Multi-Locus Sequencing Reveals Putative Novel Anaplasmataceae Agents, ‘Candidatus Ehrlichia dumleri’ and Anaplasma sp., in Ring-Tailed Coatis (Carnivora: Nasua nasua) from Urban Forested Fragments at Midwestern Brazil
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethical Statement
2.2. Blood Sampling
2.3. DNA Extraction from Coatis’ Blood Samples and Conventional PCR (PCR) for Mammalian Endogenous Control
2.4. Screening and Characterization of Anaplasmataceae Agents
2.5. Bioinformatic Analyses
3. Results
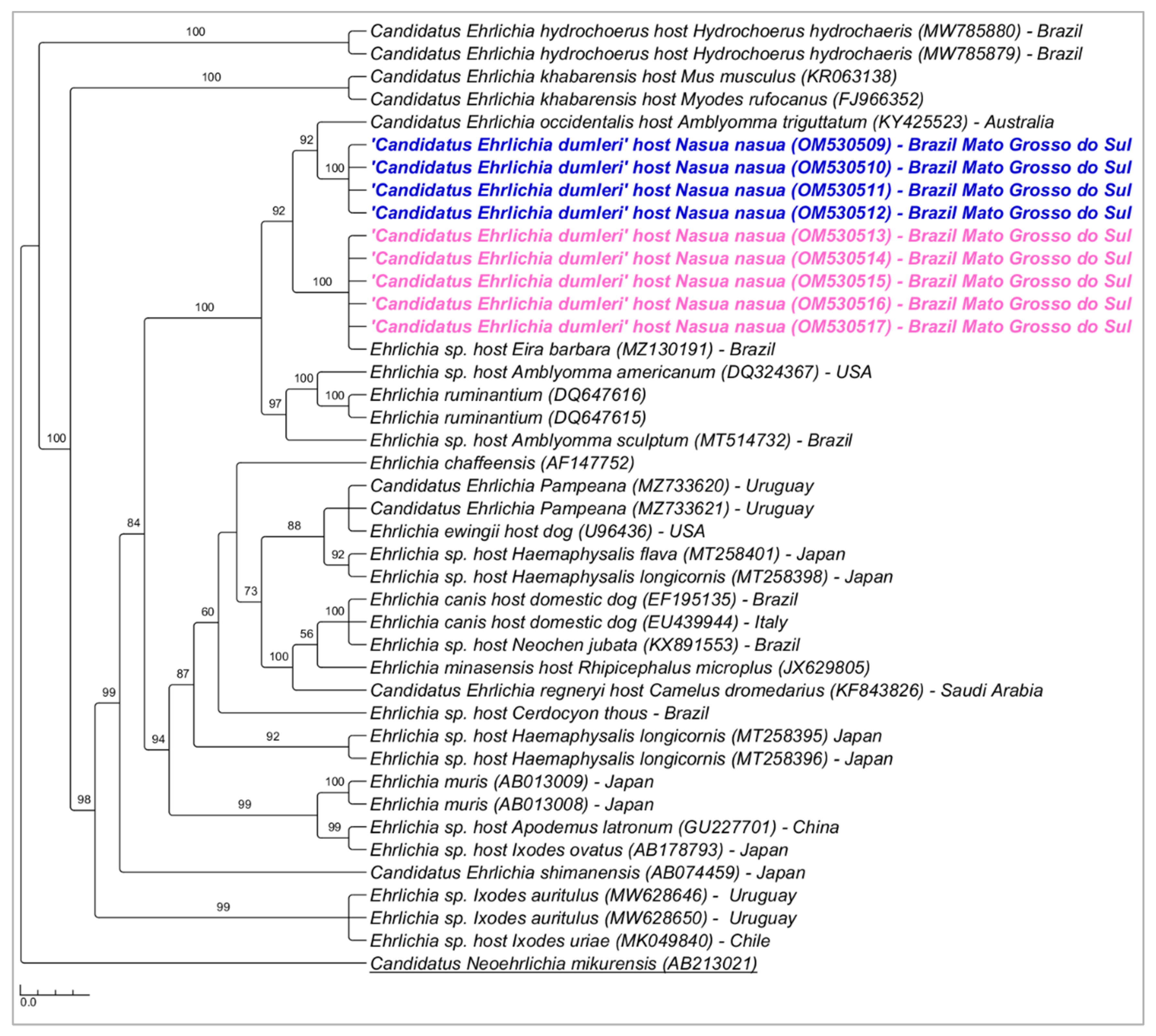
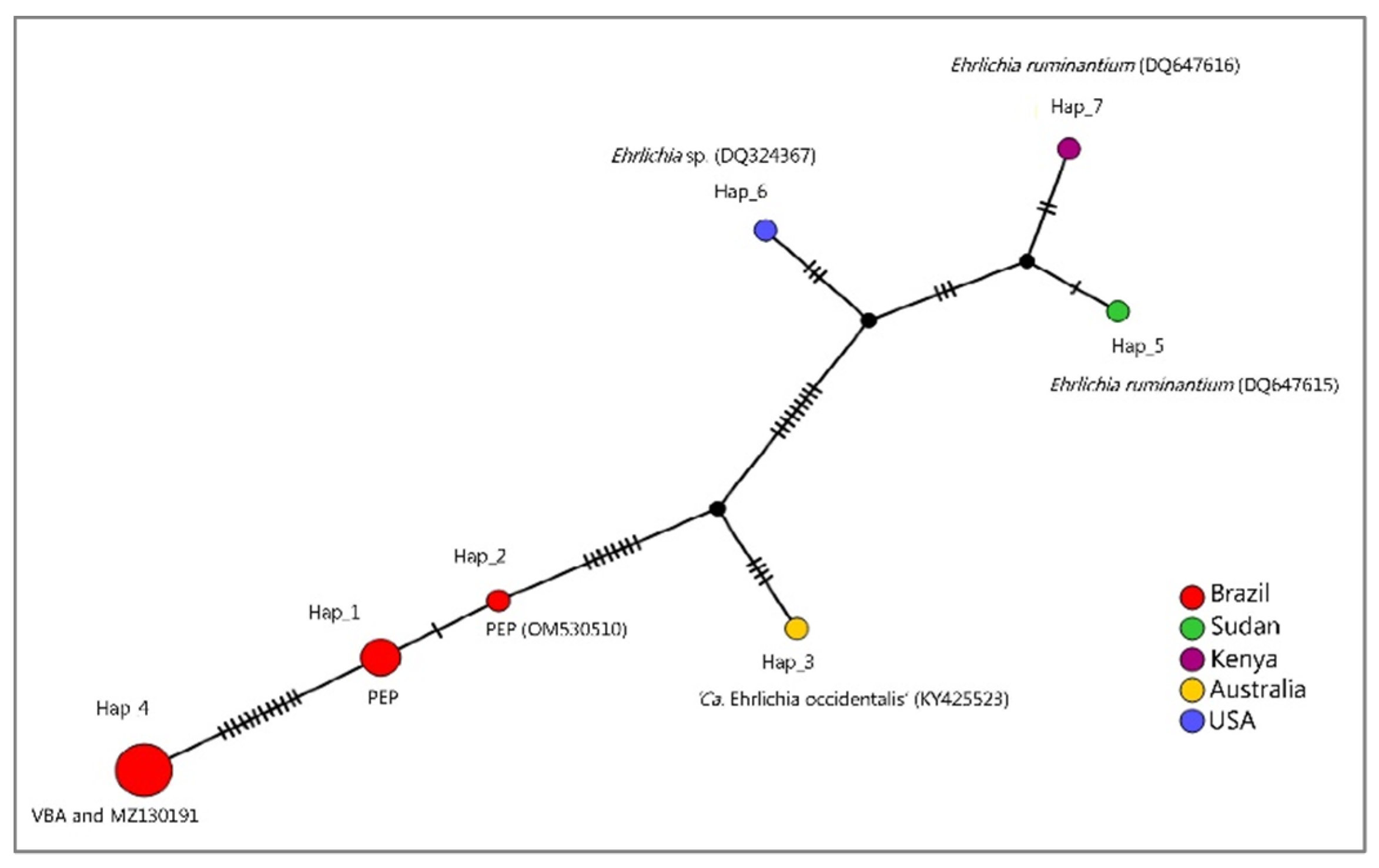
3.1. PCR Assays for Ehrlichia spp. Screening and Characterization
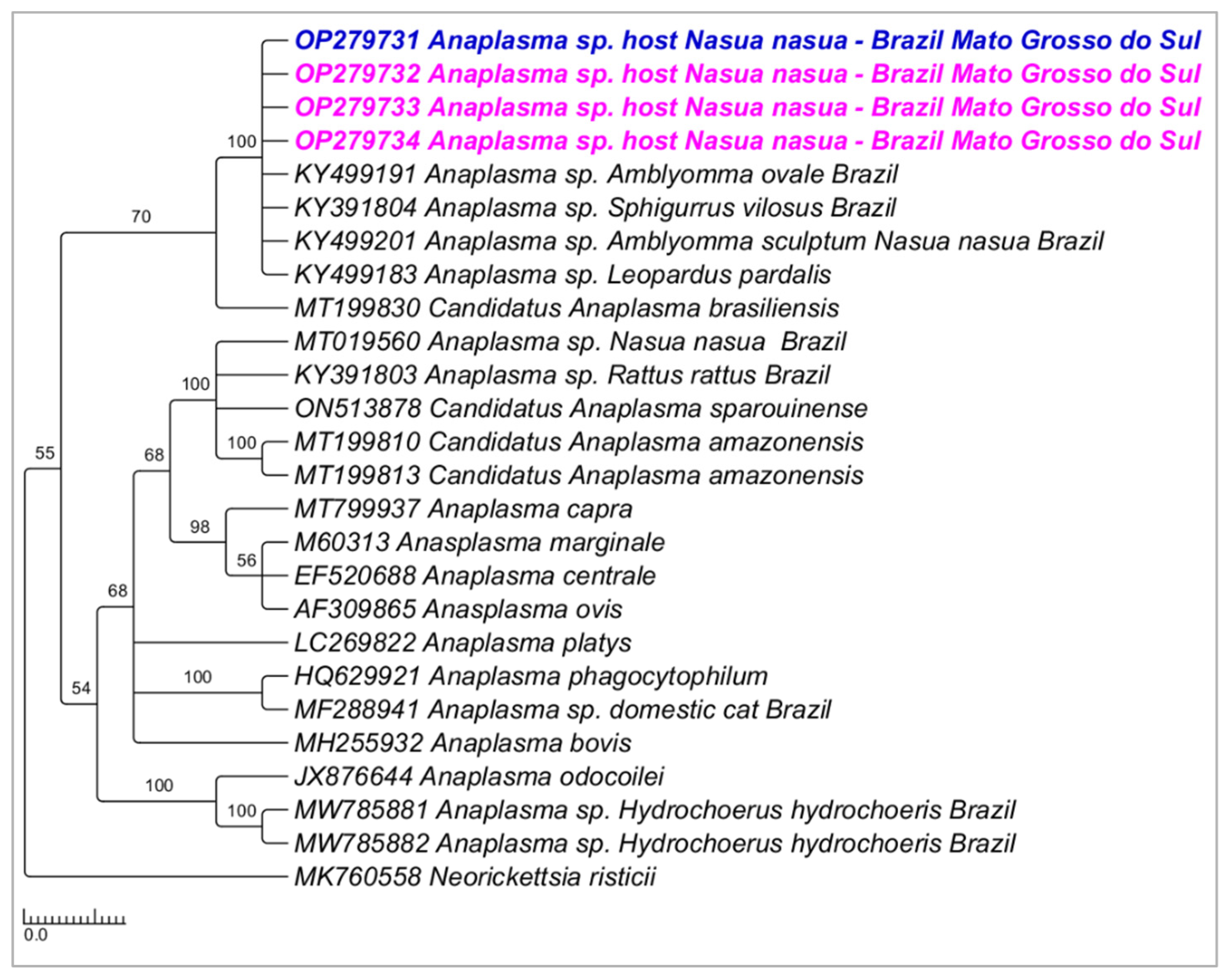
3.2. PCR Assays for Anaplasma sp. Screening and Characterization
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- André, M.R. Diversity of Anaplasma and Ehrlichia/Neoehrlichia agents in terrestrial wild carnivorous worldwide: Implications for human and domestic animal health and wildlife conservation. Frontiers 2018, 5, 293. [Google Scholar]
- Duron, O.; Koual, R.; Musset, L.; Buysse, M.; Lambert, Y.; Jaulhac, B.; Blanchet, D.; Alsibai, K.D.; Lazrek, Y.; Epelboin, L.; et al. Novel Chronic Anaplasmosis in Splenectomized Patient, Amazon Rainforest. Emerg. Infect. Dis. 2022, 28, 1673–1676. [Google Scholar] [CrossRef] [PubMed]
- Estevam, L.G.T.M.; Junior, A.A.F.; Silvestre, B.T.; Hemetrio, N.S.; Almeida, L.R.; Oliveira, M.M.; Silva, S.M.; Ribeiro, M.F.B.; Silveira, J.A.G. Seven years of evaluation of ectoparasites and vector-borne pathogens among ring-tailed coatis in an urban park in southeastern Brazil. Vet. Parasitol. Reg. Stud. Rep. 2020, 21, 100442. [Google Scholar] [CrossRef]
- Barreto, W.T.G.; Miraglia, H.H.; Macedo, G.C.; Rucco, A.C.; Assis, W.O.; Oliveira-Santos, L.G.; Porfírio, G.E.O. Density and survivorship of the South American coati (Nasua nasua) in urban areas in Central–Western Brazil. HYSTRIX 2021, 32, 82–88. [Google Scholar] [CrossRef]
- Levin, M.L.; Nicholson, W.L.; Massung, R.F.; Sumner, J.W.; Fish, D. Comparison of the reservoir competence of medium-sized mammals and Peromyscus leucopus for Anaplasma phagocytophilum in Connecticut. Vector Borne Zoonotic Dis. 2002, 2, 125–136. [Google Scholar] [CrossRef]
- Yabsley, M.J.; Murphy, S.M.; Luttrell, M.P.; Little, S.E.; Massung, R.F.; Stallknecht, D.E.; Conti, L.A.; Blackmore, C.G.M.; Durden, L.A. Experimental and field studies on the suitability of raccoons (Procyon lotor) as hosts for tick-borne pathogens. Vector Borne Zoonotic Dis. 2008, 8, 491–503. [Google Scholar] [CrossRef]
