Organelle Engineering in Yeast: Enhanced Production of Protopanaxadiol through Manipulation of Peroxisome Proliferation in Saccharomyces cerevisiae
Abstract
1. Introduction
2. Materials and Methods
2.1. Strains and Media
2.2. Plasmid Construction
2.3. Strain Construction
2.4. Isolation of Peroxisomes and Quantification of Proteins Present in the Isolated Peroxisomes
2.5. Analysis of the Sensitivity of Strains to Oxidative Stress
2.6. Bioreactor Fermentation
2.7. Fluorescence Microscopy and Image Processing
2.8. FACS Analysis
2.9. TEM Analysis
2.10. Extraction and Quantification of Dammarenediol II and Protopanaxadiol
2.11. Statistical Analysis
3. Results
3.1. Engineering of Peroxisome Proliferation in S.cerevisiae
3.2. Sensitivity of Peroxisome-Engineered S.cerevisiae to Oxidative Stress
3.3. Optimization of Peroxisome Proliferation in S.cerevisiae
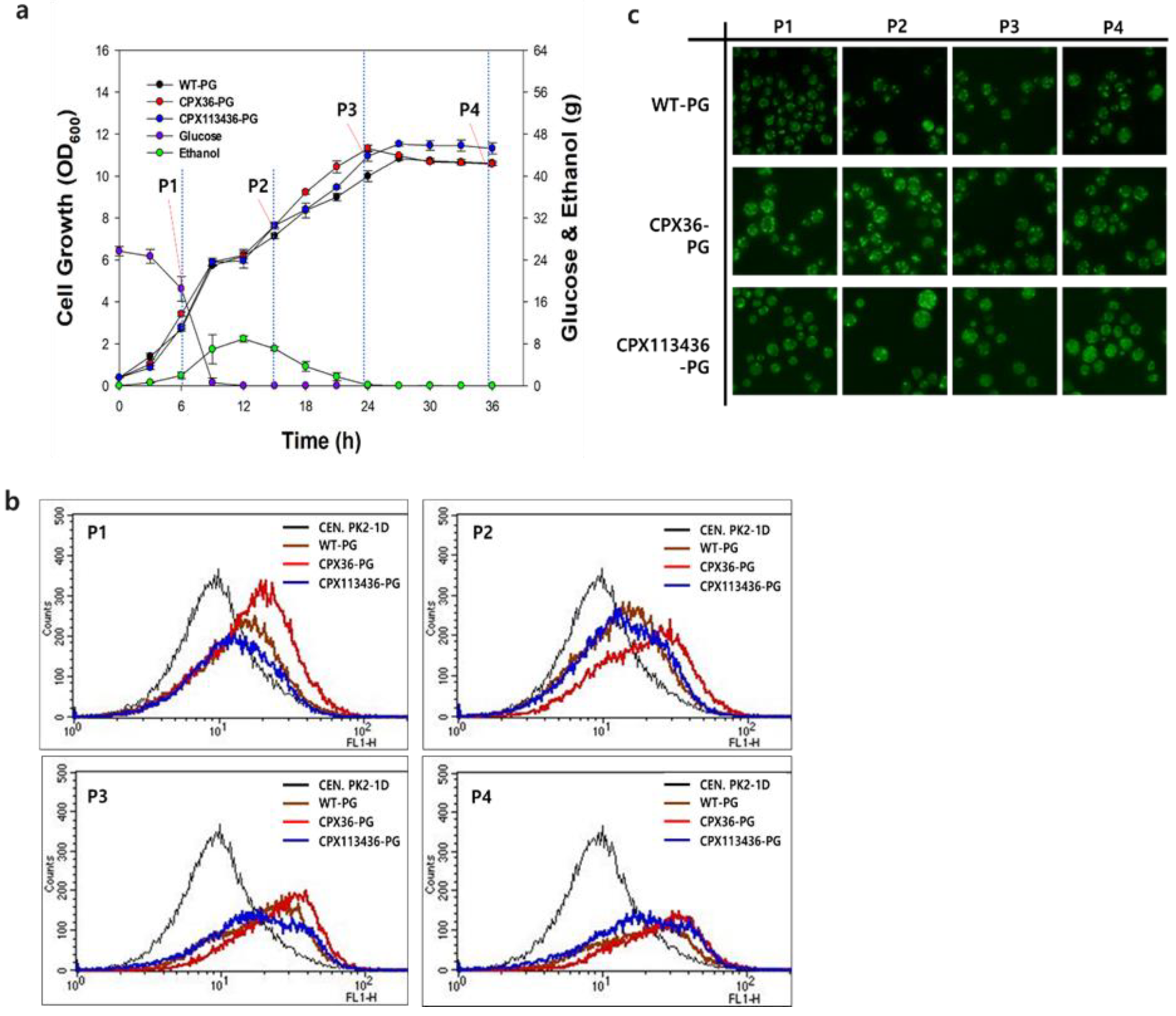
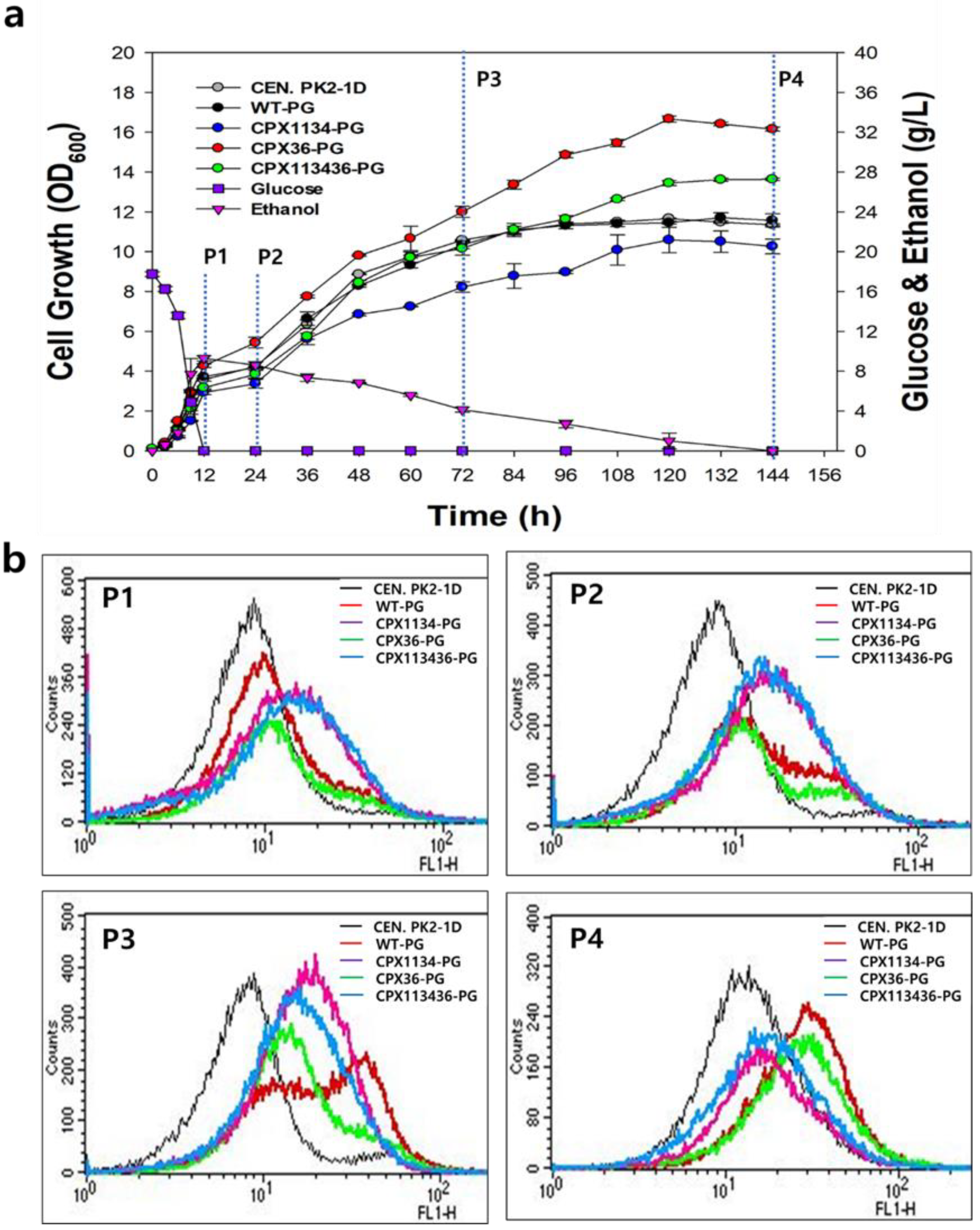
3.4. Peroxisome Stability in Engineered Strains
3.5. Effect of Oleic Acid Supplementation on Peroxisome Biogenesis in Engineered Strains
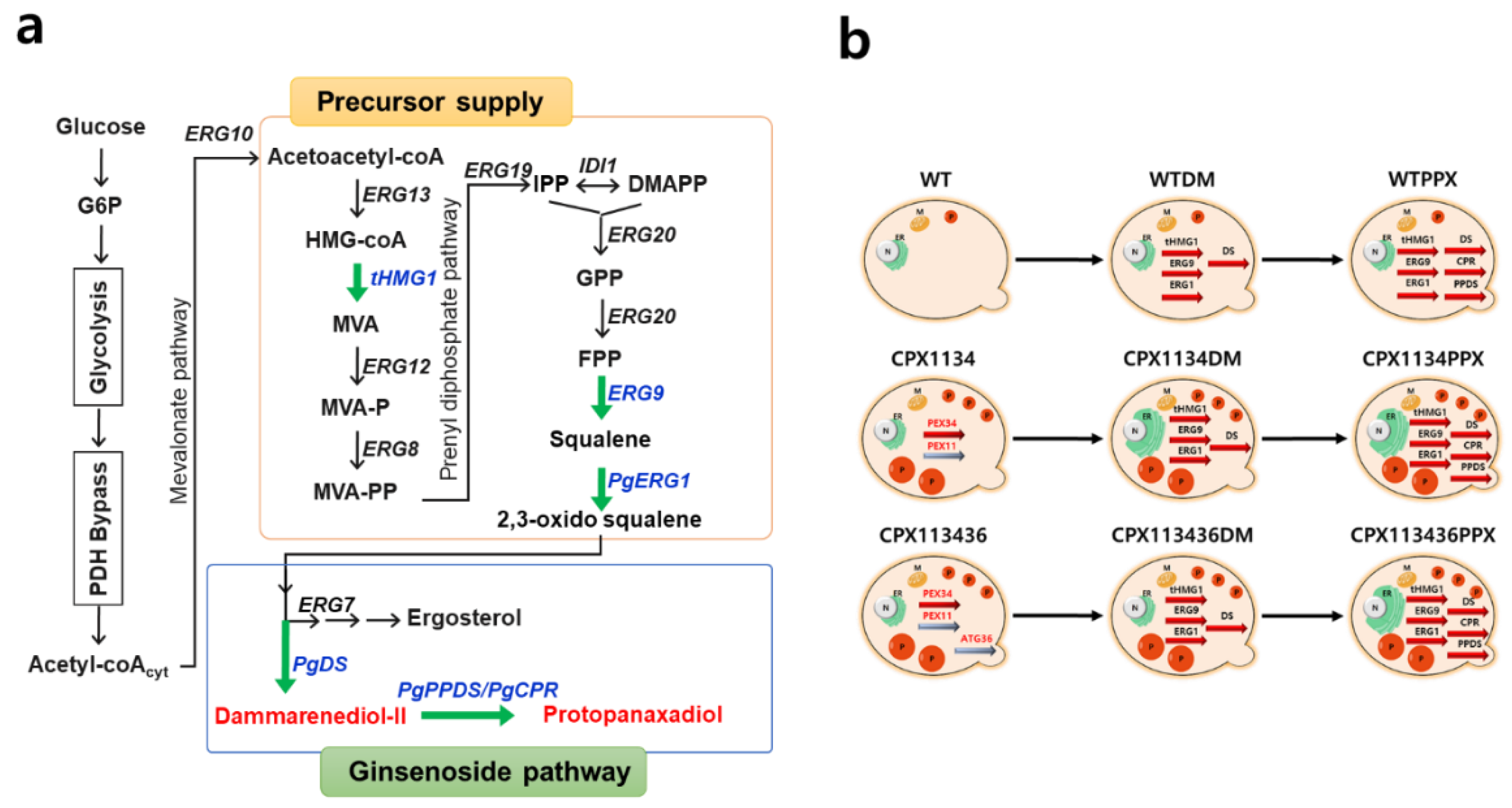
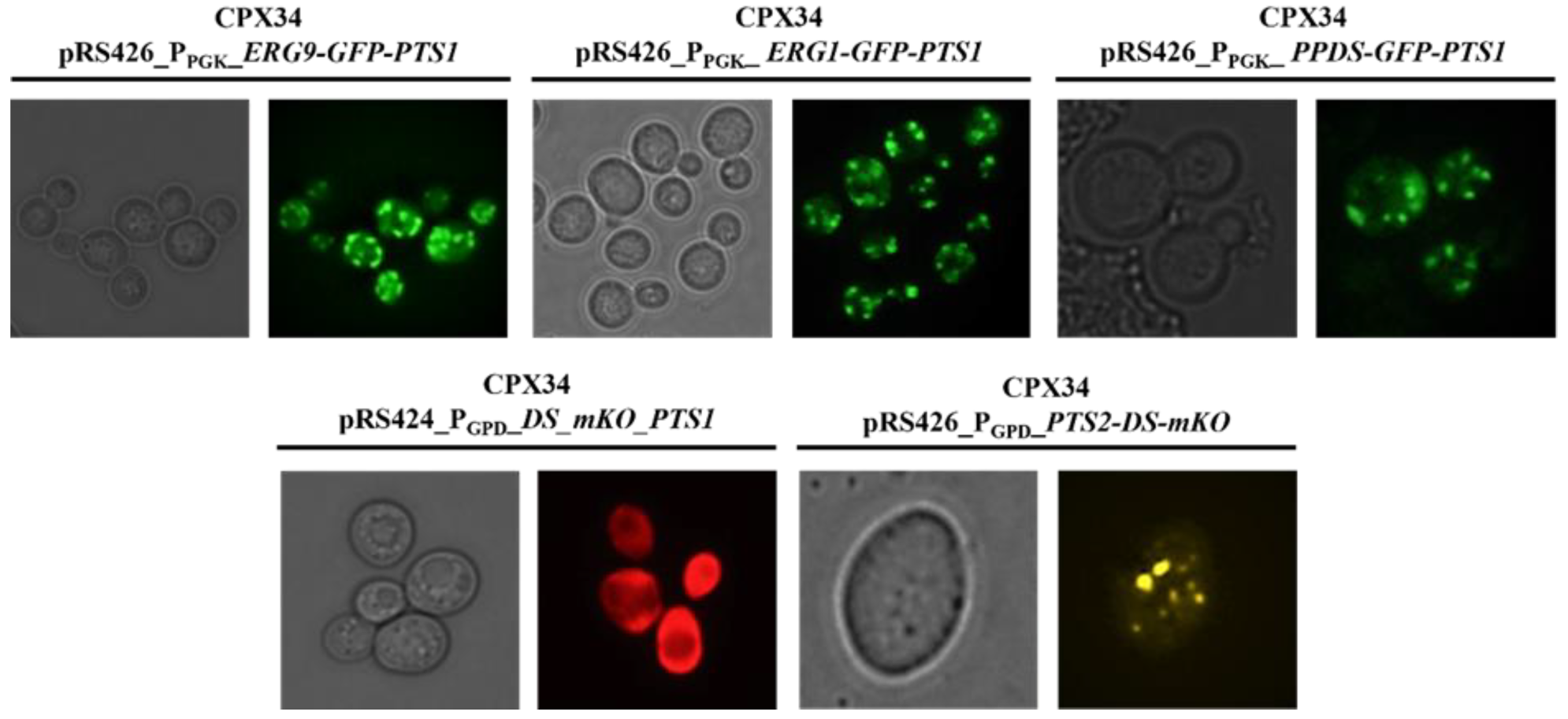
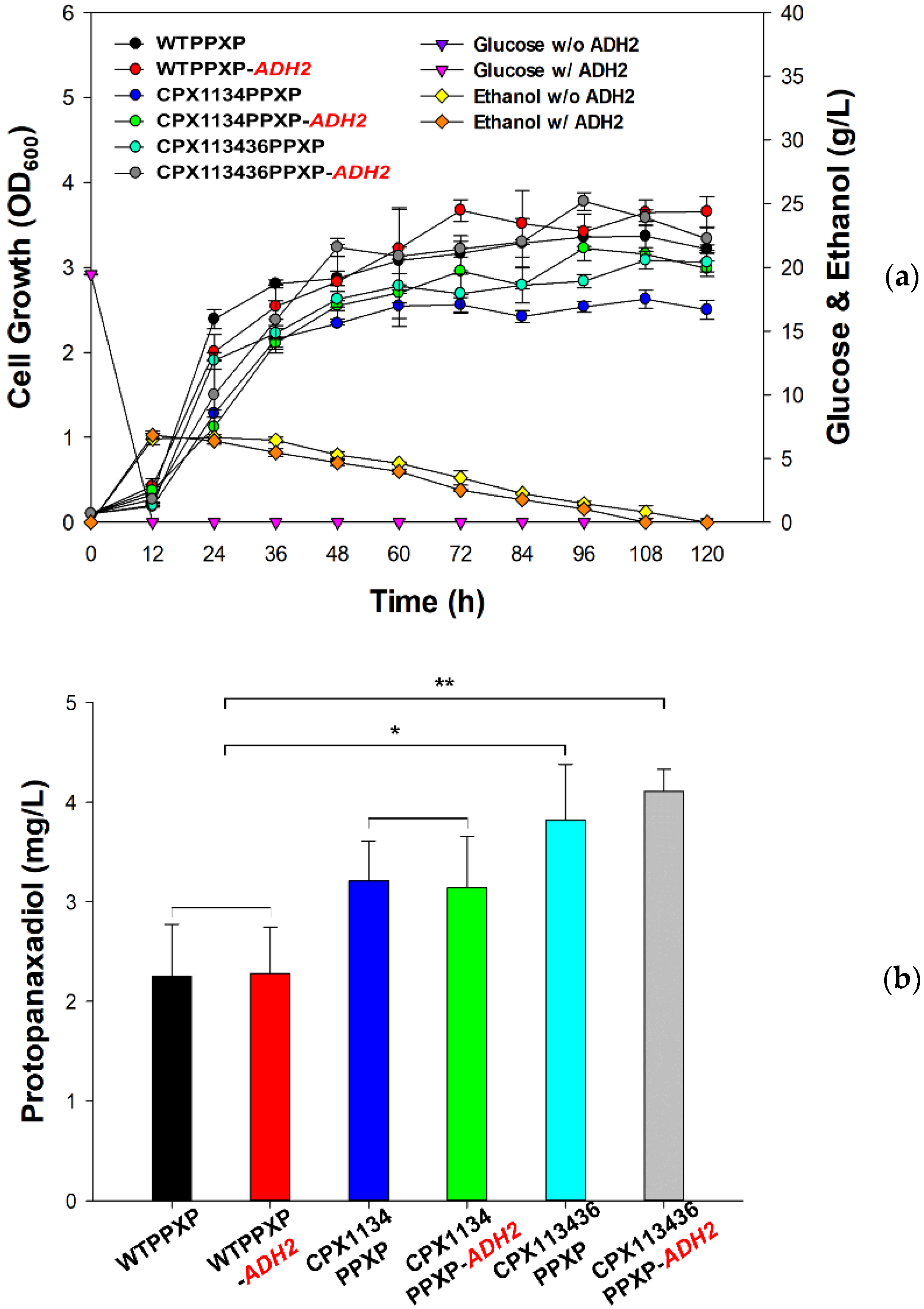
3.6. Construction of the Protopanaxadiol Pathway in Peroxisome-Engineered Strains
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Huccetogullari, D.; Luo, Z.W.; Lee, S.Y. Metabolic engineering of microorganisms for production of aromatic compounds. Microb. Cell Fact. 2019, 18, 41. [Google Scholar] [CrossRef] [PubMed]
- Keasling, J.; Martin, H.G.; Lee, T.S.; Mukhopadhyay, A.; Singer, S.W.; Sundstrom, E. Microbial production of advanced biofuels. Nat. Rev. Microbiol. 2021, 19, 701–715. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Nielsen, J. Recent trends in metabolic engineering of microbial chemical factories. Curr. Opin. 2019, 60, 188–197. [Google Scholar] [CrossRef] [PubMed]
- Choi, K.R.; Jang, W.D.; Yang, D.; Cho, J.S.; Park, D.; Lee, S.Y. Systems Metabolic Engineering Strategies: Integrating Systems and Synthetic Biology with Metabolic Engineering. Trends Biotechnol. 2019, 37, 817–837. [Google Scholar] [CrossRef] [PubMed]
- Eisenreich, W.; Bacher, A.; Arigoni, D.; Rohdich, F. Biosynthesis of isoprenoids via the non-mevalonate pathway. Cell. Mol. Life Sci. 2004, 61, 1401–1426. [Google Scholar] [CrossRef] [PubMed]
- Wilding, E.I.; Brown, J.R.; Bryant, A.P.; Chalker, A.F.; Holmes, D.J.; Ingraham, K.A.; Iordanescu, S.; So, C.Y.; Rosenberg, M.; Gwynn, M.N. Identification, evolution, and essentiality of the mevalonate pathway for isopentenyl diphosphate biosynthesis in gram-positive cocci. J. Bacteriol. 2000, 182, 4319–4327. [Google Scholar] [CrossRef]
- Wang, Z.; Zhang, R.; Yang, Q.; Zhang, J.; Zhao, Y.; Zheng, Y.; Yang, J. Recent advances in the biosynthesis of isoprenoids in engineered Saccharomyces cerevisiae. Adv. Appl. Microbiol. 2021, 114, 1–35. [Google Scholar] [PubMed]
