Comparative Analysis of Streptococcus pneumoniae Type I Restriction-Modification Loci: Variation in hsdS Gene Target Recognition Domains
Abstract
1. Importance
2. Introduction
3. Results
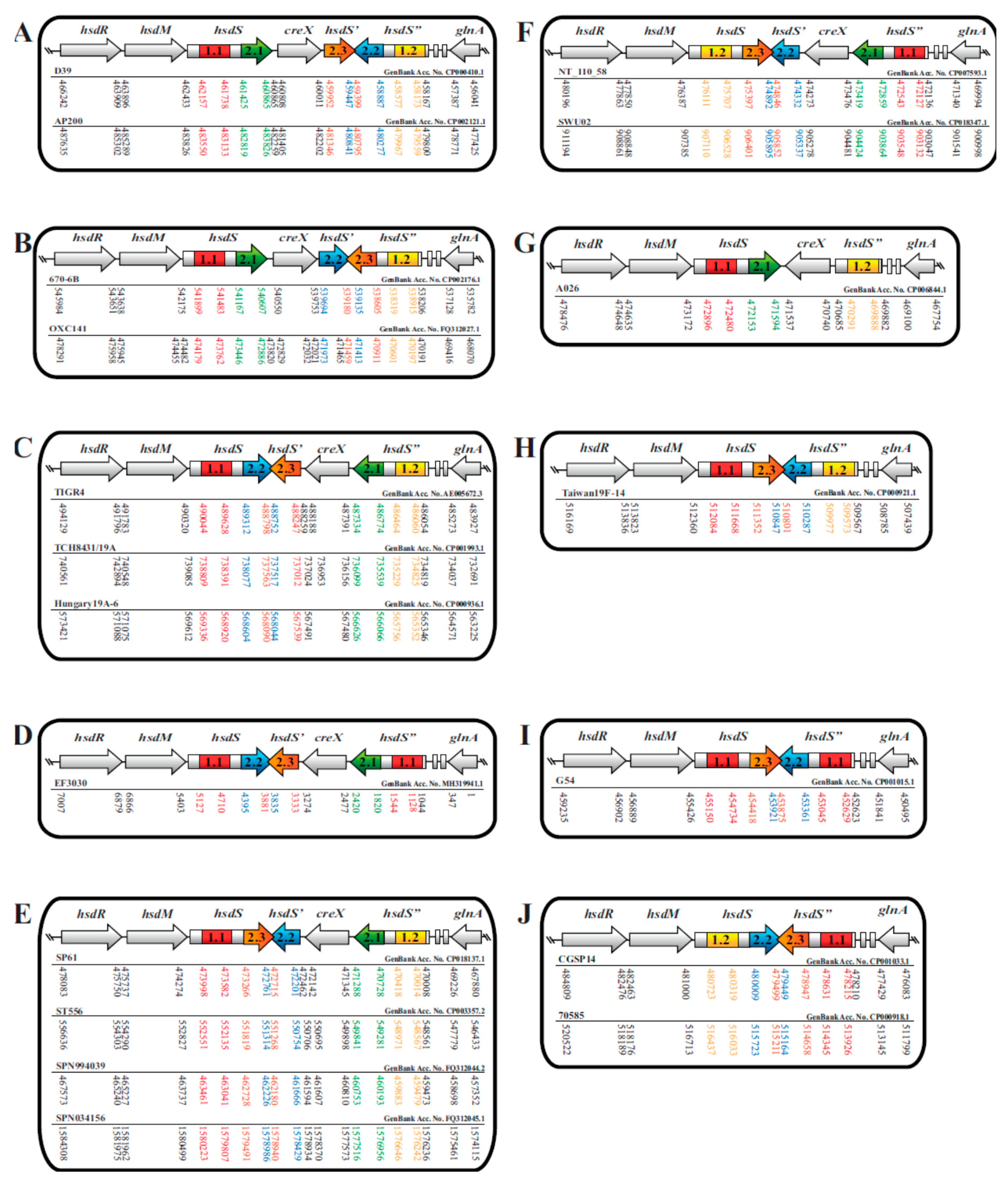
3.1. S. pneumoniae hsd Type I Restriction-Modification (R-M) Loci Are Genetically Diverse
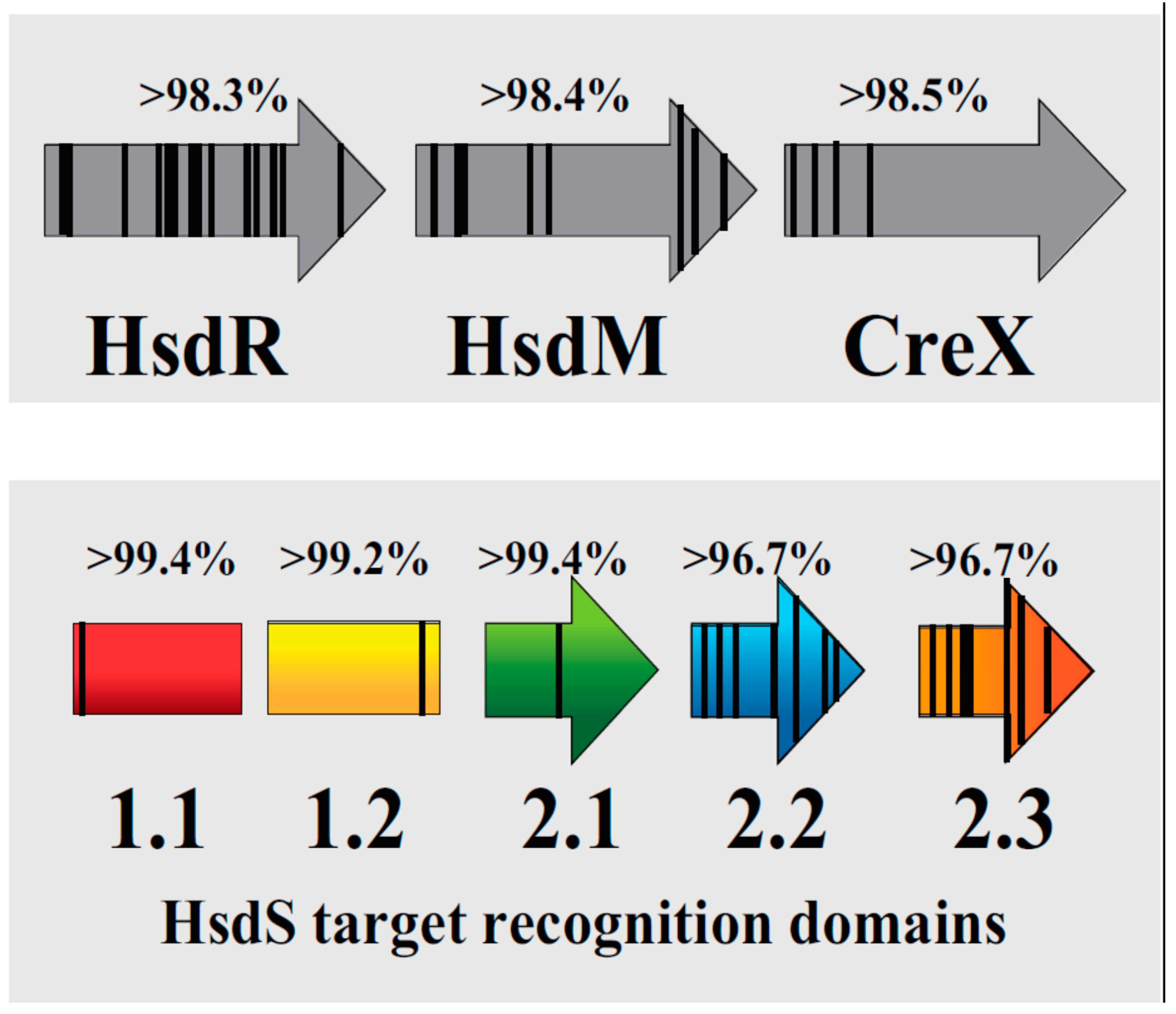
3.2. S. pneumoniae hsdS Coding Sequences Were Highly Homologous
4. Discussion
5. Materials and Methods
5.1. Sequence Analysis
5.2. Accession Numbers
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Musher, D.M. Infections caused by Streptococcus pneumoniae: Clinical spectrum, pathogenesis, immunity, and treatment. Clin. Infect. Dis. 1992, 14, 801–807. [Google Scholar] [CrossRef]
- van der Linden, M.; Winkel, N.; Kuntzel, S.; Farkas, A.; Perniciaro, S.R.; Reinert, R.R.; Imohl, M. Epidemiology of Streptococcus pneumoniae serogroup 6 isolates from IPD in children and adults in Germany. PLoS ONE 2013, 8, e60848. [Google Scholar] [CrossRef] [PubMed]
- Walker, C.L.; Rudan, I.; Liu, L.; Nair, H.; Theodoratou, E.; Bhutta, Z.A.; O’Brien, K.L.; Campbell, H.; Black, R.E. Global burden of childhood pneumonia and diarrhoea. Lancet 2013, 381, 1405–1416. [Google Scholar] [CrossRef]
- Green, M.C.; Mason, E.O.; Kaplan, S.L.; Lamberth, L.B.; Stovall, S.H.; Givner, L.B.; Bradley, J.S.; Tan, T.Q.; Barson, W.J.; Hoffman, J.A.; et al. Increase in prevalence of Streptococcus pneumoniae serotype 6C at eight children’s hospitals in the United States from 1993 to 2009. J. Clin. Microbiol. 2011, 49, 2097–2101. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Talaga, P.; Bellamy, L.; Moreau, M. Quantitative determination of C-polysaccharide in Streptococcus pneumoniae capsular polysaccharides by use of high-performance anion-exchange chromatography with pulsed amperometric detection. Vaccine 2001, 19, 2987–2994. [Google Scholar] [CrossRef]
- Bogaert, D.; De Groot, R.; Hermans, P.W. Streptococcus pneumoniae colonisation: The key to pneumococcal disease. Lancet Infect. Dis. 2004, 4, 144–154. [Google Scholar] [CrossRef]
- Simell, B.; Auranen, K.; Kayhty, H.; Goldblatt, D.; Dagan, R.; O’Brien, K.L.; Pneumococcal Carriage, G. The fundamental link between pneumococcal carriage and disease. Expert Rev. Vaccines 2012, 11, 841–855. [Google Scholar] [CrossRef]
- Regev-Yochay, G.; Raz, M.; Dagan, R.; Porat, N.; Shainberg, B.; Pinco, E.; Keller, N.; Rubinstein, E. Nasopharyngeal carriage of Streptococcus pneumoniae by adults and children in community and family settings. Clin. Infect. Dis. 2004, 38, 632–639. [Google Scholar] [CrossRef]
