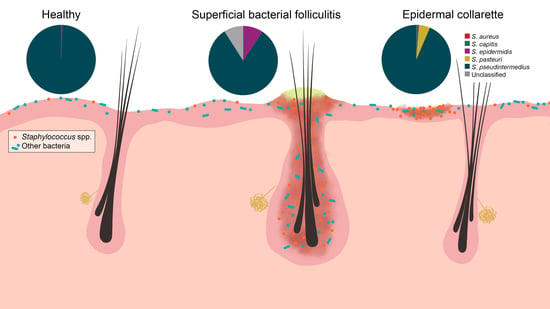
Characterization of Cutaneous Bacterial Microbiota from Superficial Pyoderma Forms in Atopic Dogs
Abstract
1. Introduction
2. Results
3. Discussion
4. Materials and Methods
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bloom, P. Canine superficial bacterial folliculitis: Current understanding of its etiology, diagnosis and treatment. Vet. J. 2014, 199, 217–222. [Google Scholar] [CrossRef]
- Hill, P.B.; Lo, A.; Eden, C.A.N.; Huntley, S.; Morey, V.; Ramsey, S.; Richardson, C.; Smith, D.J.; Sutton, C.; Taylor, M.D.; et al. Survey of the prevalence, diagnosis and treatment of dermatological conditions in small animals in general practice. Vet. Rec. 2006, 158, 533–539. [Google Scholar] [CrossRef]
- Lund, E.M.; Armstrong, P.J.; Kirk, C.A.; Kolar, L.M.; Klausner, J.S. Health status and population characteristics of dogs and cats examined at private veterinary practices in the United States. J. Am. Vet. Med. Assoc. 1999, 214, 1336–1341. [Google Scholar]
- Mason, I.S. Canine pyoderma. J. Small Anim. Pr. 1991, 32, 381–386. [Google Scholar] [CrossRef]
- Miller, W.H.; Griffin, C.E.; Campbell, K.L. Muller & Kirk’s Small Animal Dermatology, 7th ed.; Elsevier: St. Louis, MO, USA, 2013. [Google Scholar]
- Gross, T.L.; Ihrke, P.J.; Walder, E.J.; Affolter, V.K. Pustular Diseases of the Epidermis in SKIN Diseases of the Dog and Cat: Clinical and Histopathologic Diagnosis, 2nd ed.; Blackwell Science Ltd.: Oxford, UK, 2005; Volume 2, pp. 4–26. [Google Scholar]
- Banovic, F.; Linder, K.; Olivry, T. Clinical, microscopic and microbial characterization of exfoliative superficial pyoderma-associated epidermal collarettes in dogs. Vet. Dermatol. 2016, 28, 107-e123. [Google Scholar] [CrossRef] [PubMed]
- Fazakerley, J.; Nuttall, T.; Sales, D.; Schmidt, V.; Carter, S.; Hart, C.A.; McEwan, N.A. Staphylococcal colonization of mucosal and lesional skin sites in atopic and healthy dogs. Vet. Dermatol. 2009, 20, 179–184. [Google Scholar] [CrossRef] [PubMed]
- Bannoehr, J.; Guardabassi, L. Staphylococcus pseudintermedius in the dog: Taxonomy, diagnostics, ecology, epidemiology and pathogenicity. Vet. Dermatol. 2012, 23, 253–266, e251–252. [Google Scholar] [CrossRef] [PubMed]
- Frank, L.A.; Kania, S.A.; Hnilica, K.A.; Wilkes, R.P.; Bemis, D.A. Isolation of Staphylococcus schleiferi from dogs with pyoderma. J. Am. Vet. Med. Assoc. 2003, 222, 451–454. [Google Scholar] [CrossRef] [PubMed]
