An Unbiased Approach to Mapping the Signaling Network of the Pseudorabies Virus US3 Protein
Abstract
1. Introduction
2. Results
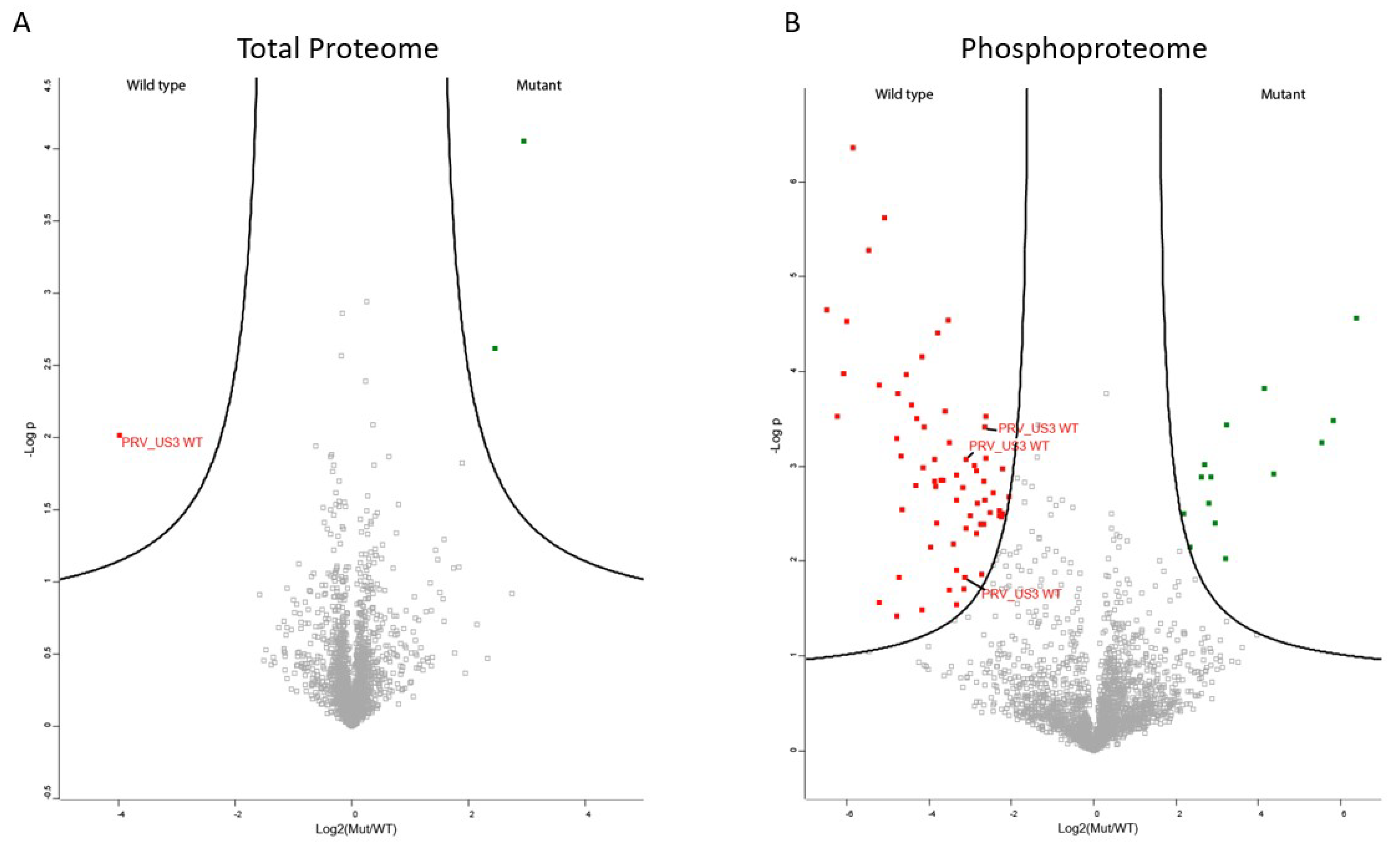
2.1. The Phosphoproteome of US3-Transfected Cells
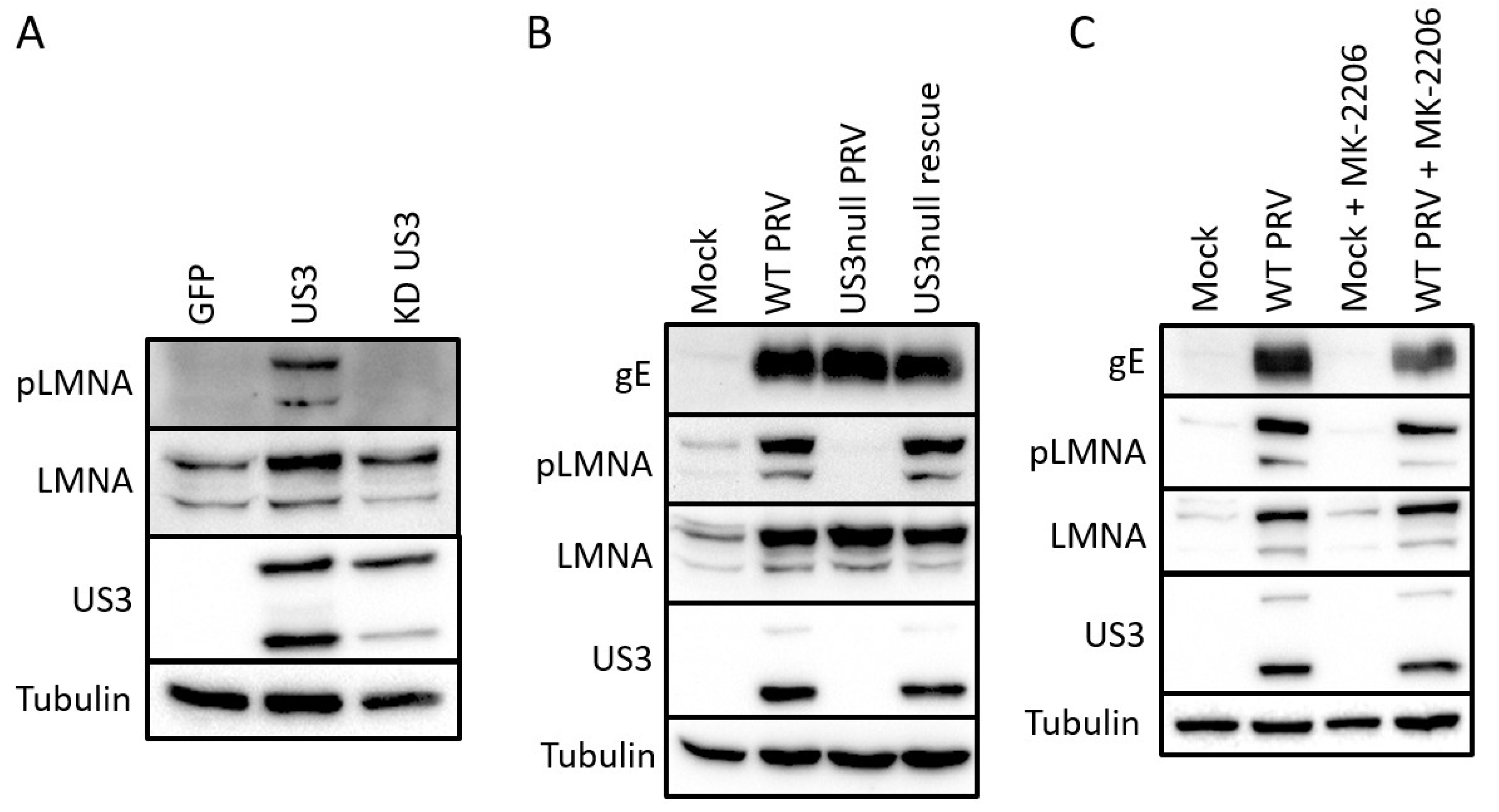
2.2. Validation of Identified S404 Phosphosite in Lamin A/C
2.3. Gene Enrichment Analysis
3. Discussion
4. Materials and Methods
4.1. Cells, Inhibitors and Viruses
4.2. Transfection
4.3. Flow Cytometry
4.4. Mass Spectrometry Sample Preparation
4.5. LC-MS/MS Analysis
4.6. Data Analysis
4.7. Western Blotting
4.8. Gene Ontology Enrichment Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Jacob, T.; Broeke, C.V.D.; Favoreel, H.W. Viral Serine/Threonine Protein Kinases. J. Virol. 2010, 85, 1158–1173. [Google Scholar] [CrossRef] [PubMed]
- Deruelle, M.J.; Favoreel, H.W. Keep it in the subfamily: The conserved alphaherpesvirus US3 protein kinase. J. Gen. Virol. 2011, 92, 18–30. [Google Scholar] [CrossRef] [PubMed]
- Kimman, T.G.; De Wind, N.; De Bruin, T.; De Visser, Y.; Voermans, J. Inactivation of Glycoprotein gE and Thymidine Kinase or the US3-Encoded Protein Kinase Synergistically Decreases In Vivo Replication of Pseudorabies Virus and the Induction of Protective Immunity. Virology 1994, 205, 511–518. [Google Scholar] [CrossRef] [PubMed]
- Olsen, L.M.; Ch’Ng, T.H.; Card, J.P.; Enquist, L.W. Role of Pseudorabies Virus Us3 Protein Kinase during Neuronal Infection. J. Virol. 2006, 80, 6387–6398. [Google Scholar] [CrossRef] [PubMed]
- De Wind, N.; Domen, J.; Berns, A. Herpesviruses encode an unusual protein-serine/threonine kinase which is nonessential for growth in cultured cells. J. Virol. 1992, 66, 5200–5209. [Google Scholar] [CrossRef]
- Koyanagi, N.; Kato, A.; Takeshima, K.; Maruzuru, Y.; Kozuka-Hata, H.; Oyama, M.; Arii, J.; Kawaguchi, Y. Regulation of Herpes Simplex Virus 2 Protein Kinase UL13 by Phosphorylation and Its Role in Viral Pathogenesis. J. Virol. 2018, 92, 1–19. [Google Scholar] [CrossRef]
- Favoreel, H.W.; van Minnebruggen, G.; Adriaensen, D.; Nauwynck, H.J. Cytoskeletal rearrangements and cell extensions induced by the US3 kinase of an alphaherpesvirus are associated with enhanced spread. Proc. Natl. Acad. Sci. USA 2005, 102, 8990–8995. [Google Scholar] [CrossRef]