- Hausdorff, W.P.; Bryant, J.; Kloek, C.; Paradiso, P.R.; Siber, G.R. The contribution of specific pneumococcal serogroups to different disease manifestations: Implications for conjugate vaccine formulation and use, part II. Clin. Infect. Dis. 2000, 30, 122–140. [Google Scholar] [CrossRef]
- Hausdorff, W.P.; Bryant, J.; Paradiso, P.R.; Siber, G.R. Which pneumococcal serogroups cause the most invasive disease: Implications for conjugate vaccine formulation and use, part I. Clin. Infect. Dis. 2000, 30, 100–121. [Google Scholar] [CrossRef]
- Jin, P.; Kong, F.; Xiao, M.; Oftadeh, S.; Zhou, F.; Liu, C.; Russell, F.; Gilbert, G.L. First report of putative Streptococcus pneumoniae serotype 6D among nasopharyngeal isolates from Fijian children. J. Infect. Dis. 2009, 200, 1375–1380. [Google Scholar] [CrossRef] [PubMed]
- Robbins, J.B.; Austrian, R.; Lee, C.J.; Rastogi, S.C.; Schiffman, G.; Henrichsen, J.; Makela, P.H.; Broome, C.V.; Facklam, R.R.; Tiesjema, R.H.; et al. Considerations for formulating the second-generation pneumococcal capsular polysaccharide vaccine with emphasis on the cross-reactive types within groups. J. Infect. Dis. 1983, 148, 1136–1159. [Google Scholar] [CrossRef] [PubMed]
- Weiser, J.N.; Austrian, R.; Sreenivasan, P.K.; Masure, H.R. Phase variation in pneumococcal opacity: Relationship between colonial morphology and nasopharyngeal colonization. Infect. Immun. 1994, 62, 2582–2589. [Google Scholar] [CrossRef]
- Briles, D.E.; Novak, L.; Hotomi, M.; van Ginkel, F.W.; King, J. Nasal colonization with Streptococcus pneumoniae includes subpopulations of surface and invasive pneumococci. Infect. Immun. 2005, 73, 6945–6951. [Google Scholar] [CrossRef]
- Arai, J.; Hotomi, M.; Hollingshead, S.K.; Ueno, Y.; Briles, D.E.; Yamanaka, N. Streptococcus pneumoniae isolates from middle ear fluid and nasopharynx of children with acute otitis media exhibit phase variation. J. Clin. Microbiol. 2011, 49, 1646–1649. [Google Scholar] [CrossRef][Green Version]
- Kim, J.O.; Weiser, J.N. Association of intrastrain phase variation in quantity of capsular polysaccharide and teichoic acid with the virulence of Streptococcus pneumoniae. J. Infect. Dis. 1998, 177, 368–377. [Google Scholar] [CrossRef]
- Avery, O.T.; Macleod, C.M.; McCarty, M. Studies on the chemical nature of the substance inducing transformation of pneumococcal types: Induction of transformation by a desoxyribonucleic acid fraction isolated from pneumococcus Type III. J. Exp. Med. 1944, 79, 137–158. [Google Scholar] [CrossRef] [PubMed]
- Austrian, R. Morphologic variation in pneumococcus. I. An analysis of the bases for morphologic variation in pneumococcus and description of a hitherto undefined morphologic variant. J. Exp. Med. 1953, 98, 21–34. [Google Scholar] [CrossRef]