- May, E.R.; Hnilica, K.A.; Frank, L.A.; Jones, R.D.; Bemis, D.A. Isolation of Staphylococcus schleiferi from healthy dogs and dogs with otitis, pyoderma, or both. J. Am. Vet. Med. Assoc. 2005, 227, 928–931. [Google Scholar] [CrossRef] [PubMed]
- Ravens, P.; Vogelnest, L.; Ewen, E.; Bosward, K.; Norris, J.; Norris, J. Canine superficial bacterial pyoderma: Evaluation of skin surface sampling methods and antimicrobial susceptibility of causalStaphylococcusisolates. Aust. Vet. J. 2014, 92, 149–155. [Google Scholar] [CrossRef]
- Bean, D.C.; Wigmore, S. Carriage rate and antibiotic susceptibility of coagulase-positive staphylococci isolated from healthy dogs in Victoria, Australia. Aust. Vet. J. 2016, 94, 456–460. [Google Scholar] [CrossRef] [PubMed]
- Bradley, C.W.; Morris, D.O.; Rankin, S.C.; Cain, C.L.; Misic, A.M.; Houser, T.; Mauldin, E.A.; Grice, E.A. Longitudinal Evaluation of the Skin Microbiome and Association with Microenvironment and Treatment in Canine Atopic Dermatitis. J. Investig. Dermatol. 2016, 136, 1182–1190. [Google Scholar] [CrossRef]
- Pierezan, F.; Olivry, T.; Paps, J.S.; Lawhon, S.D.; Wu, J.; Steiner, J.M.; Suchodolski, J.S.; Rodrigues-Hoffman, A. The skin microbiome in allergen-induced canine atopic dermatitis. Vet. Dermatol. 2016, 27, 332-e82. [Google Scholar] [CrossRef] [PubMed]
- Lozupone, C.; Knight, R. UniFrac: A New Phylogenetic Method for Comparing Microbial Communities. Appl. Environ. Microbiol. 2005, 71, 8228–8235. [Google Scholar] [CrossRef]
- Byrd, A.L.; Belkaid, Y.; Segre, J.A. The human skin microbiome. Nat. Rev. Genet. 2018, 16, 143–155. [Google Scholar] [CrossRef]
- Kobayashi, T.; Glatz, M.; Horiuchi, K.; Kawasaki, H.; Akiyama, H.; Kaplan, D.H.; Kong, H.H.; Amagai, M.; Nagao, K. Dysbiosis and Staphylococcus aureus Colonization Drives Inflammation in Atopic Dermatitis. Immunity 2015, 42, 756–766. [Google Scholar] [CrossRef] [PubMed]
- Brugger, S.D.; Bomar, L.; Lemon, K.P. Commensal–Pathogen Interactions along the Human Nasal Passages. PLoS. Pathog. 2016, 12, e1005633. [Google Scholar] [CrossRef]
- Kong, H.H.; Oh, J.; Deming, C.; Conlan, S.; Grice, E.A.; Beatson, M.A.; Nomicos, E.; Polley, E.C.; Komarow, H.D.; Murray, P.R.; et al. Temporal shifts in the skin microbiome associated with disease flares and treatment in children with atopic dermatitis. Genome. Res. 2012, 22, 850–859. [Google Scholar] [CrossRef]
- Rodrigues-Hoffman, A.; Patterson, A.P.; Diesel, A.; Lawhon, S.D.; Ly, H.J.; Stephenson, C.E.; Mansell, J.; Steiner, J.M.; Dowd, S.E.; Olivry, T.; et al. The Skin Microbiome in Healthy and Allergic Dogs. PLoS ONE 2014, 9, e83197. [Google Scholar] [CrossRef]