- Luo, Y.; Li, B.Z.; Liu, D.; Zhang, L.; Chen, Y.; Jia, B.; Zeng, B.X.; Zhao, H.; Yuan, Y.J. Engineered biosynthesis of natural products in heterologous hosts. Chem. Soc. 2015, 44, 5265–5290. [Google Scholar] [CrossRef] [PubMed]
- Singh, B.P.; Rateb, M.E.; Couto, S.R.; Polizeli, M.L.T.M.; Li, W.J. Editorial Microbial Secondary Metabolites Recent Developments and Technological Challenges. Front. Microbiol. 2019, 26, 914. [Google Scholar] [CrossRef]
- Ko, Y.S.; Kim, J.W.; Lee, J.A.; Han, T.H.; Kim, G.B.; Park, J.W.; Lee, S.Y. Tools and strategies of systems metabolic engineering for the development of microbial cell factories for chemical production. Chem. Soc. 2020, 49, 4615–4636. [Google Scholar] [CrossRef]
- Hong, K.K.; Nielsen, J. Metabolic engineering of Saccharomyces cerevisiae: A key cell factory platform for future biorefineries. Cell. Mol. Life Sci. 2012, 69, 2671–2690. [Google Scholar] [CrossRef] [PubMed]
- Buchholz, K.; Collins, J. The roots—A short history of industrial microbiology and biotechnology. Appl. Microbiol. Biotechnol. 2013, 97, 3747–3762. [Google Scholar] [CrossRef] [PubMed]
- Farhi, M.; Marhevka, E.; Masci, T.; Marcos, E.; Eyal, Y.; Ovadis, M.; Abeliovich, H.; Vainstein, A. Harnessing yeast subcellular compartments for the production of plant terpenoids. Metab. Eng. 2011, 13, 474–481. [Google Scholar] [CrossRef] [PubMed]
- Dueber, J.E.; Wu, G.C.; Malmirchegini, G.R.; Moon, T.S.; Petzold, C.J.; Ullal, A.V.; Prather, K.L.; Keasling, J.D. Synthetic protein scaffolds provide modular control over metabolic flux. Nat. Biotechnol. 2009, 27, 753–759. [Google Scholar] [CrossRef] [PubMed]
- Lv, X.; Wang, F.; Zhou, P.; Ye, L.; Xie, W.; Xu, H.; Yu, H. Dual regulation of cytoplasmic and mitochondrial acetyl-CoA utilization for improved isoprene production in Saccharomyces cerevisiae. Nat. Commun. 2016, 7, 12851. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.; Ching, C.B. Mitochondrial acetyl-CoA utilization pathway for terpenoid productions. Metab. Eng. 2016, 38, 303–309. [Google Scholar] [CrossRef]
- Szczebara, F.M.; Chandelier, C.; Villeret, C.; Masurel, A.; Bourot, S.; Duport, C.; Blanchard, S.; Groisillier, A.; Testet, E.; Costaglioli, P.; et al. Total biosynthesis of hydrocortisone from a simple carbon source in yeast. Nat. Biotechnol. 2003, 21, 143–149. [Google Scholar] [CrossRef]
- Gidijala, L.; Kiel, J.A.K.W.; Douma, R.D.; Seifar, R.M.; van Gulik, W.M.; Bovenberg, R.A.L.; Veenhuis, M.; van der Klei, I.J. An engineered yeast efficiently secreting penicillin. PLoS ONE 2009, 4, e8317. [Google Scholar] [CrossRef]
- Zhou, Y.J.; Buijs, N.A.; Zhu, Z.; Gomez, D.O.; Boonsombuti, A.; Siewers, V.; Nielsen, J. Harnessing Yeast Peroxisomes for Biosynthesis of Fatty-Acid-Derived Biofuels and Chemicals with Relieved Side-Pathway Competition. J. Am. Chem. Soc. 2016, 138, 15368–15377. [Google Scholar] [CrossRef]
- Haddouche, R.; Delessert, S.; Sabirova, J.; Neuveglise, C.; Poirier, Y.; Nicaud, J.M. Roles of multiple acyl-CoA oxidases in the routing of carbon flow towards beta-oxidation and polyhydroxyalkanoate biosynthesis in Yarrowia lipolytica. FEMS Yeast Res. 2010, 10, 917–927. [Google Scholar] [CrossRef]