- Austrian, R. Morphologic variation in pneumococcus. II. Control of pneumococcal morphology through transformation reactions. J. Exp. Med. 1953, 98, 35–40. [Google Scholar] [CrossRef]
- Dawson, M.H. The interconvertibility of “R” and “S” forms of pneumococcus. J. Exp. Med. 1928, 47, 577–591. [Google Scholar] [CrossRef]
- Paul, J.R. Pneumococcus variants: I. Intermediate forms and the influence of environment in their production during in-vitro S to R and R to S transitions. J. Bacteriol. 1934, 28, 45–67. [Google Scholar] [CrossRef] [PubMed]
- Manso, A.S.; Chai, M.H.; Atack, J.M.; Furi, L.; De Ste Croix, M.; Haigh, R.; Trappetti, C.; Ogunniyi, A.D.; Shewell, L.K.; Boitano, M.; et al. A random six-phase switch regulates pneumococcal virulence via global epigenetic changes. Nat. Commun. 2014, 5, 5055. [Google Scholar] [CrossRef]
- Li, J.; Li, J.W.; Feng, Z.; Wang, J.; An, H.; Liu, Y.; Wang, Y.; Wang, K.; Zhang, X.; Miao, Z.; et al. Epigenetic switch driven by DNA inversions dictates phase variation in Streptococcus pneumoniae. PLoS Pathog. 2016, 12, e1005762. [Google Scholar] [CrossRef] [PubMed]
- Oliver, M.B.; Basu Roy, A.; Kumar, R.; Lefkowitz, E.J.; Swords, W.E. Streptococcus pneumoniae TIGR4 phase-locked opacity variants differ in virulence phenotypes. MSphere 2017, 2, e00386-17. [Google Scholar] [CrossRef] [PubMed]
- Weiserova, M.; Ryu, J. Characterization of a restriction modification system from the commensal Escherichia coli strain A0 34/86 (O83:K24:H31). BMC Microbiol. 2008, 8, 106. [Google Scholar] [CrossRef] [PubMed]
- Hilty, M.; Wuthrich, D.; Salter, S.J.; Engel, H.; Campbell, S.; Sa-Leao, R.; de Lencastre, H.; Hermans, P.; Sadowy, E.; Turner, P.; et al. Global phylogenomic analysis of nonencapsulated Streptococcus pneumoniae reveals a deep-branching classic lineage that is distinct from multiple sporadic lineages. Genome Biol. Evol. 2014, 6, 3281–3294. [Google Scholar] [CrossRef]
- Lanie, J.A.; Ng, W.L.; Kazmierczak, K.M.; Andrzejewski, T.M.; Davidsen, T.M.; Wayne, K.J.; Tettelin, H.; Glass, J.I.; Winkler, M.E. Genome sequence of Avery’s virulent serotype 2 strain D39 of Streptococcus pneumoniae and comparison with that of unencapsulated laboratory strain R6. J. Bacteriol. 2007, 189, 38–51. [Google Scholar] [CrossRef]
- Donati, C.; Hiller, N.L.; Tettelin, H.; Muzzi, A.; Croucher, N.J.; Angiuoli, S.V.; Oggioni, M.; Dunning Hotopp, J.C.; Hu, F.Z.; Riley, D.R.; et al. Structure and dynamics of the pan-genome of Streptococcus pneumoniae and closely related species. Genome Biol. 2010, 11, R107. [Google Scholar] [CrossRef]
