Report and Comparative Genomics of an NDM-5-Producing Escherichia coli in a Portuguese Hospital: Complex Class 1 Integrons as Important Players in blaNDM Spread
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Isolation, Antibiotic Susceptibility Testing and blaNDM Detection
2.2. Whole-Genome Sequencing and Analysis
2.3. Plasmid Analysis
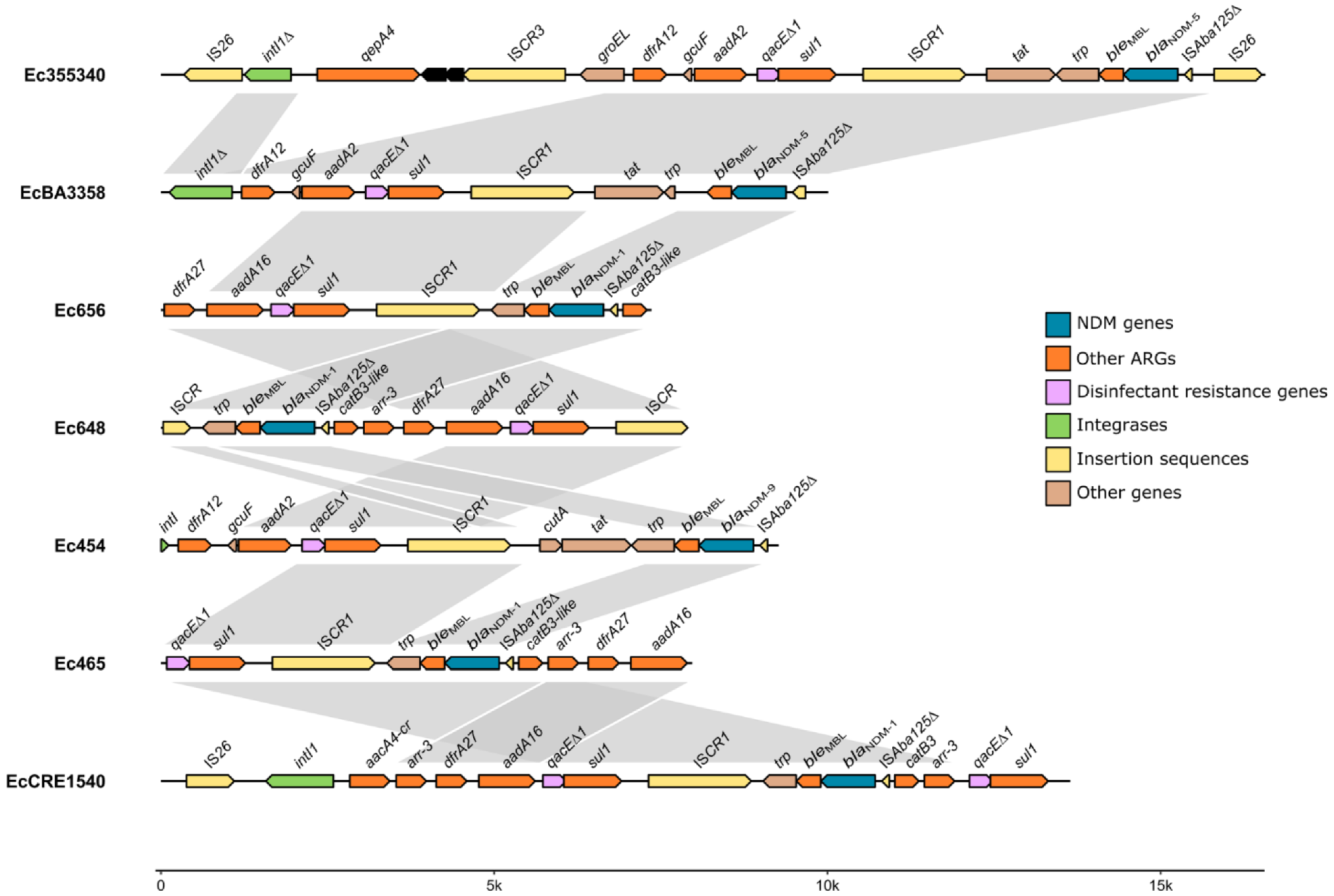
2.4. Comparative Analyses of ST156 and blaNDM-5-Harbouring Genomes
3. Results
3.1. Genotype and Phenotype of Ec355340
3.2. Characterization and Comparative Analysis of the pEc355340_NDM-5 Plasmid
3.3. Clonal Background Analysis
4. Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- O’Neill, J. Review on Antimicrobial Resistance. Antimicrobial Resistance: Tackling a Crisis for the Health and Wealth of Nations. 2014. Available online: https://amr-review.org/Publications.html (accessed on 1 January 2022).
