Early Events in Japanese Encephalitis Virus Infection: Viral Entry
Abstract
1. Introduction: JEV Is a Mosquito-Borne Neurotropic Flavivirus
2. JEV Is a Zoonotic Pathogen Capable of Infecting a Wide Range of Animal Species
3. JEV Is a Small Enveloped Positive-Strand RNA Virus
3.1. Genome Structure and Gene Expression
3.2. Viral Replication Cycle
4. Viral Entry Is the First Step in the Infection Process
4.1. Virus Structure
4.2. Attachment
4.2.1. Viral Components
4.2.2. Cellular Components
Glycosaminoglycans (GAGs)
C-Type Lectins
Integrins
Other Host Factors
4.3. Endocytosis
4.4. Membrane Fusion and Uncoating
5. Conclusions: JEV Entry Is an Area of High Interest for Future Research
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Simmonds, P.; Becher, P.; Bukh, J.; Gould, E.A.; Meyers, G.; Monath, T.; Muerhoff, S.; Pletnev, A.; Rico-Hesse, R.; Smith, D.B.; et al. ICTV virus taxonomy profile: Flaviviridae. J. Gen. Virol. 2017, 98, 2–3. [Google Scholar] [CrossRef] [PubMed]
- Simmonds, P.; Becher, P.; Collett, M.S.; Gould, E.A.; Heinz, F.X.; Meyers, G.; Monath, T.; Pletnev, A.; Rice, C.M.; Stiasny, K.; et al. Family flaviviridae. In Virus Taxonomy: 9th Report of the International Committee on Taxonomy of Viruses; King, A.M.Q., Adams, M.J., Carstens, E.B., Lefkowitz, E.J., Eds.; Elsevier Academic Press: San Diego, CA, USA, 2012. [Google Scholar]
- Gould, E.A.; de Lamballerie, X.; Zanotto, P.M.; Holmes, E.C. Origins, evolution, and vector/host coadaptations within the genus Flavivirus. Adv. Virus Res. 2003, 59, 277–314. [Google Scholar] [PubMed]
- Weaver, S.C.; Barrett, A.D. Transmission cycles, host range, evolution and emergence of arboviral disease. Nat. Rev. Microbiol. 2004, 2, 789–801. [Google Scholar] [CrossRef] [PubMed]
- Valarcher, J.F.; Hagglund, S.; Juremalm, M.; Blomqvist, G.; Renstrom, L.; Zohari, S.; Leijon, M.; Chirico, J. Tick-borne encephalitis. Rev. Sci. Tech. 2015, 34, 453–466. [Google Scholar] [CrossRef] [PubMed]
- Weissenbock, H.; Hubalek, Z.; Bakonyi, T.; Nowotny, N. Zoonotic mosquito-borne flaviviruses: Worldwide presence of agents with proven pathogenicity and potential candidates of future emerging diseases. Vet. Microbiol. 2010, 140, 271–280. [Google Scholar] [CrossRef] [PubMed]
- Kuno, G.; Mackenzie, J.S.; Junglen, S.; Hubalek, Z.; Plyusnin, A.; Gubler, D.J. Vertebrate reservoirs of arboviruses: Myth, synonym of amplifier, or reality? Viruses 2017, 9, 185. [Google Scholar] [CrossRef] [PubMed]
- Billoir, F.; de Chesse, R.; Tolou, H.; de Micco, P.; Gould, E.A.; de Lamballerie, X. Phylogeny of the genus Flavivirus using complete coding sequences of arthropod-borne viruses and viruses with no known vector. J. Gen. Virol. 2000, 81, 781–790. [Google Scholar] [CrossRef] [PubMed]
- Alkan, C.; Zapata, S.; Bichaud, L.; Moureau, G.; Lemey, P.; Firth, A.E.; Gritsun, T.S.; Gould, E.A.; de Lamballerie, X.; Depaquit, J.; et al. Ecuador Paraiso Escondido virus, a new flavivirus isolated from New World sand flies in Ecuador, is the first representative of a novel clade in the genus Flavivirus. J. Virol. 2015, 89, 11773–11785. [Google Scholar] [CrossRef] [PubMed]
- Blitvich, B.J.; Firth, A.E. A review of flaviviruses that have no known arthropod vector. Viruses 2017, 9, 154. [Google Scholar] [CrossRef] [PubMed]
- Blitvich, B.J.; Firth, A.E. Insect-specific flaviviruses: A systematic review of their discovery, host range, mode of transmission, superinfection exclusion potential and genomic organization. Viruses 2015, 7, 1927–1959. [Google Scholar] [CrossRef] [PubMed]
- Bolling, B.G.; Weaver, S.C.; Tesh, R.B.; Vasilakis, N. Insect-specific virus discovery: Significance for the arbovirus community. Viruses 2015, 7, 4911–4928. [Google Scholar] [CrossRef] [PubMed]
- Mackenzie, J.S.; Williams, D.T. The zoonotic flaviviruses of southern, south-eastern and eastern Asia, and Australasia: The potential for emergent viruses. Zoonoses Public Health 2009, 56, 338–356. [Google Scholar] [CrossRef] [PubMed]
- Mackenzie, J.S.; Gubler, D.J.; Petersen, L.R. Emerging flaviviruses: The spread and resurgence of Japanese encephalitis, West Nile and dengue viruses. Nat. Med. 2004, 10, S98–S109. [Google Scholar] [CrossRef] [PubMed]
- Sips, G.J.; Wilschut, J.; Smit, J.M. Neuroinvasive flavivirus infections. Rev. Med. Virol. 2012, 22, 69–87. [Google Scholar] [CrossRef] [PubMed]
- Daep, C.A.; Munoz-Jordan, J.L.; Eugenin, E.A. Flaviviruses, an expanding threat in public health: Focus on dengue, West Nile, and Japanese encephalitis virus. J. Neurovirol. 2014, 20, 539–560. [Google Scholar] [CrossRef] [PubMed]
- Mackenzie, J.S.; Barrett, A.D.; Deubel, V. The Japanese encephalitis serological group of flaviviruses: A brief introduction to the group. Curr. Top. Microbiol. Immunol. 2002, 267, 1–10. [Google Scholar] [PubMed]
- Smith, D.R. Waiting in the wings: The potential of mosquito transmitted flaviviruses to emerge. Crit. Rev. Microbiol. 2017, 43, 405–422. [Google Scholar] [CrossRef] [PubMed]
- Song, B.H.; Yun, S.I.; Woolley, M.; Lee, Y.M. Zika virus: History, epidemiology, transmission, and clinical presentation. J. Neuroimmunol. 2017, 308, 50–64. [Google Scholar] [CrossRef] [PubMed]
- Vasilakis, N.; Weaver, S.C. The history and evolution of human dengue emergence. Adv. Virus Res. 2008, 72, 1–76. [Google Scholar] [PubMed]
- Wasserman, S.; Tambyah, P.A.; Lim, P.L. Yellow fever cases in Asia: Primed for an epidemic. Int. J. Infect. Dis. 2016, 48, 98–103. [Google Scholar] [CrossRef] [PubMed]
- Dobler, G. Zoonotic tick-borne flaviviruses. Vet. Microbiol. 2010, 140, 221–228. [Google Scholar] [CrossRef] [PubMed]
- Solomon, T. Control of Japanese encephalitis—Within our grasp? N. Engl. J. Med. 2006, 355, 869–871. [Google Scholar] [CrossRef] [PubMed]
- Misra, U.K.; Kalita, J. Overview: Japanese encephalitis. Prog. Neurobiol. 2010, 91, 108–120. [Google Scholar] [CrossRef] [PubMed]
- Endy, T.P.; Nisalak, A. Japanese encephalitis virus: Ecology and epidemiology. Curr. Top. Microbiol. Immunol. 2002, 267, 11–48. [Google Scholar] [PubMed]
- Erlanger, T.E.; Weiss, S.; Keiser, J.; Utzinger, J.; Wiedenmayer, K. Past, present, and future of Japanese encephalitis. Emerg. Infect. Dis. 2009, 15, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Mackenzie, J.S.; Williams, D.T.; Smith, D.W. Japanese encephalitis virus: The geographic distribution, incidence, and spread of a virus with a propensity to emerge in new areas. In Emerging Viruses in Human Populations; Tabor, E., Ed.; Elsevier: Amsterdam, The Netherlands, 2007. [Google Scholar]
- Van den Hurk, A.F.; Ritchie, S.A.; Mackenzie, J.S. Ecology and geographical expansion of Japanese encephalitis virus. Annu. Rev. Entomol. 2009, 54, 17–35. [Google Scholar] [CrossRef] [PubMed]
- Mackenzie, J.S.; Johansen, C.A.; Ritchie, S.A.; van den Hurk, A.F.; Hall, R.A. Japanese encephalitis as an emerging virus: The emergence and spread of Japanese encephalitis virus in Australasia. Curr. Top. Microbiol. Immunol. 2002, 267, 49–73. [Google Scholar] [PubMed]
- Tiroumourougane, S.V.; Raghava, P.; Srinivasan, S. Japanese viral encephalitis. Postgrad. Med. J. 2002, 78, 205–215. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization (WHO). Japanese encephalitis vaccines. Wkly. Epidemiol. Rec. 2006, 81, 331–340. [Google Scholar]
- Yun, S.I.; Lee, Y.M. Japanese encephalitis: The virus and vaccines. Hum. Vaccin. Immunother. 2014, 10, 263–279. [Google Scholar] [CrossRef] [PubMed]
- Johansen, C.A.; van den Hurk, A.F.; Ritchie, S.A.; Zborowski, P.; Nisbet, D.J.; Paru, R.; Bockarie, M.J.; Macdonald, J.; Drew, A.C.; Khromykh, T.I.; et al. Isolation of Japanese encephalitis virus from mosquitoes (Diptera: Culicidae) collected in the Western Province of Papua New Guinea, 1997–1998. Am. J. Trop. Med. Hyg. 2000, 62, 631–638. [Google Scholar] [CrossRef] [PubMed]
- Hanson, J.P.; Taylor, C.T.; Richards, A.R.; Smith, I.L.; Boutlis, C.S. Japanese encephalitis acquired near Port Moresby: Implications for residents and travellers to Papua New Guinea. Med. J. Aust. 2004, 181, 282–283. [Google Scholar] [PubMed]
- Hanna, J.N.; Ritchie, S.A.; Phillips, D.A.; Lee, J.M.; Hills, S.L.; van den Hurk, A.F.; Pyke, A.T.; Johansen, C.A.; Mackenzie, J.S. Japanese encephalitis in north Queensland, Australia, 1998. Med. J. Aust. 1999, 170, 533–536. [Google Scholar] [PubMed]
- Hanna, J.N.; Ritchie, S.A.; Phillips, D.A.; Shield, J.; Bailey, M.C.; Mackenzie, J.S.; Poidinger, M.; McCall, B.J.; Mills, P.J. An outbreak of Japanese encephalitis in the Torres Strait, Australia, 1995. Med. J. Aust. 1996, 165, 256–260. [Google Scholar] [PubMed]
- Ritchie, S.A.; Phillips, D.; Broom, A.; Mackenzie, J.; Poidinger, M.; van den Hurk, A. Isolation of Japanese encephalitis virus from Culex annulirostris in Australia. Am. J. Trop. Med. Hyg. 1997, 56, 80–84. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, S.A.; Rochester, W. Wind-blown mosquitoes and introduction of Japanese encephalitis into Australia. Emerg. Infect. Dis. 2001, 7, 900–903. [Google Scholar] [CrossRef] [PubMed]
- Van den Hurk, A.F.; Montgomery, B.L.; Northill, J.A.; Smith, I.L.; Zborowski, P.; Ritchie, S.A.; Mackenzie, J.S.; Smith, G.A. Short report: The first isolation of Japanese encephalitis virus from mosquitoes collected from mainland Australia. Am. J. Trop. Med. Hyg. 2006, 75, 21–25. [Google Scholar] [PubMed]
- Van den Hurk, A.F.; Ritchie, S.A.; Johansen, C.A.; Mackenzie, J.S.; Smith, G.A. Domestic pigs and Japanese encephalitis virus infection, Australia. Emerg. Infect. Dis. 2008, 14, 1736–1738. [Google Scholar] [CrossRef] [PubMed]
- Paul, W.S.; Moore, P.S.; Karabatsos, N.; Flood, S.P.; Yamada, S.; Jackson, T.; Tsai, T.F. Outbreak of Japanese encephalitis on the island of Saipan, 1990. J. Infect. Dis. 1993, 167, 1053–1058. [Google Scholar] [CrossRef] [PubMed]
- Igarashi, A.; Tanaka, M.; Morita, K.; Takasu, T.; Ahmed, A.; Ahmed, A.; Akram, D.S.; Waqar, M.A. Detection of West Nile and Japanese encephalitis viral genome sequences in cerebrospinal fluid from acute encephalitis cases in Karachi, Pakistan. Microbiol. Immunol. 1994, 38, 827–830. [Google Scholar] [CrossRef] [PubMed]
- Li, M.H.; Fu, S.H.; Chen, W.X.; Wang, H.Y.; Guo, Y.H.; Liu, Q.Y.; Li, Y.X.; Luo, H.M.; Da, W.; Duo Ji, D.Z.; et al. Genotype V Japanese encephalitis virus is emerging. PLoS Negl. Trop. Dis. 2011, 5. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.X.; Li, M.H.; Fu, S.H.; Chen, W.X.; Liu, Q.Y.; Zhang, H.L.; Da, W.; Hu, S.L.; Mu, S.D.; Bai, J.; et al. Japanese encephalitis, Tibet, China. Emerg. Infect. Dis. 2011, 17, 934–936. [Google Scholar] [CrossRef] [PubMed]
- Ravanini, P.; Huhtamo, E.; Ilaria, V.; Crobu, M.G.; Nicosia, A.M.; Servino, L.; Rivasi, F.; Allegrini, S.; Miglio, U.; Magri, A.; et al. Japanese encephalitis virus RNA detected in Culex pipiens mosquitoes in Italy. Euro Surveill 2012, 17. [Google Scholar] [CrossRef]
- Platonov, A.; Rossi, G.; Karan, L.; Mironov, K.; Busani, L.; Rezza, G. Does the Japanese encephalitis virus (JEV) represent a threat for human health in Europe? Detection of JEV RNA sequences in birds collected in Italy. Euro Surveill 2012, 17. [Google Scholar] [CrossRef]
- Zeller, H. Is Japanese encephalitis emerging in Europe? Euro Surveill 2012, 17. [Google Scholar] [CrossRef]
- Nett, R.J.; Campbell, G.L.; Reisen, W.K. Potential for the emergence of Japanese encephalitis virus in California. Vector Borne Zoonotic Dis. 2009, 9, 511–517. [Google Scholar] [CrossRef] [PubMed]
- Scherer, W.F.; Buescher, E.L.; Flemings, M.B.; Noguchi, A.; Scanlon, J. Ecologic studies of Japanese encephalitis virus in Japan. III. Mosquito factors. Zootropism and vertical flight of Culex tritaeniorhynchus with observations on variations in collections from animal-baited traps in different habitats. Am. J. Trop. Med. Hyg. 1959, 8, 665–677. [Google Scholar] [CrossRef] [PubMed]
- Su, C.L.; Yang, C.F.; Teng, H.J.; Lu, L.C.; Lin, C.; Tsai, K.H.; Chen, Y.Y.; Chen, L.Y.; Chang, S.F.; Shu, P.Y. Molecular epidemiology of Japanese encephalitis virus in mosquitoes in Taiwan during 2005–2012. PLoS Negl. Trop. Dis. 2014, 8. [Google Scholar] [CrossRef] [PubMed]
- Gajanana, A.; Rajendran, R.; Samuel, P.P.; Thenmozhi, V.; Tsai, T.F.; Kimura-Kuroda, J.; Reuben, R. Japanese encephalitis in south Arcot district, Tamil Nadu, India: A three-year longitudinal study of vector abundance and infection frequency. J. Med. Entomol. 1997, 34, 651–659. [Google Scholar] [CrossRef] [PubMed]
- Lindahl, J.F.; Stahl, K.; Chirico, J.; Boqvist, S.; Thu, H.T.; Magnusson, U. Circulation of Japanese encephalitis virus in pigs and mosquito vectors within Can Tho city, Vietnam. PLoS Negl. Trop. Dis. 2013, 7. [Google Scholar] [CrossRef] [PubMed]