- Woldehiwet, Z. The natural history of Anaplasma phagocytophilum. Vet. Parasitol. 2010, 8, 491–503. [Google Scholar] [CrossRef]
- Perles, L.; Martins, T.F.; Barreto, W.T.G.; Carvalho de Macedo, G.; Herrera, H.M.; Mathias, L.A.; Labruna, M.B.; Barros-Battesti, D.M.; Machado, R.Z.; André, M.R. Diversity and Seasonal Dynamics of Ticks on Ring-Tailed Coatis Nasua nasua (Carnivora: Procyonidae) in Two Urban Areas from Midwestern Brazil. Animals 2022, 12, 293. [Google Scholar] [CrossRef]
- Porfirio, G.E.O.; Santos, F.M.; Macedo, G.C.; Barreto, W.T.G.; Campos, J.B.V.; Meyers, A.C.; André, M.R.; Perles, L.; Oliveira, C.E.; Xavier, S.C.C.; et al. Maintenance of Trypanosoma cruzi, T. evansi and Leishmania spp. by domestic dogs and wild mammals in a rural settlement in Brazil-Bolivian border. Int. J. Parasitol. Parasites Wildl. 2018, 7, 398–404. [Google Scholar] [CrossRef]
- Santos, F.M.; Macedo, G.C.; Barreto, W.T.G.; Oliveira-Santos, L.G.R.; Garcia, C.M.; Mourão, G.M.; Porfírio, G.E.O.; Marino, E.D.; André, M.R.; Perles, L.; et al. Outcomes of Trypanosoma cruzi and Trypanosoma evansi infections on health of Southern coati (Nasua nasua), crab-eating fox (Cerdocyon thous), and ocelot (Leopardus pardalis) in the Brazilian Pantanal. PLoS ONE 2018, 13, e0201357. [Google Scholar] [CrossRef]
- Perles, L.; Macedo, G.G.; Barreto, W.T.G.; Francisco, G.V.; Herrera, H.M.; Barros-Battesti, D.M.; Machado, R.Z.; André, M.R. Longitudinal dynamics and health impact of Hepatozoon procyonis (Apicomplexa: Hepatozoidae) on naturally infected ring-tailed coatis Nasua nasua (Carnivora: Procyonidae) from Midwestern Brazil. Ticks Tick-Borne Dis. 2022, 13, 101982. [Google Scholar] [CrossRef] [PubMed]
- Olifiers, N.; Bianchi, R.C.; D’Andrea, P.S.; Mourão, G.; Gompper, M.E. Estimating age of carnivores from the Pantanal region of Brazil. Wildl. Biol. 2010, 16, 389–399. [Google Scholar] [CrossRef] [PubMed]
- Birkenheuer, A.J.; Levy, M.G.; Breitschwerdt, E.B. Development and evaluation of a seminested PCR for detection and differentiation of Babesia gibsoni (Asian genotype) and B. canis DNA in canine blood samples. J. Clin. Microbiol. 2003, 41, 4172–4177. [Google Scholar] [CrossRef] [PubMed]
- Doyle, C.; Labruna, M.B.; Breitschwerdt, E.B.; Tang, Y.; Corstvet, R.E.; Hegarty, B.C.; Bloch, K.C.; Li, P.; Walker, D.H.; McBride, J.W. Detection of medically important Ehrlichia by quantitative multicolor TaqMan real-time polymerase chain reaction of the dsb gene. J. Mol. Diagn. 2005, 7, 504–510. [Google Scholar] [CrossRef] [PubMed]