- Deruelle, M.; Geenen, K.; Nauwynck, H.J.; Favoreel, H.W. A point mutation in the putative ATP binding site of the pseudorabies virus US3 protein kinase prevents Bad phosphorylation and cell survival following apoptosis induction. Virus Res. 2007, 128, 65–70. [Google Scholar] [CrossRef][Green Version]
- Kato, A.; Kawaguchi, Y. Us3 Protein Kinase Encoded by HSV: The Precise Function and Mechanism on Viral Life Cycle. Adv. Exp. Med. Biol. 2018, 1045, 45–62. [Google Scholar] [CrossRef]
- Wang, S.; Wang, K.; Lin, R.; Zheng, C. Herpes Simplex Virus 1 Serine/Threonine Kinase US3 Hyperphosphorylates IRF3 and Inhibits Beta Interferon Production. J. Virol. 2013, 87, 12814–12827. [Google Scholar] [CrossRef]
- Mou, F.; Forest, T.; Baines, J.D. US3 of Herpes Simplex Virus Type 1 Encodes a Promiscuous Protein Kinase That Phosphorylates and Alters Localization of Lamin A/C in Infected Cells. J. Virol. 2007, 81, 6459–6470. [Google Scholar] [CrossRef] [PubMed]
- Chuluunbaatar, U.; Roller, R.; Feldman, M.E.; Brown, S.; Shokat, K.M.; Mohr, I. Constitutive mTORC1 activation by a herpesvirus Akt surrogate stimulates mRNA translation and viral replication. Genes Dev. 2010, 24, 2627–2639. [Google Scholar] [CrossRef]
- Cartier, A.; Komai, T.; Masucci, M.G. The Us3 protein kinase of herpes simplex virus 1 blocks apoptosis and induces phosporylation of the Bcl-2 family member Bad. Exp. Cell Res. 2003, 291, 242–250. [Google Scholar] [CrossRef]
- Kato, A.; Yamamoto, M.; Ohno, T.; Kodaira, H.; Nishiyama, Y.; Kawaguchi, Y. Identification of Proteins Phosphorylated Directly by the Us3 Protein Kinase Encoded by Herpes Simplex Virus 1. J. Virol. 2005, 79, 9325–9331. [Google Scholar] [CrossRef]
- Xiong, R.; Rao, P.; Kim, S.; Li, M.; Wen, X.; Yuan, W. Herpes Simplex Virus 1 US3 Phosphorylates Cellular KIF3A To Downregulate CD1d Expression. J. Virol. 2015, 89, 6646–6655. [Google Scholar] [CrossRef] [PubMed]
- Benetti, L.; Roizman, B. Herpes simplex virus protein kinase US3 activates and functionally overlaps protein kinase A to block apoptosis. Proc. Natl. Acad. Sci. USA 2004, 101, 9411–9416. [Google Scholar] [CrossRef] [PubMed]