- Seo, H.J.; Kim, H.C.; Klein, T.A.; Ramey, A.M.; Lee, J.H.; Kyung, S.G.; Park, J.Y.; Cho, Y.S.; Cho, I.S.; Yeh, J.Y. Molecular detection and genotyping of Japanese encephalitis virus in mosquitoes during a 2010 outbreak in the Republic of Korea. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Tao, Z.; Liu, G.; Wang, M.; Wang, H.; Lin, X.; Song, L.; Wang, S.; Wang, H.; Liu, X.; Cui, N.; et al. Molecular epidemiology of Japanese encephalitis virus in mosquitoes during an outbreak in China, 2013. Sci. Rep. 2014, 4. [Google Scholar] [CrossRef] [PubMed]
- Hall-Mendelin, S.; Jansen, C.C.; Cheah, W.Y.; Montgomery, B.L.; Hall, R.A.; Ritchie, S.A.; Van den Hurk, A.F. Culex annulirostris (Diptera: Culicidae) host feeding patterns and Japanese encephalitis virus ecology in northern Australia. J. Med. Entomol. 2012, 49, 371–377. [Google Scholar] [CrossRef] [PubMed]
- Okuno, T.; Tseng, P.T.; Liu, S.Y.; Hsu, S.Y.; Huang, C.T. Rates of infection with Japanese encephalitis virus of two culicine species of mosquito in Taiwan. Bull. World Health Organ. 1971, 44, 599–604. [Google Scholar] [PubMed]
- Detels, R.; Cross, J.H.; Huang, W.C.; Lien, J.C.; Chen, S. Japanese encephalitis virus in Northern Taiwan, 1969–1973. Am. J. Trop. Med. Hyg. 1976, 25, 477–485. [Google Scholar] [CrossRef] [PubMed]
- Gingrich, J.B.; Nisalak, A.; Latendresse, J.R.; Sattabongkot, J.; Hoke, C.H.; Pomsdhit, J.; Chantalakana, C.; Satayaphanta, C.; Uechiewcharnkit, K.; Innis, B.L. Japanese encephalitis virus in Bangkok: Factors influencing vector infections in three suburban communities. J. Med. Entomol. 1992, 29, 436–444. [Google Scholar] [CrossRef] [PubMed]
- Peiris, J.S.; Amerasinghe, F.P.; Arunagiri, C.K.; Perera, L.P.; Karunaratne, S.H.; Ratnayake, C.B.; Kulatilaka, T.A.; Abeysinghe, M.R. Japanese encephalitis in Sri Lanka: Comparison of vector and virus ecology in different agro-climatic areas. Trans. R. Soc. Trop. Med. Hyg. 1993, 87, 541–548. [Google Scholar] [CrossRef]
- Vythilingam, I.; Oda, K.; Mahadevan, S.; Abdullah, G.; Thim, C.S.; Hong, C.C.; Vijayamalar, B.; Sinniah, M.; Igarashi, A. Abundance, parity, and Japanese encephalitis virus infection of mosquitoes (Diptera: Culicidae) in Sepang District, Malaysia. J. Med. Entomol. 1997, 34, 257–262. [Google Scholar] [CrossRef] [PubMed]
- Gould, D.J.; Edelman, R.; Grossman, R.A.; Nisalak, A.; Sullivan, M.F. Study of Japanese encephalitis virus in Chiangmai Valley, Thailand. IV. Vector studies. Am. J. Epidemiol. 1974, 100, 49–56. [Google Scholar] [CrossRef] [PubMed]
- Leake, C.J.; Ussery, M.A.; Nisalak, A.; Hoke, C.H.; Andre, R.G.; Burke, D.S. Virus isolations from mosquitoes collected during the 1982 Japanese encephalitis epidemic in northern Thailand. Trans. R. Soc. Trop. Med. Hyg. 1986, 80, 831–837. [Google Scholar] [CrossRef]
- Reuben, R.; Thenmozhi, V.; Samuel, P.P.; Gajanana, A.; Mani, T.R. Mosquito blood feeding patterns as a factor in the epidemiology of Japanese encephalitis in southern India. Am. J. Trop. Med. Hyg. 1992, 46, 654–663. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Cha, G.W.; Jeong, Y.E.; Lee, W.G.; Chang, K.S.; Roh, J.Y.; Yang, S.C.; Park, M.Y.; Park, C.; Shin, E.H. Detection of Japanese encephalitis virus genotype V in Culex orientalis and Culex pipiens (Diptera: Culicidae) in Korea. PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [PubMed]
- Huber, K.; Jansen, S.; Leggewie, M.; Badusche, M.; Schmidt-Chanasit, J.; Becker, N.; Tannich, E.; Becker, S.C. Aedes japonicus japonicus (Diptera: Culicidae) from Germany have vector competence for Japanese encephalitis virus but are refractory to infection with West Nile virus. Parasitol. Res. 2014, 113, 3195–3199. [Google Scholar] [CrossRef] [PubMed]
- Mackenzie-Impoinvil, L.; Impoinvil, D.E.; Galbraith, S.E.; Dillon, R.J.; Ranson, H.; Johnson, N.; Fooks, A.R.; Solomon, T.; Baylis, M. Evaluation of a temperate climate mosquito, Ochlerotatus detritus (=Aedes detritus), as a potential vector of Japanese encephalitis virus. Med. Vet. Entomol. 2015, 29, 1–9. [Google Scholar] [CrossRef] [PubMed]
- De Wispelaere, M.; Despres, P.; Choumet, V. European Aedes albopictus and Culex pipiens are competent vectors for Japanese encephalitis virus. PLoS Negl. Trop. Dis. 2017, 11. [Google Scholar] [CrossRef] [PubMed]
- Weng, M.H.; Lien, J.C.; Wang, Y.M.; Wu, H.L.; Chin, C. Susceptibility of three laboratory strains of Aedes albopictus (Diptera: Culicidae) to Japanese encephalitis virus from Taiwan. J. Med. Entomol. 1997, 34, 745–747. [Google Scholar] [CrossRef] [PubMed]
- Sucharit, S.; Surathin, K.; Shrestha, S.R. Vectors of Japanese encephalitis virus (JEV): Species complexes of the vectors. Southeast Asian J. Trop. Med. Public Health 1989, 20, 611–621. [Google Scholar] [PubMed]
- Dhanda, V.; Mourya, D.T.; Mishra, A.C.; Ilkal, M.A.; Pant, U.; Jacob, P.G.; Bhat, H.R. Japanese encephalitis virus infection in mosquitoes reared from field-collected immatures and in wild-caught males. Am. J. Trop. Med. Hyg. 1989, 41, 732–736. [Google Scholar] [CrossRef] [PubMed]
- Rosen, L.; Lien, J.C.; Shroyer, D.A.; Baker, R.H.; Lu, L.C. Experimental vertical transmission of Japanese encephalitis virus by Culex tritaeniorhynchus and other mosquitoes. Am. J. Trop. Med. Hyg. 1989, 40, 548–556. [Google Scholar] [CrossRef] [PubMed]
- Rosen, L.; Lien, J.C.; Lu, L.C. A longitudinal study of the prevalence of Japanese encephalitis virus in adult and larval Culex tritaeniorhynchus mosquitoes in northern Taiwan. Am. J. Trop. Med. Hyg. 1989, 40, 557–560. [Google Scholar] [CrossRef] [PubMed]
- Rosen, L.; Tesh, R.B.; Lien, J.C.; Cross, J.H. Transovarial transmission of Japanese encephalitis virus by mosquitoes. Science 1978, 199, 909–911. [Google Scholar] [CrossRef] [PubMed]
- Cheng, V.C.C.; Sridhar, S.; Wong, S.C.; Wong, S.C.Y.; Chan, J.F.W.; Yip, C.C.Y.; Chau, C.H.; Au, T.W.K.; Hwang, Y.Y.; Yau, C.S.W.; et al. Japanese encephalitis virus transmitted via blood transfusion, Hong Kong, China. Emerg. Infect. Dis. 2018, 24, 49–57. [Google Scholar] [CrossRef] [PubMed]
- Buescher, E.L.; Scherer, W.F.; Rosenberg, M.Z.; Gresser, I.; Hardy, J.L.; Bullock, H.R. Ecologic studies of Japanese encephalitis virus in Japan. II. Mosquito infection. Am. J. Trop. Med. Hyg. 1959, 8, 651–664. [Google Scholar] [CrossRef] [PubMed]
- Buescher, E.L.; Scherer, W.F. Ecologic studies of Japanese encephalitis virus in Japan. IX. Epidemiologic correlations and conclusions. Am. J. Trop. Med. Hyg. 1959, 8, 719–722. [Google Scholar] [CrossRef] [PubMed]
- Scherer, W.F.; Moyer, J.T.; Izumi, T.; Gresser, I.; Mc, C.J. Ecologic studies of Japanese encephalitis virus in Japan. VI. Swine infection. Am. J. Trop. Med. Hyg. 1959, 8, 698–706. [Google Scholar] [CrossRef] [PubMed]
- Ilkal, M.A.; Prasanna, Y.; Jacob, P.G.; Geevarghese, G.; Banerjee, K. Experimental studies on the susceptibility of domestic pigs to West Nile virus followed by Japanese encephalitis virus infection and vice versa. Acta Virol. 1994, 38, 157–161. [Google Scholar] [PubMed]
- Nemeth, N.; Bosco-Lauth, A.; Oesterle, P.; Kohler, D.; Bowen, R. North American birds as potential amplifying hosts of Japanese encephalitis virus. Am. J. Trop. Med. Hyg. 2012, 87, 760–767. [Google Scholar] [CrossRef] [PubMed]
- Buescher, E.L.; Scherer, W.F.; Mc, C.H.; Moyer, J.T.; Rosenberg, M.Z.; Yoshii, M.; Okada, Y. Ecologic studies of Japanese encephalitis virus in Japan. IV. Avian infection. Am. J. Trop. Med. Hyg. 1959, 8, 678–688. [Google Scholar] [CrossRef] [PubMed]
- Hale, J.H.; Lim, K.A.; Colless, D.H. Investigation of domestic pigs as a potential reservoir of Japanese B encephalitis virus on Singapore Island. Ann. Trop. Med. Parasitol. 1957, 51, 374–379. [Google Scholar] [CrossRef] [PubMed]
- Pond, W.L.; Russ, S.B.; Lancaster, W.E.; Audy, J.R.; Smadel, J.E. Japanese encephalitis in Malaya. II. Distribution of neutralizing antibodies in man and animals. Am. J. Hyg. 1954, 59, 17–25. [Google Scholar] [PubMed]
- Takashima, I.; Watanabe, T.; Ouchi, N.; Hashimoto, N. Ecological studies of Japanese encephalitis virus in Hokkaido: Interepidemic outbreaks of swine abortion and evidence for the virus to overwinter locally. Am. J. Trop. Med. Hyg. 1988, 38, 420–427. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, T.; Kawakami, Y.; Fukuhara, S.; Matumoto, M. Experimental stillbirth in pregnant swine infected with Japanese encephalitis virus. Jpn. J. Exp. Med. 1954, 24, 363–375. [Google Scholar] [PubMed]
- Nah, J.J.; Yang, D.K.; Kim, H.H.; Song, J.Y. The present and future of veterinary vaccines for Japanese encephalitis in Korea. Clin. Exp. Vaccine Res. 2015, 4, 130–136. [Google Scholar] [CrossRef] [PubMed]
- Hsu, S.T.; Chang, L.C.; Lin, S.Y.; Chuang, T.Y.; Ma, C.H.; Inoue, Y.K.; Okuno, T. The effect of vaccination with a live attenuated strain of Japanese encephalitis virus on stillbirths in swine in Taiwan. Bull. World Health Organ. 1972, 46, 465–471. [Google Scholar] [PubMed]
- Sulkin, S.E.; Allen, R.; Miura, T.; Toyokawa, K. Studies of arthropod-borne virus infections in Chiroptera. VI. Isolation of Japanese B encephalitis virus from naturally infected bats. Am. J. Trop. Med. Hyg. 1970, 19, 77–87. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.L.; Pan, X.L.; Zhang, H.L.; Fu, S.H.; Wang, H.Y.; Tang, Q.; Wang, L.F.; Liang, G.D. Japanese encephalitis viruses from bats in Yunnan, China. Emerg. Infect. Dis. 2009, 15, 939–942. [Google Scholar] [CrossRef] [PubMed]
- Van den Hurk, A.F.; Smith, C.S.; Field, H.E.; Smith, I.L.; Northill, J.A.; Taylor, C.T.; Jansen, C.C.; Smith, G.A.; Mackenzie, J.S. Transmission of Japanese encephalitis virus from the black flying fox, Pteropus alecto, to Culex annulirostris mosquitoes, despite the absence of detectable viremia. Am. J. Trop. Med. Hyg. 2009, 81, 457–462. [Google Scholar] [PubMed]
- Miura, T.; Toyokawa, K.; Allen, R.; Sulkin, S.E. Studies of arthropod-borne virus infections in Chiroptera. VII. Serologic evidence of natural Japanese B encephalitis virus infection in bats. Am. J. Trop. Med. Hyg. 1970, 19, 88–93. [Google Scholar] [CrossRef] [PubMed]
- Sulkin, S.E.; Allen, R.; Sims, R.; Singh, K.V. Studies of arthropod-borne virus infections in Chiroptera. IV. The immune response of the big brown bat (Eptesicus f. fuscus) maintained at various environmental temperatures to experimental Japanese B encephalitis virus infection. Am. J. Trop. Med. Hyg. 1966, 15, 418–427. [Google Scholar] [CrossRef] [PubMed]
- La Motte, L.C., Jr. Japanese B encephalitis in bats during simulated hibernation. Am. J. Hyg. 1958, 67, 101–108. [Google Scholar] [PubMed]
- Lam, K.H.; Ellis, T.M.; Williams, D.T.; Lunt, R.A.; Daniels, P.W.; Watkins, K.L.; Riggs, C.M. Japanese encephalitis in a racing thoroughbred gelding in Hong Kong. Vet. Rec. 2005, 157, 168–173. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, H. Japanese encephalitis in horses in Japan. Equine Vet. J. 1972, 4, 155–156. [Google Scholar] [CrossRef] [PubMed]
- Sugiura, T.; Shimada, K. Seroepizootiological survey of Japanese encephalitis virus and Getah virus in regional horse race tracks from 1991 to 1997 in Japan. J. Vet. Med. Sci. 1999, 61, 877–881. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yamanaka, T.; Tsujimura, K.; Kondo, T.; Yasuda, W.; Okada, A.; Noda, K.; Okumura, T.; Matsumura, T. Isolation and genetic analysis of Japanese encephalitis virus from a diseased horse in Japan. J. Vet. Med. Sci. 2006, 68, 293–295. [Google Scholar] [CrossRef] [PubMed]
- Gould, D.J.; Byrne, R.J.; Hayes, D.E. Experimental infection of horses with Japanese encephalitis virus by mosquito bite. Am. J. Trop. Med. Hyg. 1964, 13, 742–746. [Google Scholar] [CrossRef] [PubMed]
- Konishi, E.; Shoda, M.; Kondo, T. Analysis of yearly changes in levels of antibodies to Japanese encephalitis virus nonstructural 1 protein in racehorses in central Japan shows high levels of natural virus activity still exist. Vaccine 2006, 24, 516–524. [Google Scholar] [CrossRef] [PubMed]
- Gulati, B.R.; Singha, H.; Singh, B.K.; Virmani, N.; Kumar, S.; Singh, R.K. Isolation and genetic characterization of Japanese encephalitis virus from equines in India. J. Vet. Sci. 2012, 13, 111–118. [Google Scholar] [CrossRef] [PubMed]
- Kheng, C.S.; Chee, T.K.; Marchette, N.J.; Garcia, R.; Rudnick, A.; Coughlan, R.F. Japanese B encephalitis in a horse. Aust. Vet. J. 1968, 44, 23–25. [Google Scholar] [CrossRef]
- Yang, D.K.; Kim, B.H.; Kweon, C.H.; Nah, J.J.; Kim, H.J.; Lee, K.W.; Yang, Y.J.; Mun, K.W. Serosurveillance for Japanese encephalitis, Akabane, and Aino viruses for Thoroughbred horses in Korea. J. Vet. Sci. 2008, 9, 381–385. [Google Scholar] [CrossRef] [PubMed]