- Nordmann, P.; Dortet, L.; Poirel, L. Carbapenem resistance in Enterobacteriaceae: Here is the storm! Trends Mol. Med. 2012, 18, 263–272. [Google Scholar] [CrossRef] [PubMed]
- WHO (World Health Organization). Global Priority List of Antibiotic-Resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics. 2017. Available online: https://www.who.int/medicines/publications/en/ (accessed on 1 January 2022).
- Martin, A.; Fahrbach, K.; Zhao, Q.; Lodise, T. Association between carbapenem resistance and mortality among adult, hospitalized patients with serious infections due to Enterobacteriaceae: Results of a systematic literature review and meta-analysis. Open Forum Infect. Dis. 2018, 5, ofy150. [Google Scholar] [CrossRef] [PubMed]
- Yong, D.; Toleman, M.A.; Giske, C.G.; Cho, H.S.; Sundman, K.; Lee, K.; Walsh, T.R. Characterization of a new metallo-beta-lactamase gene, blaNDM-1, and a novel erythromycin esterase gene carried on a unique genetic structure in Klebsiella pneumoniae sequence type 14 from India. Antimicrob. Agents Chemother. 2009, 53, 5046–5054. [Google Scholar] [CrossRef] [PubMed]
- Nordmann, P.; Poirel, L.; Walsh, T.R.; Livermore, D.M. The emerging NDM carbapenemases. Trends Microbiol. 2011, 19, 588–595. [Google Scholar] [CrossRef] [PubMed]
- Nordmann, P.; Poirel, L. Epidemiology and diagnostics of carbapenem resistance in gram-negative bacteria. Clin. Infect. Dis. 2019, 69, S521–S528. [Google Scholar] [CrossRef]
- Wu, W.; Feng, Y.; Tang, G.; Qiao, F.; McNally, A.; Zong, Z. NDM metallo-beta-lactamases and their bacterial producers in health care settings. Clin. Microbiol. Rev. 2019, 32, e00115–e00118. [Google Scholar] [CrossRef]
- Karlowsky, J.A.; Lob, S.H.; Kazmierczak, K.M.; Badal, R.E.; Young, K.; Motyl, M.R.; Sahm, D.F. In vitro activity of imipenem against carbapenemase-positive Enterobacteriaceae isolates collected by the SMART Global surveillance program from 2008 to 2014. J. Clin. Microbiol. 2017, 55, 1638–1649. [Google Scholar] [CrossRef]
- Farhat, N.; Khan, A.U. Evolving trends of New Delhi Metallo-beta-lactamase (NDM) variants: A threat to antimicrobial resistance. Infect. Genet. Evol. 2020, 86, 104588. [Google Scholar] [CrossRef]
- Dadashi, M.; Yaslianifard, S.; Hajikhani, B.; Kabir, K.; Owlia, P.; Goudarzi, M.; Hakemivala, M.; Darban-Sarokhalil, D. Frequency distribution, genotypes and prevalent sequence types of New Delhi metallo-beta-lactamase-producing Escherichia coli among clinical isolates around the world: A review. J. Glob. Antimicrob. Resist. 2019, 19, 284–293. [Google Scholar] [CrossRef]
- Manageiro, V.; Sampaio, D.A.; Pereira, P.; Rodrigues, P.; Vieira, L.; Palos, C.; Caniça, M. Draft genome sequence of the first NDM-1-producing Providencia stuartii strain isolated in Portugal. Genome Announc. 2015, 3, e01077-15. [Google Scholar] [CrossRef]