- Hotopp, J.D.; Censini, S.; Masignani, V.; Covacci, A.; Tettelin, H. Direct Submission of Streptococcus Pneumoniae Strain 70585, Complete Genome (Accession No. CP000918.1); J Craig Venter Institute: Rockville, MD, USA, 2007; Available online: https://www.ncbi.nlm.nih.gov/nuccore/CP000918.1 (accessed on 29 August 2020).
- Tettelin, H.; Nelson, K.E.; Paulsen, I.T.; Eisen, J.A.; Read, T.D.; Peterson, S.; Heidelberg, J.; DeBoy, R.T.; Haft, D.H.; Dodson, R.J.; et al. Complete genome sequence of a virulent isolate of Streptococcus pneumoniae. Science 2001, 293, 498–506. [Google Scholar] [CrossRef]
- Neuendorf, B.; Radune, D.; Fedorova, N.B.; Khouri, H.M.; Dodson, R.J.; Daugherty, S.C.; Hollingshead, S.; Tettelin, H. Direct Submission of Streptococcus Pneumoniae Strain 670-6B, Complete Genome, DNA (Accession No. CP002176.1); Institute for Genome Sciences, University of Maryland School of Medicine: Baltimore, MD, USA, 2010; Available online: https://www.ncbi.nlm.nih.gov/nuccore/CP002176.1 (accessed on 29 August 2020).
- Camilli, R.; Del Grosso, M.; Iannelli, F.; Pantosti, A. New genetic element carrying the erythromycin resistance determinant erm(TR) in Streptococcus pneumoniae. Antimicrob Agents Chemother 2008, 52, 619–625. [Google Scholar] [CrossRef][Green Version]
- Ding, F.; Tang, P.; Hsu, M.H.; Cui, P.; Hu, S.; Yu, J.; Chiu, C.H. Genome evolution driven by host adaptations results in a more virulent and antimicrobial-resistant Streptococcus pneumoniae serotype 14. BMC Genom. 2009, 10, 158. [Google Scholar] [CrossRef] [PubMed]
- Muzny, D.M.; Qin, X.; Buhay, C.J.; Dugan-Rocha, S.; Ding, Y.; Chen, G.; Hawes, A.C.; Holder, M.; Jhangiani, S.N.; Johnson, A.J.; et al. Direct Submission of Streptococcus Pneumoniae Strain TCH8431/19A, Complete Genome, DNA (Accession No. CP001993.1); Human Genome Sequencing Center, Baylor College of Medicine: Houston, TX, USA, 2010; Available online: https://www.ncbi.nlm.nih.gov/nuccore/CP001993.1 (accessed on 29 August 2020).
- Hotopp, J.D.; Censini, S.; Masignani, V.; Covacci, A.; Tettelin, H. Direct Submission of Streptococcus Pneumoniae Strain Hungary19A-6, Complete Genome, DNA (Accession No.CP000936.1); J. Craig Venter Institute: Rockville, MD, USA, 2008; Available online: https://www.ncbi.nlm.nih.gov/nuccore/CP000936.1 (accessed on 29 August 2020).
- Donner, J.; Bunk, B.; Schober, I.; Jarek, M.; Overmann, J.; Wagner-Doebler, I. Data from “Full Genome Sequence of Streptococcus Pneumoniae Clinical Isolate SP61,”(Accession No. CP018137.1); Departemet Molecular Infection Biology, Helmholtz-Centre for Infection Research: Braunschweig, Germany, 2016; Available online: https://www.ncbi.nlm.nih.gov/nuccore/CP018137.1 (accessed on 29 August 2020).