- Greiman, S.E.; Tkach, V.V.; Pulis, E.; Fayton, T.J.; Curran, S.S. Large Scale Screening of Digeneans for Neorickettsia Endosymbionts Using Real-Time PCR Reveals New Neorickettsia Genotypes, Host Associations and Geographic Records. PLoS ONE 2014, 9, e98453. [Google Scholar] [CrossRef]
- Müller, A.; Monti, G.; Otth, C.; Sepúlveda, P.; Bittencourt, P.; Nachum-Biala, Y.; Gutierrez, R.; Harrus, S. “Candidatus Neoehrlichia chilensis” sp. nov.: Molecular detection and characterization of a novel Anaplasmataceae in wild rodents from Valdivia, southern Chile. Transb. Emerg. Dis. 2018, 65, 357–362. [Google Scholar] [CrossRef]
- Victoria, L.O.; Montilla, H.J.; Qurollo, B.A.; Maggi, R.G.; Hegarty, B.C.; Tornquist, S.J.; Breitschwerdt, E.B. Potentially novel Ehrlichia species in horses, Nicaragua. Emerg. Infect. Dis. 2015, 21, 335–338. [Google Scholar] [CrossRef]
- Li, H.; Zheng, Y.C.; Ma, L.; Jia, N.; Jiang, B.G.; Jiang, R.R. Human infection with a novel tick-borne Anaplasma species in China: A surveillance study. Lancet Infect. Dis. 2015, 15, 663–670. [Google Scholar] [CrossRef]
- Rejmanek, D.; Bradburd, G.; Foley, J. Molecular characterization reveals distinct genospecies of Anaplasma phagocytophilum from diverse North American hosts. J. Med. Microbiol. 2012, 61, 204–212. [Google Scholar] [CrossRef]
- Massung, R.F.; Slater, K.; Owens, J.H.; Nicholson, W.L.; Mather, T.N.; Solberg, V.B.; Olson, J.G. Nested PCR assay for detection of granulocytic ehrlichiae. J. Clin. Microbiol. 1998, 36, 1090–1095. [Google Scholar] [CrossRef]
- Zhang, Y.; Lv, Y.; Zhang, F.; Zhang, W.; Wang, J.; Cui, Y.; Wang, R.; Jian, F.; Zhang, L.; Ning, C. Molecular and phylogenetic analysis of Anaplasma spp. in sheep and goats from six provinces of China. J. Vet. Sci. 2016, 17, 523–529. [Google Scholar] [CrossRef] [PubMed]
- Sanger, F.; Nicklen, S.; Coulson, A.R. DNA sequencing with chain-terminating inhibitors. Proc. Natl. Acad. Sci. USA 1997, 74, 5463–5467. [Google Scholar] [CrossRef]
- Ewing, B.; Hillier, L.; Wendl, M.C.; Green, P. Base-calling of automated sequencer traces usingPhred. I. Accuracy assessment. Genome Res. 1998, 8, 175–185. [Google Scholar] [CrossRef] [PubMed]
- Ewing, B.; Green, P. Base-calling of automated sequencer traces using phred. II. Error probabilities. Genome Res. 1998, 8, 186–194. [Google Scholar] [CrossRef] [PubMed]
