Importance of Microbiome of Fecal Samples Obtained from Adolescents with Different Weight Conditions on Resistance Gene Transfer
Abstract
:1. Introduction
2. Materials and Methods
2.1. Design and Study Population
2.2. Selection Criteria
2.3. Receipt of Information and Biological Material
2.4. Genomic DNA Extraction from the Fecal Samples
2.5. Ribosomal 16s Gene Sequencing
2.6. Analysis of Antimicrobial Resistance
2.7. Bioinformatic Analysis
2.8. Statistical Analysis
3. Results
3.1. Adolescent Cohort
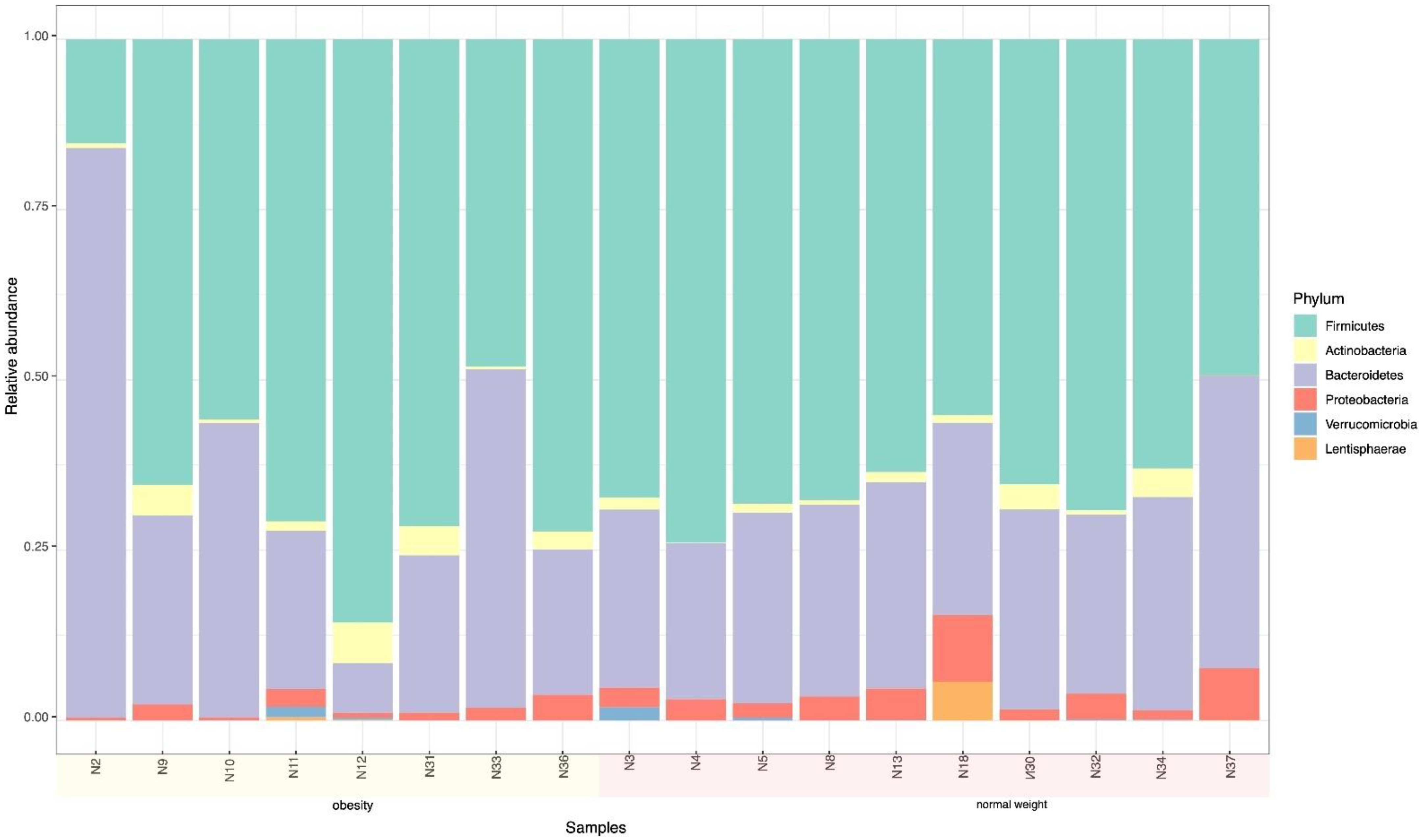
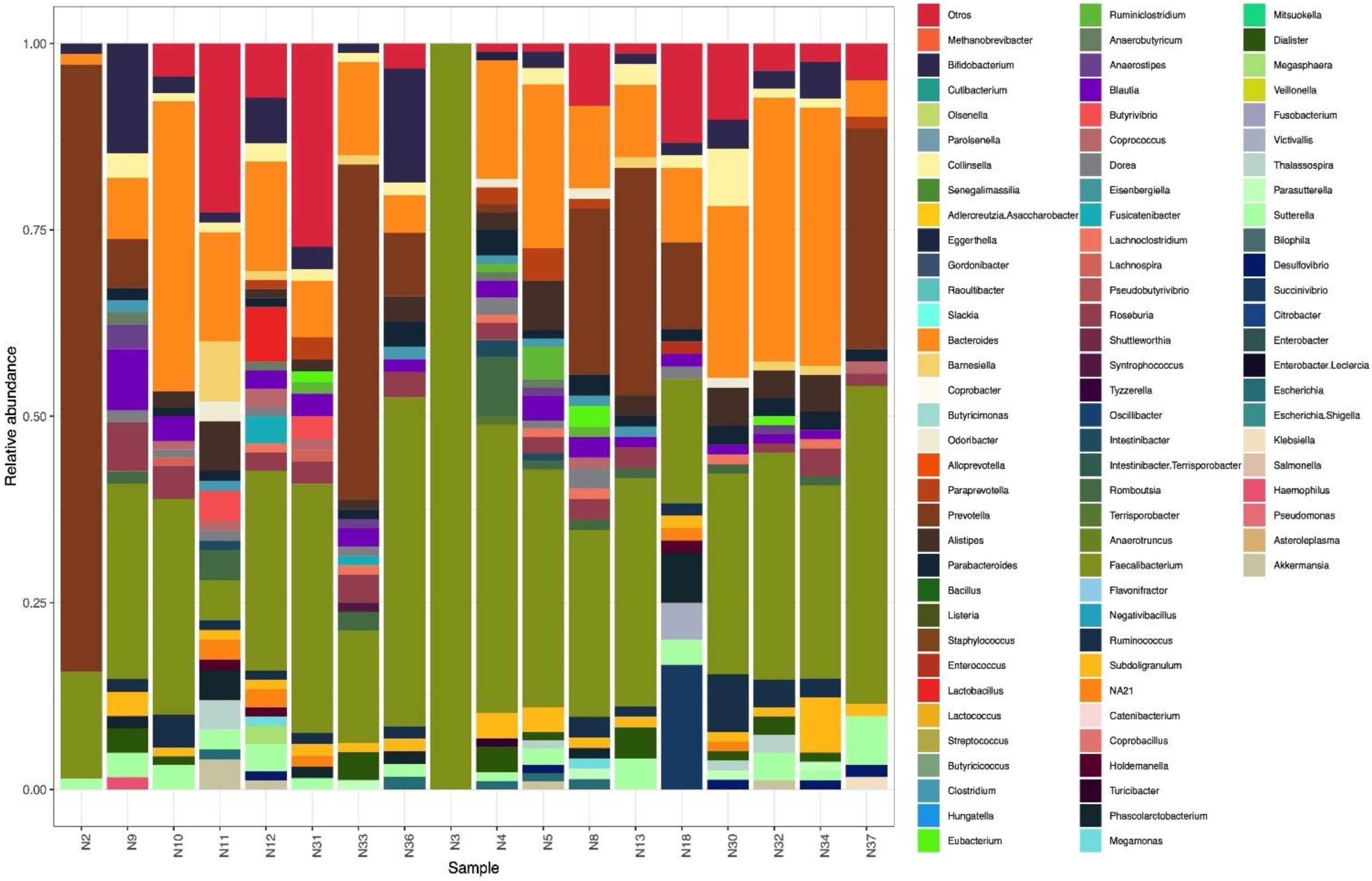
3.2. Microbiome
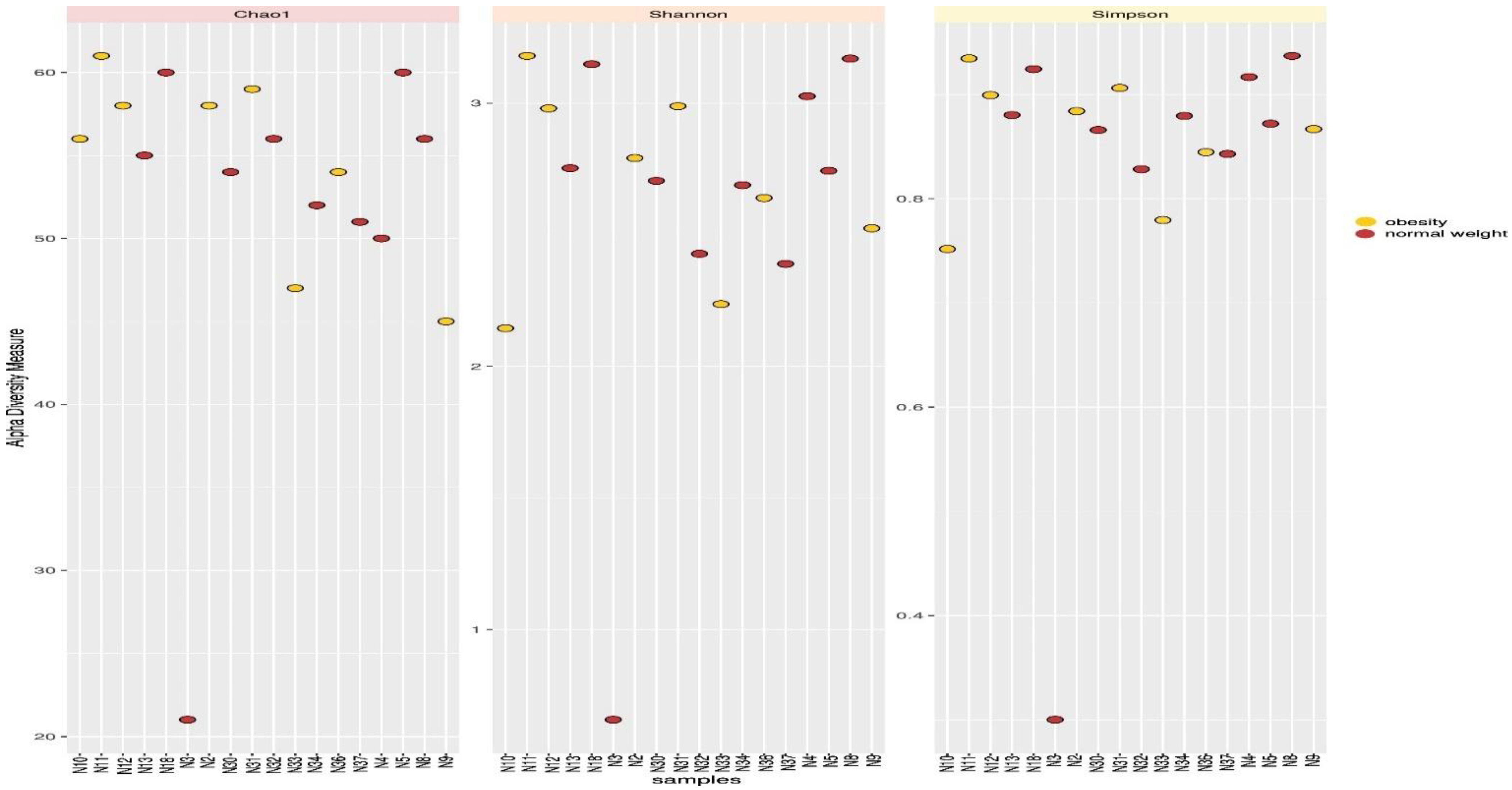
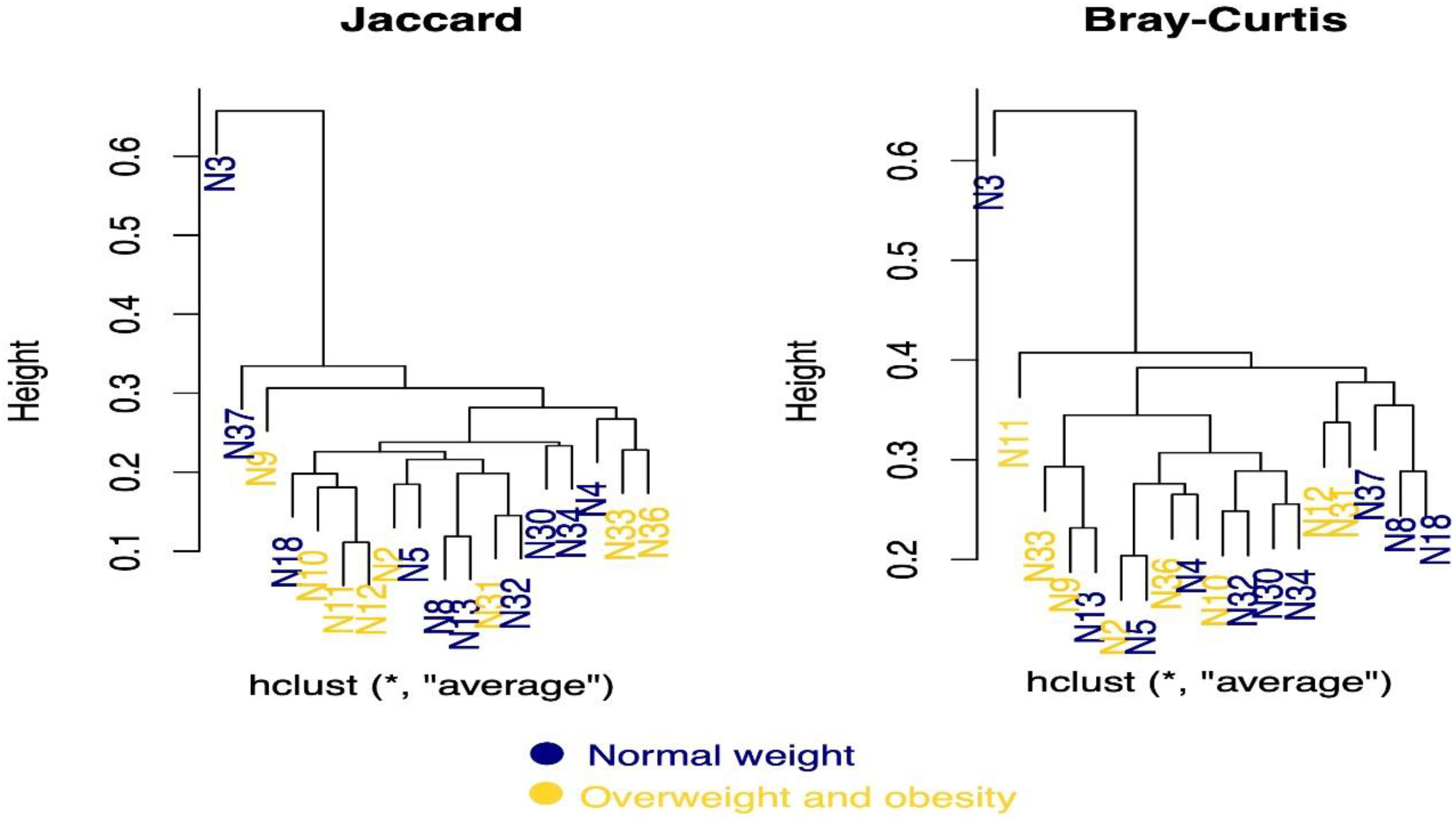
3.3. Microbial Diversity
3.4. Presence of Beneficial and Pathogenic Bacteria in the Intestinal Microbiota in Adolescents
3.5. Enterotypes
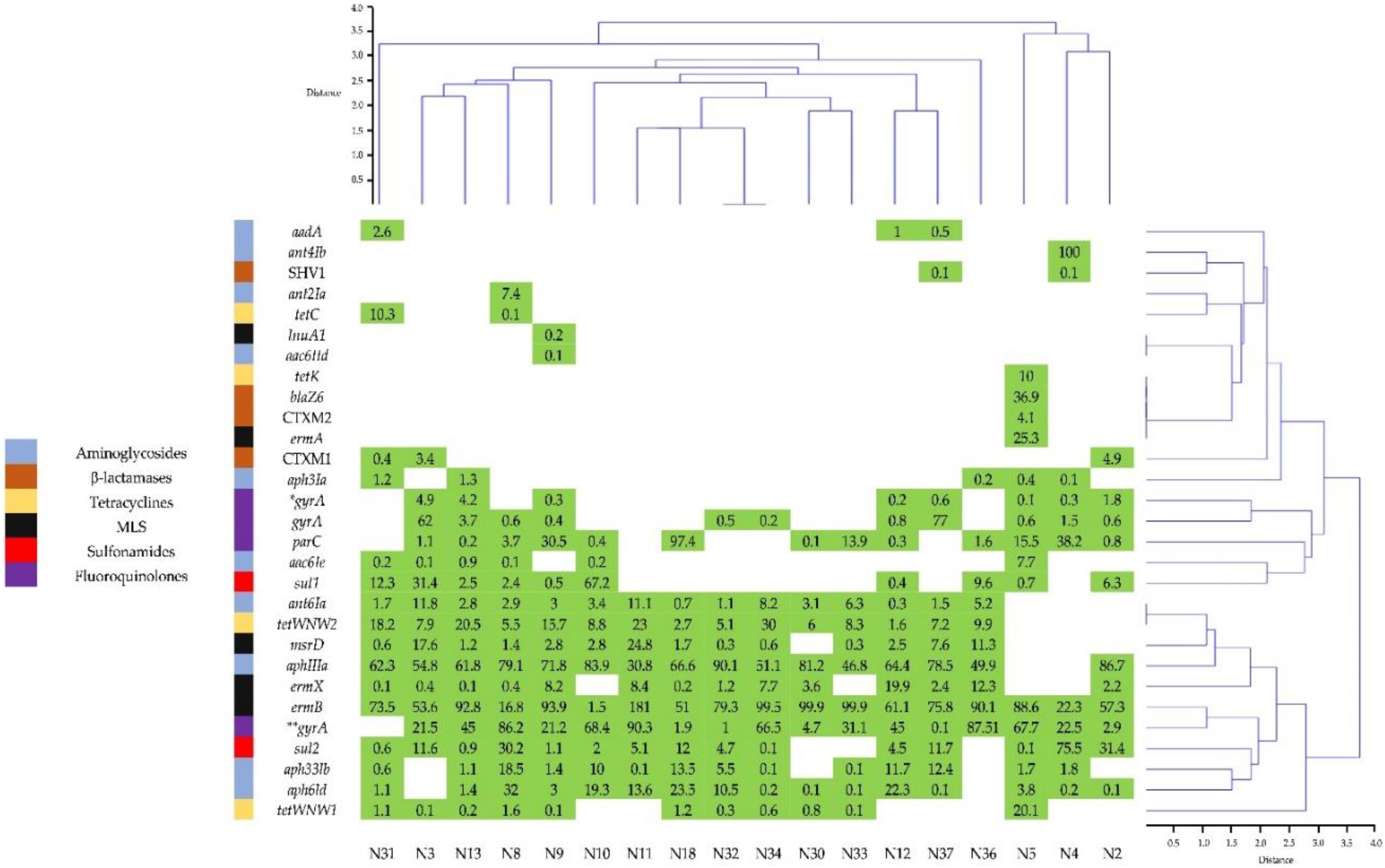
3.6. Presence of Resistance Genes to Antimicrobials
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Murray, C.J.; Ikuta, K.S.; Sharara, F.; Swetschinski, L.; Robles Aguilar, G.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E.; et al. Global Burden of Bacterial Antimicrobial Resistance in 2019: A Systematic Analysis. Lancet 2022, 399, 629–655. [Google Scholar] [CrossRef]
- O’Neill, J. Tackling Drug-Resistant Infections Globally: Final Report and Recommendations. Lon-Don: Review on Antimicrobial Resistance; Government of the United Kingdom: London, UK, 2016. [Google Scholar]
- WHO Regional Office for Europe. WHO Regional Office for Europe/European Centre for Disease Prevention and Control. Anti-Microbial Resistance Surveillance in Europe 2022–2020 Data; WHO Regional Office for Europe: Copenhagen, Denmark, 2022. [Google Scholar]
- Klevens, R.M.; Morrison, M.A.; Nadle, J.; Petit, S.; Gershman, K.; Ray, S.; Harrison, L.H.; Lynfield, R.; Dumyati, G.; Townes, J.M.; et al. Invasive Methicillin-Resistant Staphylococcus aureus Infections in the United States. JAMA 2007, 298, 1763–1771. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Centers for Disease Control and Prevention (U.S.). Antibiotic Resistance Threats in the United States, 2019; Centers for Disease Control and Prevention (U.S.): Atlanta, GA, USA, 2019. [Google Scholar]