- Rubio, R.M.; Mohr, I. Inhibition of ULK1 and Beclin1 by an α-herpesvirus Akt-like Ser/Thr kinase limits autophagy to stimulate virus replication. Proc. Natl. Acad. Sci. USA 2019, 116, 26941–26950. [Google Scholar] [CrossRef]
- Broeke, C.V.D.; Radu, M.; Deruelle, M.; Nauwynck, H.; Hofmann, C.; Jaffer, Z.M.; Chernoff, J.; Favoreel, H.W. Alphaherpesvirus US3-mediated reorganization of the actin cytoskeleton is mediated by group A p21-activated kinases. Proc. Natl. Acad. Sci. USA 2009, 106, 8707–8712. [Google Scholar] [CrossRef]
- Benetti, L.; Roizman, B. Protein kinase B/Akt is present in activated form throughout the entire replicative cycle of deltaU(S)3 mutant virus but only at early times after infection with wild-type herpes simplex virus 1. J. Virol. 2006, 80, 3341–3348. [Google Scholar] [CrossRef]
- Jacob, T.; Broeke, C.V.D.; Van Waesberghe, C.; Van Troys, L.; Favoreel, H.W. Pseudorabies virus US3 triggers RhoA phosphorylation to reorganize the actin cytoskeleton. J. Gen. Virol. 2015, 96, 2328–2335. [Google Scholar] [CrossRef]
- Yang, S.; Pei, Y.; Zhao, A. iTRAQ-based Proteomic Analysis of Porcine Kidney Epithelial PK15 cells Infected with Pseudorabies virus. Sci. Rep. 2017, 7, 45922. [Google Scholar] [CrossRef]
- Soh, T.K.; Davies, C.T.; Muenzner, J.; Hunter, L.M.; Barrow, H.G.; Connor, V.; Bouton, C.R.; Smith, C.; Emmott, E.; Antrobus, R.; et al. Temporal Proteomic Analysis of Herpes Simplex Virus 1 Infection Reveals Cell-Surface Remodeling via pUL56-Mediated GOPC Degradation. Cell Rep. 2020, 33, 108235. [Google Scholar] [CrossRef]
- Kulej, K.; Avgousti, D.C.; Sidoli, S.; Herrmann, C.; Della Fera, A.N.; Kim, E.T.; Garcia, B.A.; Weitzman, M.D. Time-resolved Global and Chromatin Proteomics during Herpes Simplex Virus Type 1 (HSV-1) Infection. Mol. Cell. Proteom. 2017, 16, S92–S107. [Google Scholar] [CrossRef]
- Magalhães-Junior, M.J.; Baracat-Pereira, M.C.; Pereira, L.K.J.; Vital, C.E.; Santos, M.R.; Cunha, P.S.; Fernandes, K.M.; Bressan, G.C.; Fietto, J.L.R.; Silva-Júnior, A.; et al. Proteomic and phosphoproteomic analyses reveal several events involved in the early stages of bovine herpesvirus 1 infection. Arch. Virol. 2019, 165, 69–85. [Google Scholar] [CrossRef]