- Benson, D.A.; Cavanaugh, M.; Clark, K.; Karsch-Mizrachi, I.; Lipman, D.J.; Ostell, J.; Sayers, E.W. GenBank. Nucleic Acids Res. 2012, 41, 36–42. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT online service: Multiple sequence alignment, interactive, sequence choice and visualization. Briefings in Bioinform. 2019, 20, 1160–1166. [Google Scholar] [CrossRef] [PubMed]
- Posada, D.; Buckley, T.R.; Thorne, J. Model selection and model averaging in phylogenetics: Advantages of Akaike information criterion and Bayesian approaches over likelihood ratio tests. Syst. Biol. 2004, 53, 793–808. [Google Scholar] [CrossRef]
- Trifinopoulus, J.; Nguyen, L.T.; Haeseler, V.A.; Minh, B.Q. W-IQ-TREE: A fast-online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 2016, 44, 232–235. [Google Scholar] [CrossRef]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 12, 1572–1574. [Google Scholar] [CrossRef]
- Stover, B.C.; Muller, K.F. TreeGraph 2: Combining and visualizing evidence from different phylogenetic analyses. BMC Bioinform. 2010, 11, 7. [Google Scholar] [CrossRef]
- Librado, P.; Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analyses across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- André, M.R.; Adania, C.H.; Machado, R.Z.; Allegretti, S.M.; Felippe, P.A.N.; Silva, K.F.; Kagaghi, A.C. Molecular and serologic detection of Ehrlichia spp. in endangered Brazilian wild capitve felids. J. Wildl. Dis. 2010, 3, 1017–1023. [Google Scholar] [CrossRef] [PubMed]
- André, M.R.; Dumler, J.S.; Scorpio, D.G.; Teixeira, R.H.F.; Allegretti, S.M.; Machado, R.Z. Molecular detection of tick-borne bacterial agents in Brazilian and exotic captive carnivores. Ticks Tick-Borne Dis. 2012, 3, 247–253. [Google Scholar] [CrossRef]
- André, M.R.; Calchi, A.C.; Perles, L.; Gonçalves, L.R.; Uccella, L.; Lemes, J.R.B.; Nantes, W.A.G.; Santos, F.M.; Porfírio, G.E.O.; Barros-Battesti, D.M.; et al. Novel Ehrlichia and Hepatozoon genotypes in white-eared opossums (Didelphis albiventris) and associated ticks from Brazil. Ticks Tick-Borne Dis. 2022, 13, 102022. [Google Scholar] [CrossRef]
- Sacchi, A.B.V.; Duarte, J.M.B.; André, M.R.; Machado, R.Z. Prevalence and molecular characterization of Anaplasmataceae agents in free-ranging Brazilian marsh deer (Blastocerus dichotomus). Comp. Immunol. Microbiol. Infect. Dis. 2012, 4, 325–334. [Google Scholar] [CrossRef]