- Hameed, M.F.; Chen, Y.; Wang, Y.; Shafiq, M.; Bilal, H.; Liu, L.; Ma, J.; Gu, P.; Ge, H. Epidemiological Characterization of Colistin and Carbapenem Resistant Enterobacteriaceae in a Tertiary: A Hospital from Anhui Province. Infect. Drug Resist. 2021, 14, 1325. [Google Scholar] [CrossRef]
- Bilal, H.; Zhang, G.; Rehman, T.; Han, J.; Khan, S.; Shafiq, M.; Yang, X.; Yan, Z.; Yang, X. First Report of blaNDM-1. Bearing IncX3 Plasmid in Clinically Isolated ST11 Klebsiella pneumoniae from Pakistan. Microorganisms 2021, 9, 951. [Google Scholar] [CrossRef]
- Sánchez-Huesca, R.; Lerma, A.; Guzmán-Saldaña, R.M.E.; Lerma, C. Prevalence of Antibiotics Prescription and Assessment of Prescribed Daily Dose in Outpatients from Mexico City. Antibiotics 2020, 9, 38. [Google Scholar] [CrossRef] [Green Version]
- Garza-González, E.; Morfín-Otero, R.; Mendoza-Olazarán, S.; Bocanegra-Ibarias, P.; Flores-Treviño, S.; Rodríguez-Noriega, E.; Ponce-de-León, A.; Sanchez-Francia, D.; Franco-Cendejas, R.; Arroyo-Escalante, S.; et al. A Snapshot of Antimicrobial Resistance in Mexico. Results from 47 Centers from 20 States during a Six-Month Period. PLoS ONE 2019, 14, e0209865. [Google Scholar] [CrossRef] [PubMed]
- Universidad Nacional Autónoma de México. Ciudad de México Programa Universitario de Investigación En Salud Plan Universitario de Control de La Resistencia Antimicrobiana; Universidad Nacional Autónoma de México: Ciudad de Mexico, Mexico, 2018. [Google Scholar]
- Ballesteros-Monrreal, M.G.; Arenas-Hernández, M.M.P.; Enciso-Martínez, Y.; Martinez de la Peña, C.F.; Rocha-Gracia, R.; del, C.; Lozano-Zarain, P.; Navarro-Ocaña, A.; Martínez-Laguna, Y.; de la Rosa-López, R. Virulence and Resistance Determinants of Uropathogenic Escherichia coli Strains Isolated from Pregnant and Non-Pregnant Women from Two States in Mexico. Infect. Drug Resist. 2020, 13, 295–310. [Google Scholar] [CrossRef] [Green Version]
- Yatsunenko, T.; Rey, F.E.; Manary, M.J.; Trehan, I.; Dominguez-Bello, M.G.; Contreras, M.; Magris, M.; Hidalgo, G.; Baldassano, R.N.; Anokhin, A.P.; et al. Human Gut Microbiome Viewed across Age and Geography. Nature 2012, 486, 222–227. [Google Scholar] [CrossRef] [Green Version]
