among atypical enteropathogenic Escherichia coli strains carrying common intimin subtypes. Pathog. Dis. 2014, 70, 167–175. [Google Scholar] [CrossRef]
- Wieler, L.H.; McDaniel, T.K.; Whittam, T.S.; Kaper, J.B. Insertion site of the locus of enterocyte effacement in enteropathogenic and enterohemorrhagic Escherichia coli differs in relation to the clonal phylogeny of the strains. FEMS Microbiol. Lett. 1997, 156, 49–53. [Google Scholar] [CrossRef]
- Lacher, D.W.; Steinsland, H.; Blank, T.E.; Donnenberg, M.S.; Whittam, T.S. Molecular Molecular evolution of typical enteropathogenic Escherichia coli: Clonal analysis by multilocus sequence typing and virulence gene allelic profiling. J. Bacteriol. 2007, 189, 342–350. [Google Scholar] [CrossRef]
- Lima, M.P.; Yamamoto, D.; Santos, A.C.M.; Ooka, T.; Hernandes, R.T.; Vieira, M.A.M.; Santos, F.F.; Silva, R.M.; Hayashi, T.; Gomes, T. Phenotypic characterization and virulence-related properties of Escherichia albertii strains isolated from children with diarrhea in Brazil. Pathog. Dis. 2019, 77, ftz014. [Google Scholar] [CrossRef]
- Hinenoya, A.; Yasuda, N.; Hibino, T.; Shima, A.; Nagita, A.; Tsukamoto, T.; Yamasaki, S. Isolation and Isolation and characterization of an Escherichia albertii strain producing three different toxins from a child with diarrhea. Jpn. J. Infect. Dis. 2017, 70, 252–257. [Google Scholar] [CrossRef]
- Jinadasa, R.N.; Bloom, S.E.; Weiss, R.S.; Duhamel, G.E. Cytolethal distending toxin: A conserved bacterial genotoxin that blocks cell cycle progression, leading to apoptosis of a broad range of mammalian cell lineages. Microbiology 2011, 157, 1851–1875. [Google Scholar] [CrossRef]
- Fedor, Y.; Vignard, J.; Nicolau-Travers, M.-L.; Boutet-Robinet, E.; Watrin, C.; Salles, B.; Mirey, G. From single-strand breaks to double-strand breaks during S-phase: A new mode of action of the Escherichia coli Cytolethal Distending Toxin. Cell. Microbiol. 2013, 15, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Taieb, F.; Petit, C.; Nougayrede, J.-P.; Oswald, E. The enterobacterial genotoxins: Cytolethal distending toxin and colibactin. EcoSal Plus 2016, 7. [Google Scholar] [CrossRef]
- Ge, Z.; Feng, Y.; Whary, M.T.; Nambiar, P.R.; Xu, S.; Ng, V.; Taylor, N.S.; Fox, J.G. Cytolethal distending toxin is essential for Helicobacter hepaticus colonization in outbred Swiss Webster mice. Infect. Immun. 2005, 73, 3559–3567. [Google Scholar] [CrossRef]
- Scuron, M.D.; Boesze-Battaglia, K.; Dlakic, M.; Shenker, B.J. The Cytolethal Distending Toxin Contributes to Microbial Virulence and Disease Pathogenesis by Acting As a Tri-Perditious Toxin. Front. Cell. Infect. Microbiol. 2016, 6, 168. [Google Scholar] [CrossRef] [PubMed]
- Iyoda, S.; Ishijima, N.; Lee, K.; Ishihara, T.; Ohnishi, M. Stx2f-positive Escherichia albertii isolated from a patient with hemolytic uremic syndrome, August 2016. Infect. Agents Surveill. Rep. 2016, 37, 255. [Google Scholar]
- De Rauw, K.; Jacobs, S.; Piérard, D. Twenty-seven years of screening for Shiga toxin-producing Escherichia coli in a university hospital. Brussels, Belgium, 1987–2014. PLoS ONE 2018, 13, e0199968. [Google Scholar] [CrossRef] [PubMed]