- Geenen, K.; Favoreel, H.W.; Olsen, L.; Enquist, L.W.; Nauwynck, H.J. The pseudorabies virus US3 protein kinase possesses anti-apoptotic activity that protects cells from apoptosis during infection and after treatment with sorbitol or staurosporine. Virolology 2005, 331, 144–150. [Google Scholar] [CrossRef]
- Jansens, R.J.J.; Broeck, W.V.D.; De Pelsmaeker, S.; Lamote, J.A.S.; Van Waesberghe, C.; Couck, L.; Favoreel, H.W. Pseudorabies Virus US3-Induced Tunneling Nanotubes Contain Stabilized Microtubules, Interact with Neighboring Cells via Cadherins, and Allow Intercellular Molecular Communication. J. Virol. 2017, 91, e00749-17. [Google Scholar] [CrossRef]
- Cenni, V.; Bertacchini, J.; Beretti, F.; Lattanzi, G.; Bavelloni, A.; Riccio, M.; Ruzzene, M.; Marin, O.; Arrigoni, G.; Parnaik, V.; et al. Lamin A Ser404 Is a Nuclear Target of Akt Phosphorylation in C2C12 Cells. J. Proteome Res. 2008, 7, 4727–4735. [Google Scholar] [CrossRef]
- Bertacchini, J.; Beretti, F.; Cenni, V.; Guida, M.; Gibellini, F.; Mediani, L.; Marin, O.; Maraldi, N.M.; De Pol, A.; Lattanzi, G.; et al. The protein kinase Akt/PKB regulates both prelamin A degradation and Lmna gene expression. FASEB J. 2013, 27, 2145–2155. [Google Scholar] [CrossRef]
- Naeem, A.S.; Zhu, Y.; Di, W.L.; Marmiroli, S.; O’Shaughnessy, R.F.L. AKT1-mediated Lamin A/C degradation is required for nuclear degradation and normal epidermal terminal differentiation. Cell Death Differ. 2015, 22, 2123–2132. [Google Scholar] [CrossRef] [PubMed]
- Chuluunbaatar, U.; Mohr, I. A herpesvirus kinase that masquerades as Akt. Cell Cycle 2011, 10, 2064–2068. [Google Scholar] [CrossRef]
- Reynolds, A.E.; Liang, L.; Baines, J.D. Conformational Changes in the Nuclear Lamina Induced by Herpes Simplex Virus Type 1 Require Genes U L 31 and U L 34. J. Virol. 2004, 78, 5564–5575. [Google Scholar] [CrossRef]
- Leach, N.R.; Roller, R.J. Significance of host cell kinases in herpes simplex virus type 1 egress and lamin-associated protein disassembly from the nuclear lamina. Virology 2010, 406, 127–137. [Google Scholar] [CrossRef]