- Aires-de-Sousa, M.; Ortiz de la Rosa, J.M.; Goncalves, M.L.; Costa, A.; Nordmann, P.; Poirel, L. Occurrence of NDM-1-producing Morganella morganii and Proteus mirabilis in a single patient in Portugal: Probable in vivo transfer by conjugation. J. Antimicrob. Chemother. 2020, 75, 903–906. [Google Scholar] [CrossRef]
- Teixeira, P.; Tacão, M.; Pureza, L.; Gonçalves, J.; Silva, A.; Cruz-Schneider, M.P.; Henriques, I. Occurrence of carbapenemase-producing Enterobacteriaceae in a Portuguese river: blaNDM, blaKPC and blaGES among the detected genes. Environ. Pollut. 2020, 260, 113913. [Google Scholar] [CrossRef]
- Mendes, G.; Ramalho, J.F.; Duarte, A.; Pedrosa, A.; Silva, A.C.; Méndez, L.; Caneiras, C. First outbreak of NDM-1-Producing Klebsiella pneumoniae ST11 in a Portuguese Hospital Centre during the COVID-19 pandemic. Microorganisms 2022, 10, 251. [Google Scholar] [CrossRef]
- Novais, Â.; Ferraz, R.V.; Viana, M.; da Costa, P.M.; Peixe, L. NDM-1 introduction in Portugal through a ST11 KL105 Klebsiella pneumoniae widespread in Europe. Antibiotics 2022, 11, 92. [Google Scholar] [CrossRef]
- Hornsey, M.; Phee, L.; Wareham, D.W. A novel variant, NDM-5, of the New Delhi metallo-β-lactamase in a multidrug-resistant Escherichia coli ST648 isolate recovered from a patient in the United Kingdom. Antimicrob. Agents Chemother. 2011, 55, 5952–5954. [Google Scholar] [CrossRef]
- Zhang, Z.; Guo, H.; Li, X.; Li, W.; Yang, G.; Ni, W.; Zhan, M.; Lu, L.; Zhang, Z.; Li, X.; et al. Genetic diversity and characteristics of blaNDM-positive plasmids in Escherichia coli. Front. Microbiol. 2021, 12, 729952. [Google Scholar] [CrossRef]
- Tavares, R.D.S.; Tacão, M.; Figueiredo, A.S.; Duarte, A.S.; Esposito, F.; Lincopan, N.; Manaia, C.M.; Henriques, I. Genotypic and phenotypic traits of blaCTX-M-carrying Escherichia coli strains from an UV-C-treated wastewater effluent. Water Res. 2020, 184, 116079. [Google Scholar] [CrossRef]
- Fernandez-Lopez, R.; de Toro, M.; Moncalian, G.; Garcillan-Barcia, M.P.; de la Cruz, F. Comparative genomics of the conjugation region of F-like plasmids: Five shades of F. Front. Mol. Biosci. 2016, 3, 71. [Google Scholar] [CrossRef]
- Miajlovic, H.; Smith, S.G. Bacterial self-defence: How Escherichia coli evades serum killing. FEMS Microbiol. Lett. 2014, 354, 1–9. [Google Scholar] [CrossRef]
- Sarowska, J.; Futoma-Koloch, B.; Jama-Kmiecik, A.; Frej-Madrzak, M.; Ksiazczyk, M.; Bugla-Ploskonska, G.; Choroszy-Krol, I. Virulence factors, prevalence and potential transmission of extraintestinal pathogenic Escherichia coli isolated from different sources: Recent reports. Gut Pathog. 2019, 11, 10. [Google Scholar] [CrossRef]
- Kieffer, N.; Aires-de-Sousa, M.; Nordmann, P.; Poirel, L. High rate of MCR-1-producing Escherichia coli and Klebsiella pneumoniae among pigs, Portugal. Emerg. Infect. Dis. 2017, 23, 2023–2029. [Google Scholar] [CrossRef] [PubMed]