- Benevenute, J.L.; Dumler, J.S.; Ogrzewalska, M.; Roque, A.L.R.; Mello, V.V.; Sousa, K.C.M.; Gonçalves, L.R.; D’Andrea, P.S.; Sampaio Lemos, E.R.; Machado, R.Z.; et al. Assessment of a quantitative 5’ nuclease real-time polymerase chain reaction using groEl gene for Ehrlichia and Anaplasma species in rodents in Brazil. Ticks Tick-Born Dis. 2017, 8, 646–656. [Google Scholar] [CrossRef]
- Soares, H.S.; Marcili, A.; Barbieri, A.R.M.; Minervino, A.H.H.; Malheiros, A.F.; Gennari, S.M.; Labruna, M.B. Novel Anaplasma and Ehrlichia organisms infecting the wildlife of two regions of the Brazilian Amazon. Acta Trop. 2017, 174, 82–87. [Google Scholar] [CrossRef]
- Sousa, K.C.M.; Calchi, A.C.; Herrera, H.M.; Dumler, M.; Barros-Battesti, D.M.; Machado, R.Z.; André, M.R. Anaplasmataceae agents among wild mammals and ectoparasites in Brazil. Epidemiol. Infect. 2017, 16, 3424–3437. [Google Scholar] [CrossRef]
- Guimarães, A.; Raimundo, J.M.; Silva, A.T.D.; Carpintero, F.M.; Pires, J.R.; Benevenute, J.L.; Machado, R.Z.; André, M.R.; Baldani, C.D. Detection of a puntative novel of Ehrlichia sp. from opossums (Didelphis aurita) from Brazil. Rev. Bras. Parasitol. Vet. 2019, 28, 140–144. [Google Scholar] [CrossRef]
- Calchi, A.C.; Vultão, J.G.; Alves, M.H.; Yogui, D.R.; Desbiez, A.L.J.; Santi, M.D.; de Souza Santana, M.; da Silva, T.M.V.; Werther, K.; Teixeira, M.M.G.; et al. Ehrlichia spp. and Anaplasma spp. in Xenarthra mammals from Brazil, with evidence of novel ‘Candidatus Anaplasma spp.’. Sci. Rep. 2020, 10, 12615. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, P.; Torres, J.M.; Placa, A.J.V.; Mello, V.V.C.D.; Lourenço, E.C.; Herrera, H.M.; Oliveira, C.E.; Hemsley, C.; Titball, R.W.; Machado, R.Z.; et al. Molecular Survey of Anaplasmataceae Agents and Coxiellaceae in Non-Hematophagous Bats and Associated Ectoparasites from Brazil. Parasitologia 2021, 1, 197–209. [Google Scholar] [CrossRef]
- Collere, F.C.M.; Delai, R.M.; Ferrari, L.D.R.; da Silva, L.H.; Fogaça, P.L.; Rodrigues, A.N.; Gonçalves, D.D.; Baggio, R.A.; Moraes, M.F.D.; Hoppe, E.G.L.; et al. ‘Candidatus Mycoplasma haematonasua’ and tick-borne pathogens in ring-tailed coatis (Nasua nasua, Linnaeus, 1976) from the Iguaçu National Park, Paraná State, southern Brazil. Transboud. Emerg. Dis. 2021, 68, 3222–3229. [Google Scholar] [CrossRef] [PubMed]
- Barros-Battesti, D.M.; Baggio, D. Ectoparasites Ixodida Leach, 1817 on wild mammals in the State of Paraná, Brazil. Mem. Inst. Oswaldo Cruz 1992, 87, 291–296. [Google Scholar] [CrossRef][Green Version]
- Labruna, M.B.; Jorge, R.S.P.; Sana, D.A.; Ja’como, A.T.A.; Kashivakura, C.K.; Furtado, M.M.; Ferro, C.; Perez, S.A.; Silveira, L.; Santos, T.S.; et al. Ticks (Acari: Ixodida) on wild carnivores in Brazil. Exp. Appl. Acarol. 2005, 36, 149–163. [Google Scholar] [CrossRef] [PubMed]