- Li, G.; Hu, F.Z.; Yang, X.; Cui, Y.; Yang, J.; Qu, F.; Gao, G.F.; Zhang, J.R. Complete genome sequence of Streptococcus pneumoniae strain ST556, a multidrug-resistant isolate from an otitis media patient. J. Bacteriol. 2012, 194, 3294–3295. [Google Scholar] [CrossRef] [PubMed]
- Dopazo, J.; Mendoza, A.; Herrero, J.; Caldara, F.; Humbert, Y.; Friedli, L.; Guerrier, M.; Grand-Schenk, E.; Gandin, C.; de Francesco, M.; et al. Annotated draft genomic sequence from a Streptococcus pneumoniae type 19F clinical isolate. Microb. Drug Resist. 2001, 7, 99–125. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.S. Direct Submission of Streptococcus Pneumoniae Strain SWU2, Complete Genome, DNA (Accession No. CP018347.1); Department of Pathogenic Biology, School of Basic Medical Sciences: Jinan, China; Southwest Medical University: Luzhou, China, 2016; Available online: https://www.ncbi.nlm.nih.gov/nuccore/CP018347.1 (accessed on 29 August 2020).
- Sui, Z.; Zhou, W.; Yao, K.; Liu, L.; Zhang, G.; Yang, Y.; Feng, J. Complete genome sequence of Streptococcus pneumoniae strain A026, a clinical multidrug-resistant isolate carrying Tn2010. Genome Announc. 2013, 1, e01034-13. [Google Scholar] [CrossRef]
- Didelot, X.; Walker, A.S.; Peto, T.E.; Crook, D.W.; Wilson, D.J. Within-host evolution of bacterial pathogens. Nat. Rev. Microbiol. 2016, 14, 150–162. [Google Scholar] [CrossRef]
- Weiser, J.N.; Pan, N.; McGowan, K.L.; Musher, D.; Martin, A.; Richards, J. Phosphorylcholine on the lipopolysaccharide of Haemophilus influenzae contributes to persistence in the respiratory tract and sensitivity to serum killing mediated by C-reactive protein. J. Exp. Med. 1998, 187, 631–640. [Google Scholar] [CrossRef]
- VanWagoner, T.M.; Atack, J.M.; Nelson, K.L.; Smith, H.K.; Fox, K.L.; Jennings, M.P.; Stull, T.L.; Smith, A.L. The modA10 phasevarion of nontypeable Haemophilus influenzae R2866 regulates multiple virulence-associated traits. Microb. Pathog. 2016, 92, 60–67. [Google Scholar] [CrossRef]
- Atack, J.M.; Srikhanta, Y.N.; Fox, K.L.; Jurcisek, J.A.; Brockman, K.L.; Clark, T.A.; Boitano, M.; Power, P.M.; Jen, F.E.; McEwan, A.G.; et al. A biphasic epigenetic switch controls immunoevasion, virulence and niche adaptation in non-typeable Haemophilus influenzae. Nat. Commun. 2015, 6, 7828. [Google Scholar] [CrossRef]
- Srikhanta, Y.N.; Maguire, T.L.; Stacey, K.J.; Grimmond, S.M.; Jennings, M.P. The phasevarion: A genetic system controlling coordinated, random switching of expression of multiple genes. Proc. Natl. Acad Sci. USA 2005, 102, 5547–5551. [Google Scholar] [CrossRef]
- Srikhanta, Y.N.; Gorrell, R.J.; Steen, J.A.; Gawthorne, J.A.; Kwok, T.; Grimmond, S.M.; Robins-Browne, R.M.; Jennings, M.P. Phasevarion mediated epigenetic gene regulation in Helicobacter pylori. PLoS ONE 2011, 6, e27569. [Google Scholar] [CrossRef]
- Torres-Cruz, J.; van der Woude, M.W. Slipped-strand mispairing can function as a phase variation mechanism in Escherichia coli. J. Bacteriol. 2003, 185, 6990–6994. [Google Scholar] [CrossRef]
- Brooks, J.L.; Jefferson, K.K. Phase variation of poly-N-acetylglucosamine expression in Staphylococcus aureus. PLoS Pathog. 2014, 10, e1004292. [Google Scholar] [CrossRef] [PubMed]
- Waldron, D.E.; Lindsay, J.A. Sau1: A novel lineage-specific type I restriction-modification system that blocks horizontal gene transfer into Staphylococcus aureus and between S. aureus isolates of different lineages. J. Bacteriol. 2006, 188, 5578–5585. [Google Scholar] [CrossRef] [PubMed]
- Zieg, J.; Silverman, M.; Hilmen, M.; Simon, M. Recombinational switch for gene expression. Science 1977, 196, 170–172. [Google Scholar] [CrossRef]