- Liu, G.S.; Li, T.; Zhou, W.; Jiang, M.; Tao, X.Y.; Liu, M.; Zhao, M.; Ren, Y.H.; Gao, B.; Wang, F.Q.; et al. The yeast peroxisome: A dynamic storage depot and subcellular factory for squalene overproduction. Metab. Eng. 2020, 57, 151–161. [Google Scholar] [CrossRef] [PubMed]
- Kate Thodey, S.G.; Christina, D.S. A microbial biomanufacturing platform for natural and semisynthetic opioids. Nat. Chem. Biol. 2014, 10, 837–844. [Google Scholar] [CrossRef] [PubMed]
- Bayer, T.S.; Widmaier, D.M.; Karsten, T.; Ethan, A.M.; Daniel, V.S.; Christopher, A.V. Synthesis of methyl halides from biomass using engineered microbes. J. Am. Chem. Soc. 2009, 131, 6508–6515. [Google Scholar] [CrossRef] [PubMed]
- Ayer, A.; Sanwald, J.; Pillay, B.A.; Meyer, A.J.; Perrone, G.G.; Dawes, I.W. Distinct redox regulation in sub-cellular compartments in response to various stress conditions in Saccharomyces cerevisiae. PLoS ONE 2013, 8, e65240. [Google Scholar] [CrossRef] [PubMed]
- Veenhuis, M.; Mateblowski, M.; Kunau, W.H.; Harder, W. Proliferation of microbodies in Saccharomyces cerevisiae. Yeast 1987, 3, 77–84. [Google Scholar] [CrossRef] [PubMed]
- Kunau, W.-H.; Dommes, V.; Schulz, H. β-Oxidation of fatty acids in mitochondria, peroxisomes, and bacteria: A century of continued progress. Prog. Lipids Res. 1995, 34, 267–342. [Google Scholar] [CrossRef]
- Liang, Y.L.; Zhao, S.J.; Xu, L.X.; Zhang, X.Y. Heterologous expression of dammarenediol synthase gene in an engineered Saccharomyces cerevisiae. Lett. Appl. Microbiol. 2012, 55, 323–329. [Google Scholar] [CrossRef]
- Chu, L.L.; Montecillo, J.A.V.; Bae, H. Recent Advances in the Metabolic Engineering of Yeasts for Ginsenoside Biosynthesis. Front. Bioeng. Biotechnol. 2020, 8, 139. [Google Scholar] [CrossRef]
- Bradford, M.M. A rapid and sensitive method for the quantification of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Liu, J.; Wisniewski, M.; Droby, S.; Vero, S.; Tian, S.; Hershkovitz, V. Glycine betaine improves oxidative stress tolerance and biocontrol efficacy of the antagonistic yeast Cystofilobasidium infirmominiatum. Int. J. Food Microbiol. 2011, 146, 76–83. [Google Scholar] [CrossRef]
- Vidhya, B.; Amandeep, G.; Archana, P.; Meenkshi, V.; Vibha, T.; Basant, K.P. Use of ade1 and ade2 mutations for development of a versatile red/white colour assay of amyloid-induced oxidative stress in saccharomyces cerevisiae. Yeast 2016, 33, 607–620. [Google Scholar]
- Wang, G.S.; Grammel, H.; Abou-Aisha, K.; Sagesser, R.; Ghosh, R. High-level production of the industrial product lycopene by the photosynthetic bacterium Rhodospirillum rubrum. Appl. Environ. Microbiol. 2012, 78, 7205–7215. [Google Scholar] [CrossRef]
- Yu, H.; Braun, P.; Yildirim, M.A.; Lemmens, I.; Venkatesan, K.; Sahalie, J.; Hirozane-Kishikawa, T.; Gebreab, F.; Li, N.; Simonis, N.; et al. High-quality binary protein interaction map of the yeast interactome network. Science 2008, 322, 104–110. [Google Scholar] [CrossRef] [PubMed]