- Vredenburg, J.; Varela, A.R.; Hasan, B.; Bertilsson, S.; Olsen, B.; Narciso-da-Rocha, C.; Bonnedahl, J.; Stedt, J.; Da Costa, P.M.; Manaia, C.M. Quinolone-resistant Escherichia coli isolated from birds of prey in Portugal are genetically distinct from those isolated from water environments and gulls in Portugal, Spain and Sweden. Environ. Microbiol. 2014, 16, 995–1004. [Google Scholar] [CrossRef] [PubMed]
- Manageiro, V.; Félix, D.; Jones-Dias, D.; Sampaio, D.A.; Vieira, L.; Sancho, L.; Ferreira, E.; Caniça, M. Genetic background and expression of the new qepA4 gene variant recovered in clinical TEM-1- and CMY-2-producing Escherichia coli. Front. Microbiol. 2017, 8, 1899. [Google Scholar] [CrossRef]
- Ramaloko, W.T.; Osei Sekyere, J. Phylogenomics, epigenomics, virulome, and mobilome of Gram-negative bacteria co-resistant to carbapenems and polymyxins: A One-Health systematic review and meta-analyses. Environ. Microbiol. 2022, 24, 1518–1542. [Google Scholar] [CrossRef] [PubMed]
- Naseer, U.; Sundsfjord, A. The CTX-M conundrum: Dissemination of plasmids and Escherichia coli clones. Microb. Drug Resist. 2011, 17, 83–97. [Google Scholar] [CrossRef]
- Douarre, P.E.; Mallet, L.; Radomski, N.; Felten, A.; Mistou, M.Y. Analysis of COMPASS, a new comprehensive plasmid database revealed prevalence of multireplicon and extensive diversity of IncF plasmids. Front. Microbiol. 2020, 11, 483. [Google Scholar] [CrossRef]
- Villa, L.; García-Fernández, A.; Fortini, D.; Carattoli, A. Replicon sequence typing of IncF plasmids carrying virulence and resistance determinants. J. Antimicrob. Chemother. 2010, 65, 2518–2529. [Google Scholar] [CrossRef]
- Baloch, Z.; Lv, L.; Yi, L.; Wan, M.; Aslam, B.; Yang, J.; Liu, J.H. Emergence of almost identical F36:A-:B32 plasmids carrying blaNDM-5 and qepA in Escherichia coli from both Pakistan and Canada. Infect. Drug Resist. 2019, 12, 3981–3985. [Google Scholar] [CrossRef]
- Li, J.; Lan, R.; Xiong, Y.; Ye, C.; Yuan, M.; Liu, X.; Chen, X.; Yu, D.; Liu, B.; Lin, W.; et al. Sequential isolation in a patient of Raoultella planticola and Escherichia coli bearing a novel ISCR1 element carrying blaNDM-1. PLoS ONE 2014, 9, e89893. [Google Scholar] [CrossRef]
- Le-Vo, H.N.; Tran, P.T.; Le, L.; Matsumoto, Y.; Motooka, D.; Nakamura, S.; Jones, J.W.; Iida, T.; Cao, V. Complex class 1 integron in a clinical Escherichia coli strain from Vietnam carrying both mcr-1 and blaNDM-1. Front. Microbiol. 2019, 10, 2472. [Google Scholar] [CrossRef]
- Li, Z.; Lin, Y.; Lu, L.; Wang, K.; Yang, L.; Li, P.; Li, J.; Jia, L.; Li, P.; Song, H. Genetic characterisation of a complex class 1 integron in an NDM-1-producing Citrobacter freundii ST396 clinical strain isolated from a urine sample. J. Glob. Antimicrob. Resist. 2020, 23, 64–66. [Google Scholar] [CrossRef]