- Otto, M. Next-Generation Sequencing to Monitor the Spread of Antimicrobial Resistance. Genome Med. 2017, 9, 68. [Google Scholar] [CrossRef] [Green Version]
- Bervoets, L.; Van Hoorenbeeck, K.; Kortleven, I.; Van Noten, C.; Hens, N.; Vael, C.; Goossens, H.; Desager, K.N.; Vankerckhoven, V. Differences in Gut Microbiota Composition between Obese and Lean Children: A Cross-Sectional Study. Gut Pathog. 2013, 5, 10. [Google Scholar] [CrossRef]
- Rosas-Plaza, S.; Hernández-Terán, A.; Navarro-Díaz, M.; Escalante, A.E.; Morales-Espinosa, R.; Cerritos, R. Human Gut Microbiome Across Different Lifestyles: From Hunter-Gatherers to Urban Populations. Front. Microbiol. 2022, 13, 843170. [Google Scholar] [CrossRef] [PubMed]
- Tsang, K.K.; Maguire, F.; Zubyk, H.L.; Chou, S.; Edalatmand, A.; Wright, G.D.; Beiko, R.G.; McArthur, A.G. Identifying Novel β-Lactamase Substrate Activity through in Silico Prediction of Antimicrobial Resistance. Microb. Genomics 2021, 7, 000500. [Google Scholar] [CrossRef]
- Johnson, J.S.; Spakowicz, D.J.; Hong, B.-Y.; Petersen, L.M.; Demkowicz, P.; Chen, L.; Leopold, S.R.; Hanson, B.M.; Agresta, H.O.; Gerstein, M. Evaluation of 16S RRNA Gene Sequencing for Species and Strain-Level Microbiome Analysis. Nat. Commun. 2019, 10, 5029. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-Resolution Sample Inference from Illumina Amplicon Data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, S.; Prem, A.; Tjokrosurjo, J.; Sary, M.; Van Bel, M.A.; Rodrigues-Hoffmann, A.; Kavanagh, M.; Wu, G.; Van Eden, M.E.; Krumbeck, J.A. The Canine Skin and Ear Microbiome: A Comprehensive Survey of Pathogens Implicated in Canine Skin and Ear Infections Using a Novel next-Generation-Sequencing-Based Assay. Vet. Microbiol. 2020, 247, 108764. [Google Scholar] [CrossRef]
- Lloyd-Price, J.; Abu-Ali, G.; Huttenhower, C. The Healthy Human Microbiome. Genome Med. 2016, 8, 51. [Google Scholar] [CrossRef] [Green Version]