- Katayama, T.; Saito, S.; Horiuchi, S.; Maruta, T.; Kato, T.; Yanase, T.; Yamakawa, M.; Shirafuji, H. Nonsuppurative encephalomyelitis in a calf in Japan and isolation of Japanese encephalitis virus genotype 1 from the affected calf. J. Clin. Microbiol. 2013, 51, 3448–3453. [Google Scholar] [CrossRef] [PubMed]
- Kako, N.; Suzuki, S.; Sugie, N.; Kato, T.; Yanase, T.; Yamakawa, M.; Shirafuji, H. Japanese encephalitis in a 114-month-old cow: Pathological investigation of the affected cow and genetic characterization of Japanese encephalitis virus isolate. BMC Vet. Res. 2014, 10. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, T.; Mochizuki, H.; Sugawa, Y.; Okazaki, K.; Matumoto, M. Studies on Japanese encephalitis of cattle—1. Bovine encephalitis caused by natural infection with Japanese encephalitis virus. Natl. Inst. Anim. Health Q. 1951, 23, 111–118. [Google Scholar]
- Shimizu, T.; Mochizuki, H.; Sugawa, Y.; Okazaki, K.; Matumoto, M. Studies on Japanese encephalitis of cattle—2. Experimental infection of calves with virus of Japanese encephalitis. Natl. Inst. Anim. Health Q. 1951, 23, 119–128. [Google Scholar]
- Lim, S.I.; Kweon, C.H.; Tark, D.S.; Kim, S.H.; Yang, D.K. Sero-survey on Aino, Akabane, Chuzan, bovine ephemeral fever and Japanese encephalitis virus of cattle and swine in Korea. J. Vet. Sci. 2007, 8, 45–49. [Google Scholar] [CrossRef] [PubMed]
- Ilkal, M.A.; Dhanda, V.; Rao, B.U.; George, S.; Mishra, A.C.; Prasanna, Y.; Gopalkrishna, S.; Pavri, K.M. Absence of viraemia in cattle after experimental infection with Japanese encephalitis virus. Trans. R. Soc. Trop. Med. Hyg. 1988, 82, 628–631. [Google Scholar] [CrossRef]
- Mall, M.P.; Kumar, A.; Malik, S.V. Sero-positivity of domestic animals against Japanese encephalitis in Bareilly area, U.P. J. Commun. Dis. 1995, 27, 242–246. [Google Scholar] [PubMed]
- Kalaiyarasu, S.; Mishra, N.; Khetan, R.K.; Singh, V.P. Serological evidence of widespread West Nile virus and Japanese encephalitis virus infection in native domestic ducks (Anas platyrhynchos var domesticus) in Kuttanad region, Kerala, India. Comp. Immunol. Microbiol. Infect. Dis. 2016, 48, 61–68. [Google Scholar] [CrossRef] [PubMed]
- Dhanda, V.; Banerjee, K.; Deshmukh, P.K.; Ilkal, M.A. Experimental viraemia and transmission of Japanese encephalitis virus by mosquitoes in domestic ducks. Indian J. Med. Res. 1977, 66, 881–888. [Google Scholar] [PubMed]
- Nidaira, M.; Taira, K.; Itokazu, K.; Kudaka, J.; Nakamura, M.; Ohno, A.; Takasaki, T. Survey of the antibody against Japanese encephalitis virus in Ryukyu wild boars (Sus scrofa riukiuanus) in Okinawa, Japan. Jpn. J. Infect. Dis. 2007, 60, 309–311. [Google Scholar] [PubMed]
- Ohno, Y.; Sato, H.; Suzuki, K.; Yokoyama, M.; Uni, S.; Shibasaki, T.; Sashika, M.; Inokuma, H.; Kai, K.; Maeda, K. Detection of antibodies against Japanese encephalitis virus in raccoons, raccoon dogs and wild boars in Japan. J. Vet. Med. Sci. 2009, 71, 1035–1039. [Google Scholar] [CrossRef] [PubMed]
- Hamano, M.; Lim, C.K.; Takagi, H.; Sawabe, K.; Kuwayama, M.; Kishi, N.; Kurane, I.; Takasaki, T. Detection of antibodies to Japanese encephalitis virus in the wild boars in Hiroshima prefecture, Japan. Epidemiol. Infect. 2007, 135, 974–977. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.K.; Kweon, C.H.; Kim, B.H.; Hwang, I.J.; Kang, M.I.; So, B.J.; Cho, K.O. The seroprevalence of Japanese encephalitis virus in goats raised in Korea. J. Vet. Sci. 2007, 8, 197–199. [Google Scholar] [CrossRef] [PubMed]
- Mansfield, K.L.; Hernandez-Triana, L.M.; Banyard, A.C.; Fooks, A.R.; Johnson, N. Japanese encephalitis virus infection, diagnosis and control in domestic animals. Vet. Microbiol. 2017, 201, 85–92. [Google Scholar] [CrossRef] [PubMed]
- Cleton, N.B.; Bosco-Lauth, A.; Page, M.J.; Bowen, R.A. Age-related susceptibility to Japanese encephalitis virus in domestic ducklings and chicks. Am. J. Trop. Med. Hyg. 2014, 90, 242–246. [Google Scholar] [CrossRef] [PubMed]
- Ricklin, M.E.; Garcia-Nicolas, O.; Brechbuhl, D.; Python, S.; Zumkehr, B.; Posthaus, H.; Oevermann, A.; Summerfield, A. Japanese encephalitis virus tropism in experimentally infected pigs. Vet. Res. 2016, 47. [Google Scholar] [CrossRef] [PubMed]
- Ricklin, M.E.; Garcia-Nicolas, O.; Brechbuhl, D.; Python, S.; Zumkehr, B.; Nougairede, A.; Charrel, R.N.; Posthaus, H.; Oevermann, A.; Summerfield, A. Vector-free transmission and persistence of Japanese encephalitis virus in pigs. Nat. Commun. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Miller, R.H.; Masuoka, P.; Klein, T.A.; Kim, H.C.; Somer, T.; Grieco, J. Ecological niche modeling to estimate the distribution of Japanese encephalitis virus in Asia. PLoS Negl. Trop. Dis. 2012, 6. [Google Scholar] [CrossRef] [PubMed]
- Le Flohic, G.; Porphyre, V.; Barbazan, P.; Gonzalez, J.P. Review of climate, landscape, and viral genetics as drivers of the Japanese encephalitis virus ecology. PLoS Negl. Trop. Dis. 2013, 7. [Google Scholar] [CrossRef] [PubMed]
- Lord, J.S.; Gurley, E.S.; Pulliam, J.R. Rethinking Japanese encephalitis virus transmission: A framework for implicating host and vector species. PLoS Negl. Trop. Dis. 2015, 9. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Fink, K.; Zust, R.; Lim, S.P.; Qin, C.F.; Shi, P.Y. Flavivirus RNA methylation. J. Gen. Virol. 2014, 95, 763–778. [Google Scholar] [CrossRef] [PubMed]
- Klema, V.J.; Padmanabhan, R.; Choi, K.H. Flaviviral replication complex: Coordination between RNA synthesis and 5′-RNA capping. Viruses 2015, 7, 4640–4656. [Google Scholar] [CrossRef] [PubMed]
- Ray, D.; Shah, A.; Tilgner, M.; Guo, Y.; Zhao, Y.; Dong, H.; Deas, T.S.; Zhou, Y.; Li, H.; Shi, P.Y. West Nile virus 5′-cap structure is formed by sequential guanine N-7 and ribose 2′-O methylations by nonstructural protein 5. J. Virol. 2006, 80, 8362–8370. [Google Scholar] [CrossRef] [PubMed]
- Daffis, S.; Szretter, K.J.; Schriewer, J.; Li, J.; Youn, S.; Errett, J.; Lin, T.Y.; Schneller, S.; Zust, R.; Dong, H.; et al. 2′-O methylation of the viral mRNA cap evades host restriction by IFIT family members. Nature 2010, 468, 452–456. [Google Scholar] [CrossRef] [PubMed]
- Zust, R.; Cervantes-Barragan, L.; Habjan, M.; Maier, R.; Neuman, B.W.; Ziebuhr, J.; Szretter, K.J.; Baker, S.C.; Barchet, W.; Diamond, M.S.; et al. Ribose 2′-O-methylation provides a molecular signature for the distinction of self and non-self mRNA dependent on the RNA sensor Mda5. Nat. Immunol. 2011, 12, 137–143. [Google Scholar] [CrossRef] [PubMed]
- Yun, S.I.; Choi, Y.J.; Song, B.H.; Lee, Y.M. 3′ cis-acting elements that contribute to the competence and efficiency of Japanese encephalitis virus genome replication: Functional importance of sequence duplications, deletions, and substitutions. J. Virol. 2009, 83, 7909–7930. [Google Scholar] [CrossRef] [PubMed]
- Song, B.H.; Yun, S.I.; Choi, Y.J.; Kim, J.M.; Lee, C.H.; Lee, Y.M. A complex RNA motif defined by three discontinuous 5-nucleotide-long strands is essential for flavivirus RNA replication. RNA 2008, 14, 1791–1813. [Google Scholar] [CrossRef] [PubMed]
- Brinton, M.A.; Basu, M. Functions of the 3′ and 5′ genome RNA regions of members of the genus Flavivirus. Virus Res. 2015, 206, 108–119. [Google Scholar] [CrossRef] [PubMed]
- Ng, W.C.; Soto-Acosta, R.; Bradrick, S.S.; Garcia-Blanco, M.A.; Ooi, E.E. The 5′ and 3′ untranslated regions of the flaviviral genome. Viruses 2017, 9, 137. [Google Scholar] [CrossRef] [PubMed]
- Villordo, S.M.; Carballeda, J.M.; Filomatori, C.V.; Gamarnik, A.V. RNA structure duplications and flavivirus host adaptation. Trends Microbiol. 2016, 24, 270–283. [Google Scholar] [CrossRef] [PubMed]
- Lin, K.C.; Chang, H.L.; Chang, R.Y. Accumulation of a 3′-terminal genome fragment in Japanese encephalitis virus-infected mammalian and mosquito cells. J. Virol. 2004, 78, 5133–5138. [Google Scholar] [CrossRef] [PubMed]
- Pijlman, G.P.; Funk, A.; Kondratieva, N.; Leung, J.; Torres, S.; van der Aa, L.; Liu, W.J.; Palmenberg, A.C.; Shi, P.Y.; Hall, R.A.; et al. A highly structured, nuclease-resistant, noncoding RNA produced by flaviviruses is required for pathogenicity. Cell Host Microbe 2008, 4, 579–591. [Google Scholar] [CrossRef] [PubMed]
- Silva, P.A.; Pereira, C.F.; Dalebout, T.J.; Spaan, W.J.; Bredenbeek, P.J. An RNA pseudoknot is required for production of yellow fever virus subgenomic RNA by the host nuclease XRN1. J. Virol. 2010, 84, 11395–11406. [Google Scholar] [CrossRef] [PubMed]
- Funk, A.; Truong, K.; Nagasaki, T.; Torres, S.; Floden, N.; Balmori Melian, E.; Edmonds, J.; Dong, H.; Shi, P.Y.; Khromykh, A.A. RNA structures required for production of subgenomic flavivirus RNA. J. Virol. 2010, 84, 11407–11417. [Google Scholar] [CrossRef] [PubMed]
- Akiyama, B.M.; Laurence, H.M.; Massey, A.R.; Costantino, D.A.; Xie, X.; Yang, Y.; Shi, P.Y.; Nix, J.C.; Beckham, J.D.; Kieft, J.S. Zika virus produces noncoding RNAs using a multi-pseudoknot structure that confounds a cellular exonuclease. Science 2016, 354, 1148–1152. [Google Scholar] [CrossRef] [PubMed]
- Moon, S.L.; Anderson, J.R.; Kumagai, Y.; Wilusz, C.J.; Akira, S.; Khromykh, A.A.; Wilusz, J. A noncoding RNA produced by arthropod-borne flaviviruses inhibits the cellular exoribonuclease XRN1 and alters host mRNA stability. RNA 2012, 18, 2029–2040. [Google Scholar] [CrossRef] [PubMed]
- Pijlman, G.P. Flavivirus RNAi suppression: Decoding non-coding RNA. Curr. Opin. Virol. 2014, 7, 55–60. [Google Scholar] [CrossRef] [PubMed]
- Clarke, B.D.; Roby, J.A.; Slonchak, A.; Khromykh, A.A. Functional non-coding RNAs derived from the flavivirus 3′ untranslated region. Virus Res. 2015, 206, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Charley, P.A.; Wilusz, J. Standing your ground to exoribonucleases: Function of flavivirus long non-coding RNAs. Virus Res. 2016, 212, 70–77. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.K.; Kim, J.M.; Song, B.H.; Yun, S.I.; Yun, G.N.; Byun, S.J.; Lee, Y.M. Profiling of viral proteins expressed from the genomic RNA of Japanese encephalitis virus using a panel of 15 region-specific polyclonal rabbit antisera: Implications for viral gene expression. PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [PubMed]
- Lindenbach, B.D.; Murray, C.L.; Thiel, H.J.; Rice, C.M. Flaviviridae. In Fields Virology; Knipe, D.M., Howley, P.M., Cohen, J.I., Griffin, D.E., Lamb, R.A., Martin, M.A., Racaniello, V.R., Roizman, B., Eds.; Wolters Kluwer Health: Philadelphia, PA, USA, 2013. [Google Scholar]
- Mason, P.W.; McAda, P.C.; Dalrymple, J.M.; Fournier, M.J.; Mason, T.L. Expression of Japanese encephalitis virus antigens in Escherichia coli. Virology 1987, 158, 361–372. [Google Scholar] [CrossRef]
- Chambers, T.J.; McCourt, D.W.; Rice, C.M. Production of yellow fever virus proteins in infected cells: Identification of discrete polyprotein species and analysis of cleavage kinetics using region-specific polyclonal antisera. Virology 1990, 177, 159–174. [Google Scholar] [CrossRef]
- Castle, E.; Nowak, T.; Leidner, U.; Wengler, G.; Wengler, G. Sequence analysis of the viral core protein and the membrane-associated proteins V1 and NV2 of the flavivirus West Nile virus and of the genome sequence for these proteins. Virology 1985, 145, 227–236. [Google Scholar] [CrossRef]
- Wengler, G.; Castle, E.; Leidner, U.; Nowak, T.; Wengler, G. Sequence analysis of the membrane protein V3 of the flavivirus West Nile virus and of its gene. Virology 1985, 147, 264–274. [Google Scholar] [CrossRef]
- Lobigs, M. Flavivirus premembrane protein cleavage and spike heterodimer secretion require the function of the viral proteinase NS3. Proc. Natl. Acad. Sci. USA 1993, 90, 6218–6222. [Google Scholar] [CrossRef] [PubMed]
- Stocks, C.E.; Lobigs, M. Signal peptidase cleavage at the flavivirus C-prM junction: Dependence on the viral NS2B-3 protease for efficient processing requires determinants in C, the signal peptide, and prM. J. Virol. 1998, 72, 2141–2149. [Google Scholar] [PubMed]
- Luo, D.; Vasudevan, S.G.; Lescar, J. The flavivirus NS2B-NS3 protease-helicase as a target for antiviral drug development. Antivir. Res. 2015, 118, 148–158. [Google Scholar] [CrossRef] [PubMed]
- Castle, E.; Leidner, U.; Nowak, T.; Wengler, G.; Wengler, G. Primary structure of the West Nile flavivirus genome region coding for all nonstructural proteins. Virology 1986, 149, 10–26. [Google Scholar] [CrossRef]
- Speight, G.; Coia, G.; Parker, M.D.; Westaway, E.G. Gene mapping and positive identification of the non-structural proteins NS2A, NS2B, NS3, NS4B and NS5 of the flavivirus Kunjin and their cleavage sites. J. Gen. Virol. 1988, 69, 23–34. [Google Scholar] [CrossRef] [PubMed]