- Acman, M.; Wang, R.; van Dorp, L.; Shaw, L.P.; Wang, Q.; Luhmann, N.; Yin, Y.; Sun, S.; Chen, H.; Wang, H.; et al. Role of mobile genetic elements in the global dissemination of the carbapenem resistance gene blaNDM. Nat. Commun. 2022, 13, 1131. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, R.; Li, J.; Wu, Z.; Yin, W.; Schwarz, S.; Tyrrell, J.M.; Zheng, Y.; Wang, S.; Shen, Z.; et al. Comprehensive resistome analysis reveals the prevalence of NDM and MCR-1 in Chinese poultry production. Nat. Microbiol. 2017, 2, 16260. [Google Scholar] [CrossRef]
- Lacotte, Y.; Ploy, M.C.; Raherison, S. Class 1 integrons are low-cost structures in Escherichia coli. ISME J. 2017, 11, 1535–1544. [Google Scholar] [CrossRef]
- Zhang, Y.; Liao, K.; Gao, H.; Wang, Q.; Wang, X.; Li, H.; Wang, R.; Wang, H. Decreased fitness and virulence in ST10 Escherichia coli harboring blaNDM-5 and mcr-1 against a ST4981 strain with blaNDM-5. Front. Cell Infect. Microbiol. 2017, 7, 242. [Google Scholar] [CrossRef]
- Wang, R.; Liu, Y.; Zhang, Q.; Jin, L.; Wang, Q.; Zhang, Y.; Wang, X.; Hu, M.; Li, L.; Qi, J.; et al. The prevalence of colistin resistance in Escherichia coli and Klebsiella pneumoniae isolated from food animals in China: Coexistence of mcr-1 and blaNDM with low fitness cost. Int. J. Antimicrob. Agents 2018, 51, 739–744. [Google Scholar] [CrossRef]
- Ma, T.; Fu, J.; Xie, N.; Ma, S.; Lei, L.; Zhai, W.; Shen, Y.; Sun, C.; Wang, S.; Shen, Z.; et al. Fitness cost of blaNDM-5-carrying p3R-IncX3 plasmids in wild-type NDM-free Enterobacteriaceae. Microorganisms 2020, 8, 377. [Google Scholar] [CrossRef]
- Pitout, J.D.; Nordmann, P.; Poirel, L. Carbapenemase-producing Klebsiella pneumoniae, a key pathogen set for global nosocomial dominance. Antimicrob. Agents Chemother. 2015, 59, 5873–5884. [Google Scholar] [CrossRef]
- Poirel, L.; Walsh, T.R.; Cuvillier, V.; Nordmann, P. Multiplex PCR for detection of acquired carbapenemase genes. Diagn. Microbiol. Infect. Dis. 2011, 70, 119–123. [Google Scholar] [CrossRef]
- Versalovic, J.; Schneider, M.; De Bruijn, F.J.; Lupski, J.R. Genomic fingerprinting of bacteria using repetitive sequence-based polymerase chain reaction. Methods Mol. Cell. Biol. 1994, 5, 25–40. [Google Scholar]
- Dallenne, C.; Da Costa, A.; Decré, D.; Favier, C.; Arlet, G. Development of a set of multiplex PCR assays for the detection of genes encoding important beta-lactamases in Enterobacteriaceae. J. Antimicrob. Chemother. 2010, 65, 490–495. [Google Scholar] [CrossRef] [PubMed]
- Nawaz, M.; Sung, K.; Khan, S.A.; Khan, A.A.; Steele, R. Biochemical and molecular characterization of tetracycline-resistant Aeromonas veronii isolates from catfish. Appl. Environ. Microbiol. 2006, 72, 6461–6466. [Google Scholar] [CrossRef] [PubMed]