- Jacob, T.; Broeke, C.V.D.; Van Troys, M.; Waterschoot, D.; Ampe, C.; Favoreel, H.W. Alphaherpesviral US3 Kinase Induces Cofilin Dephosphorylation To Reorganize the Actin Cytoskeleton. J. Virol. 2013, 87, 4121–4126. [Google Scholar] [CrossRef] [PubMed]
- Toshima, J. Cofilin Phosphorylation and Actin Reorganization Activities of Testicular Protein Kinase 2 and Its Predominant Expression in Testicular Sertoli Cells. J. Biol. Chem. 2001, 276, 31449–31458. [Google Scholar] [CrossRef]
- Røsok, Ø.; Pédeutour, F.; Ree, A.H.; Aasheim, H.-C. Identification and Characterization of TESK2, a Novel Member of the LIMK/TESK Family of Protein Kinases, Predominantly Expressed in Testis. Genomics 1999, 61, 44–54. [Google Scholar] [CrossRef] [PubMed]
- Edwards, D.C.; Sanders, L.C.; Bokoch, G.M.; Gill, G.N. Activation of LIM-kinase by Pak1 couples Rac/Cdc42 GTPase signalling to actin cytoskeletal dynamics. Nat. Cell Biol. 1999, 1, 253–259. [Google Scholar] [CrossRef]
- Krugmann, S.; Jordens, I.; Gevaert, K.; Driessens, M.; Vandekerckhove, J.; Hall, A. Cdc42 induces filopodia by promoting the formation of an IRSp53: Mena complex. Curr. Biol. 2001, 11, 1645–1655. [Google Scholar] [CrossRef]
- Loureiro, J.J.; Rubinson, D.A.; Bear, J.E.; Baltus, G.A.; Kwiatkowski, A.V.; Gertler, F.B. Critical Roles of Phosphorylation and Actin Binding Motifs, but Not the Central Proline-rich Region, for Ena/Vasodilator-stimulated Phosphoprotein (VASP) Function during Cell Migration. Mol. Biol. Cell 2002, 13, 2533–2546. [Google Scholar] [CrossRef] [PubMed]
- Lambrechts, A.; Kwiatkowski, A.V.; Lanier, L.M.; Bear, J.E.; Vandekerckhove, J.; Ampe, C.; Gertler, F.B. cAMP-dependent Protein Kinase Phosphorylation of EVL, a Mena/VASP Relative, Regulates Its Interaction with Actin and SH3 Domains. J. Biol. Chem. 2000, 275, 36143–36151. [Google Scholar] [CrossRef]
- Kawauchi, T.; Chihama, K.; Nishimura, Y.V.; Nabeshima, Y.I.; Hoshino, M. MAP1B phosphorylation is differentially regulated by Cdk5/p35, Cdk5/p25, and JNK. Biochem. Biophys. Res. Commun. 2005, 331, 50–55. [Google Scholar] [CrossRef]