- Tower, R.J.; Fagarasanu, A.; Aitchison, J.D.; Rachubinski, R.A. The peroxin Pex34p functions with the Pex11 family of peroxisomal divisional proteins to regulate the peroxisome population in yeast. Mol. Biol. Cell. 2011, 22, 1727–1738. [Google Scholar] [CrossRef] [PubMed]
- Tam, Y.Y.; Torres-Guzman, J.C.; Vizeacoumar, F.J.; Smith, J.J.; Marelli, M.; Aitchison, J.D.; Rachubinski, R.A. Pex11-related proteins in peroxisome dynamics: A role for the novel peroxin Pex27p in controlling peroxisome size and number in Saccharomyces cerevisiae. Mol. Biol. Cell. 2003, 14, 4089–4102. [Google Scholar] [CrossRef] [PubMed]
- Motley, A.M.; Nuttall, J.M.; Hettema, E.H. Pex3-anchored Atg36 tags peroxisomes for degradation in Saccharomyces cerevisiae. EMBO J. 2012, 31, 2852–2868. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Singh, R.; Williams, C.P.; van der Klei, I.J. Stress exposure results in increased peroxisomal levels of yeast Pnc1 and Gpd1, which are imported via a piggy-backing mechanism. Biochim. Biophys. Acta Bioenerg. 2016, 1863, 148–156. [Google Scholar] [CrossRef]
- Saraya, R.; Veenhuis, M.; van der Klei, I.J. Peroxisomes as dynamic organelles: Peroxisome abundance in yeast. FEBS J. 2010, 277, 3279–3288. [Google Scholar] [CrossRef]
- Veenhuis, M.; van der Klei, I.J. A critical reflection on the principles of peroxisome formation in yeast. Front. Physiol. 2014, 5, 110. [Google Scholar] [CrossRef]
- Dansen, T.B.; Wirtz, K.W. The Peroxisome in Oxidative Stress. IUBMB Life 2001, 51, 223–230. [Google Scholar]
- Titorenko, V.I.; Terlecky, S.R. Peroxisome metabolism and cellular aging. Traffic 2011, 12, 252–259. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Zhang, H.; Yang, Q.; Komla, M.G.; Zhang, X.; Zhu, S. Ascorbic acid enhances oxidative stress tolerance and biological control efficacy of Pichia caribbica against postharvest blue mold decay of apples. J. Agric. Food Chem. 2014, 62, 7612–7621. [Google Scholar] [CrossRef]
- Campbell, K.; Vowinckel, J.; Keller, M.A.; Ralser, M. Methionine Metabolism Alters Oxidative Stress Resistance via the Pentose Phosphate Pathway. Antioxid. Redox Signal. 2016, 24, 543–547. [Google Scholar] [CrossRef] [PubMed]
- Hughes, A.L.; Gottschling, D.E. An early age increase in vacuolar pH limits mitochondrial function and lifespan in yeast. Nature 2012, 492, 261–265. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.J.; Brown, T.W.; Eitzen, G.A.; Rachubinski, R.A. Regulation of peroxisome size and number by fatty acid beta -oxidation in the yeast yarrowia lipolytica. J. Biol. Chem. 2000, 275, 20168–20178. [Google Scholar] [CrossRef]
- Erdmann, R.; Blobel, G. Giant peroxisomes in oleic acid-induced Saccharomyces cerevisiae lacking the peroxisomal membrane protein Pmp27p. J. Cell Biol. 1995, 1238, 509–523. [Google Scholar] [CrossRef] [PubMed]