- Wen, B.; Rikihisa, Y.; Mott, J.M.; Greene, R.; Kim, H.; Zhi, N.; Couto, G.C.; Unver, A.; Bartsch, R. Comparison of nested PCR with immunofluorescentantibody assay for detection of Ehrlichia canis infection in dogs treated with doxycicline. J. Clin. Microbiol. 1997, 35, 1852–1855. [Google Scholar] [CrossRef]
- Carelli, G.; Decaro, N.; Lorusso, A.; Elia, G.; Lorusso, E.; Mari, V.; Buonavoglia, C. Detection and quantification of Anaplasma marginale DNA in blood samples of cattle by real-time PCR. Vet. Microbiol. 2007, 124, 107–114. [Google Scholar] [CrossRef]
- Costa, E.M.J.; Mauro, R.A. Secondary dispersion in coatis’ feces Nasua nasua e (Linnaeus, 1766) (Mammalia: Procyonidae) in a fragment of Cerrado, Mato Grosso do Sul, Brazil. Neotrop. Biol. Conserv. 2008, 3, 66–72. [Google Scholar]
- Gonçalves, L.R.; Paludo, G.; Bisol, T.B.; Perles, L.; de Oliveira, L.B.; de Oliveira, C.M.; da Silva, T.M.V.; Nantes, W.A.G.; Duarte, M.A.; Santos, F.M.; et al. Molecular detection of piroplasmids in synanthropic rodents, marsupials, and associated ticks from Brazil, with phylogenetic inference of a putative novel Babesia sp. from white-eared opossum (Didelphis albiventris). Parasitol. Res. 2021, 120, 3537–3546. [Google Scholar] [CrossRef]
- Nantes, W.A.G.; Santos, F.M.; de Macedo, G.C.; Barreto, W.T.G.; Gonçalves, L.R.; Rodrigues, M.S.; Chulli, J.V.M.; Rucco, A.C.; Assis, W.O.; Porfírio, G.E.O.; et al. Trypanosomatid species in Didelphis albiventris from urban forest fragments. Parasitol Res. 2021, 120, 223–231. [Google Scholar] [CrossRef]
- Nantes, W.A.G.; Barreto, W.T.G.; Santos, F.M.; Macedo, G.C.; Rucco, A.C.; Assis, W.O.; Porfírio, G.E.O.; Andrade, G.B.; Jansen, A.M.; Herrera, H.M. The influence of parasitism by Trypanosoma cruzi in the hematological parameters of the white ear opossum (Didelphis albiventris) from Campo Grande, Mato Grosso do Sul, Brazil. Int. J. Parasitol. Parasites Wildl. 2019, 9, 16–20. [Google Scholar] [CrossRef]
- Matuschka, F.R.; Fischer, P.; Heiler, M.; Richter, D.; Spielman, A. Hosts on which nymphal Ixodes ricinus most abundantly feed. Am. J. Trop. Med. Hyg. 1991, 44, 100–107. [Google Scholar] [CrossRef] [PubMed]
- Matuschka, F.R.; Fischer, P.; Heiler, M.; Richter, D.; Spielman, A. Capacity of European animals as reservoir hosts for the Lyme disease spirochete. J. Infect. Dis. 1992, 156, 479–483. [Google Scholar] [CrossRef] [PubMed]