- Trivedi, N.; Marsh, P.; Goold, R.G.; Wood-Kaczmar, A.; Gordon-Weeks, P.R. Glycogen synthase kinase-3β phosphorylation of MAP1B at Ser1260 and Thr1265 is spatially restricted to growing axons. J. Cell Sci. 2005, 118, 993–1005. [Google Scholar] [CrossRef]
- Fischer, I.; Romano-Clarke, G. Changes in Microtubule-Associated Protein MAP1B Phosphorylation During Rat Brain Development. J. Neurochem. 1990, 55, 328–333. [Google Scholar] [CrossRef]
- Rontogianni, S.; Iskit, S.; Van Doorn, S.; Peeper, D.S.; Altelaar, A.F.M. Combined EGFR and ROCK Inhibition in Triple-negative Breast Cancer Leads to Cell Death Via Impaired Autophagic Flux. Mol. Cell. Proteom. 2019, 19, 261–277. [Google Scholar] [CrossRef] [PubMed]
- Naghavi, M.H.; Gundersen, G.G.; Walsh, D. Plus-end tracking proteins, CLASPs, and a viral Akt mimic regulate herpesvirus-induced stable microtubule formation and virus spread. Proc. Natl. Acad. Sci. USA 2013, 110, 18268–18273. [Google Scholar] [CrossRef]
- Schaffer, B.E.; Levin, R.S.; Hertz, N.T.; Maures, T.J.; Schoof, M.L.; Hollstein, P.E.; Benayoun, B.A.; Banko, M.R.; Shaw, R.J.; Shokat, K.M.; et al. Identification of AMPK Phosphorylation Sites Reveals a Network of Proteins Involved in Cell Invasion and Facilitates Large-Scale Substrate Prediction. Cell Metab. 2015, 22, 907–921. [Google Scholar] [CrossRef]
- Miranda, L.M.; Carpentier, S.; Platek, A.; Hussain, N.; Gueuning, M.-A.; Vertommen, D.; Ozkan, Y.; Sid, B.; Hue, L.; Courtoy, P.J.; et al. AMP-activated protein kinase induces actin cytoskeleton reorganization in epithelial cells. Biochem. Biophys. Res. Commun. 2010, 396, 656–661. [Google Scholar] [CrossRef]
- Van Minnebruggen, G.; Favoreel, H.W.; Jacobs, L.; Nauwynck, H.J.; Minnebruggen, G.V. Pseudorabies Virus US3 Protein Kinase Mediates Actin Stress Fiber Breakdown. J. Virol. 2003, 77, 9074–9080. [Google Scholar] [CrossRef]
- McEwen, A.E.; Maher, M.T.; Mo, R.; Gottardi, C.J. E-cadherin phosphorylation occurs during its biosynthesis to promote its cell surface stability and adhesion. Mol. Biol. Cell 2014, 25, 2365–2374. [Google Scholar] [CrossRef]
- Lickert, H.; Bauer, A.; Kemler, R.; Stappert, J. Casein Kinase II Phosphorylation of E-cadherin Increases E-cadherin/β-Catenin Interaction and Strengthens Cell-Cell Adhesion. J. Biol. Chem. 2000, 275, 5090–5095. [Google Scholar] [CrossRef]