- López-Contreras, B.E.; Morán-Ramos, S.; Villarruel-Vázquez, R.; Macías-Kauffer, L.; Villamil-Ramírez, H.; León-Mimila, P.; Vega-Badillo, J.; Sánchez-Muñoz, F.; Llanos-Moreno, L.E.; Canizalez-Román, A.; et al. Composition of Gut Microbiota in Obese and Normal-Weight Mexican School-Age Children and Its Association with Metabolic Traits: Gut Microbiota in Mexican Obese Children. Pediatr. Obes. 2018, 13, 381–388. [Google Scholar] [CrossRef]
- Méndez-Salazar, E.O.; Ortiz-López, M.G.; Granados-Silvestre, M.A.; Palacios-González, B.; Menjivar, M. Altered Gut Microbiota and Compositional Changes in Firmicutes and Proteobacteria in Mexican Undernourished and Obese Children. Front. Microbiol. 2018, 9, 2494. [Google Scholar] [CrossRef] [Green Version]
- Lopez-Siles, M.; Duncan, S.H.; Garcia-Gil, L.J.; Martinez-Medina, M. Faecalibacterium prausnitzii: From Microbiology to Diagnostics and Prognostics. ISME J. 2017, 11, 841–852. [Google Scholar] [CrossRef]
- Peters, B.A.; Shapiro, J.A.; Church, T.R.; Miller, G.; Trinh-Shevrin, C.; Yuen, E.; Friedlander, C.; Hayes, R.B.; Ahn, J. A Taxonomic Signature of Obesity in a Large Study of American Adults. Sci. Rep. 2018, 8, 9749. [Google Scholar] [CrossRef]
- Deleu, S.; Machiels, K.; Raes, J.; Verbeke, K.; Vermeire, S. Short Chain Fatty Acids and Its Producing Organisms: An Overlooked Therapy for IBD? EBioMedicine 2021, 66, 103293. [Google Scholar] [CrossRef] [PubMed]
- Sokol, H.; Pigneur, B.; Watterlot, L.; Lakhdari, O.; Bermúdez-Humarán, L.G.; Gratadoux, J.-J.; Blugeon, S.; Bridonneau, C.; Furet, J.-P.; Corthier, G.; et al. Faecalibacterium prausnitzii Is an Anti-Inflammatory Commensal Bacterium Identified by Gut Microbiota Analysis of Crohn Disease Patients. Proc. Natl. Acad. Sci. USA 2008, 105, 16731–16736. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mayorga Reyes, L.; González Vázquez, R.; Cruz Arroyo, S.M.; Melendez Avalos, A.; Reyes Castillo, P.A.; Chavaro Pérez, D.A.; Ramos Terrones, I.; Ramos Ibáñez, N.; Rodríguez Magallanes, M.M.; Langella, P.; et al. Correlation between Diet and Gut Bacteria in a Population of Young Adults. Int. J. Food Sci. Nutr. 2016, 67, 470–478. [Google Scholar] [CrossRef] [PubMed]
- Huang, F.; Sardari, R.R.; Jasilionis, A.; Böök, O.; Öste, R.; Rascón, A.; Heyman-Lindén, L.; Holst, O.; Karlsson, E.N. Cultivation of the Gut Bacterium Prevotella Copri DSM 18205T Using Glucose and Xylose as Carbon Sources. MicrobiologyOpen 2021, 10, e1213. [Google Scholar] [CrossRef] [PubMed]