- Torres, S.; Clayton, J.B.; Danzeisen, J.L.; Ward, T.; Huang, H.; Knights, D.; Johnson, T.J. Diverse bacterial communities exist on canine skin and are impacted by cohabitation and time. PeerJ 2017, 5, e3075. [Google Scholar] [CrossRef]
- Cuscó, A.; Sánchez, A.; Altet, L.; Ferrer, L.; Francino, O. Individual Signatures Define Canine Skin Microbiota Composition and Variability. Front. Vet. Sci. 2017, 4, 6. [Google Scholar] [CrossRef]
- Webster, G.F.; Ruggieri, M.R.; McGinley, K.J. Correlation of Propionibacterium acnes Populations with the Presence of Triglycerides on Nonhuman Skin. Appl. Environ. Microbiol. 1981, 41, 1269–1270. [Google Scholar] [CrossRef] [PubMed]
- Leyden, J.J.; Marples, R.R.; Kligman, A.M.; And, R.R.M. Staphylococcus aureus in the lesions of atopic dermatitis. Br. J. Dermatol. 1974, 90, 525. [Google Scholar] [CrossRef] [PubMed]
- Nakatsuji, T.; Chen, T.H.; Narala, S.; Chun, K.A.; Two, A.M.; Tong, Y.; Shafiq, F.; Kotol, P.F.; Bouslimani, A.; Melnik, A.V.; et al. Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Sci. Transl. Med. 2017, 9, eaah4680. [Google Scholar] [CrossRef] [PubMed]
- Iwase, T.; Uehara, Y.; Shinji, H.; Tajima, A.; Seo, H.; Takada, K.; Agata, T.; Mizunoe, Y. Staphylococcus epidermidis Esp inhibits Staphylococcus aureus biofilm formation and nasal colonization. Nature 2010, 465, 346–349. [Google Scholar] [CrossRef]
- Sugimoto, S.; Iwamoto, T.; Takada, K.; Okuda, K.-I.; Tajima, A.; Iwase, T.; Mizunoe, Y. Staphylococcus epidermidis Esp Degrades Specific Proteins Associated with Staphylococcus aureus Biofilm Formation and Host-Pathogen Interaction. J. Bacteriol. 2013, 195, 1645–1655. [Google Scholar] [CrossRef]
- Zipperer, A.; Konnerth, M.C.; Laux, C.; Berscheid, A.; Janek, D.; Weidenmaier, C.; Burian, M.; Schilling, N.A.; Slavetinsky, C.; Marschal, M.; et al. Human commensals producing a novel antibiotic impair pathogen colonization. Nature 2016, 535, 511–516. [Google Scholar] [CrossRef]
- Myles, I.; Earland, N.J.; Anderson, E.D.; Moore, I.N.; Kieh, M.D.; Williams, K.W.; Saleem, A.; Fontecilla, N.M.; Welch, P.A.; Darnell, D.A.; et al. First-in-human topical microbiome transplantation with Roseomonas mucosa for atopic dermatitis. JCI Insight 2018, 3, 3. [Google Scholar] [CrossRef]
- Ramsey, M.M.; Freire, M.; Gabrilska, R.A.; Rumbaugh, K.P.; Lemon, K.P. Staphylococcus aureus Shifts toward Commensalism in Response to Corynebacterium Species. Front. Microbiol. 2016, 7, 1230. [Google Scholar] [CrossRef]