- Preugschat, F.; Strauss, J.H. Processing of nonstructural proteins NS4A and NS4B of dengue 2 virus In vitro and in vivo. Virology 1991, 185, 689–697. [Google Scholar] [CrossRef]
- Lin, C.; Amberg, S.M.; Chambers, T.J.; Rice, C.M. Cleavage at a novel site in the NS4A region by the yellow fever virus NS2B-3 proteinase is a prerequisite for processing at the downstream 4A/4B signalase site. J. Virol. 1993, 67, 2327–2335. [Google Scholar] [PubMed]
- Amberg, S.M.; Nestorowicz, A.; McCourt, D.W.; Rice, C.M. NS2B-3 proteinase-mediated processing in the yellow fever virus structural region: In vitro and in vivo studies. J. Virol. 1994, 68, 3794–3802. [Google Scholar] [PubMed]
- Yamshchikov, V.F.; Compans, R.W. Processing of the intracellular form of the West Nile virus capsid protein by the viral NS2B-NS3 protease: An In vitro study. J. Virol. 1994, 68, 5765–5771. [Google Scholar] [PubMed]
- Stadler, K.; Allison, S.L.; Schalich, J.; Heinz, F.X. Proteolytic activation of tick-borne encephalitis virus by furin. J. Virol. 1997, 71, 8475–8481. [Google Scholar] [PubMed]
- Falgout, B.; Chanock, R.; Lai, C.J. Proper processing of dengue virus nonstructural glycoprotein NS1 requires the N-terminal hydrophobic signal sequence and the downstream nonstructural protein NS2A. J. Virol. 1989, 63, 1852–1860. [Google Scholar] [PubMed]
- Falgout, B.; Markoff, L. Evidence that flavivirus NS1-NS2A cleavage is mediated by a membrane-bound host protease in the endoplasmic reticulum. J. Virol. 1995, 69, 7232–7243. [Google Scholar] [PubMed]
- Hori, H.; Lai, C.J. Cleavage of dengue virus NS1-NS2A requires an octapeptide sequence at the C terminus of NS1. J. Virol. 1990, 64, 4573–4577. [Google Scholar] [PubMed]
- Firth, A.E.; Atkins, J.F. A conserved predicted pseudoknot in the NS2A-encoding sequence of West Nile and Japanese encephalitis flaviviruses suggests NS1′ may derive from ribosomal frameshifting. Virol. J. 2009, 6. [Google Scholar] [CrossRef] [PubMed]
- Melian, E.B.; Hinzman, E.; Nagasaki, T.; Firth, A.E.; Wills, N.M.; Nouwens, A.S.; Blitvich, B.J.; Leung, J.; Funk, A.; Atkins, J.F.; et al. NS1′ of flaviviruses in the Japanese encephalitis virus serogroup is a product of ribosomal frameshifting and plays a role in viral neuroinvasiveness. J. Virol. 2010, 84, 1641–1647. [Google Scholar] [CrossRef] [PubMed]
- Ye, Q.; Li, X.F.; Zhao, H.; Li, S.H.; Deng, Y.Q.; Cao, R.Y.; Song, K.Y.; Wang, H.J.; Hua, R.H.; Yu, Y.X.; et al. A single nucleotide mutation in NS2A of Japanese encephalitis-live vaccine virus (SA14-14-2) ablates NS1′ formation and contributes to attenuation. J. Gen. Virol. 2012, 93, 1959–1964. [Google Scholar] [CrossRef] [PubMed]
- Yun, S.I.; Song, B.H.; Polejaeva, I.A.; Davies, C.J.; White, K.L.; Lee, Y.M. Comparison of the live-attenuated Japanese encephalitis vaccine SA14-14-2 strain with its pre-attenuated virulent parent SA14 strain: Similarities and differences In vitro and in vivo. J. Gen. Virol. 2016, 97, 2575–2591. [Google Scholar] [CrossRef] [PubMed]
- Yun, S.I.; Kim, S.Y.; Choi, W.Y.; Nam, J.H.; Ju, Y.R.; Park, K.Y.; Cho, H.W.; Lee, Y.M. Molecular characterization of the full-length genome of the Japanese encephalitis viral strain K87P39. Virus Res. 2003, 96, 129–140. [Google Scholar] [CrossRef]
- Yun, S.I.; Kim, S.Y.; Rice, C.M.; Lee, Y.M. Development and application of a reverse genetics system for Japanese encephalitis virus. J. Virol. 2003, 77, 6450–6465. [Google Scholar] [CrossRef] [PubMed]
- Xie, X.; Gayen, S.; Kang, C.; Yuan, Z.; Shi, P.Y. Membrane topology and function of dengue virus NS2A protein. J. Virol. 2013, 87, 4609–4622. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Li, Q.; Wong, Y.L.; Liew, L.S.; Kang, C. Membrane topology of NS2B of dengue virus revealed by NMR spectroscopy. Biochim. Biophys. Acta 2015, 1848, 2244–2252. [Google Scholar] [CrossRef] [PubMed]
- Zou, J.; Xie, X.; Wang, Q.Y.; Dong, H.; Lee, M.Y.; Kang, C.; Yuan, Z.; Shi, P.Y. Characterization of dengue virus NS4A and NS4B protein interaction. J. Virol. 2015, 89, 3455–3470. [Google Scholar] [CrossRef] [PubMed]
- Miller, S.; Sparacio, S.; Bartenschlager, R. Subcellular localization and membrane topology of the dengue virus type 2 non-structural protein 4B. J. Biol. Chem. 2006, 281, 8854–8863. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, E.R.A.; Mohana-Borges, R.; de Alencastro, R.B.; Horta, B.A.C. The flavivirus capsid protein: Structure, function and perspectives towards drug design. Virus Res. 2017, 227, 115–123. [Google Scholar] [CrossRef] [PubMed]
- Dokland, T.; Walsh, M.; Mackenzie, J.M.; Khromykh, A.A.; Ee, K.H.; Wang, S. West Nile virus core protein; tetramer structure and ribbon formation. Structure 2004, 12, 1157–1163. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Jones, C.T.; Groesch, T.D.; Kuhn, R.J.; Post, C.B. Solution structure of dengue virus capsid protein reveals another fold. Proc. Natl. Acad. Sci. USA 2004, 101, 3414–3419. [Google Scholar] [CrossRef] [PubMed]
- Shang, Z.; Song, H.; Shi, Y.; Qi, J.; Gao, G.F. Crystal structure of the capsid protein from Zika virus. J. Mol. Biol. 2018, 430, 948–962. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, R.J.; Zhang, W.; Rossmann, M.G.; Pletnev, S.V.; Corver, J.; Lenches, E.; Jones, C.T.; Mukhopadhyay, S.; Chipman, P.R.; Strauss, E.G.; et al. Structure of dengue virus: Implications for flavivirus organization, maturation, and fusion. Cell 2002, 108, 717–725. [Google Scholar] [CrossRef]
- Mukhopadhyay, S.; Kim, B.S.; Chipman, P.R.; Rossmann, M.G.; Kuhn, R.J. Structure of West Nile virus. Science 2003, 302. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Li, S.H.; Zhu, L.; Nian, Q.G.; Yuan, S.; Gao, Q.; Hu, Z.; Ye, Q.; Li, X.F.; Xie, D.Y.; et al. Near-atomic structure of Japanese encephalitis virus reveals critical determinants of virulence and stability. Nat. Commun. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Kostyuchenko, V.A.; Lim, E.X.; Zhang, S.; Fibriansah, G.; Ng, T.S.; Ooi, J.S.; Shi, J.; Lok, S.M. Structure of the thermally stable Zika virus. Nature 2016, 533, 425–428. [Google Scholar] [CrossRef] [PubMed]
- Sirohi, D.; Chen, Z.; Sun, L.; Klose, T.; Pierson, T.C.; Rossmann, M.G.; Kuhn, R.J. The 3.8 A resolution cryo-EM structure of Zika virus. Science 2016, 352, 467–470. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Ge, P.; Yu, X.; Brannan, J.M.; Bi, G.; Zhang, Q.; Schein, S.; Zhou, Z.H. Cryo-EM structure of the mature dengue virus at 3.5-A resolution. Nat. Struct. Mol. Biol. 2013, 20, 105–110. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Maguire, T.; Hileman, R.E.; Fromm, J.R.; Esko, J.D.; Linhardt, R.J.; Marks, R.M. Dengue virus infectivity depends on envelope protein binding to target cell heparan sulfate. Nat. Med. 1997, 3, 866–871. [Google Scholar] [CrossRef] [PubMed]
- Davis, C.W.; Nguyen, H.Y.; Hanna, S.L.; Sanchez, M.D.; Doms, R.W.; Pierson, T.C. West Nile virus discriminates between DC-SIGN and DC-SIGNR for cellular attachment and infection. J. Virol. 2006, 80, 1290–1301. [Google Scholar] [CrossRef] [PubMed]
- Navarro-Sanchez, E.; Altmeyer, R.; Amara, A.; Schwartz, O.; Fieschi, F.; Virelizier, J.L.; Arenzana-Seisdedos, F.; Despres, P. Dendritic-cell-specific ICAM3-grabbing non-integrin is essential for the productive infection of human dendritic cells by mosquito-cell-derived dengue viruses. EMBO Rep. 2003, 4, 723–728. [Google Scholar] [CrossRef] [PubMed]
- Pokidysheva, E.; Zhang, Y.; Battisti, A.J.; Bator-Kelly, C.M.; Chipman, P.R.; Xiao, C.; Gregorio, G.G.; Hendrickson, W.A.; Kuhn, R.J.; Rossmann, M.G. Cryo-EM reconstruction of dengue virus in complex with the carbohydrate recognition domain of DC-SIGN. Cell 2006, 124, 485–493. [Google Scholar] [CrossRef] [PubMed]
- Tassaneetrithep, B.; Burgess, T.H.; Granelli-Piperno, A.; Trumpfheller, C.; Finke, J.; Sun, W.; Eller, M.A.; Pattanapanyasat, K.; Sarasombath, S.; Birx, D.L.; et al. DC-SIGN (CD209) mediates dengue virus infection of human dendritic cells. J. Exp. Med. 2003, 197, 823–829. [Google Scholar] [CrossRef] [PubMed]
- Hamel, R.; Dejarnac, O.; Wichit, S.; Ekchariyawat, P.; Neyret, A.; Luplertlop, N.; Perera-Lecoin, M.; Surasombatpattana, P.; Talignani, L.; Thomas, F.; et al. Biology of Zika virus infection in human skin cells. J. Virol. 2015, 89, 8880–8896. [Google Scholar] [CrossRef] [PubMed]
- Perera-Lecoin, M.; Meertens, L.; Carnec, X.; Amara, A. Flavivirus entry receptors: An update. Viruses 2013, 6, 69–88. [Google Scholar] [CrossRef] [PubMed]
- Pierson, T.C.; Kielian, M. Flaviviruses: Braking the entering. Curr. Opin. Virol. 2013, 3, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Nain, M.; Abdin, M.Z.; Kalia, M.; Vrati, S. Japanese encephalitis virus invasion of cell: Allies and alleys. Rev. Med. Virol. 2016, 26, 129–141. [Google Scholar] [CrossRef] [PubMed]
- Acosta, E.G.; Castilla, V.; Damonte, E.B. Functional entry of dengue virus into Aedes albopictus mosquito cells is dependent on clathrin-mediated endocytosis. J. Gen. Virol. 2008, 89, 474–484. [Google Scholar] [CrossRef] [PubMed]
- Chu, J.J.; Leong, P.W.; Ng, M.L. Analysis of the endocytic pathway mediating the infectious entry of mosquito-borne flavivirus West Nile into Aedes albopictus mosquito (C6/36) cells. Virology 2006, 349, 463–475. [Google Scholar] [CrossRef] [PubMed]
- Chu, J.J.; Ng, M.L. Infectious entry of West Nile virus occurs through a clathrin-mediated endocytic pathway. J. Virol. 2004, 78, 10543–10555. [Google Scholar] [CrossRef] [PubMed]
- Mosso, C.; Galvan-Mendoza, I.J.; Ludert, J.E.; del Angel, R.M. Endocytic pathway followed by dengue virus to infect the mosquito cell line C6/36 HT. Virology 2008, 378, 193–199. [Google Scholar] [CrossRef] [PubMed]
- Van der Schaar, H.M.; Rust, M.J.; Chen, C.; van der Ende-Metselaar, H.; Wilschut, J.; Zhuang, X.; Smit, J.M. Dissecting the cell entry pathway of dengue virus by single-particle tracking in living cells. PLoS Pathog. 2008, 4. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.C.; Zhang, Y.N.; Li, Z.Y.; Hou, J.X.; Zhou, J.; Kan, L.; Zhou, B.; Chen, P.Y. Rab5 and Rab11 are required for clathrin-dependent endocytosis of Japanese encephalitis virus in BHK-21 cells. J. Virol. 2017, 91, e01113–e01117. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; He, M.; Liu, X.; Li, X.; Fan, B.; Zhao, S. Japanese encephalitis virus infects porcine kidney epithelial PK15 cells via clathrin- and cholesterol-dependent endocytosis. Virol. J. 2013, 10. [Google Scholar] [CrossRef] [PubMed]
- Chuang, C.K.; Yang, T.H.; Chen, T.H.; Yang, C.F.; Chen, W.J. Heat shock cognate protein 70 isoform D is required for clathrin-dependent endocytosis of Japanese encephalitis virus in C6/36 cells. J. Gen. Virol. 2015, 96, 793–803. [Google Scholar] [CrossRef] [PubMed]
- Das, S.; Chakraborty, S.; Basu, A. Critical role of lipid rafts in virus entry and activation of phosphoinositide 3′ kinase/Akt signaling during early stages of Japanese encephalitis virus infection in neural stem/progenitor cells. J. Neurochem. 2010, 115, 537–549. [Google Scholar] [CrossRef] [PubMed]
- Nawa, M.; Takasaki, T.; Yamada, K.; Kurane, I.; Akatsuka, T. Interference in Japanese encephalitis virus infection of Vero cells by a cationic amphiphilic drug, chlorpromazine. J. Gen. Virol. 2003, 84, 1737–1741. [Google Scholar] [CrossRef] [PubMed]
- Ang, F.; Wong, A.P.; Ng, M.M.; Chu, J.J. Small interference RNA profiling reveals the essential role of human membrane trafficking genes in mediating the infectious entry of dengue virus. Virol. J. 2010, 7. [Google Scholar] [CrossRef] [PubMed]
- Suksanpaisan, L.; Susantad, T.; Smith, D.R. Characterization of dengue virus entry into HepG2 cells. J. Biomed. Sci. 2009, 16. [Google Scholar] [CrossRef] [PubMed]
- Kalia, M.; Khasa, R.; Sharma, M.; Nain, M.; Vrati, S. Japanese encephalitis virus infects neuronal cells through a clathrin-independent endocytic mechanism. J. Virol. 2013, 87, 148–162. [Google Scholar] [CrossRef] [PubMed]
- Xu, Q.; Cao, M.; Song, H.; Chen, S.; Qian, X.; Zhao, P.; Ren, H.; Tang, H.; Wang, Y.; Wei, Y.; et al. Caveolin-1-mediated Japanese encephalitis virus entry requires a two-step regulation of actin reorganization. Future Microbiol. 2016, 11, 1227–1248. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.Z.; Xu, Q.Q.; Wu, D.G.; Ren, H.; Zhao, P.; Lao, W.G.; Wang, Y.; Tao, Q.Y.; Qian, X.J.; Wei, Y.H.; et al. Japanese encephalitis virus enters rat neuroblastoma cells via a pH-dependent, dynamin and caveola-mediated endocytosis pathway. J. Virol. 2012, 86, 13407–13422. [Google Scholar] [CrossRef] [PubMed]
- Piccini, L.E.; Castilla, V.; Damonte, E.B. Dengue-3 virus entry into Vero cells: Role of clathrin-mediated endocytosis in the outcome of infection. PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [PubMed]