- Kingsman, S.M.; Kingsman, A.J.; Dobson, M.J.; Mellor, J.; Roberts, N.A. Heterologous gene expression in Saccharomyces cerevisiae. Biotechnol. Genet. Eng. Rev. 1985, 3, 377–416. [Google Scholar] [CrossRef] [PubMed]
- Okumoto, K.; Noda, H.; Fujiki, Y. Distinct modes of ubiquitination of peroxisome-targeting signal type 1 (PTS1) receptor Pex5p regulate PTS1 protein import. J. Biol. Chem. 2014, 289, 14089–14108. [Google Scholar] [CrossRef] [PubMed]
- Schafer, A.; Kerssen, D.; Veenhuis, M.; Kunau, W.H.; Schliebs, W. Functional similarity between the peroxisomal PTS2 receptor binding protein Pex18p and the N-terminal half of the PTS1 receptor Pex5p. Mol. Cell. Biol. 2004, 24, 8895–8906. [Google Scholar] [CrossRef] [PubMed]
- Sibirny, A.A. Yeast peroxisomes: Structure, functions and biotechnological opportunities. FEMS Yeast Res. 2016, 16. [Google Scholar] [CrossRef] [PubMed]
- Kunze, M. The type-2 peroxisomal targeting signal. Biochim. Biophys. Acta (BBA)-Mol. Cell Res. 2020, 1867, 118609. [Google Scholar] [CrossRef]
- Maestre, O.; Garcia-Martinez, T.; Peinado, R.A.; Mauricio, J.C. Effects of ADH2 overexpression in Saccharomyces bayanus during alcoholic fermentation. Appl. Environ. Microbiol. 2008, 74, 702–707. [Google Scholar] [CrossRef]
- Martínez, J.L.; Petranovic, D.; Nielsen, J. Heme metabolism in stress regulation and protein production: From Cinderella to a key player. Bioengineered 2016, 7, 112–115. [Google Scholar] [CrossRef]
- Ferreira, R.; Teixeira, P.G.; Gossing, M.; David, F.; Siewers, V.; Nielsen, J. Metabolic engineering of Saccharomyces cerevisiae for overproduction of triacylglycerols. Metab. Eng. Commun. 2018, 6, 22–27. [Google Scholar] [CrossRef]
- Ma, T.; Shi, B.; Ye, Z.; Li, X.; Lu, M.; Chen, Y.; Xia, J.; Nielsen, J.; Deng, Z.; Liu, T. Lipid engineering combined with systematic metabolic engineering of Saccharomyces cerevisiae for high-yield production of lycopene. Metab. Eng. 2019, 52, 134–142. [Google Scholar] [CrossRef]
- Zhou, Y.J.; Buijs, N.A.; Siewers, V.; Nielsen, J. Fatty acid-derived biofuels and chemicals production in Saccharomyces cerevisiae. Bioeng. Biotechnol. 2014, 2, 32. [Google Scholar] [CrossRef]
- Grewal, P.S.; Samson, J.A.; Baker, J.J.; Choi, B.; Dueber, J.E. Peroxisome compartmentalization of a toxic enzyme improves alkaloid production. Nat. Chem. Biol. 2021, 17, 96–103. [Google Scholar] [CrossRef]
- Dusseaux, S.; Wajn, W.T.; Liu, Y.; Kampranis, S.C. Transforming yeast peroxisomes into microfactories for the efficient production of high-value isoprenoids. Proc. Natl. Acad. Sci. USA 2020, 117, 31789–31799. [Google Scholar] [CrossRef]
- Cao, X.; Yang, S.; Cao, C.; Zhou, Y.J. Harnessing sub-organelle metabolism for biosynthesis of isoprenoids in yeast. Synth. Syst. Biotechnol. 2020, 5, 179–186. [Google Scholar] [CrossRef]
- Meadows, A.L.; Hawkins, K.M.; Tsegaye, Y.; Antipov, E.; Kim, Y.; Raetz, L.; Dahl, R.H.; Tai, A.; Mahatdejkul-Meadows, T.; Xu, L.; et al. Rewriting yeast central carbon metabolism for industrial isoprenoid production. Nature 2016, 537, 694–697. [Google Scholar] [CrossRef]