- Conlan, S.; Kong, H.H.; Segre, J.A. Species-Level Analysis of DNA Sequence Data from the NIH Human Microbiome Project. PLoS ONE 2012, 7, e47075. [Google Scholar] [CrossRef]
- Meason-Smith, C.; Older, C.E.; Ocana, R.; Dominguez, B.; Lawhon, S.D.; Wu, J.; Patterson, A.P.; Rodrigues Hoffmann, A. Novel association of Psychrobacter and Pseudomonas with malodor in Bloodhound dogs and effects of a topical product composed of essential oils and plant-derived essential fatty acids in a randomized, blinded, placebo-controlled study. Vet. Dermatol. 2018, 29, 465-e158. [Google Scholar] [CrossRef] [PubMed]
- Bannoehr, J.; Ben Zakour, N.L.; Waller, A.; Guardabassi, L.; Thoday, K.L.; Broek, A.H.M.V.D.; Fitzgerald, J.R. Population Genetic Structure of the Staphylococcus intermedius Group: Insights into agr Diversification and the Emergence of Methicillin-Resistant Strains. J. Bacteriol. 2007, 189, 8685–8692. [Google Scholar] [CrossRef]
- Sasaki, T.; Tsubakishita, S.; Tanaka, Y.; Sakusabe, A.; Ohtsuka, M.; Hirotaki, S.; Kawakami, T.; Fukata, T.; Hiramatsu, K. Multiplex-PCR Method for Species Identification of Coagulase-Positive Staphylococci. J. Clin. Microbiol. 2010, 48, 765–769. [Google Scholar] [CrossRef] [PubMed]
- Misic, A.M.; Davis, M.F.; Tyldsley, A.S.; Hodkinson, B.P.; Tolomeo, P.; Hu, B.; Nachamkin, I.; Lautenbach, E.; Morris, D.O.; Grice, E.A. The shared microbiota of humans and companion animals as evaluated from Staphylococcus carriage sites. Microbiome 2015, 3, 2. [Google Scholar] [CrossRef] [PubMed]
- Salter, S.J.; Cox, M.J.; Turek, E.; Calus, S.; Cookson, W.O.C.; Moffatt, M.F.; Turner, P.; Parkhill, J.; Loman, N.J.; Walker, A. Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol. 2014, 12, 87. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; I Gordon, J.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef]
- Rognes, T.; Flouri, T.; Nichols, B.; Quince, C.; Mahé, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ 2016, 4, e2584. [Google Scholar] [CrossRef]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef]
- McDonald, D.; Price, M.N.; Goodrich, J.; Nawrocki, E.P.; DeSantis, T.Z.; Probst, A.J.; Andersen, G.L.; Knight, R.; Hugenholtz, P. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 2011, 6, 610–618. [Google Scholar] [CrossRef]
- Abraham, A.; Pedregosa, F.; Eickenberg, M.; Gervais, P.; Mueller, A.; Kossaifi, J.; Gramfort, A.; Thirion, B.; Varoquaux, G. Machine learning for neuroimaging with scikit-learn. Front. Aging Neurosci. 2014, 8, 14. [Google Scholar] [CrossRef]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.S.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef] [PubMed]