- Walters, M.S.; Kinchington, P.R.; Banfield, B.W.; Silverstein, S. Hyperphosphorylation of Histone Deacetylase 2 by Alphaherpesvirus US3 Kinases. J. Virol. 2010, 84, 9666–9676. [Google Scholar] [CrossRef]
- Poon, A.P.W.; Gu, H.; Roizman, B. ICP0 and the US3 protein kinase of herpes simplex virus 1 independently block histone deacetylation to enable gene expression. Proc. Natl. Acad. Sci. USA 2006, 103, 9993–9998. [Google Scholar] [CrossRef]
- De Wind, N.; Zijderveld, A.; Glazenburg, K.; Gielkens, A.; Berns, A. Linker insertion mutagenesis of herpesviruses: Inactivation of single genes within the Us region of pseudorabies virus. J. Virol. 1990, 64, 4691–4696. [Google Scholar] [CrossRef]
- Kimman, T.G.; De Wind, N.; Oei-Lie, N.; Pol, J.M.A.; Berns, A.J.M.; Gielkens, A.L.J. Contribution of single genes within the unique short region of Aujeszky’s disease virus (suid herpesvirus type 1) to virulence, pathogenesis and immunogenicity. J. Gen. Virol. 1992, 73, 243–251. [Google Scholar] [CrossRef]
- Maia, T.M.; Staes, A.; Plasman, K.; Pauwels, J.; Boucher, K.; Argentini, A.; Martens, L.; Montoye, T.; Gevaert, K.; Impens, F. Simple Peptide Quantification Approach for MS-Based Proteomics Quality Control. ACS Omega 2020, 5, 6754–6762. [Google Scholar] [CrossRef]
- Chiva, C.; Olivella, R.; Borràs, E.; Espadas, G.; Pastor, O.; Solé, A.; Sabidó, E. QCloud: A cloud-based quality control system for mass spectrometry-based proteomics laboratories. PLoS ONE 2018, 13, e0189209. [Google Scholar] [CrossRef]
- Cox, J.; Hein, M.Y.; Luber, C.A.; Paron, I.; Nagaraj, N.; Mann, M. Accurate Proteome-wide Label-free Quantification by Delayed Normalization and Maximal Peptide Ratio Extraction, Termed MaxLFQ. Mol. Cell. Proteom. 2014, 13, 2513–2526. [Google Scholar] [CrossRef] [PubMed]
- Perez-Riverol, Y.; Csordas, A.; Bai, J.; Bernal-Llinares, M.; Hewapathirana, S.; Kundu, D.J.; Inuganti, A.; Griss, J.; Mayer, G.; Eisenacher, M.; et al. The PRIDE database and related tools and resources in 2019: Improving support for quantification data. Nucleic Acids Res. 2018, 47, D442–D450. [Google Scholar] [CrossRef]