- Ostfeld, K.A.; Keesing, R.S. Biodiversity and the dilution effect in disease ecology. ESA 2000, 82, 609–619. [Google Scholar]
- Bastos, A.D.S.; Mohammed, O.B.; Bennett, N.C.; Petevinos, C.; Alagaili, A.N. Molecular detection of novel Anaplasmataceae closely related to Anaplasma platys and Ehrlichia canis in the dromedary camel (Camelus dromedarius). Vet. Microbiol. 2015, 4, 310–314. [Google Scholar] [CrossRef]
- Feliz, M.L.; Munoz-Leal, S.; Carvalho, L.A.; Queirolo, D.; Remesar, S.; Armua-Fernandez, M.T.; Venzal, J.M. Characterization of ‘Candidatus Ehrlichia pampenana’ in Haemaphysalis juxtakochi ticks and gray brocket deer (Mazama gouazoubira) from Uruguay. Microorganisms 2021, 9, 2165. [Google Scholar] [CrossRef]
- Vieira, T.S.W.J.; Collere, F.C.M.; Ferrari, L.D.R.; Baggio, R.A.; Lange, R.R.; Ferrari, M.V.; Duque, J.C.M.; Sanches, G.S.; Pereira, N.A.; Aguiar, D.M.; et al. Novel Anaplasmataceae agents Candidatus Ehrlichia hydrochoerus and Anaplasma spp. Infecting Capybaras, Brazil. Emerg. Infect. Dis. 2022, 28, 480–482. [Google Scholar] [CrossRef] [PubMed]






| Captures | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| ID | Sex | Age | Local | 1° Sampling | 2° Sampling | 3° Sampling | 4° Sampling | 5° Sampling | 6° Sampling |
| PEP05 | M 1 | A 3 | PEP 6 | Negative | Positive | Negative | NR | NR | NR |
| PEP32 | M | A | PEP | Negative | Positive | NR | NR | NR | NR |
| PEP43 | M | A | PEP | Negative | Positive | NR | NR | NR | NR |
| PEP49 | M | I | PEP | Positive | NR | NR | NR | NR | NR |
| VBA01 | M | A | VBA 7 | Positive | Negative | NR | NR | NR | NR |
| VBA03 | F 2 | A | VBA | Negative | Positive | Positive | Negative | Positive | Positive |
| VBA04 | F | A | VBA | Positive | NR | NR | NR | NR | NR |
| VBA08 | F | A | VBA | Negative | Negative | Positive | NR | NR | NR |
| VBA09 | M | SA 4 | VBA | Positive | Negative | Negative | Negative | NR | NR |
| VBA11 | M | A | VBA | Negative | Negative | Positive | NR | NR | NR |
| VBA16 | F | A | VBA | Negative | Negative | Negative | Positive | NR | NR |
| VBA19 | F | A | VBA | Positive | Negative | NR | NR | NR | NR |
| VBA21 | M | I 5 | VBA | Negative | Negative | Negative | Negative | Positive | Positive |
| VBA23 | F | A | VBA | Positive | Negative | NR | NR | NR | NR |
| VBA29 | M | F | VBA | Negative | Positive | NR | NR | NR | NR |
| VBA32 | F | A | VBA | Positive | NR | NR | NR | NR | NR |
| VBA34 | M | A | VBA | Positive | NR | NR | NR | NR | NR |