- Martineau, F.; Picard, F.J.; Ke, D.; Paradis, S.; Roy, P.H.; Ouellette, M.; Bergeron, M.G. Development of a PCR Assay for Identification of Staphylococci at Genus and Species Levels. J. Clin. Microbiol. 2001, 39, 2541–2547. [Google Scholar] [CrossRef] [PubMed]
- Segata, N.; Izard, J.; Waldron, L.; Gevers, D.; Miropolsky, L.; Garrett, W.S.; Huttenhower, C. Metagenomic biomarker discovery and explanation. Genome Biol. 2011, 12, R60. [Google Scholar] [CrossRef] [PubMed]
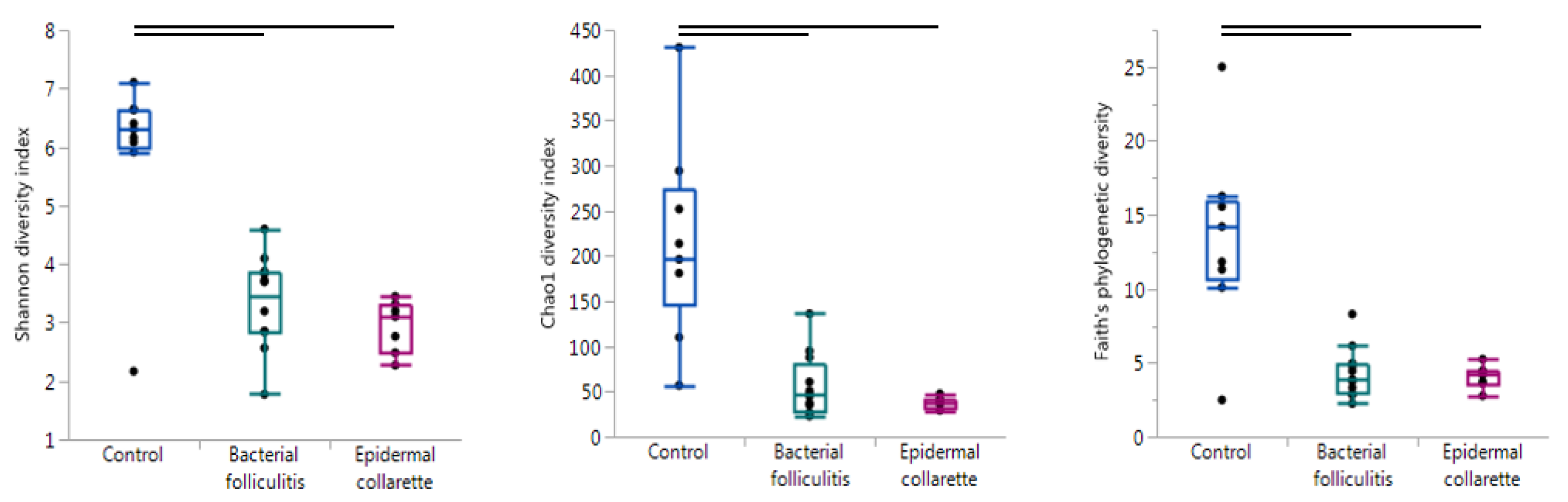
| Control vs. Pyoderma | Control vs. EC | Control vs. SBF | EC vs. SBF | |
|---|---|---|---|---|
| Alpha diversity | ||||
| Chao1 diversity index | <0.001 | 0.001 | <0.001 | 0.337 |
| Faith’s phylogenetic diversity | <0.001 | 0.009 | 0.008 | 0.915 |
| Observed OTUs | <0.001 | 0.007 | 0.003 | 1.000 |
| Pielou’s evenness | <0.001 | 0.007 | 0.012 | 0.145 |
| Shannon diversity index | <0.001 | 0.001 | <0.001 | 0.337 |
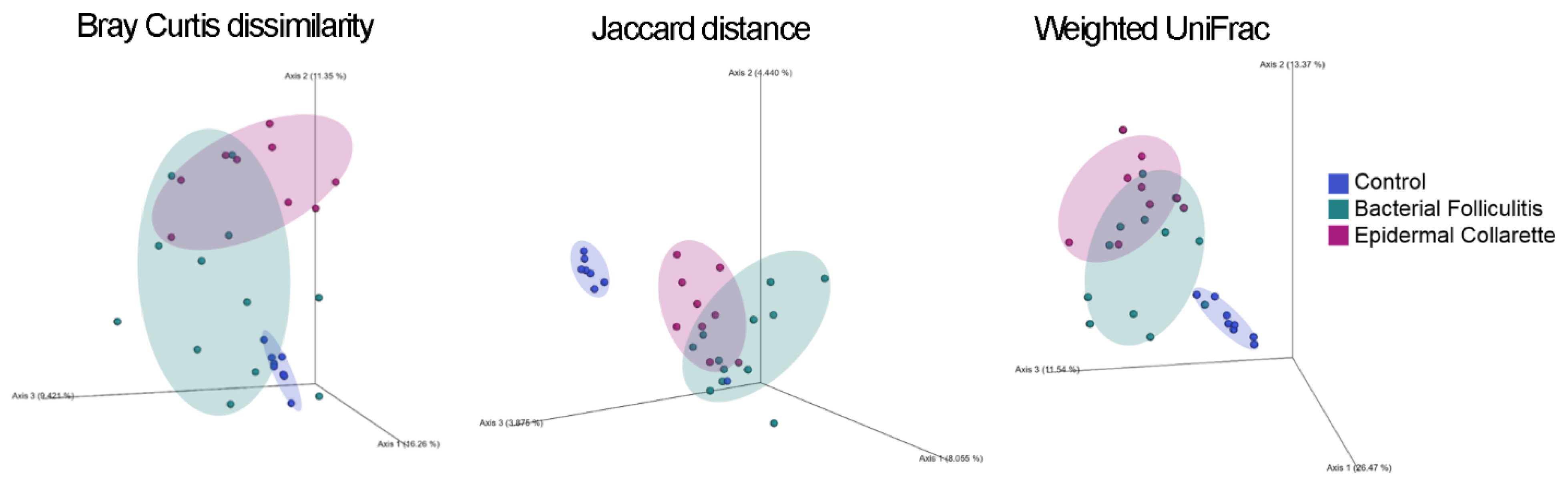
| Beta diversity | ||||
| Bray-Curtis | R = 0.908, p = 0.001 | R = 1.000, p = 0.002 | R = 0.923, p = 0.001 | R = 0.168, p = 0.030 |
| Jaccard | R = 0.923, p = 0.001 | R = 0.959, p = 0.001 | R = 0.983, p = 0.001 | R = 0.154, p = 0.023 |
| Unweighted UniFrac | R = 0.956, p = 0.001 | R = 0.974, p = 0.001 | R = 0.966, p = 0.001 | R = 0.023, p = 0.321 |
| Weighted UniFrac | R = 0.866, p = 0.001 | R = 1.000, p = 0.001 | R = 0.854, p = 0.001 | R = 0.135, p = 0.055 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Older, C.E.; Rodrigues Hoffmann, A.; Hoover, K.; Banovic, F. Characterization of Cutaneous Bacterial Microbiota from Superficial Pyoderma Forms in Atopic Dogs. Pathogens 2020, 9, 638. https://doi.org/10.3390/pathogens9080638
Older CE, Rodrigues Hoffmann A, Hoover K, Banovic F. Characterization of Cutaneous Bacterial Microbiota from Superficial Pyoderma Forms in Atopic Dogs. Pathogens. 2020; 9(8):638. https://doi.org/10.3390/pathogens9080638
Chicago/Turabian StyleOlder, Caitlin E., Aline Rodrigues Hoffmann, Kathleen Hoover, and Frane Banovic. 2020. "Characterization of Cutaneous Bacterial Microbiota from Superficial Pyoderma Forms in Atopic Dogs" Pathogens 9, no. 8: 638. https://doi.org/10.3390/pathogens9080638
APA StyleOlder, C. E., Rodrigues Hoffmann, A., Hoover, K., & Banovic, F. (2020). Characterization of Cutaneous Bacterial Microbiota from Superficial Pyoderma Forms in Atopic Dogs. Pathogens, 9(8), 638. https://doi.org/10.3390/pathogens9080638