- Ori, E.L.; Takagi, E.H.; Andrade, T.S.; Miguel, B.T.; Cergole-Novella, M.C.; Guth, B.E.C.; Hernandes, R.T.; Dias, R.C.B.; Pinheiro, S.R.S.; Camargo, C.H.; et al. Diarrhoeagenic Escherichia coli and Escherichia albertii in Brazil: Pathotypes and serotypes over a 6-year period of surveillance. Epidemiol. Infect. 2019, 147, e10. [Google Scholar] [CrossRef] [PubMed]
- Boerlin, P.; McEwen, S.A.; Boerlin-Petzold, F.; Wilson, J.B.; Johnson, R.P.; Gyles, C.L. Associations between virulence factors of Shiga toxin-producing Escherichia coli and disease in humans. J. Clin. Microbiol. 1999, 37, 497–503. [Google Scholar] [CrossRef]
- Tesh, V.L.; Burris, J.A.; Owens, J.W.; Gordon, V.M.; Wadolkowski, E.A.; O’Brien, A.D.; Samuel, J.E. Comparison of the relative toxicities of Shiga-like toxins type I and type II for mice. Infect. Immun. 1993, 61, 3392–3402. [Google Scholar] [CrossRef]
- Friedrich, A.W.; Bielaszewska, M.; Zhang, W.-L.; Pulz, M.; Kuczius, T.; Ammon, A.; Karch, H. Escherichia coli harboring Shiga toxin 2 gene variants: Frequency and association with clinical symptoms. J. Infect. Dis. 2002, 185, 74–84. [Google Scholar] [CrossRef]
- Fuller, C.A.; Pellino, C.A.; Flagler, M.J.; Strasser, J.E.; Weiss, A.A. Shiga toxin subtypes display dramatic differences in potency. Infect. Immun. 2011, 79, 1329–1337. [Google Scholar] [CrossRef] [PubMed]
- Bielaszewska, M.; Idelevich, E.A.; Zhang, W.; Bauwens, A.; Schaumburg, F.; Mellmann, A.; Peters, G.; Karch, H. Effects of antibiotics on Shiga toxin 2 production and bacteriophage induction by epidemic Escherichia coli O104:H4 strain. Antimicrob. Agents Chemother. 2012, 56, 3277–3282. [Google Scholar] [CrossRef] [PubMed]
- Kenny, B.; Lai, L.-C.; Finlay, B.B.; Donnenberg, M.S. EspA, a protein secreted by enteropathogenic Escherichia coli, is required to induce signals in epithelial cells. Mol. Microbiol. 1996, 20, 313–323. [Google Scholar] [CrossRef] [PubMed]
- O’Connell, C.B.; Creasey, E.A.; Knutton, S.; Elliott, S.; Crowther, L.J.; Luo, W.; Albert, M.J.; Kaper, J.B.; Frankel, G.; Donnenberg, M.S. SepL, a protein required for enteropathogenic Escherichia coli type III translocation, interacts with secretion component SepD. Mol. Microbiol. 2004, 52, 1613–1625. [Google Scholar] [CrossRef]
- Yen, H.; Sugimoto, N.; Tobe, T. Enteropathogenic Escherichia coli Uses NleA to Inhibit NLRP3 Inflammasome Activation. PLoS Pathog. 2015, 11, e1005121. [Google Scholar] [CrossRef]
- Bhatt, S.; Egan, M.; Jenkins, V.; Muche, S.; El-Fenej, J. The tip of the iceberg: On the roles of regulatory small RNAs in the virulence of enterohemorrhagic and enteropathogenic Escherichia coli. Front. Cell. Infect. Microbiol. 2016, 6, 105. [Google Scholar] [CrossRef]
- Egan, M.; Ramirez, J.; Xander, C.; Upreti, C.; Bhatt, S. Lambda red-mediated recombineering in the attaching and effacing pathogen Escherichia albertii. Biol. Proced. Online 2016, 18, 3. [Google Scholar] [CrossRef]
- Schroeder, M.R.; Juieng, P.; Batra, D.; Knipe, K.; Rowe, L.A.; Sheth, M.; Smith, P.; Garcia-Toledo, L.; Loparev, V.N.; Lindsey, R.L. High-quality complete and draft genome sequences for three Escherichia spp. and three Shigella spp. generated with Pacific Biosciences and Illumina sequencing and optical mapping. Genome Announc. 2018, 6, e01384-17. [Google Scholar] [CrossRef]
- Lindsey, R.L.; Rowe, L.A.; Batra, D.; Smith, P.; Strockbine, N.A. PacBio Genome Sequences of Eight Escherichia albertii Strains Isolated from Humans in the United States. Microbiol. Resour. Announc. 2019, 8, e01663-18. [Google Scholar] [CrossRef]