- Kraft, C.A.; Timbury, M.C.; Platt, D.J. Distribution and genetic location of Tn7 in trimethoprim-resistant Escherichia coli. J. Med. Microbiol. 1986, 22, 125–131. [Google Scholar] [CrossRef] [PubMed]

| Antibiotic Class | Antibiotics | Ec355340∆pNDM-5 | Ec355340::pNDM-5 | TOP10 | TOP10::pNDM-5 | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| 3rd generation penicillins | Ampicillin | ≥32 | ≥32 | 8 | ≥32 | |||||
| Amoxicillin/clavulanic acid | ≥32 | ≥32 | 4 | ≥32 | ||||||
| 4th generation penicillins | Piperacillin/tazobactam | ≤4 | ≥128 | ≤4 | ≥128 | |||||
| 2nd generation cephalosporins | Cefuroxime | 16 | ≥64 | 8 | ≥64 | |||||
| Cefuroxime axetil | 16 | ≥64 | 8 | ≥64 | ||||||
| 3rd generation cephalosporins | Cefotaxime | ≤1 | ≥64 | ≤1 | ≥64 | |||||
| Ceftazidime | ≤1 | ≥64 | ≤1 | ≥64 | Resistant | |||||
| 4th generation cephalosporins | Cefepime | ≤1 | 8 | ≤1 | 16 | Intermediate | ||||
| Carbapenems | Ertapenem | ≤0.12 | ≥8 | ≤0.12 | ≥8 | Susceptible | ||||
| Meropenem | ≤0.25 | ≥16 | ≤0.25 | ≥16 | ||||||
| Aminoglycosides | Amikacin | ≤2 | ≤2 | ≤2 | ≤2 | |||||
| Gentamicin | ≤1 | ≤1 | ≤1 | ≤1 | ||||||
| Fluoroquinolones | Ciprofloxacin | ≥4 | ≥4 | ≤0.25 | ≤0.25 | |||||
| - | Fosfomycin | ≤16 | ≤16 | ≤16 | ≤16 | |||||
| - | Nitrofurantoin | 32 | ≤16 | ≤16 | ≤16 | |||||
| Polymyxins | Colistin | ≤0.5 | ≤0.5 | ≤0.5 | ≤0.5 | |||||
| Sulphonamides | Trimethoprim/sulphamethoxazole | ≤20 | ≥320 | ≤20 | ≥320 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tavares, R.D.S.; Tacão, M.; Ramalheira, E.; Ferreira, S.; Henriques, I. Report and Comparative Genomics of an NDM-5-Producing Escherichia coli in a Portuguese Hospital: Complex Class 1 Integrons as Important Players in blaNDM Spread. Microorganisms 2022, 10, 2243. https://doi.org/10.3390/microorganisms10112243
Tavares RDS, Tacão M, Ramalheira E, Ferreira S, Henriques I. Report and Comparative Genomics of an NDM-5-Producing Escherichia coli in a Portuguese Hospital: Complex Class 1 Integrons as Important Players in blaNDM Spread. Microorganisms. 2022; 10(11):2243. https://doi.org/10.3390/microorganisms10112243
Chicago/Turabian StyleTavares, Rafael D. S., Marta Tacão, Elmano Ramalheira, Sónia Ferreira, and Isabel Henriques. 2022. "Report and Comparative Genomics of an NDM-5-Producing Escherichia coli in a Portuguese Hospital: Complex Class 1 Integrons as Important Players in blaNDM Spread" Microorganisms 10, no. 11: 2243. https://doi.org/10.3390/microorganisms10112243
APA StyleTavares, R. D. S., Tacão, M., Ramalheira, E., Ferreira, S., & Henriques, I. (2022). Report and Comparative Genomics of an NDM-5-Producing Escherichia coli in a Portuguese Hospital: Complex Class 1 Integrons as Important Players in blaNDM Spread. Microorganisms, 10(11), 2243. https://doi.org/10.3390/microorganisms10112243