- Acosta, E.G.; Castilla, V.; Damonte, E.B. Differential requirements in endocytic trafficking for penetration of dengue virus. PLoS ONE 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Acosta, E.G.; Piccini, L.E.; Talarico, L.B.; Castilla, V.; Damonte, E.B. Changes in antiviral susceptibility to entry inhibitors and endocytic uptake of dengue-2 virus serially passaged in Vero or C6/36 cells. Virus Res. 2014, 184, 39–43. [Google Scholar] [CrossRef] [PubMed]
- Allison, S.L.; Schalich, J.; Stiasny, K.; Mandl, C.W.; Kunz, C.; Heinz, F.X. Oligomeric rearrangement of tick-borne encephalitis virus envelope proteins induced by an acidic pH. J. Virol. 1995, 69, 695–700. [Google Scholar] [PubMed]
- Bressanelli, S.; Stiasny, K.; Allison, S.L.; Stura, E.A.; Duquerroy, S.; Lescar, J.; Heinz, F.X.; Rey, F.A. Structure of a flavivirus envelope glycoprotein in its low-pH-induced membrane fusion conformation. EMBO J. 2004, 23, 728–738. [Google Scholar] [CrossRef] [PubMed]
- Liao, M.; Sanchez-San Martin, C.; Zheng, A.; Kielian, M. In vitro reconstitution reveals key intermediate states of trimer formation by the dengue virus membrane fusion protein. J. Virol. 2010, 84, 5730–5740. [Google Scholar] [CrossRef] [PubMed]
- Modis, Y.; Ogata, S.; Clements, D.; Harrison, S.C. Structure of the dengue virus envelope protein after membrane fusion. Nature 2004, 427, 313–319. [Google Scholar] [CrossRef] [PubMed]
- Nayak, V.; Dessau, M.; Kucera, K.; Anthony, K.; Ledizet, M.; Modis, Y. Crystal structure of dengue virus type 1 envelope protein in the postfusion conformation and its implications for membrane fusion. J. Virol. 2009, 83, 4338–4344. [Google Scholar] [CrossRef] [PubMed]
- Stiasny, K.; Allison, S.L.; Schalich, J.; Heinz, F.X. Membrane interactions of the tick-borne encephalitis virus fusion protein E at low pH. J. Virol. 2002, 76, 3784–3790. [Google Scholar] [CrossRef] [PubMed]
- Stiasny, K.; Kossl, C.; Lepault, J.; Rey, F.A.; Heinz, F.X. Characterization of a structural intermediate of flavivirus membrane fusion. PLoS Pathog. 2007, 3. [Google Scholar] [CrossRef] [PubMed]
- Corver, J.; Ortiz, A.; Allison, S.L.; Schalich, J.; Heinz, F.X.; Wilschut, J. Membrane fusion activity of tick-borne encephalitis virus and recombinant subviral particles in a liposomal model system. Virology 2000, 269, 37–46. [Google Scholar] [CrossRef] [PubMed]
- Despres, P.; Frenkiel, M.P.; Deubel, V. Differences between cell membrane fusion activities of two dengue type-1 isolates reflect modifications of viral structure. Virology 1993, 196, 209–219. [Google Scholar] [CrossRef] [PubMed]
- Gollins, S.W.; Porterfield, J.S. pH-dependent fusion between the flavivirus West Nile and liposomal model membranes. J. Gen. Virol. 1986, 67, 157–166. [Google Scholar] [CrossRef] [PubMed]
- Guirakhoo, F.; Hunt, A.R.; Lewis, J.G.; Roehrig, J.T. Selection and partial characterization of dengue 2 virus mutants that induce fusion at elevated pH. Virology 1993, 194, 219–223. [Google Scholar] [CrossRef] [PubMed]
- McMinn, P.C.; Weir, R.C.; Dalgarno, L. A mouse-attenuated envelope protein variant of Murray Valley encephalitis virus with altered fusion activity. J. Gen. Virol. 1996, 77, 2085–2088. [Google Scholar] [CrossRef] [PubMed]
- Randolph, V.B.; Stollar, V. Low pH-induced cell fusion in flavivirus-infected Aedes albopictus cell cultures. J. Gen. Virol. 1990, 71, 1845–1850. [Google Scholar] [CrossRef] [PubMed]
- Stiasny, K.; Koessl, C.; Heinz, F.X. Involvement of lipids in different steps of the flavivirus fusion mechanism. J. Virol. 2003, 77, 7856–7862. [Google Scholar] [CrossRef] [PubMed]
- Summers, P.L.; Cohen, W.H.; Ruiz, M.M.; Hase, T.; Eckels, K.H. Flaviviruses can mediate fusion from without in Aedes albopictus mosquito cell cultures. Virus Res. 1989, 12, 383–392. [Google Scholar] [CrossRef]
- Fernandez-Garcia, M.D.; Mazzon, M.; Jacobs, M.; Amara, A. Pathogenesis of flavivirus infections: Using and abusing the host cell. Cell Host Microbe 2009, 5, 318–328. [Google Scholar] [CrossRef] [PubMed]
- Gillespie, L.K.; Hoenen, A.; Morgan, G.; Mackenzie, J.M. The endoplasmic reticulum provides the membrane platform for biogenesis of the flavivirus replication complex. J. Virol. 2010, 84, 10438–10447. [Google Scholar] [CrossRef] [PubMed]
- Hsu, N.Y.; Ilnytska, O.; Belov, G.; Santiana, M.; Chen, Y.H.; Takvorian, P.M.; Pau, C.; van der Schaar, H.; Kaushik-Basu, N.; Balla, T.; et al. Viral reorganization of the secretory pathway generates distinct organelles for RNA replication. Cell 2010, 141, 799–811. [Google Scholar] [CrossRef] [PubMed]
- Welsch, S.; Miller, S.; Romero-Brey, I.; Merz, A.; Bleck, C.K.; Walther, P.; Fuller, S.D.; Antony, C.; Krijnse-Locker, J.; Bartenschlager, R. Composition and three-dimensional architecture of the dengue virus replication and assembly sites. Cell Host Microbe 2009, 5, 365–375. [Google Scholar] [CrossRef] [PubMed]
- Neufeldt, C.J.; Cortese, M.; Acosta, E.G.; Bartenschlager, R. Rewiring cellular networks by members of the Flaviviridae family. Nat. Rev. Microbiol. 2018, 16, 125–142. [Google Scholar] [CrossRef] [PubMed]
- Harak, C.; Lohmann, V. Ultrastructure of the replication sites of positive-strand RNA viruses. Virology 2015, 479–480, 418–433. [Google Scholar] [CrossRef] [PubMed]
- Brand, C.; Bisaillon, M.; Geiss, B.J. Organization of the flavivirus RNA replicase complex. Wiley Interdiscip. Rev. RNA 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Bollati, M.; Alvarez, K.; Assenberg, R.; Baronti, C.; Canard, B.; Cook, S.; Coutard, B.; Decroly, E.; de Lamballerie, X.; Gould, E.A.; et al. Structure and functionality in flavivirus NS-proteins: Perspectives for drug design. Antivir. Res. 2010, 87, 125–148. [Google Scholar] [CrossRef] [PubMed]
- Selisko, B.; Wang, C.; Harris, E.; Canard, B. Regulation of flavivirus RNA synthesis and replication. Curr. Opin. Virol. 2014, 9, 74–83. [Google Scholar] [CrossRef] [PubMed]
- Konishi, E.; Mason, P.W. Proper maturation of the Japanese encephalitis virus envelope glycoprotein requires cosynthesis with the premembrane protein. J. Virol. 1993, 67, 1672–1675. [Google Scholar] [PubMed]
- Li, L.; Lok, S.M.; Yu, I.M.; Zhang, Y.; Kuhn, R.J.; Chen, J.; Rossmann, M.G. The flavivirus precursor membrane-envelope protein complex: Structure and maturation. Science 2008, 319, 1830–1834. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Corver, J.; Chipman, P.R.; Zhang, W.; Pletnev, S.V.; Sedlak, D.; Baker, T.S.; Strauss, J.H.; Kuhn, R.J.; Rossmann, M.G. Structures of immature flavivirus particles. EMBO J. 2003, 22, 2604–2613. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Kaufmann, B.; Chipman, P.R.; Kuhn, R.J.; Rossmann, M.G. Structure of immature West Nile virus. J. Virol. 2007, 81, 6141–6145. [Google Scholar] [CrossRef] [PubMed]
- Lorenz, I.C.; Allison, S.L.; Heinz, F.X.; Helenius, A. Folding and dimerization of tick-borne encephalitis virus envelope proteins prM and E in the endoplasmic reticulum. J. Virol. 2002, 76, 5480–5491. [Google Scholar] [CrossRef] [PubMed]
- Elshuber, S.; Allison, S.L.; Heinz, F.X.; Mandl, C.W. Cleavage of protein prM is necessary for infection of BHK-21 cells by tick-borne encephalitis virus. J. Gen. Virol. 2003, 84, 183–191. [Google Scholar] [CrossRef] [PubMed]
- Guirakhoo, F.; Heinz, F.X.; Mandl, C.W.; Holzmann, H.; Kunz, C. Fusion activity of flaviviruses: Comparison of mature and immature (prM-containing) tick-borne encephalitis virions. J. Gen. Virol. 1991, 72, 1323–1329. [Google Scholar] [CrossRef] [PubMed]
- Yu, I.M.; Holdaway, H.A.; Chipman, P.R.; Kuhn, R.J.; Rossmann, M.G.; Chen, J. Association of the pr peptides with dengue virus at acidic pH blocks membrane fusion. J. Virol. 2009, 83, 12101–12107. [Google Scholar] [CrossRef] [PubMed]
- Yu, I.M.; Zhang, W.; Holdaway, H.A.; Li, L.; Kostyuchenko, V.A.; Chipman, P.R.; Kuhn, R.J.; Rossmann, M.G.; Chen, J. Structure of the immature dengue virus at low pH primes proteolytic maturation. Science 2008, 319, 1834–1837. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, W.; Ogata, S.; Clements, D.; Strauss, J.H.; Baker, T.S.; Kuhn, R.J.; Rossmann, M.G. Conformational changes of the flavivirus E glycoprotein. Structure 2004, 12, 1607–1618. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, S.; Dowd, K.A.; Manhart, C.J.; Ledgerwood, J.E.; Durbin, A.P.; Whitehead, S.S.; Pierson, T.C. Mechanism and significance of cell type-dependent neutralization of flaviviruses. J. Virol. 2014, 88, 7210–7220. [Google Scholar] [CrossRef] [PubMed]
- Nelson, S.; Jost, C.A.; Xu, Q.; Ess, J.; Martin, J.E.; Oliphant, T.; Whitehead, S.S.; Durbin, A.P.; Graham, B.S.; Diamond, M.S.; et al. Maturation of West Nile virus modulates sensitivity to antibody-mediated neutralization. PLoS Pathog. 2008, 4. [Google Scholar] [CrossRef] [PubMed]
- Goo, L.; DeMaso, C.R.; Pelc, R.S.; Ledgerwood, J.E.; Graham, B.S.; Kuhn, R.J.; Pierson, T.C. The Zika virus envelope protein glycan loop regulates virion antigenicity. Virology 2018, 515, 191–202. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, S.; Lin, T.Y.; Dowd, K.A.; Manhart, C.J.; Pierson, T.C. The infectivity of prM-containing partially mature West Nile virus does not require the activity of cellular furin-like proteases. J. Virol. 2011, 85, 12067–12072. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, S.C.; Wu, Y.C.; Zou, G.; Nerurkar, V.R.; Shi, P.Y.; Wang, W.K. Highly conserved residues in the helical domain of dengue virus type 1 precursor membrane protein are involved in assembly, precursor membrane (prM) protein cleavage, and entry. J. Biol. Chem. 2014, 289, 33149–33160. [Google Scholar] [CrossRef] [PubMed]
- Mori, Y.; Okabayashi, T.; Yamashita, T.; Zhao, Z.; Wakita, T.; Yasui, K.; Hasebe, F.; Tadano, M.; Konishi, E.; Moriishi, K.; et al. Nuclear localization of Japanese encephalitis virus core protein enhances viral replication. J. Virol. 2005, 79, 3448–3458. [Google Scholar] [CrossRef] [PubMed]
- Sangiambut, S.; Keelapang, P.; Aaskov, J.; Puttikhunt, C.; Kasinrerk, W.; Malasit, P.; Sittisombut, N. Multiple regions in dengue virus capsid protein contribute to nuclear localization during virus infection. J. Gen. Virol. 2008, 89, 1254–1264. [Google Scholar] [CrossRef] [PubMed]
- Netsawang, J.; Noisakran, S.; Puttikhunt, C.; Kasinrerk, W.; Wongwiwat, W.; Malasit, P.; Yenchitsomanus, P.T.; Limjindaporn, T. Nuclear localization of dengue virus capsid protein is required for DAXX interaction and apoptosis. Virus Res. 2010, 147, 275–283. [Google Scholar] [CrossRef] [PubMed]
- Bhuvanakantham, R.; Cheong, Y.K.; Ng, M.L. West Nile virus capsid protein interaction with importin and HDM2 protein is regulated by protein kinase C-mediated phosphorylation. Microbes Infect. 2010, 12, 615–625. [Google Scholar] [CrossRef] [PubMed]
- Bhuvanakantham, R.; Chong, M.K.; Ng, M.L. Specific interaction of capsid protein and importin-α/β influences West Nile virus production. Biochem. Biophys. Res. Commun. 2009, 389, 63–69. [Google Scholar] [CrossRef] [PubMed]
- Uchil, P.D.; Kumar, A.V.; Satchidanandam, V. Nuclear localization of flavivirus RNA synthesis in infected cells. J. Virol. 2006, 80, 5451–5464. [Google Scholar] [CrossRef] [PubMed]
- Buckley, A.; Gaidamovich, S.; Turchinskaya, A.; Gould, E.A. Monoclonal antibodies identify the NS5 yellow fever virus non-structural protein in the nuclei of infected cells. J. Gen. Virol. 1992, 73, 1125–1130. [Google Scholar] [CrossRef] [PubMed]
- Forwood, J.K.; Brooks, A.; Briggs, L.J.; Xiao, C.Y.; Jans, D.A.; Vasudevan, S.G. The 37-amino-acid interdomain of dengue virus NS5 protein contains a functional NLS and inhibitory CK2 site. Biochem. Biophys. Res. Commun. 1999, 257, 731–737. [Google Scholar] [CrossRef] [PubMed]
- Tay, M.Y.; Smith, K.; Ng, I.H.; Chan, K.W.; Zhao, Y.; Ooi, E.E.; Lescar, J.; Luo, D.; Jans, D.A.; Forwood, J.K.; et al. The C-terminal 18 amino acid region of dengue virus NS5 regulates its subcellular localization and contains a conserved arginine residue essential for infectious virus production. PLoS Pathog. 2016, 12. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Buhler, S.; Selisko, B.; Davidson, A.; Mulder, K.; Canard, B.; Miller, S.; Bartenschlager, R. Nuclear localization of dengue virus nonstructural protein 5 does not strictly correlate with efficient viral RNA replication and inhibition of type I interferon signaling. J. Virol. 2013, 87, 4545–4557. [Google Scholar] [CrossRef] [PubMed]
- Pryor, M.J.; Rawlinson, S.M.; Butcher, R.E.; Barton, C.L.; Waterhouse, T.A.; Vasudevan, S.G.; Bardin, P.G.; Wright, P.J.; Jans, D.A.; Davidson, A.D. Nuclear localization of dengue virus nonstructural protein 5 through its importin α/β-recognized nuclear localization sequences is integral to viral infection. Traffic 2007, 8, 795–807. [Google Scholar] [CrossRef] [PubMed]