- Verbrugghe, P.; Brynjólfsson, J.; Jing, X.; Björck, I.; Hållenius, F.; Nilsson, A. Evaluation of Hypoglycemic Effect, Safety and Immunomodulation of Prevotella copri in Mice. Sci. Rep. 2021, 11, 21279. [Google Scholar] [CrossRef]
- Duranti, S.; Ruiz, L.; Lugli, G.A.; Tames, H.; Milani, C.; Mancabelli, L.; Mancino, W.; Longhi, G.; Carnevali, L.; Sgoifo, A.; et al. Bifidobacterium Adolescentis as a Key Member of the Human Gut Microbiota in the Production of GABA. Sci. Rep. 2020, 10, 14112. [Google Scholar] [CrossRef]
- Nirmalkar, K.; Murugesan, S.; Pizano-Zárate, M.; Villalobos-Flores, L.; García-González, C.; Morales-Hernández, R.; Nuñez-Hernández, J.; Hernández-Quiroz, F.; Romero-Figueroa, M.; Hernández-Guerrero, C.; et al. Gut Microbiota and Endothelial Dysfunction Markers in Obese Mexican Children and Adolescents. Nutrients 2018, 10, 2009. [Google Scholar] [CrossRef] [Green Version]
- Mena-Vázquez, N.; Ruiz-Limón, P.; Moreno-Indias, I.; Manrique-Arija, S.; Tinahones, F.J.; Fernández-Nebro, A. Expansion of Rare and Harmful Lineages Is Associated with Established Rheumatoid Arthritis. J. Clin. Med. 2020, 9, 1044. [Google Scholar] [CrossRef] [Green Version]
- Arumugam, M.; Raes, J.; Pelletier, E.; Le Paslier, D.; Yamada, T.; Mende, D.R.; Fernandes, G.R.; Tap, J.; Bruls, T.; Bato, J.-M.; et al. Enterotypes of the Human Gut Microbiome. Nature 2011, 473, 174–180. [Google Scholar] [CrossRef] [Green Version]
- Maya-Lucas, O.; Murugesan, S.; Nirmalkar, K.; Alcaraz, L.D.; Hoyo-Vadillo, C.; Pizano-Zárate, M.L.; García-Mena, J. The Gut Microbiome of Mexican Children Affected by Obesity. Anaerobe 2019, 55, 11–23. [Google Scholar] [CrossRef]
- Van Schaik, W. The Human Gut Resistome. Philos. Trans. R. Soc. B Biol. Sci. 2015, 370, 20140087. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Takayama, Y.; Tanaka, T.; Oikawa, K.; Fukano, N.; Goto, M.; Takahashi, T. Prevalence of BlaZ Gene and Performance of Phenotypic Tests to Detect Penicillinase in Staphylococcus aureus Isolates from Japan. Ann. Lab. Med. 2018, 38, 155–159. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kareem, S.M.; Al-Kadmy, I.M.; Kazaal, S.S.; Mohammed Ali, A.N.; Aziz, S.N.; Makharita, R.R.; Algammal, A.M.; Al-Rejaie, S.; Behl, T.; Batiha, G.E.-S.; et al. Detection of GyrA and ParC Mutations and Prevalence of Plasmid-Mediated Quinolone Resistance Genes in Klebsiella pneumoniae. Infect. Drug Resist. 2021, 14, 555–563. [Google Scholar] [CrossRef] [PubMed]
- Wright, G.D. The Antibiotic Resistome: The Nexus of Chemical and Genetic Diversity. Nat. Rev. Microbiol. 2007, 5, 175–186. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.-K.; Kim, D.; Moon, D.-C.; Cho, Y.; Rho, M. Antibiotic Resistomes Discovered in the Gut Microbiomes of Korean Swine and Cattle. GigaScience 2020, 9, giaa043. [Google Scholar] [CrossRef]
- Woegerbauer, M.; Zeinzinger, J.; Springer, B.; Hufnagl, P.; Indra, A.; Korschineck, I.; Hofrichter, J.; Kopacka, I.; Fuchs, R.; Steinwider, J.; et al. Prevalence of the Aminoglycoside Phosphotransferase Genes Aph(3′)-IIIa and Aph(3′)-IIa in Escherichia coli, Enterococcus faecalis, Enterococcus faecium, Pseudomonas aeruginosa, Salmonella enterica Subsp. enterica and Staphylococcus aureus Isolates in Austria. J. Med. Microbiol. 2014, 63, 210–217. [Google Scholar] [CrossRef] [Green Version]
- Machado, D.; Barbosa, J.C.; Domingos, M.; Almeida, D.; Andrade, J.C.; Freitas, A.C.; Gomes, A.M. Revealing Antimicrobial Resistance Profile of the Novel Probiotic Candidate Faecalibacterium prausnitzii DSM 17677. Int. J. Food Microbiol. 2022, 363, 109501. [Google Scholar] [CrossRef]
- Szczuka, E.; Makowska, N.; Bosacka, K.; Słotwińska, A.; Kaznowski, A. Molecular Basis of Resistance to Macrolides, Lincosamides and Streptogramins in Staphylococcus hominis Strains Isolated from Clinical Specimens. Folia Microbiol. 2016, 61, 143–147. [Google Scholar] [CrossRef] [Green Version]
- Ammor, M.S.; Florez, A.B.; Alvarez-Martin, P.; Margolles, A.; Mayo, B. Analysis of Tetracycline Resistance Tet(W) Genes and Their Flanking Sequences in Intestinal Bifidobacterium Species. J. Antimicrob. Chemother. 2008, 62, 688–693. [Google Scholar] [CrossRef] [Green Version]