- Reddy, S.P.; Rasmussen, W.G.; Baseman, J.B. Molecular cloning and characterization of an adherence-related operon of Mycoplasma genitalium. J. Bacteriol. 1995, 177, 5943–5951. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sitaraman, R.; Dybvig, K. The hsd loci of Mycoplasma pulmonis: Organization, rearrangements and expression of genes. Mol. Microbiol. 1997, 26, 109–120. [Google Scholar] [CrossRef] [PubMed]
- Alamro, M.; Bidmos, F.A.; Chan, H.; Oldfield, N.J.; Newton, E.; Bai, X.; Aidley, J.; Care, R.; Mattick, C.; Turner, D.P.; et al. Phase variation mediates reductions in expression of surface proteins during persistent meningococcal carriage. Infect. Immun. 2014, 82, 2472–2484. [Google Scholar] [CrossRef] [PubMed]
- Gawthorne, J.A.; Beatson, S.A.; Srikhanta, Y.N.; Fox, K.L.; Jennings, M.P. Origin of the diversity in DNA recognition domains in phasevarion associated modA genes of pathogenic Neisseria and Haemophilus influenzae. PLoS ONE 2012, 7, e32337. [Google Scholar] [CrossRef] [PubMed]
- Guymon, L.F.; Esser, M.; Shafer, W.M. Pyocin-resistant lipopolysaccharide mutants of Neisseria gonorrhoeae: Alterations in sensitivity to normal human serum and polymyxin B. Infect. Immun. 1982, 36, 541–547. [Google Scholar] [CrossRef]
- Markussen, T.; Marvig, R.L.; Gomez-Lozano, M.; Aanaes, K.; Burleigh, A.E.; Hoiby, N.; Johansen, H.K.; Molin, S.; Jelsbak, L. Environmental heterogeneity drives within-host diversification and evolution of Pseudomonas aeruginosa. mBio 2014, 5, e01592-14. [Google Scholar] [CrossRef]

| Strain | GenBank No. | Serotype | Body Site Origin | Country of Origin | Reference |
|---|---|---|---|---|---|
| NT_110_58 | CP007593.1 | NT | Nasopharynx | Switzerland | [26] |
| D39 | CP000410.1 | 2 | Blood | ND | [27] |
| OXC141 | FQ312027.1 | 3 | Carriage | UK | [28] |
| SPN034156 | FQ312045.1 | 3 | ND | Italy | [28] |
| SPN994039 | FQ312044.2 | 3 | ND | ND | [28] |
| 70585 | CP000918.1 | 4 | ND | ND | [29] |
| TIGR4 | AE005672.3 | 4 | Blood | Norway | [30] |
| 670-6B | CP002176.1 | 6B | ND | Spain | [31] |
| AP200 | CP002121.1 | 11A | Meningitis | Italy | [32] |
| CGSP14 | CP001033.1 | 14 | ND | China | [33] |
| TCH8431/19A | CP001993.1 | 19A | ND | USA | [34] |
| Hungary19A-6 | CP000936.1 | 19A | ND | Hungary | [35] |
| SP61 | CP018137.1 | 19A | Thorax | Germany | [36] |
| ST556 | CP003357.2 | 19F | Otitis media | USA | [37] |
| Taiwan19F-14 | CP000921.1 | 19F | ND | Taiwan | [28] |
| G54 | CP001015.1 | 19F | Respiratory | Italy | [38] |
| SWU02 | CP018347.1 | 19F | Sputum | China | [39] |
| A026 | CP006844.1 | 19F | ND | China | [40] |
| EF3030 | MH319941.1 | 19F | Otitis media | USA | This study |
| hsdS Target Recognition Domain | ||||||||
|---|---|---|---|---|---|---|---|---|
| S. pneumoniae Strain | hsdR | hsdM | creX | 1.1 | 1.2 | 2.1 | 2.2 | 2.3 |
| D39 | + | + | + | + | + | + | + | + |
| TIGR4 | + | + | + | + | + | + | + | + |
| TCH8431/19A | + | + | + | + | + | + | + | + |
| Hungary19A-6 | + | + | + | + | + | + | + | + |
| SP61 | + | + | + | + | + | + | + | + |
| ST556 | + | + | + | + | + | + | + | + |
| SPN034156 | + | + | + | + | + | + | + | + |
| SPN994039 | + | + | + | + | + | + | + | + |
| NT_110_58 | + | + | + | + | + | + | + | + |
| 670-6B | + | + | + | + | + | + | + | + |
| OXC141 | + | + | + | + a | + | + | + | + |
| AP200 | + | + | + | + | + a | + | + a | + |
| SWU02 | + | + | + | + | + a | + | + | + a |
| 70585 | + | + | + | + | + | - | + | + |
| SP14 | + | + | - | + | + | - | + | + |
| Taiwan19F-14 | + | + | - | + | + | - | + | + |
| A026 | + | + | - | + | + | + | - | - |
| G54 | + | + | - | + b | - | - | + | + a |
| EF3030 | + | + | + | + b | - | + | + | + a |
| Sum | 19 | 19 | 15 | 19 | 17 | 15 | 18 | 18 |
| Included for analysis | 18 | 18 | 15 | 18 | 15 | 15 | 17 | 15 |
| Length (amino acid) | 777 | 487 | 265 | 179 | 135 | 186 | 186 | 183 |
| Matching residues | 763 | 479 | 261 | 178 | 134 | 185 | 180 | 177 |
| Similarity (%) | 98.3 | 98.4 | 98.5 | 99.4 | 99.2 | 99.4 | 96.7 | 96.7 |