- Ajikumar, P.K.; Xiao, W.H.; Tyo, K.E.J.; Wang, Y.; Simeon, F.; Leonard, E.; Mucha, O.; Phon, T.H.; Pfeifer, B.; Stephanopoulos, G. Isoprenoid pathway optimization for Taxol precursor overproduction in Escherichia coli. Science 2010, 330, 70–74. [Google Scholar] [CrossRef]
- Yoshikuni, Y.; Dietrich, J.A.; Noweoozi, F.F.; Babbitt, P.C.; Keasling, J.D. Redesigning enzymes based on adaptive evolution for optimal function in synthetic metabolic pathway. Chem. Biol. 2008, 15, 607–618. [Google Scholar] [CrossRef]


| Strains | Relevant Properties | Source or Reference |
|---|---|---|
| Saccharomyces cerevisiae | ||
| CEN.PK2-1D | MATa/αura3-52 trp1-289 leu2-3_112 his3∆1 MAL2-8C SUC2 | This study |
| CEN-P11 | CEN.PK2-1D, ΔPEX11::PTRP1-TRP1-TTRP1 | This study |
| CEN-P30 | CEN.PK2-1D, ΔPEX30::PTRP1-TRP1-TTRP1 | This study |
| CEN-P5 | CEN.PK2-1D, TRP1::PPGK1-PEX5-TCYC1 | This study |
| CEN-P34-5 | CEN-P34, Leu2::PPGK1-PEX5-TCYC1 | This study |
| CPX34 | CEN.PK2-1D, TRP1::PPGK1-PEX34-TCYC1 | This study |
| CPX1134 | CEN.PK2-1D, ΔPEX11:: 3MYC-PPGK1-PEX34-TCYC1-3MYC | This study |
| CPX36 | CPX36, ΔPEX11:: 3MYC-PPGK1-PEX34-TCYC1-3MYC | This study |
| CPX113436 | CPX36, ΔPEX11:: 3MYC-PPGK1-PEX34-TCYC1-3MYC | This study |
| WT-PG | CEN.PK2-1D, POT1::PPOT1-POT1-GFP-TPOT1 | This study |
| CPX34-PG | CPX34, POT1::PPOT1-POT1-GFP-TPOT1 | This study |
| CPX1134-PG | CPX1134, POT1::PPOT1-POT1-GFP-TPOT1 | This study |
| CPX36-PG | CPX36, POT1::PPOT1-POT1-GFP-TPOT1 | This study |
| CPX113436-PG | CPX113436, POT1::PPOT1-POT1-GFP-TPOT1 | This study |
| WTDM | CEN.PK2-1D, Leu2::PGPD-ERG9-TCYC1-PPGK1-ERG1pg-TCYC1, TRP1::PTEF1-DSpg-TCYC1-PGPD-tHMG1-TCYC1 | This study |
| WTDMP | CEN.PK2-1D, Leu2::PGPD-ERG9-PTS1-TCYC1-PPGK1-ERG1pg-PTS1-TCYC1, TRP1::PTEF1-DS-PTS2-TCYC-PGPD-tHMG1-TCYC1 | This study |
| CPX1134DM | CPX1134, Leu2::PGPD-ERG9-TCYC1-PPGK1-ERG1pg-TCYC1, TRP1::PTEF1-DSpg-TCYC1-PGPD-tHMG1-TCYC1 | This study |
| CPX1134DMP | CPX1134, Leu2::PGPD-ERG9-PTS1-TCYC1-PPGK1-ERG1pg-PTS1-TCYC1, TRP1::PTEF1-DS-PTS2-TCYC-PGPD-tHMG1-TCYC1 | This study |
| WTPPXP | WTDM, URA3:: PPGK1-PPDSpg-PTS1-TCYC1-PTEF1-CPRpg-TCYC1 | This study |
| CPX1134PPXP | CPX1134DMP, URA3:: PPGK1-PPDSpg-PTS1-TCYC1-PTEF1-CPRpg-TCYC1 | This study |
| CPX113436PPXP | CPX1134DMP, ΔATG36::3MYC-PURA3-URA3-TURA3-3MYC, URA3:: PPGK1-PPDSpg-PTS1-TCYC1-PTEF1-CPRpg-TCYC1 | This study |
| Escherichia coli | ||
| XL1-Blue | endA1 gyrA96(nalR) thi-1 recA1 relA1 lac glnV44 F’[::Tn10 proAB+ lacIq Δ(lacZ)M15 Amy CmR] hsdR17(rK-mK+) | Stratagene |
| Strains | Relevant Properties | Source or Reference |
|---|---|---|
| pRS424_GPD | YX-type shuttle vector, T7, lac, GPD promoter, 2micron, f1, pMB1 replicon, ampR, TRP1 | ATCC 87357 |
| pRS426_GPD | YX-type shuttle vector, T7, lac, PGK1 promoter, 2micron, f1, pMB1 replicon, ampR, URA3 | ATCC 87359 |
| pRS426_PGK1 | pRS426-GPD, GPD promoter is replaced with PGK1 promoter, 2micron, f1, pMB1 replicon, ampR, URA3 | This study |
| pRS424_GPD_ERG9 | Constitutively expressed ERG9 gene from S.cerevisiae CEN. PK2-1D | This study |