- Sulaiman, M.A.; Aminu, M.; Ella, E.E.; Abdullahi, I.O. Prevalence and risks factors of the novel Escherichia albertii among gastroenteritis patients in Kano State, Nigeria. J. Med. Trop. 2021, 23, 39–45. [Google Scholar] [CrossRef]
- Afshin, N.; Bahareh, R.; Azadeh, F. Detection of Escherichia albertii in urinary and gastrointestinal infections in Kermanshah, Iran Research square. Int. J. Enteric Pathog. 2020, 9, 37–42. [Google Scholar] [CrossRef]
- Zaki, M.E.S.; Eid, A.E.; El-Kazzaz, S.S.; El-Sabbagh, A.M. Molecular Study of Escherichia albertii in Pediatric Urinary Tract Infections. Open Microbiol. J. 2021, 15, 139–144. [Google Scholar] [CrossRef]
- La Ragione, R.M.; McLaren, I.M.; Foster, G.; Cooley, W.A.; Woodward, M.J. Phenotypic and genotypic characterization of avian Escherichia coli O86:K61 isolates possessing a gamma-like intimin. Appl. Environ. Microbiol. 2002, 68, 4932–4942. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Wang, H.; Xu, Y.; Bai, X.; Wang, J.; Zhang, Z.; Liu, X.; Miao, Y.; Zhang, L.; Li, X.; et al. Multidrug-resistant Escherichia albertii: Co-occurrence of beta-lactamase and MCR-1 encoding genes. Front. Microbiol. 2018, 9, 258. [Google Scholar] [CrossRef] [PubMed]
- Kotloff, K.L.; Nataro, J.P.; Blackwelder, W.C.; Nasrin, D.; Farag, T.H.; Panchalingam, S.; Wu, Y.; Sow, S.O.; Sur, D.; Breiman, R.F.; et al. Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global Enteric Multicenter Study, GEMS): A prospective, case-control study. Lancet 2013, 382, 209–222. [Google Scholar] [CrossRef]
- Guerrant, R.L.; Oriá, R.B.; Moore, S.R.; Oriá, M.O.; Lima, A.A. Malnutrition as an enteric infectious disease with long-term effects on child development. Nutr. Rev. 2008, 66, 487–505. [Google Scholar] [CrossRef]
- Proulx, F.; Seidman, E.G.; Karpman, D. Pathogenesis of Shiga toxin-associated hemolytic uremic syndrome. Pediatr. Res. 2001, 50, 163–171. [Google Scholar] [CrossRef]
- Hernandes, R.T.; De la Cruz, M.A.; Yamamoto, D.; Girón, J.A.; Gomes, T.A. Dissection of the role of pili and type 2 and 3 secretion systems in adherence and biofilm formation of an atypical enteropathogenic Escherichia coli strain. Infect. Immun. 2013, 81, 3793–3802. [Google Scholar] [CrossRef]
- Costerton, J.W.; Stewart, P.S.; Greenberg, E.P. Bacterial biofilms: A common cause of persistent infections. Science 1999, 284, 1318–1322. [Google Scholar] [CrossRef]
- Beloin, C.; Roux, A.; Ghigo, J.M. Escherichia coli biofilms. Curr. Top. Microbiol. Immunol. 2008, 322, 249–289. [Google Scholar] [CrossRef]

| Condition | Phenotype | Condition | Phenotype |
|---|---|---|---|
| Lactose | -v | Acid from: Glycerol * | +v |
| D-Xylose | -v | D-Glucose (+gas) | + |
| Melibiose | -v | Adonitol | - |
| Sucrose | -v | Amygdalin | - |
| L-Rhamnose | -v | D-Arabinose | v |
| D-Sorbitol | v | Indole | +v |
| D-Arabitol | - | Oxidase | - |
| Cellobiose ** | - | Catalase | + |
| Acetate | +v | Voges-Proskauer (25 °C) | - |
| D-Mannitol | +v | Voges-Proskauer (35 °C) | - |
| Trehalose | +v | Lysine decarboxylase | +v |
| Maltose | v | Arginine dihydrolase (4 days) | -v |
| Lactulose | v | Ornithine decarboxylase (4 days) | +v |
| D-Arabinose | +v | Glutamate decarboxylase | + |