| S. pneumoniae Strain GenBank | D39 CP000410.1 | AP200 CP002121.1 | 670-6B CP002176.1 | OXC141 FQ312027.1 | TIGR4 AE005672.3 | Hungary19A-6 CP000936.1 | TCH8431/19A CP001993.1 | SP61 CP018137.1 | ST556 CP003357.2 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Start | End | Start | End | Start | End | Start | End | Start | End | Start | End | Start | End | Start | End | Start | End | |
| Locus | 456,041 | 466,242 | 477,425 | 487,635 | 535,782 | 545,984 | 468,070 | 478,291 | 483,927 | 494,129 | 563,225 | 573,421 | 732,691 | 742,894 | 467,880 | 478,083 | 546,433 | 556,636 |
| hsdR | 466,242 | 463,909 | 487,635 | 485,302 | 545,984 | 543,651 | 478,291 | 475,958 | 494,129 | 491,796 | 573,421 | 571,088 | 742,894 | 740,561 | 478,083 | 475,750 | 556,636 | 554,303 |
| hsdM | 463,896 | 462,433 | 485,289 | 483,826 | 543,638 | 542,175 | 475,945 | 474,455 | 491,783 | 490,320 | 571,075 | 569,612 | 740,548 | 739,085 | 475,737 | 474,274 | 554,290 | 552,827 |
| hsdS | 462,433 | 460,865 | 483,826 | 482,259 | 542,175 | 540,607 | 474,482 | 473,820 | 490,320 | 488,752 | 569,612 | 568,044 | 739,085 | 737,517 | 474,274 | 472,709 | 552,827 | 551,262 |
| creX | 460,808 | 460,011 | 482,202 | 481,405 | 540,550 | 539,753 | 472,032 | 472,829 | 487,391 | 488,188 | 566,683 | 567,480 | 736,156 | 736,953 | 471,345 | 472,142 | 549,898 | 550,695 |
| hsdS’ | 460,000 | 459,401 | 481,295 | 480,789 | 539,742 | 539,134 | 471,413 | 472,021 | 488,259 | 488,798 | 567,491 | 568,096 | 737,024 | 737,569 | 472,462 | 472,761 | 550,706 | 551,314 |
| hsdS’’ | 458,167 | 459,447 | 479,800 | 480,693 | 538,206 | 539,180 | 470,191 | 471,465 | 486,054 | 487,334 | 565,346 | 566,626 | 734,819 | 736,099 | 470,008 | 471,288 | 548,561 | 549,841 |
| Hypothetical | 457,860 | 457,997 | 479,245 | 479,382 | 537,224 | 537,343 | 469,883 | 470,020 | 485,746 | 485,883 | 565,038 | 565,175 | 734,366 | 734,217 | 469,322 | 469,441 | 548,253 | 548,390 |
| Hypothetical | 457,483 | 457,602 | 478,951 | 479,100 | 537,601 | 537,738 | 469,506 | 469,625 | 485,488 | 485,369 | 564,780 | 564,661 | 734,133 | 734,252 | 469,406 | 469,555 | 547,959 | 548,108 |
| glnA | 456,041 | 457,387 | 477,425 | 478,771 | 535,782 | 537,128 | 468,070 | 469,416 | 483,927 | 485,273 | 563,225 | 564,571 | 732,691 | 734,037 | 467,880 | 469,226 | 546,433 | 547,779 |
| TRD 1.1 | 461,738 | 462,157 | 483,133 | 483,550 | 541,483 | 541,899 | 473,762 | 474,179 | 489,628 | 490,044 | 568,920 | 569,336 | 738,391 | 738,809 | 473,582 | 473,998 | 552,135 | 552,551 |
| TRD 1.2 | 458,173 | 458,577 | 479,559 | 479,967 | 537,915 | 538,319 | 470,197 | 470,601 | 486,060 | 486,464 | 565,352 | 565,756 | 734,825 | 735,229 | 470,014 | 470,418 | 548,567 | 548,971 |
| TRD 2.1 | 460,865 | 461,425 | 482,259 | 482,819 | 540,607 | 541,167 | 472,886 | 473,446 | 486,774 | 487,334 | 566,066 | 566,626 | 735,539 | 736,099 | 470,728 | 471,288 | 549,281 | 549,841 |
| TRD 2.2 | 458,887 | 459,447 | 480,277 | 480,841 | 539,134 | 539,694 | 471,413 | 471,973 | 488,752 | 489,312 | 568,044 | 568,604 | 737,517 | 738,077 | 472,201 | 472,761 | 550,754 | 551,314 |
| TRD 2.3 | 459,399 | 459,952 | 480,795 | 481,346 | 538,605 | 539,180 | 470,911 | 471,459 | 488,247 | 488,798 | 567,539 | 568,090 | 737,012 | 737,563 | 472,715 | 473,266 | 551,268 | 551,819 |
| S. pneumoniae Strain | n SPN994039 | SPN034156 | NT_110_58 | SWU02 a | A026 | Taiwan19F-14 | G54 | CGSP14 | 70585 | |||||||||
| GenBank | FQ312044.2 | FQ312 | 045.1 | CP007593.1 | CP018347.1 | CP006844.1 | CP000921.1 | CP001015.1 | CP001033.1 | CP000918.1 | ||||||||
| start | End | start | End | start | End | start | End | start | End | start | End | start | End | start | End | start | End | |
| Locus | 457,352 | 467,573 | 1,574,115 | 1,584,308 | 469,994 | 480,196 | 900,998 | 911,194 | 467,754 | 476,981 | 507,439 | 516,169 | 450,495 | 459,235 | 476,083 | 484,809 | 511,799 | 520,522 |