| pRS426_PGK1_ERG1 | Constitutively expressed ERG1 gene from Panax ginseng | This study |
| pRS424_GPD_DS | Constitutively expressed DS gene from Panax ginseng | This study |
| pRS424_GPD_PPDS | Constitutively expressed PPDS gene from Panax ginseng | This study |
| pRS424_GPD_ERG9P1 | Constitutively expressed ERG9 gene with PTS1 at C-terminal from S.cerevisiae CEN. PK2-1D | This study |
| pRS426_PGK1_ERG1p1 | Constitutively expressed ERG1 gene with PTS1 at C-terminal from Panax ginseng | This study |
| pRS424_GPD_DSP1 | Constitutively expressed DS gene with PTS1 at C-terminal from Panax ginseng | This study |
| pRS424_GPD_DSP2 | Constitutively expressed DS gene with PTS2 at N-terminal from Panax ginseng | This study |
| pRS424_GPD_PPDSP1 | Constitutively expressed PPDS gene with PTS1 at C-terminal from Panax ginseng | This study |
| pRS424_GPD_EGFP | Constitutively expressed EGFP gene | This study |
| pRS424_GPD_EGFPP1 | Constitutively expressed EGFP gene with PTS1 at C-terminal | This study |
| pRS424_GPD_ERG9_EGFPp1 | Constitutively expressed ERG9 and EGFP fusion gene with PTS1 at C-terminal from S.cerevisiae CEN. PK2-1D | This study |
| pRS424_GPD_ERG1_EGFPp1 | Constitutively expressed ERG1 and EGFP fusion gene with PTS1 at C-terminal from Panax ginseng | This study |
| pRS425_GPD_DS_mKOp1 | Constitutively expressed DS and mKO fusion gene with PTS1 at C-terminal from Panax ginseng | This study |
| pRS425_GPD_DS_mKOp2 | Constitutively expressed DS and mKO fusion gene with PTS2 at N-terminal from Panax ginseng | This study |
| pRS424_GPS_PPDS_EGFPp1 | Constitutively expressed PPDS and EGFP fusion gene with PTS1 at C-terminal from Panax ginseng | This study |
| pCEV-G1 | pSP-G1-type shuttle vector, TEF1 and PGK1 duel promoter, G418/kanamycin/neomycin resistance | Addgene #46813 |
| pCEV-G1-TEF1-tHMG1 | Constitutively expressed truncated HMG1 gene from S.cerevisiae CEN. PK2-1D | This study |
| pCEV-G1-TEF1_DS | Constitutively expressed DS gene from Panax ginseng by TEF1 promoter | This study |
| pCEV-G1-TEF1_DSP2 | Constitutively expressed DS gene with PTS2 at N-terminal from Panax ginseng by TEF1 promoter | This study |
| pCEV-G1-PGK1_PPDS | Constitutively expressed PPDS gene from Panax ginseng by PGK1 promoter | This study |
| pCEV-G1-PGK1_PPDSP1 | Constitutively expressed PPDS gene with PTS1 at C-terminal from Panax ginseng by PGK1 promoter | This study |
| pCEV-G1-TEF1_CPR | Constitutively expressed CPR gene from Panax ginseng by TEF1 promoter | This study |
| pRS426-PGK1_ADH2 | Constitutively expressed ADH2 gene from Saccharomyces cerevisiae CEN PK2-1D by PGK1 promoter | This study |
| YIplac128 | YI-type shuttle vector, lac promoter, pBR322 origin, ampR, LEU2 | ATCC 87592 |
| YIplac204 | YI-type shuttle vector, lac promoter, pBR322 origin, ampR, TRP1 | ATCC 87591 |
| YIplac211 | YI-type shuttle vector, lac promoter, pBR322 origin, ampR, URA3 | ATCC 87593 |