- Brooks, A.J.; Johansson, M.; John, A.V.; Xu, Y.; Jans, D.A.; Vasudevan, S.G. The interdomain region of dengue NS5 protein that binds to the viral helicase NS3 contains independently functional importin β1 and importin α/β-recognized nuclear localization signals. J. Biol. Chem. 2002, 277, 36399–36407. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Denman, A.J.; Mackenzie, J.M. The IMPORTance of the nucleus during flavivirus replication. Viruses 2017, 9, 14. [Google Scholar] [CrossRef] [PubMed]
- Marsh, M.; Helenius, A. Virus entry: Open sesame. Cell 2006, 124, 729–740. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.E.; Helenius, A. How viruses enter animal cells. Science 2004, 304, 237–242. [Google Scholar] [CrossRef] [PubMed]
- Schweighardt, B.; Atwood, W.J. Virus receptors in the human central nervous system. J. Neurovirol. 2001, 7, 187–195. [Google Scholar] [PubMed]
- Altmeyer, R. Virus attachment and entry offer numerous targets for antiviral therapy. Curr. Pharm. Des. 2004, 10, 3701–3712. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.Y.; Shi, P.Y. Flavivirus entry inhibitors. ACS Infect. Dis. 2015, 1, 428–434. [Google Scholar] [CrossRef] [PubMed]
- Allison, S.L.; Stiasny, K.; Stadler, K.; Mandl, C.W.; Heinz, F.X. Mapping of functional elements in the stem-anchor region of tick-borne encephalitis virus envelope protein E. J. Virol. 1999, 73, 5605–5612. [Google Scholar] [PubMed]
- Stiasny, K.; Allison, S.L.; Marchler-Bauer, A.; Kunz, C.; Heinz, F.X. Structural requirements for low-pH-induced rearrangements in the envelope glycoprotein of tick-borne encephalitis virus. J. Virol. 1996, 70, 8142–8147. [Google Scholar] [PubMed]
- Zhang, W.; Chipman, P.R.; Corver, J.; Johnson, P.R.; Zhang, Y.; Mukhopadhyay, S.; Baker, T.S.; Strauss, J.H.; Rossmann, M.G.; Kuhn, R.J. Visualization of membrane protein domains by cryo-electron microscopy of dengue virus. Nat. Struct. Biol. 2003, 10, 907–912. [Google Scholar] [CrossRef] [PubMed]
- Allison, S.L.; Schalich, J.; Stiasny, K.; Mandl, C.W.; Heinz, F.X. Mutational evidence for an internal fusion peptide in flavivirus envelope protein E. J. Virol. 2001, 75, 4268–4275. [Google Scholar] [CrossRef] [PubMed]
- Beasley, D.W.; Barrett, A.D. Identification of neutralizing epitopes within structural domain III of the West Nile virus envelope protein. J. Virol. 2002, 76, 13097–13100. [Google Scholar] [CrossRef] [PubMed]
- Bhardwaj, S.; Holbrook, M.; Shope, R.E.; Barrett, A.D.; Watowich, S.J. Biophysical characterization and vector-specific antagonist activity of domain III of the tick-borne flavivirus envelope protein. J. Virol. 2001, 75, 4002–4007. [Google Scholar] [CrossRef] [PubMed]
- Chu, J.J.; Rajamanonmani, R.; Li, J.; Bhuvanakantham, R.; Lescar, J.; Ng, M.L. Inhibition of West Nile virus entry by using a recombinant domain III from the envelope glycoprotein. J. Gen. Virol. 2005, 86, 405–412. [Google Scholar] [CrossRef] [PubMed]
- Crill, W.D.; Roehrig, J.T. Monoclonal antibodies that bind to domain III of dengue virus E glycoprotein are the most efficient blockers of virus adsorption to Vero cells. J. Virol. 2001, 75, 7769–7773. [Google Scholar] [CrossRef] [PubMed]
- Halstead, S.B.; Heinz, F.X.; Barrett, A.D.; Roehrig, J.T. Dengue virus: Molecular basis of cell entry and pathogenesis, 25–27 June 2003, Vienna, Austria. Vaccine 2005, 23, 849–856. [Google Scholar] [CrossRef] [PubMed]
- Kaufmann, B.; Nybakken, G.E.; Chipman, P.R.; Zhang, W.; Diamond, M.S.; Fremont, D.H.; Kuhn, R.J.; Rossmann, M.G. West Nile virus in complex with the Fab fragment of a neutralizing monoclonal antibody. Proc. Natl. Acad. Sci. USA 2006, 103, 12400–12404. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.; Lobigs, M. Mechanism of virulence attenuation of glycosaminoglycan-binding variants of Japanese encephalitis virus and Murray Valley encephalitis virus. J. Virol. 2002, 76, 4901–4911. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.W.; Chu, J.J.; Ng, M.L. Quantifying the specific binding between West Nile virus envelope domain III protein and the cellular receptor αvβ3 integrin. J. Biol. Chem. 2006, 281, 1352–1360. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Barrett, A.D.; Beasley, D.W. Differential expression of domain III neutralizing epitopes on the envelope proteins of West Nile virus strains. Virology 2005, 335, 99–105. [Google Scholar] [CrossRef] [PubMed]
- Pierson, T.C.; Xu, Q.; Nelson, S.; Oliphant, T.; Nybakken, G.E.; Fremont, D.H.; Diamond, M.S. The stoichiometry of antibody-mediated neutralization and enhancement of West Nile virus infection. Cell Host Microbe 2007, 1, 135–145. [Google Scholar] [CrossRef] [PubMed]
- Rey, F.A.; Heinz, F.X.; Mandl, C.; Kunz, C.; Harrison, S.C. The envelope glycoprotein from tick-borne encephalitis virus at 2 A resolution. Nature 1995, 375, 291–298. [Google Scholar] [CrossRef] [PubMed]
- Stiasny, K.; Kiermayr, S.; Holzmann, H.; Heinz, F.X. Cryptic properties of a cluster of dominant flavivirus cross-reactive antigenic sites. J. Virol. 2006, 80, 9557–9568. [Google Scholar] [CrossRef] [PubMed]
- Sukupolvi-Petty, S.; Austin, S.K.; Purtha, W.E.; Oliphant, T.; Nybakken, G.E.; Schlesinger, J.J.; Roehrig, J.T.; Gromowski, G.D.; Barrett, A.D.; Fremont, D.H.; et al. Type- and subcomplex-specific neutralizing antibodies against domain III of dengue virus type 2 envelope protein recognize adjacent epitopes. J. Virol. 2007, 81, 12816–12826. [Google Scholar] [CrossRef] [PubMed]
- Wu, K.P.; Wu, C.W.; Tsao, Y.P.; Kuo, T.W.; Lou, Y.C.; Lin, C.W.; Wu, S.C.; Cheng, J.W. Structural basis of a flavivirus recognized by its neutralizing antibody: Solution structure of the domain III of the Japanese encephalitis virus envelope protein. J. Biol. Chem. 2003, 278, 46007–46013. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.Y.; Butrapet, S.; Moss, K.J.; Childers, T.; Erb, S.M.; Calvert, A.E.; Silengo, S.J.; Kinney, R.M.; Blair, C.D.; Roehrig, J.T. The dengue virus type 2 envelope protein fusion peptide is essential for membrane fusion. Virology 2010, 396, 305–315. [Google Scholar] [CrossRef] [PubMed]
- Beltramello, M.; Williams, K.L.; Simmons, C.P.; Macagno, A.; Simonelli, L.; Quyen, N.T.; Sukupolvi-Petty, S.; Navarro-Sanchez, E.; Young, P.R.; de Silva, A.M.; et al. The human immune response to dengue virus is dominated by highly cross-reactive antibodies endowed with neutralizing and enhancing activity. Cell Host Microbe 2010, 8, 271–283. [Google Scholar] [CrossRef] [PubMed]
- Chiou, S.S.; Fan, Y.C.; Crill, W.D.; Chang, R.Y.; Chang, G.J. Mutation analysis of the cross-reactive epitopes of Japanese encephalitis virus envelope glycoprotein. J. Gen. Virol. 2012, 93, 1185–1192. [Google Scholar] [CrossRef] [PubMed]
- Fan, W.; Qian, P.; Wang, D.; Zhi, X.; Wei, Y.; Chen, H.; Li, X. Integrin αvβ3 promotes infection by Japanese encephalitis virus. Res. Vet. Sci. 2017, 111, 67–74. [Google Scholar] [CrossRef] [PubMed]
- Shimojima, M.; Takenouchi, A.; Shimoda, H.; Kimura, N.; Maeda, K. Distinct usage of three C-type lectins by Japanese encephalitis virus: DC-SIGN, DC-SIGNR, and LSECtin. Arch. Virol. 2014, 159, 2023–2031. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Hu, K.; Luo, S.; Zhang, M.; Deng, X.; Li, C.; Jin, W.; Hu, B.; He, S.; Li, M.; et al. DC-SIGN as an attachment factor mediates Japanese encephalitis virus infection of human dendritic cells via interaction with a single high-mannose residue of viral E glycoprotein. Virology 2016, 488, 108–119. [Google Scholar] [CrossRef] [PubMed]
- Chu, J.J.; Ng, M.L. Interaction of West Nile virus with αvβ3 integrin mediates virus entry into cells. J. Biol. Chem. 2004, 279, 54533–54541. [Google Scholar] [CrossRef] [PubMed]
- Hurrelbrink, R.J.; McMinn, P.C. Attenuation of Murray Valley encephalitis virus by site-directed mutagenesis of the hinge and putative receptor-binding regions of the envelope protein. J. Virol. 2001, 75, 7692–7702. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.; Lobigs, M. Substitutions at the putative receptor-binding site of an encephalitic flavivirus alter virulence and host cell tropism and reveal a role for glycosaminoglycans in entry. J. Virol. 2000, 74, 8867–8875. [Google Scholar] [CrossRef] [PubMed]
- Lobigs, M.; Usha, R.; Nestorowicz, A.; Marshall, I.D.; Weir, R.C.; Dalgarno, L. Host cell selection of Murray Valley encephalitis virus variants altered at an RGD sequence in the envelope protein and in mouse virulence. Virology 1990, 176, 587–595. [Google Scholar] [CrossRef]
- Van der Most, R.G.; Corver, J.; Strauss, J.H. Mutagenesis of the RGD motif in the yellow fever virus 17D envelope protein. Virology 1999, 265, 83–95. [Google Scholar] [CrossRef] [PubMed]
- Barba-Spaeth, G.; Longman, R.S.; Albert, M.L.; Rice, C.M. Live attenuated yellow fever 17D infects human DCs and allows for presentation of endogenous and recombinant T cell epitopes. J. Exp. Med. 2005, 202, 1179–1184. [Google Scholar] [CrossRef] [PubMed]
- Fontes-Garfias, C.R.; Shan, C.; Luo, H.; Muruato, A.E.; Medeiros, D.B.A.; Mays, E.; Xie, X.; Zou, J.; Roundy, C.M.; Wakamiya, M.; et al. Functional analysis of glycosylation of Zika virus envelope protein. Cell Rep. 2017, 21, 1180–1190. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Liu, Y.; Wang, S.; Zhang, Y.; Zu, X.; Zhou, Z.; Zhang, B.; Xiao, G. Structure-based mutational analysis of several sites in the E protein: Implications for understanding the entry mechanism of Japanese encephalitis virus. J. Virol. 2015, 89, 5668–5686. [Google Scholar] [CrossRef] [PubMed]
- Luca, V.C.; AbiMansour, J.; Nelson, C.A.; Fremont, D.H. Crystal structure of the Japanese encephalitis virus envelope protein. J. Virol. 2012, 86, 2337–2346. [Google Scholar] [CrossRef] [PubMed]
- Chiou, S.S.; Liu, H.; Chuang, C.K.; Lin, C.C.; Chen, W.J. Fitness of Japanese encephalitis virus to Neuro-2a cells is determined by interactions of the viral envelope protein with highly sulfated glycosaminoglycans on the cell surface. J. Med. Virol. 2005, 76, 583–592. [Google Scholar] [CrossRef] [PubMed]
- Su, C.M.; Liao, C.L.; Lee, Y.L.; Lin, Y.L. Highly sulfated forms of heparin sulfate are involved in Japanese encephalitis virus infection. Virology 2001, 286, 206–215. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Chiou, S.S.; Chen, W.J. Differential binding efficiency between the envelope protein of Japanese encephalitis virus variants and heparan sulfate on the cell surface. J. Med. Virol. 2004, 72, 618–624. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.; Hall, R.A.; Lobigs, M. Common E protein determinants for attenuation of glycosaminoglycan-binding variants of Japanese encephalitis and West Nile viruses. J. Virol. 2004, 78, 8271–8280. [Google Scholar] [CrossRef] [PubMed]
- Nickells, J.; Cannella, M.; Droll, D.A.; Liang, Y.; Wold, W.S.; Chambers, T.J. Neuroadapted yellow fever virus strain 17D: A charged locus in domain III of the E protein governs heparin binding activity and neuroinvasiveness in the SCID mouse model. J. Virol. 2008, 82, 12510–12519. [Google Scholar] [CrossRef] [PubMed]
- Germi, R.; Crance, J.M.; Garin, D.; Guimet, J.; Lortat-Jacob, H.; Ruigrok, R.W.; Zarski, J.P.; Drouet, E. Heparan sulfate-mediated binding of infectious dengue virus type 2 and yellow fever virus. Virology 2002, 292, 162–168. [Google Scholar] [CrossRef] [PubMed]
- Hilgard, P.; Stockert, R. Heparan sulfate proteoglycans initiate dengue virus infection of hepatocytes. Hepatology 2000, 32, 1069–1077. [Google Scholar] [CrossRef] [PubMed]
- Kroschewski, H.; Allison, S.L.; Heinz, F.X.; Mandl, C.W. Role of heparan sulfate for attachment and entry of tick-borne encephalitis virus. Virology 2003, 308, 92–100. [Google Scholar] [CrossRef]
- Okamoto, K.; Kinoshita, H.; Parquet Mdel, C.; Raekiansyah, M.; Kimura, D.; Yui, K.; Islam, M.A.; Hasebe, F.; Morita, K. Dengue virus strain DEN2 16681 utilizes a specific glycochain of syndecan-2 proteoglycan as a receptor. J. Gen. Virol. 2012, 93, 761–770. [Google Scholar] [CrossRef] [PubMed]
- Mandl, C.W.; Kroschewski, H.; Allison, S.L.; Kofler, R.; Holzmann, H.; Meixner, T.; Heinz, F.X. Adaptation of tick-borne encephalitis virus to BHK-21 cells results in the formation of multiple heparan sulfate binding sites in the envelope protein and attenuation in vivo. J. Virol. 2001, 75, 5627–5637. [Google Scholar] [CrossRef] [PubMed]
- Roehrig, J.T.; Butrapet, S.; Liss, N.M.; Bennett, S.L.; Luy, B.E.; Childers, T.; Boroughs, K.L.; Stovall, J.L.; Calvert, A.E.; Blair, C.D.; et al. Mutation of the dengue virus type 2 envelope protein heparan sulfate binding sites or the domain III lateral ridge blocks replication in Vero cells prior to membrane fusion. Virology 2013, 441, 114–125. [Google Scholar] [CrossRef] [PubMed]