| Gene Name | Protein Name | Site | Log2(KD/WT) | -Log2(p-Value) |
|---|---|---|---|---|
| CDS2 | Phosphatidate cytidylyltransferase | S33 | 6.38 | 4.56 |
| BCKDHA | 2-oxoisovalerate dehydrogenase alpha | S313 | 5.83 | 3.48 |
| BCKDHA | 2-oxoisovalerate dehydrogenase alpha | S303 | 5.54 | 3.25 |
| PPP6R1 | S/T-protein phosphatase 6 subunit 1 | S531 | 4.39 | 2.92 |
| WDR20 | WD repeat-containing protein 20 | S348 | 4.16 | 3.83 |
| Gene Name | Protein Name | Site | Log2(KD/WT) | -Log2(p-Value) |
|---|---|---|---|---|
| TSSC1 | EARP and GARP complex interacting protein 1 | S320 | −6.50 | 4.65 |
| RAB11FIP5 | Rab11 family-interacting protein 5 | T162 | −6.24 | 3.53 |
| RAB11FIP5 | Rab11 family-interacting protein 5 | S164 | −6.09 | 3.98 |
| TOMM70 | Mitochondrial import receptor subunit TOM70 | S97 | −6.01 | 4.52 |
| TESK2 | Dual specificity testis-specific protein kinase 2 | S8 | −5.85 | 6.37 |
| SZRD1 | SUZ domain-containing protein 1 | S17 | −5.48 | 2.28 |
| DDX17 | Probable ATP-dependent RNA helicase DDX17 | S575 | −5.23 | 3.85 |
| LMNA | Prelamin-A/C | S12 | −5.21 | 1.56 |
| SZRD1 | SUZ domain-containing protein 1 | S19 | −5.09 | 5.62 |
| PALMD | Palmdelphin | T255 | −4.78 | 3.29 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jansens, R.J.J.; Marmiroli, S.; Favoreel, H.W. An Unbiased Approach to Mapping the Signaling Network of the Pseudorabies Virus US3 Protein. Pathogens 2020, 9, 916. https://doi.org/10.3390/pathogens9110916
Jansens RJJ, Marmiroli S, Favoreel HW. An Unbiased Approach to Mapping the Signaling Network of the Pseudorabies Virus US3 Protein. Pathogens. 2020; 9(11):916. https://doi.org/10.3390/pathogens9110916
Chicago/Turabian StyleJansens, Robert J. J., Sandra Marmiroli, and Herman W. Favoreel. 2020. "An Unbiased Approach to Mapping the Signaling Network of the Pseudorabies Virus US3 Protein" Pathogens 9, no. 11: 916. https://doi.org/10.3390/pathogens9110916
APA StyleJansens, R. J. J., Marmiroli, S., & Favoreel, H. W. (2020). An Unbiased Approach to Mapping the Signaling Network of the Pseudorabies Virus US3 Protein. Pathogens, 9(11), 916. https://doi.org/10.3390/pathogens9110916