| YIplac128_ERG9 | Constitutively expressed ERG9 gene from S.cerevisiae CEN. PK2-1D with GDP promoter | This study |
| YIplac128_ERG9P1 | Constitutively expressed ERG9 gene with PTS1 at C-terminal from S.cerevisiae CEN. PK2-1D with GDP promoter | This study |
| YIplac128_ERG9_ERG1 | Constitutively expressed ERG9 gene from S.cerevisiae CEN. PK2-1D with GDP promoter and ERG1 gene from Panax ginseng with PGK1 promoter | This study |
| YIplac128_ERG9p1_ERG1p1 | Constitutively expressed ERG9 gene with PTS1 at C-terminal from S.cerevisiae CEN. PK2-1D with GDP promoter and ERG1 gene with PTS1 at C-terminal from Panax ginseng with PGK1 promoter | This study |
| YIplac204_tHMG1 | Constitutively expressed truncated HMG1 gene from S.cerevisiae CEN. PK2-1D with TEF1 promoter | This study |
| YIplac204_tHMG1_DS | Constitutively expressed truncated HMG1 from S.cerevisiae CEN. PK2-1D and DS gene from Panax ginseng with TEF1 promoters | This study |
| YIplac204_tHMG1_DSP2 | Constitutively expressed truncated HMG1 from S.cerevisiae CEN. PK2-1D and DS gene with PTS2 at N-terminal from Panax ginseng with TEF1 promoters | This study |
| YIplac211_CPR | Constitutively expressed CPR gene from Panax ginseng with TEF1 promoter | This study |
| YIplac211_CPR _PPDS | Constitutively expressed CPR and PPDS gene from Panax ginseng with TEF1 and PGK1 promoters | This study |
| YIplac211_CPR _PPDSP1 | Constitutively expressed CPR gene and PPDS gene with PTS1 at C-terminal from Panax ginseng with TEF1 and PGK1 promoters | This study |
| YIplac128_tHMG1 | Constitutively expressed truncated HMG1 gene from S.cerevisiae CEN. PK2-1D with TEF1 promoter | This study |
| pUC57_URA blast | Cloning vector for E.coli, URA selectable marker cassette with 3Myc site at both N-terminal and C-terminal for integration in Yeast genome | KITECH |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Choi, B.H.; Kang, H.J.; Kim, S.C.; Lee, P.C. Organelle Engineering in Yeast: Enhanced Production of Protopanaxadiol through Manipulation of Peroxisome Proliferation in Saccharomyces cerevisiae. Microorganisms 2022, 10, 650. https://doi.org/10.3390/microorganisms10030650
Choi BH, Kang HJ, Kim SC, Lee PC. Organelle Engineering in Yeast: Enhanced Production of Protopanaxadiol through Manipulation of Peroxisome Proliferation in Saccharomyces cerevisiae. Microorganisms. 2022; 10(3):650. https://doi.org/10.3390/microorganisms10030650
Chicago/Turabian StyleChoi, Bo Hyun, Hyun Joon Kang, Sun Chang Kim, and Pyung Cheon Lee. 2022. "Organelle Engineering in Yeast: Enhanced Production of Protopanaxadiol through Manipulation of Peroxisome Proliferation in Saccharomyces cerevisiae" Microorganisms 10, no. 3: 650. https://doi.org/10.3390/microorganisms10030650
APA StyleChoi, B. H., Kang, H. J., Kim, S. C., & Lee, P. C. (2022). Organelle Engineering in Yeast: Enhanced Production of Protopanaxadiol through Manipulation of Peroxisome Proliferation in Saccharomyces cerevisiae. Microorganisms, 10(3), 650. https://doi.org/10.3390/microorganisms10030650