- Watterson, D.; Kobe, B.; Young, P.R. Residues in domain III of the dengue virus envelope glycoprotein involved in cell-surface glycosaminoglycan binding. J. Gen. Virol. 2012, 93, 72–82. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.Y.; Zhao, J.; Liu, X.; Fraser, K.; Lin, L.; Zhang, X.; Zhang, F.; Dordick, J.S.; Linhardt, R.J. Interaction of Zika virus envelope protein with glycosaminoglycans. Biochemistry 2017, 56, 1151–1162. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Barragan, J.J.; del Angel, R.M. Identification of a putative coreceptor on Vero cells that participates in dengue 4 virus infection. J. Virol. 2001, 75, 7818–7827. [Google Scholar] [CrossRef] [PubMed]
- Dalrymple, N.; Mackow, E.R. Productive dengue virus infection of human endothelial cells is directed by heparan sulfate-containing proteoglycan receptors. J. Virol. 2011, 85, 9478–9485. [Google Scholar] [CrossRef] [PubMed]
- Hung, S.L.; Lee, P.L.; Chen, H.W.; Chen, L.K.; Kao, C.L.; King, C.C. Analysis of the steps involved in dengue virus entry into host cells. Virology 1999, 257, 156–167. [Google Scholar] [CrossRef] [PubMed]
- Kamhi, E.; Joo, E.J.; Dordick, J.S.; Linhardt, R.J. Glycosaminoglycans in infectious disease. Biol. Rev. Camb. Philos. Soc. 2013, 88, 928–943. [Google Scholar] [CrossRef] [PubMed]
- Belting, M. Heparan sulfate proteoglycan as a plasma membrane carrier. Trends Biochem. Sci. 2003, 28, 145–151. [Google Scholar] [CrossRef]
- Chiou, S.S.; Chen, W.J. Phenotypic changes in the Japanese encephalitis virus after one passage in Neuro-2a cells: Generation of attenuated strains of the virus. Vaccine 2007, 26, 15–23. [Google Scholar] [CrossRef] [PubMed]
- Mason, C.P.; Tarr, A.W. Human lectins and their roles in viral infections. Molecules 2015, 20, 2229–2271. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Ren, S.; Zuo, Y. DC-SIGN, DC-SIGNR and LSECtin: C-type lectins for infection. Int. Rev. Immunol. 2014, 33, 54–66. [Google Scholar] [CrossRef] [PubMed]
- Geijtenbeek, T.B.; Kwon, D.S.; Torensma, R.; van Vliet, S.J.; van Duijnhoven, G.C.; Middel, J.; Cornelissen, I.L.; Nottet, H.S.; KewalRamani, V.N.; Littman, D.R.; et al. DC-SIGN, a dendritic cell-specific HIV-1-binding protein that enhances trans-infection of T cells. Cell 2000, 100, 587–597. [Google Scholar] [CrossRef]
- Geijtenbeek, T.B.; Torensma, R.; van Vliet, S.J.; van Duijnhoven, G.C.; Adema, G.J.; van Kooyk, Y.; Figdor, C.G. Identification of DC-SIGN, a novel dendritic cell-specific ICAM-3 receptor that supports primary immune responses. Cell 2000, 100, 575–585. [Google Scholar] [CrossRef]
- Engering, A.; van Vliet, S.J.; Hebeda, K.; Jackson, D.G.; Prevo, R.; Singh, S.K.; Geijtenbeek, T.B.; van Krieken, H.; van Kooyk, Y. Dynamic populations of dendritic cell-specific ICAM-3 grabbing nonintegrin-positive immature dendritic cells and liver/lymph node-specific ICAM-3 grabbing nonintegrin-positive endothelial cells in the outer zones of the paracortex of human lymph nodes. Am. J. Pathol. 2004, 164, 1587–1595. [Google Scholar] [CrossRef]
- Pohlmann, S.; Soilleux, E.J.; Baribaud, F.; Leslie, G.J.; Morris, L.S.; Trowsdale, J.; Lee, B.; Coleman, N.; Doms, R.W. DC-SIGNR, a DC-SIGN homologue expressed in endothelial cells, binds to human and simian immunodeficiency viruses and activates infection in trans. Proc. Natl. Acad. Sci. USA 2001, 98, 2670–2675. [Google Scholar] [CrossRef] [PubMed]
- Bashirova, A.A.; Geijtenbeek, T.B.; van Duijnhoven, G.C.; van Vliet, S.J.; Eilering, J.B.; Martin, M.P.; Wu, L.; Martin, T.D.; Viebig, N.; Knolle, P.A.; et al. A dendritic cell-specific intercellular adhesion molecule 3-grabbing nonintegrin (DC-SIGN)-related protein is highly expressed on human liver sinusoidal endothelial cells and promotes HIV-1 infection. J. Exp. Med. 2001, 193, 671–678. [Google Scholar] [CrossRef] [PubMed]
- Khoo, U.S.; Chan, K.Y.; Chan, V.S.; Lin, C.L. DC-SIGN and L-SIGN: The SIGNs for infection. J. Mol. Med. 2008, 86, 861–874. [Google Scholar] [CrossRef] [PubMed]
- Davis, C.W.; Mattei, L.M.; Nguyen, H.Y.; Ansarah-Sobrinho, C.; Doms, R.W.; Pierson, T.C. The location of asparagine-linked glycans on West Nile virions controls their interactions with CD209 (dendritic cell-specific ICAM-3 grabbing nonintegrin). J. Biol. Chem. 2006, 281, 37183–37194. [Google Scholar] [CrossRef] [PubMed]
- Mondotte, J.A.; Lozach, P.Y.; Amara, A.; Gamarnik, A.V. Essential role of dengue virus envelope protein N-glycosylation at asparagine-67 during viral propagation. J. Virol. 2007, 81, 7136–7148. [Google Scholar] [CrossRef] [PubMed]
- Dejnirattisai, W.; Webb, A.I.; Chan, V.; Jumnainsong, A.; Davidson, A.; Mongkolsapaya, J.; Screaton, G. Lectin switching during dengue virus infection. J. Infect. Dis. 2011, 203, 1775–1783. [Google Scholar] [CrossRef] [PubMed]
- Lozach, P.Y.; Burleigh, L.; Staropoli, I.; Navarro-Sanchez, E.; Harriague, J.; Virelizier, J.L.; Rey, F.A.; Despres, P.; Arenzana-Seisdedos, F.; Amara, A. Dendritic cell-specific intercellular adhesion molecule 3-grabbing non-integrin (DC-SIGN)-mediated enhancement of dengue virus infection is independent of DC-SIGN internalization signals. J. Biol. Chem. 2005, 280, 23698–23708. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Ridilla, M.; Patel, P.; Betts, L.; Gallichotte, E.; Shahidi, L.; Thompson, N.L.; Jacobson, K. Beyond attachment: Roles of DC-SIGN in dengue virus infection. Traffic 2017, 18, 218–231. [Google Scholar] [CrossRef] [PubMed]
- Shah, M.; Wadood, A.; Rahman, Z.; Husnain, T. Interaction and inhibition of dengue envelope glycoprotein with mammalian receptor DC-sign, an in-silico approach. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Rey, F.A.; Stiasny, K.; Heinz, F.X. Flavivirus structural heterogeneity: Implications for cell entry. Curr. Opin. Virol. 2017, 24, 132–139. [Google Scholar] [CrossRef] [PubMed]
- Pierson, T.C.; Diamond, M.S. Degrees of maturity: The complex structure and biology of flaviviruses. Curr. Opin. Virol. 2012, 2, 168–175. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.M.; Yun, S.I.; Song, B.H.; Hahn, Y.S.; Lee, C.H.; Oh, H.W.; Lee, Y.M. A single N-linked glycosylation site in the Japanese encephalitis virus prM protein is critical for cell type-specific prM protein biogenesis, virus particle release, and pathogenicity in mice. J. Virol. 2008, 82, 7846–7862. [Google Scholar] [CrossRef] [PubMed]
- Pabalan, N.; Chaisri, S.; Tabunhan, S.; Phumyen, A.; Jarjanazi, H.; Steiner, T.S. Associations of DC-SIGN (CD209) promoter -336G/A polymorphism (rs4804803) with dengue infection: A systematic review and meta-analysis. Acta Trop. 2018, 177, 186–193. [Google Scholar] [CrossRef] [PubMed]
- Barkhash, A.V.; Perelygin, A.A.; Babenko, V.N.; Brinton, M.A.; Voevoda, M.I. Single nucleotide polymorphism in the promoter region of the CD209 gene is associated with human predisposition to severe forms of tick-borne encephalitis. Antivir. Res. 2012, 93, 64–68. [Google Scholar] [CrossRef] [PubMed]
- Miller, J.L.; de Wet, B.J.; Martinez-Pomares, L.; Radcliffe, C.M.; Dwek, R.A.; Rudd, P.M.; Gordon, S. The mannose receptor mediates dengue virus infection of macrophages. PLoS Pathog. 2008, 4. [Google Scholar] [CrossRef]
- Chen, S.T.; Liu, R.S.; Wu, M.F.; Lin, Y.L.; Chen, S.Y.; Tan, D.T.; Chou, T.Y.; Tsai, I.S.; Li, L.; Hsieh, S.L. CLEC5A regulates Japanese encephalitis virus-induced neuroinflammation and lethality. PLoS Pathog. 2012, 8. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.T.; Lin, Y.L.; Huang, M.T.; Wu, M.F.; Cheng, S.C.; Lei, H.Y.; Lee, C.K.; Chiou, T.W.; Wong, C.H.; Hsieh, S.L. CLEC5A is critical for dengue virus-induced lethal disease. Nature 2008, 453, 672–676. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Pomares, L. The mannose receptor. J. Leukoc. Biol. 2012, 92, 1177–1186. [Google Scholar] [CrossRef] [PubMed]
- Bakker, A.B.; Baker, E.; Sutherland, G.R.; Phillips, J.H.; Lanier, L.L. Myeloid DAP12-associating lectin (MDL)-1 is a cell surface receptor involved in the activation of myeloid cells. Proc. Natl. Acad. Sci. USA 1999, 96, 9792–9796. [Google Scholar] [CrossRef] [PubMed]
- Lanier, L.L.; Corliss, B.C.; Wu, J.; Leong, C.; Phillips, J.H. Immunoreceptor DAP12 bearing a tyrosine-based activation motif is involved in activating NK cells. Nature 1998, 391, 703–707. [Google Scholar] [CrossRef] [PubMed]
- Dominguez-Soto, A.; Aragoneses-Fenoll, L.; Martin-Gayo, E.; Martinez-Prats, L.; Colmenares, M.; Naranjo-Gomez, M.; Borras, F.E.; Munoz, P.; Zubiaur, M.; Toribio, M.L.; et al. The DC-SIGN-related lectin LSECtin mediates antigen capture and pathogen binding by human myeloid cells. Blood 2007, 109, 5337–5345. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Qian, Y.; Jung, Y.S.; Zhou, B.; Cao, R.; Shen, T.; Shao, D.; Wei, J.; Ma, Z.; Chen, P.; et al. mosGCTL-7, a C-type lectin protein, mediates Japanese encephalitis virus infection in mosquitoes. J. Virol. 2017, 91. [Google Scholar] [CrossRef] [PubMed]
- Cheng, G.; Cox, J.; Wang, P.; Krishnan, M.N.; Dai, J.; Qian, F.; Anderson, J.F.; Fikrig, E. A C-type lectin collaborates with a CD45 phosphatase homolog to facilitate West Nile virus infection of mosquitoes. Cell 2010, 142, 714–725. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zhang, F.; Liu, J.; Xiao, X.; Zhang, S.; Qin, C.; Xiang, Y.; Wang, P.; Cheng, G. Transmission-blocking antibodies against mosquito C-type lectins for dengue prevention. PLoS Pathog. 2014, 10. [Google Scholar] [CrossRef] [PubMed]
- Thomas, M.L. The leukocyte common antigen family. Annu. Rev. Immunol. 1989, 7, 339–369. [Google Scholar] [CrossRef] [PubMed]
- Byth, K.F.; Conroy, L.A.; Howlett, S.; Smith, A.J.; May, J.; Alexander, D.R.; Holmes, N. CD45-null transgenic mice reveal a positive regulatory role for CD45 in early thymocyte development, in the selection of CD4+CD8+ thymocytes, and B cell maturation. J. Exp. Med. 1996, 183, 1707–1718. [Google Scholar] [CrossRef] [PubMed]
- Trowbridge, I.S.; Thomas, M.L. CD45: An emerging role as a protein tyrosine phosphatase required for lymphocyte activation and development. Annu. Rev. Immunol. 1994, 12, 85–116. [Google Scholar] [CrossRef] [PubMed]
- Baldwin, T.A.; Ostergaard, H.L. Developmentally regulated changes in glucosidase II association with, and carbohydrate content of, the protein tyrosine phosphatase CD45. J. Immunol. 2001, 167, 3829–3835. [Google Scholar] [CrossRef] [PubMed]
- Luo, H.; Wang, T. Recent advances in understanding West Nile virus host immunity and viral pathogenesis. F1000Research 2018, 7. [Google Scholar] [CrossRef] [PubMed]
- Fuchs, A.; Lin, T.Y.; Beasley, D.W.; Stover, C.M.; Schwaeble, W.J.; Pierson, T.C.; Diamond, M.S. Direct complement restriction of flavivirus infection requires glycan recognition by mannose-binding lectin. Cell Host Microbe 2010, 8, 186–195. [Google Scholar] [CrossRef] [PubMed]
- Figueiredo, G.G.; Cezar, R.D.; Freire, N.M.; Teixeira, V.G.; Baptista, P.; Cordeiro, M.; Carmo, R.F.; Vasconcelos, L.R.; Moura, P. Mannose-binding lectin gene (MBL2) polymorphisms related to the mannose-binding lectin low levels are associated to dengue disease severity. Hum. Immunol. 2016, 77, 571–575. [Google Scholar] [CrossRef] [PubMed]
- Pandolfi, F.; Franza, L.; Altamura, S.; Mandolini, C.; Cianci, R.; Ansari, A.; Kurnick, J.T. Integrins: Integrating the biology and therapy of cell-cell interactions. Clin. Ther. 2017, 39, 2420–2436. [Google Scholar] [CrossRef] [PubMed]
- Chu, J.J.; Ng, M.L. Characterization of a 105-kDa plasma membrane associated glycoprotein that is involved in West Nile virus binding and infection. Virology 2003, 312, 458–469. [Google Scholar] [CrossRef]
- Medigeshi, G.R.; Hirsch, A.J.; Streblow, D.N.; Nikolich-Zugich, J.; Nelson, J.A. West Nile virus entry requires cholesterol-rich membrane microdomains and is independent of αvβ3 integrin. J. Virol. 2008, 82, 5212–5219. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, K.; Keller, M.; Bader, B.L.; Korytar, T.; Finke, S.; Ziegler, U.; Groschup, M.H. Integrins modulate the infection efficiency of West Nile virus into cells. J. Gen. Virol. 2013, 94, 1723–1733. [Google Scholar] [CrossRef] [PubMed]
- Horvath, I.; Multhoff, G.; Sonnleitner, A.; Vigh, L. Membrane-associated stress proteins: More than simply chaperones. Biochim. Biophys. Acta 2008, 1778, 1653–1664. [Google Scholar] [CrossRef] [PubMed]
- Ren, J.; Ding, T.; Zhang, W.; Song, J.; Ma, W. Does Japanese encephalitis virus share the same cellular receptor with other mosquito-borne flaviviruses on the C6/36 mosquito cells? Virol. J. 2007, 4. [Google Scholar] [CrossRef] [PubMed]