| Sedoheptulose anhydride | - | β-galactosidase | +v |
| D-Galactose | + | Pigment | - |
| D-Mannose | + | Urea | - |
| D-Ribose | + | MUG | -v |
| D-Fructose | + | PPA | - |
| Raffinose | -v | Methyl red | + |
| Salicin | - | Nitrate reductase | + |
| D-Lyxose | - | ONPG | +v |
| L-Arabinose | +v | Protease | - |
| Dulcitol | - | H2S production on: TSI GCF | -- |
| Citrate | -v | Pectinase | - |
| D-Fucose | - | Degradation of: Mucate | - |
| Palatinose | - | Elastin | - |
| L-Sorbose | +v | Gelatin | - |
| D-Tagatose | -v | Hide powder | - |
| alpha-Methyl-D-glucoside | - | Polypectate (25 °C) | - |
| Erythritol | - | Tyrosine crystals | - |
| i-Inositol | - | DNA | - |
| D-Turanose, | - | Corn oil (lipase) | - |
| Xylitol | - | Hydrolysis of: Esculin | v |
| Malonate | - | Arbutin | - |
| Growth in KCN broth | - | L-Prolineaminopeptidase | v |
| 3-Hydroxybenzoate assimilation | + | 2-Ketogluconate assimilation | - |
| Myoinositol utilization | - | Histidine assimilation | - |
| Biogroup | Indole Production | Lysine Decarboxylase | Acid Production from D-Sorbitol |
|---|---|---|---|
| 1 | - | + | + |
| 2 | + | - | - |
| 3 | + | + | NA |
| Gene | Primers a | Total Tested b | E. albertii | Missed c | non-E. albertii d |
|---|---|---|---|---|---|
| Eacdt | fw: GCTTAACTGGATGATTCTTG rv: CTATTTCCCATCCAATAGTCT | 319 | 310 | 6 | 3 |
| EAKF1_ch4033 | fw: GTAAATAATGCTGGTCAGACGTTA rv: AGTGTAGAGTATATTGGCAACTTC | 319 | 305 | 11 | 3 |
| Gene | Primers a | Total Tested e | E. albertii | Missed c | non-E. alberti f |
| Eacdt | fw: GCTTAACTGGATGATTCTTG rv: CTATTTCCCATCCAATAGTCT | 178 | 20 | 6 | 1 |
| EAKF1_ch4033 | fw: GTAAATAATGCTGGTCAGACGTTA rv: AGTGTAGAGTATATTGGCAACTTC | 178 | 25 | 1 | 0 |
| Gene | Function or Annotation | Reference |
|---|---|---|
| LEE | Locus of enterocyte effacement | [5,13,78] |
| eae | Intimin; formation of attaching-effacing lesions | [5,13,78] |
| ETT2 | E. coli type III secretion system 2 | [13,14] |
| paa | Porcine attaching-effacing associated protein | [90] |
| cdtABC | Cytolethal distending toxin | [5,6,91] |
| stx | Shiga toxin | [5,6,13,39,45,91,97,99] |
| hlyABCD | Cytotoxicity | [6] |
| iuc-ABCD | Iron acquisition and transport | [6] |
| ent and fep | Enterobactin synthesis/iron acquisition | [48,67] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Muchaamba, F.; Barmettler, K.; Treier, A.; Houf, K.; Stephan, R. Microbiology and Epidemiology of Escherichia albertii—An Emerging Elusive Foodborne Pathogen. Microorganisms 2022, 10, 875. https://doi.org/10.3390/microorganisms10050875
Muchaamba F, Barmettler K, Treier A, Houf K, Stephan R. Microbiology and Epidemiology of Escherichia albertii—An Emerging Elusive Foodborne Pathogen. Microorganisms. 2022; 10(5):875. https://doi.org/10.3390/microorganisms10050875
Chicago/Turabian StyleMuchaamba, Francis, Karen Barmettler, Andrea Treier, Kurt Houf, and Roger Stephan. 2022. "Microbiology and Epidemiology of Escherichia albertii—An Emerging Elusive Foodborne Pathogen" Microorganisms 10, no. 5: 875. https://doi.org/10.3390/microorganisms10050875
APA StyleMuchaamba, F., Barmettler, K., Treier, A., Houf, K., & Stephan, R. (2022). Microbiology and Epidemiology of Escherichia albertii—An Emerging Elusive Foodborne Pathogen. Microorganisms, 10(5), 875. https://doi.org/10.3390/microorganisms10050875