- Nøhr-Meldgaard, K.; Struve, C.; Ingmer, H.; Agersø, Y. The Tetracycline Resistance Gene, Tet(W) in Bifidobacterium Animalis Subsp. Lactis Follows Phylogeny and Differs from Tet(W) in Other Species. Front. Microbiol. 2021, 12, 658943. [Google Scholar] [CrossRef]
- Smoglica, C.; Angelucci, S.; Farooq, M.; Antonucci, A.; Marsilio, F.; Di Francesco, C.E. Microbial Community and Antimicrobial Resistance in Fecal Samples from Wild and Domestic Ruminants in Maiella National Park, Italy. One Health 2022, 15, 100403. [Google Scholar] [CrossRef] [PubMed]
- Gerzova, L.; Babak, V.; Sedlar, K.; Faldynova, M.; Videnska, P.; Cejkova, D.; Jensen, A.N.; Denis, M.; Kerouanton, A.; Ricci, A.; et al. Characterization of Antibiotic Resistance Gene Abundance and Microbiota Composition in Feces of Organic and Conventional Pigs from Four EU Countries. PLoS ONE 2015, 10, e0132892. [Google Scholar] [CrossRef] [PubMed]
- Tavella, T.; Turroni, S.; Brigidi, P.; Candela, M.; Rampelli, S. The Human Gut Resistome up to Extreme Longevity. mSphere 2021, 6, e00691-21. [Google Scholar] [CrossRef] [PubMed]

| Phylum | W + O + N | W + O | N |
|---|---|---|---|
| Relative Frequency % | |||
| None; Others | 4.4 | 5.88 | 3.32 |
| * Euryarchaeota | 0.0 | 0.00 | 0.02 |
| Melainabacteria | 0.0 | 0.13 | 0.20 |
| Actinobacteria | 3.9 | 5.00 | 2.62 |
| Bacteroidetes | 38.3 | 43.38 | 33.84 |
| Firmicutes | 49.4 | 42.13 | 55.41 |
| Lentisphaerae | 0.0 | 0.00 | 0.33 |
| Proteobacteria | 3.3 | 2.63 | 3.97 |
| Tenericutes | 0.0 | 0.25 | 0.07 |
| Verrucomicrobia | 0.6 | 0.63 | 0.19 |
| Total | 99.9 | 100.0 | 100.0 |
| Phylum | Species | Normal Weight | Overweight and Obese |
|---|---|---|---|
| Frequency % | |||
| Actinobacteria | Bifidobacterium adolescentis | 0.20 | 1.46 |
| Collinsella aerofaciens | 1.21 | 1.00 | |
| Bacteroidetes | Bacteroides stercoris | 0.90 | 3.46 |
| Bacteroides massiliensis | 0.83 | 0.91 | |
| Alistipes putredinis | 1.23 | 0.90 | |
| Barnesiella intestinihominis | 0.08 | 1.06 | |
| Bacteroides dorei | 0.30 | 1.70 | |
| Bacteroides vulgatus | 5.21 | 2.46 | |
| Prevotella copri | 3.40 | 12.78 | |
| Bacteroides uniformis | 1.23 | 0.55 | |
| Prevotella stercorea | 2.31 | 0.00 | |
| Firmicutes | Lactobacillus ruminis | 0.03 | 0.79 |
| Dorea formicigenerans | 0.40 | 0.35 | |
| Roseburia faecis | 0.49 | 1.04 | |