| VBA43 | M | A | VBA | Positive | NR | NR | NR | NR | NR |
| VBA44 | M | A | VBA | Positive | Positive | Positive | NR | NR | NR |
| VBA57 | F | A | VBA | Positive | NR | NR | NR | NR | NR |
| 1° Sampling | 2° Sampling | 3° sampling | 4° Sampling | 5° Sampling | 6° Sampling | |
|---|---|---|---|---|---|---|
| VBA03 | 2 May 2018 | 18 June 2018 | 28 August 2018 | 23 October 2018 | 23 January 2019 | 20 March 2019 |
| VBA21 | 25 June 2018 | 24 August 2018 | 29 October 2018 | 21 January 2019 | 21 March 2019 | 23 April 2019 |
| VBA44 | 7 November 2018 | 21 March 2019 | 30 April 2019 | NR | NR | NR |
| Gene | bp | N | VS | GC% | H | dh (Mean ± SD) | π (Mean ± SD) | K |
|---|---|---|---|---|---|---|---|---|
| 16S rRNA | 1066 | 14 | 34 | 49.4 | 7 | 0.802 ± 0.094 | 0.001168 ± 0.00170 | 12.37363 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Perles, L.; Herrera, H.M.; Barreto, W.T.G.; de Macedo, G.C.; Calchi, A.C.; Machado, R.Z.; André, M.R. Multi-Locus Sequencing Reveals Putative Novel Anaplasmataceae Agents, ‘Candidatus Ehrlichia dumleri’ and Anaplasma sp., in Ring-Tailed Coatis (Carnivora: Nasua nasua) from Urban Forested Fragments at Midwestern Brazil. Microorganisms 2022, 10, 2379. https://doi.org/10.3390/microorganisms10122379
Perles L, Herrera HM, Barreto WTG, de Macedo GC, Calchi AC, Machado RZ, André MR. Multi-Locus Sequencing Reveals Putative Novel Anaplasmataceae Agents, ‘Candidatus Ehrlichia dumleri’ and Anaplasma sp., in Ring-Tailed Coatis (Carnivora: Nasua nasua) from Urban Forested Fragments at Midwestern Brazil. Microorganisms. 2022; 10(12):2379. https://doi.org/10.3390/microorganisms10122379
Chicago/Turabian StylePerles, Lívia, Heitor M. Herrera, Wanessa T. G. Barreto, Gabriel C. de Macedo, Ana C. Calchi, Rosangela Z. Machado, and Marcos R. André. 2022. "Multi-Locus Sequencing Reveals Putative Novel Anaplasmataceae Agents, ‘Candidatus Ehrlichia dumleri’ and Anaplasma sp., in Ring-Tailed Coatis (Carnivora: Nasua nasua) from Urban Forested Fragments at Midwestern Brazil" Microorganisms 10, no. 12: 2379. https://doi.org/10.3390/microorganisms10122379
APA StylePerles, L., Herrera, H. M., Barreto, W. T. G., de Macedo, G. C., Calchi, A. C., Machado, R. Z., & André, M. R. (2022). Multi-Locus Sequencing Reveals Putative Novel Anaplasmataceae Agents, ‘Candidatus Ehrlichia dumleri’ and Anaplasma sp., in Ring-Tailed Coatis (Carnivora: Nasua nasua) from Urban Forested Fragments at Midwestern Brazil. Microorganisms, 10(12), 2379. https://doi.org/10.3390/microorganisms10122379