| hsdR | 467,573 | 465,240 | 1,584,308 | 1,581,975 | 480,196 | 477,863 | 911,194 | 908,861 | 476,981 | 474,648 | 516,169 | 513,836 | 459,235 | 456,902 | 484,809 | 482,476 | 520,522 | 518,189 |
| hsdM | 465,227 | 463,737 | 1,581,962 | 1,580,499 | 477,850 | 476,387 | 908,848 | 907,385 | 474,635 | 473,172 | 513,823 | 512,360 | 456,889 | 455,426 | 482,463 | 481,000 | 518,176 | 516,713 |
| hsdS | 463,764 | 462,174 | 1,580,499 | 1,578,934 | 474,840 | 476,387 | 907,385 | 905,852 | 473,172 | 471,594 | 512,360 | 510,795 | 455,426 | 454,332 | 481,000 | 479,450 | 516,713 | 515,163 |
| creX | 460,810 | 461,607 | 1,577,573 | 1,578,370 | 473,476 | 474,273 | 904,481 | 905,278 | 471,537 | 470,740 | Absent | Absent | Absent | Absent | Absent | Absent | Absent | Absent |
| hsdS’ | 461,594 | 462226 | 1,578,934 | 1,578,986 | Absent | Absent | 905,337 | 905,895 | Absent | Absent | Absent | Absent | Absent | Absent | Absent | Absent | Absent | Absent |
| hsdS’’ | 459,473 | 460,753 | 1,576,236 | 1,577,516 | 472,136 | 473,419 | 903,047 | 904,424 | 469,882 | 470,685 | 509,567 | 510,847 | 452,623 | 453,921 | 478,210 | 479,502 | 513,926 | 514,345 |
| Hypothetical | 459,165 | 459,302 | 1,575,928 | 1,576,065 | 471,813 | 471,950 | 902,014 | 902,151 | 469,574 | 469,711 | 508,965 | 509,114 | 451,937 | 452,056 | 477,902 | 478,039 | 513,319 | 513,468 |
| Hypothetical | 458,788 | 458,907 | 1,575,551 | 1,575,670 | 471,520 | 471,669 | 901,637 | 901,756 | 469,220 | 469,429 | 508,881 | 509,000 | 452,021 | 452,170 | 477,524 | 477,643 | 513,235 | 513,354 |
| glnA | 457,352 | 458,698 | 1,574,115 | 1,575,461 | 469,994 | 471,340 | 900,998 | 901,541 | 467,754 | 469,100 | 507,439 | 508,785 | 450,495 | 451,841 | 476,083 | 477,429 | 511,799 | 513,145 |
| TRD 1.1 | 463,041 | 463,461 | 1,579,807 | 1,580,223 | 472,127 | 472,543 | 903,132 | 903,548 | 472,480 | 472,896 | 511,668 | 512,084 | 454,734 | 455,150 | 478,215 | 478,631 | 513,926 | 514,345 |
| TRD 1.1’ | Absent | Absent | Absent | Absent | Absent | Absent | Absent | Absent | Absent | Absent | Absent | Absent | 452,629 | 453,045 | Absent | Absent | Absent | Absent |
| TRD 1.2 | 459,479 | 459,883 | 1,576,242 | 1,576,646 | 475,707 | 476,111 | 906,528 | 907,110 | 469,888 | 470,291 | 509,573 | 509,977 | Absent | Absent | 480,319 | 480,723 | 516,033 | 516,437 |
| TRD 2.1 | 460,193 | 460,753 | 1,576,956 | 1,577,516 | 472,859 | 473,419 | 903,864 | 904,424 | 471,594 | 472,153 | Absent | Absent | Absent | Absent | Absent | Absent | Absent | Absent |
| TRD 2.2 | 461,666 | 462,226 | 1,578,429 | 1,578,986 | 474,332 | 474,892 | 905,337 | 905,895 | Absent | Absent | 510,287 | 510,847 | 453,361 | 453,921 | 479,449 | 480,009 | 515,164 | 515,723 |
| TRD 2.3 | 462,180 | 462,728 | 1,578,940 | 1,579,491 | 474,846 | 475,397 | 905,852 | 906,401 | Absent | Absent | 510,801 | 511,352 | 453,875 | 454,418 | 478,947 | 479,499 | 514,658 | 515,211 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oliver, M.B.; Swords, W.E. Comparative Analysis of Streptococcus pneumoniae Type I Restriction-Modification Loci: Variation in hsdS Gene Target Recognition Domains. Pathogens 2020, 9, 712. https://doi.org/10.3390/pathogens9090712
Oliver MB, Swords WE. Comparative Analysis of Streptococcus pneumoniae Type I Restriction-Modification Loci: Variation in hsdS Gene Target Recognition Domains. Pathogens. 2020; 9(9):712. https://doi.org/10.3390/pathogens9090712
Chicago/Turabian StyleOliver, Melissa B., and W. Edward Swords. 2020. "Comparative Analysis of Streptococcus pneumoniae Type I Restriction-Modification Loci: Variation in hsdS Gene Target Recognition Domains" Pathogens 9, no. 9: 712. https://doi.org/10.3390/pathogens9090712
APA StyleOliver, M. B., & Swords, W. E. (2020). Comparative Analysis of Streptococcus pneumoniae Type I Restriction-Modification Loci: Variation in hsdS Gene Target Recognition Domains. Pathogens, 9(9), 712. https://doi.org/10.3390/pathogens9090712