| Ruminococcus bicirculans | 1.05 | 0.54 | |
| Faecalibacterium prausnitzii | 29.37 | 16.75 | |
| Proteobacteria | Desulfovibrio piger | 0.33 | 0.13 |
| Sutterella wadsworthensis | 1.17 | 0.86 | |
| Escherichia coli | 0.40 | 0.24 | |
| Verrucomicrobia | Akkermansia muciniphila | 0.17 | 0.45 |
| Total | 50.30 | 47.40 | |
| C. aerofaciens | Total (%) | |||
|---|---|---|---|---|
| Positive | Negative | |||
| Diarrhea | Yes | 2 | 2 | 4 (22.2) |
| No | 13 | 1 | 14 (77.8) | |
| Total | 15 | 3 | 18 (100) | |
| E. coli | Total (%) | |||
|---|---|---|---|---|
| Positive | Negative | |||
| Physical activity | Yes | 8 | 1 | 9 (50) |
| No | 4 | 5 | 9 (50) | |
| Total | 12 | 6 | 18 (100) | |
| Enterotype | ||
|---|---|---|
| Adolescents | 1 | 2 |
| Bacteroides sp. | Prevotella sp. | |
| N (%) | ||
| Boys | 1 (5.6) | 6 (33.3) |
| Girls | 9 (50.0) | 2 (11.1) |
| 10 (55.6) | 8 (44.4) | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Navarro, A.; Rodea, G.E.; Castelán-Sánchez, H.G.; Saucedo-Pastrana, H.A.; Licona-Moreno, D.; Eslava-Campos, C.; Tirado-Gómez, L.L.; Vilchis-Reyes, A.; García de la Torre, G.; Cruz-Licea, V. Importance of Microbiome of Fecal Samples Obtained from Adolescents with Different Weight Conditions on Resistance Gene Transfer. Microorganisms 2022, 10, 1995. https://doi.org/10.3390/microorganisms10101995
Navarro A, Rodea GE, Castelán-Sánchez HG, Saucedo-Pastrana HA, Licona-Moreno D, Eslava-Campos C, Tirado-Gómez LL, Vilchis-Reyes A, García de la Torre G, Cruz-Licea V. Importance of Microbiome of Fecal Samples Obtained from Adolescents with Different Weight Conditions on Resistance Gene Transfer. Microorganisms. 2022; 10(10):1995. https://doi.org/10.3390/microorganisms10101995
Chicago/Turabian StyleNavarro, Armando, Gerardo E. Rodea, Hugo G. Castelán-Sánchez, Héctor Armando Saucedo-Pastrana, Delia Licona-Moreno, Carlos Eslava-Campos, Laura L. Tirado-Gómez, Ariel Vilchis-Reyes, Guadalupe García de la Torre, and Verónica Cruz-Licea. 2022. "Importance of Microbiome of Fecal Samples Obtained from Adolescents with Different Weight Conditions on Resistance Gene Transfer" Microorganisms 10, no. 10: 1995. https://doi.org/10.3390/microorganisms10101995