- Das, S.; Laxminarayana, S.V.; Chandra, N.; Ravi, V.; Desai, A. Heat shock protein 70 on Neuro2a cells is a putative receptor for Japanese encephalitis virus. Virology 2009, 385, 47–57. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.Z.; Cao, M.M.; Wang, W.B.; Wang, W.; Ren, H.; Zhao, P.; Qi, Z.T. Association of heat shock protein 70 with lipid rafts is required for Japanese encephalitis virus infection in Huh7 cells. J. Gen. Virol. 2012, 93, 61–71. [Google Scholar] [CrossRef] [PubMed]
- Nain, M.; Mukherjee, S.; Karmakar, S.P.; Paton, A.W.; Paton, J.C.; Abdin, M.Z.; Basu, A.; Kalia, M.; Vrati, S. GRP78 is an important host factor for Japanese encephalitis virus entry and replication in mammalian cells. J. Virol. 2017, 91. [Google Scholar] [CrossRef] [PubMed]
- Hung, C.Y.; Tsai, M.C.; Wu, Y.P.; Wang, R.Y. Identification of heat-shock protein 90β in Japanese encephalitis virus-induced secretion proteins. J. Gen. Virol. 2011, 92, 2803–2809. [Google Scholar] [CrossRef] [PubMed]
- Thongtan, T.; Wikan, N.; Wintachai, P.; Rattanarungsan, C.; Srisomsap, C.; Cheepsunthorn, P.; Smith, D.R. Characterization of putative Japanese encephalitis virus receptor molecules on microglial cells. J. Med. Virol. 2012, 84, 615–623. [Google Scholar] [CrossRef] [PubMed]
- Reyes-Del Valle, J.; Chavez-Salinas, S.; Medina, F.; Del Angel, R.M. Heat shock protein 90 and heat shock protein 70 are components of dengue virus receptor complex in human cells. J. Virol. 2005, 79, 4557–4567. [Google Scholar] [CrossRef] [PubMed]
- Cabrera-Hernandez, A.; Thepparit, C.; Suksanpaisan, L.; Smith, D.R. Dengue virus entry into liver (HepG2) cells is independent of hsp90 and hsp70. J. Med. Virol. 2007, 79, 386–392. [Google Scholar] [CrossRef] [PubMed]
- Salas-Benito, J.; Reyes-Del Valle, J.; Salas-Benito, M.; Ceballos-Olvera, I.; Mosso, C.; del Angel, R.M. Evidence that the 45-kD glycoprotein, part of a putative dengue virus receptor complex in the mosquito cell line C6/36, is a heat-shock related protein. Am. J. Trop. Med. Hyg. 2007, 77, 283–290. [Google Scholar] [PubMed]
- Jindadamrongwech, S.; Thepparit, C.; Smith, D.R. Identification of GRP78 (BiP) as a liver cell expressed receptor element for dengue virus serotype 2. Arch. Virol. 2004, 149, 915–927. [Google Scholar] [CrossRef] [PubMed]
- Vega-Almeida, T.O.; Salas-Benito, M.; De Nova-Ocampo, M.A.; Del Angel, R.M.; Salas-Benito, J.S. Surface proteins of C6/36 cells involved in dengue virus 4 binding and entry. Arch. Virol. 2013, 158, 1189–1207. [Google Scholar] [CrossRef] [PubMed]
- Taguwa, S.; Maringer, K.; Li, X.; Bernal-Rubio, D.; Rauch, J.N.; Gestwicki, J.E.; Andino, R.; Fernandez-Sesma, A.; Frydman, J. Defining hsp70 subnetworks in dengue virus replication reveals key vulnerability in flavivirus infection. Cell 2015, 163, 1108–1123. [Google Scholar] [CrossRef] [PubMed]
- Upanan, S.; Kuadkitkan, A.; Smith, D.R. Identification of dengue virus binding proteins using affinity chromatography. J. Virol. Methods 2008, 151, 325–328. [Google Scholar] [CrossRef] [PubMed]
- Howe, M.K.; Speer, B.L.; Hughes, P.F.; Loiselle, D.R.; Vasudevan, S.; Haystead, T.A. An inducible heat shock protein 70 small molecule inhibitor demonstrates anti-dengue virus activity, validating Hsp70 as a host antiviral target. Antivir. Res. 2016, 130, 81–92. [Google Scholar] [CrossRef] [PubMed]
- Liang, J.J.; Yu, C.Y.; Liao, C.L.; Lin, Y.L. Vimentin binding is critical for infection by the virulent strain of Japanese encephalitis virus. Cell. Microbiol. 2011, 13, 1358–1370. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.K.; Lin, Y.L.; Liao, C.L.; Lin, C.G.; Huang, Y.L.; Yeh, C.T.; Lai, S.C.; Jan, J.T.; Chin, C. Generation and characterization of organ-tropism mutants of Japanese encephalitis virus in vivo and In vitro. Virology 1996, 223, 79–88. [Google Scholar] [CrossRef] [PubMed]
- Das, S.; Ravi, V.; Desai, A. Japanese encephalitis virus interacts with vimentin to facilitate its entry into porcine kidney cell line. Virus Res. 2011, 160, 404–408. [Google Scholar] [CrossRef] [PubMed]
- Chien, Y.J.; Chen, W.J.; Hsu, W.L.; Chiou, S.S. Bovine lactoferrin inhibits Japanese encephalitis virus by binding to heparan sulfate and receptor for low density lipoprotein. Virology 2008, 379, 143–151. [Google Scholar] [CrossRef] [PubMed]
- Kimura, T.; Kimura-Kuroda, J.; Nagashima, K.; Yasui, K. Analysis of virus-cell binding characteristics on the determination of Japanese encephalitis virus susceptibility. Arch. Virol. 1994, 139, 239–251. [Google Scholar] [CrossRef] [PubMed]
- Boonsanay, V.; Smith, D.R. Entry into and production of the Japanese encephalitis virus from C6/36 cells. Intervirology 2007, 50, 85–92. [Google Scholar] [CrossRef] [PubMed]
- Meertens, L.; Carnec, X.; Lecoin, M.P.; Ramdasi, R.; Guivel-Benhassine, F.; Lew, E.; Lemke, G.; Schwartz, O.; Amara, A. The TIM and TAM families of phosphatidylserine receptors mediate dengue virus entry. Cell Host Microbe 2012, 12, 544–557. [Google Scholar] [CrossRef] [PubMed]
- Jemielity, S.; Wang, J.J.; Chan, Y.K.; Ahmed, A.A.; Li, W.; Monahan, S.; Bu, X.; Farzan, M.; Freeman, G.J.; Umetsu, D.T.; et al. TIM-family proteins promote infection of multiple enveloped viruses through virion-associated phosphatidylserine. PLoS Pathog. 2013, 9. [Google Scholar] [CrossRef] [PubMed]
- Morizono, K.; Chen, I.S. Role of phosphatidylserine receptors in enveloped virus infection. J. Virol. 2014, 88, 4275–4290. [Google Scholar] [CrossRef] [PubMed]
- Richard, A.S.; Zhang, A.; Park, S.J.; Farzan, M.; Zong, M.; Choe, H. Virion-associated phosphatidylethanolamine promotes TIM1-mediated infection by Ebola, dengue, and West Nile viruses. Proc. Natl. Acad. Sci. USA 2015, 112, 14682–14687. [Google Scholar] [CrossRef] [PubMed]
- Savidis, G.; McDougall, W.M.; Meraner, P.; Perreira, J.M.; Portmann, J.M.; Trincucci, G.; John, S.P.; Aker, A.M.; Renzette, N.; Robbins, D.R.; et al. Identification of Zika virus and dengue virus dependency factors using functional genomics. Cell Rep. 2016, 16, 232–246. [Google Scholar] [CrossRef] [PubMed]
- Persaud, M.; Martinez-Lopez, A.; Buffone, C.; Porcelli, S.A.; Diaz-Griffero, F. Infection by Zika viruses requires the transmembrane protein AXL, endocytosis and low pH. Virology 2018, 518, 301–312. [Google Scholar] [CrossRef] [PubMed]
- Meertens, L.; Labeau, A.; Dejarnac, O.; Cipriani, S.; Sinigaglia, L.; Bonnet-Madin, L.; Le Charpentier, T.; Hafirassou, M.L.; Zamborlini, A.; Cao-Lormeau, V.M.; et al. Axl mediates Zika virus entry in human glial cells and modulates innate immune responses. Cell Rep. 2017, 18, 324–333. [Google Scholar] [CrossRef] [PubMed]
- Richard, A.S.; Shim, B.S.; Kwon, Y.C.; Zhang, R.; Otsuka, Y.; Schmitt, K.; Berri, F.; Diamond, M.S.; Choe, H. AXL-dependent infection of human fetal endothelial cells distinguishes Zika virus from other pathogenic flaviviruses. Proc. Natl. Acad. Sci. USA 2017, 114, 2024–2029. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; DeLalio, L.J.; Isakson, B.E.; Wang, T.T. AXL-mediated productive infection of human endothelial cells by Zika virus. Circ. Res. 2016, 119, 1183–1189. [Google Scholar] [CrossRef] [PubMed]
- Nowakowski, T.J.; Pollen, A.A.; Di Lullo, E.; Sandoval-Espinosa, C.; Bershteyn, M.; Kriegstein, A.R. Expression analysis highlights AXL as a candidate Zika virus entry receptor in neural stem cells. Cell Stem Cell 2016, 18, 591–596. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.Y.; Wang, Z.; Zhen, Z.D.; Feng, K.H.; Guo, J.; Gao, N.; Fan, D.Y.; Han, D.S.; Wang, P.G.; An, J. Axl is not an indispensable factor for Zika virus infection in mice. J. Gen. Virol. 2017, 98, 2061–2068. [Google Scholar] [CrossRef] [PubMed]
- Hastings, A.K.; Yockey, L.J.; Jagger, B.W.; Hwang, J.; Uraki, R.; Gaitsch, H.F.; Parnell, L.A.; Cao, B.; Mysorekar, I.U.; Rothlin, C.V.; et al. TAM receptors are not required for Zika virus infection in mice. Cell Rep. 2017, 19, 558–568. [Google Scholar] [CrossRef] [PubMed]
- Carnec, X.; Meertens, L.; Dejarnac, O.; Perera-Lecoin, M.; Hafirassou, M.L.; Kitaura, J.; Ramdasi, R.; Schwartz, O.; Amara, A. The phosphatidylserine and phosphatidylethanolamine receptor CD300a binds dengue virus and enhances infection. J. Virol. 2016, 90, 92–102. [Google Scholar] [CrossRef] [PubMed]
- Che, P.; Tang, H.; Li, Q. The interaction between claudin-1 and dengue viral prM/M protein for its entry. Virology 2013, 446, 303–313. [Google Scholar] [CrossRef] [PubMed]
- Gao, F.; Duan, X.; Lu, X.; Liu, Y.; Zheng, L.; Ding, Z.; Li, J. Novel binding between pre-membrane protein and claudin-1 is required for efficient dengue virus entry. Biochem. Biophys. Res. Commun. 2010, 391, 952–957. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Kakinami, C.; Li, Q.; Yang, B.; Li, H. Human apolipoprotein A-I is associated with dengue virus and enhances virus infection through SR-BI. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Thepparit, C.; Smith, D.R. Serotype-specific entry of dengue virus into liver cells: Identification of the 37-kilodalton/67-kilodalton high-affinity laminin receptor as a dengue virus serotype 1 receptor. J. Virol. 2004, 78, 12647–12656. [Google Scholar] [CrossRef] [PubMed]
- Sakoonwatanyoo, P.; Boonsanay, V.; Smith, D.R. Growth and production of the dengue virus in C6/36 cells and identification of a laminin-binding protein as a candidate serotype 3 and 4 receptor protein. Intervirology 2006, 49, 161–172. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.C.; Wang, S.Y.; King, C.C. Bacterial lipopolysaccharide inhibits dengue virus infection of primary human monocytes/macrophages by blockade of virus entry via a CD14-dependent mechanism. J. Virol. 1999, 73, 2650–2657. [Google Scholar] [PubMed]
- Aoki, C.; Hidari, K.I.; Itonori, S.; Yamada, A.; Takahashi, N.; Kasama, T.; Hasebe, F.; Islam, M.A.; Hatano, K.; Matsuoka, K.; et al. Identification and characterization of carbohydrate molecules in mammalian cells recognized by dengue virus type 2. J. Biochem. 2006, 139, 607–614. [Google Scholar] [CrossRef] [PubMed]
- Wichit, S.; Jittmittraphap, A.; Hidari, K.I.; Thaisomboonsuk, B.; Petmitr, S.; Ubol, S.; Aoki, C.; Itonori, S.; Morita, K.; Suzuki, T.; et al. Dengue virus type 2 recognizes the carbohydrate moiety of neutral glycosphingolipids in mammalian and mosquito cells. Microbiol. Immunol. 2011, 55, 135–140. [Google Scholar] [CrossRef] [PubMed]
- Hershkovitz, O.; Rosental, B.; Rosenberg, L.A.; Navarro-Sanchez, M.E.; Jivov, S.; Zilka, A.; Gershoni-Yahalom, O.; Brient-Litzler, E.; Bedouelle, H.; Ho, J.W.; et al. NKp44 receptor mediates interaction of the envelope glycoproteins from the West Nile and dengue viruses with NK cells. J. Immunol. 2009, 183, 2610–2621. [Google Scholar] [CrossRef] [PubMed]
- Kuadkitkan, A.; Wikan, N.; Fongsaran, C.; Smith, D.R. Identification and characterization of prohibitin as a receptor protein mediating DENV-2 entry into insect cells. Virology 2010, 406, 149–161. [Google Scholar] [CrossRef] [PubMed]
- Cruz-Oliveira, C.; Freire, J.M.; Conceicao, T.M.; Higa, L.M.; Castanho, M.A.; Da Poian, A.T. Receptors and routes of dengue virus entry into the host cells. FEMS Microbiol. Rev. 2015, 39, 155–170. [Google Scholar] [CrossRef] [PubMed]
- Mayor, S.; Pagano, R.E. Pathways of clathrin-independent endocytosis. Nat. Rev. Mol. Cell Biol. 2007, 8, 603–612. [Google Scholar] [CrossRef] [PubMed]
- Krishnan, M.N.; Sukumaran, B.; Pal, U.; Agaisse, H.; Murray, J.L.; Hodge, T.W.; Fikrig, E. Rab5 is required for the cellular entry of dengue and West Nile viruses. J. Virol. 2007, 81, 4881–4885. [Google Scholar] [CrossRef] [PubMed]
- Acosta, E.G.; Castilla, V.; Damonte, E.B. Infectious dengue-1 virus entry into mosquito C6/36 cells. Virus Res. 2011, 160, 173–179. [Google Scholar] [CrossRef] [PubMed]
- Acosta, E.G.; Castilla, V.; Damonte, E.B. Alternative infectious entry pathways for dengue virus serotypes into mammalian cells. Cell. Microbiol. 2009, 11, 1533–1549. [Google Scholar] [CrossRef] [PubMed]
- Alhoot, M.A.; Wang, S.M.; Sekaran, S.D. RNA interference mediated inhibition of dengue virus multiplication and entry in HepG2 cells. PLoS ONE 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Peng, T.; Wang, J.L.; Chen, W.; Zhang, J.L.; Gao, N.; Chen, Z.T.; Xu, X.F.; Fan, D.Y.; An, J. Entry of dengue virus serotype 2 into ECV304 cells depends on clathrin-dependent endocytosis, but not on caveolae-dependent endocytosis. Can. J. Microbiol. 2009, 55, 139–145. [Google Scholar] [CrossRef] [PubMed]
- Alhoot, M.A.; Wang, S.M.; Sekaran, S.D. Inhibition of dengue virus entry and multiplication into monocytes using RNA interference. PLoS Negl. Trop. Dis. 2011, 5. [Google Scholar] [CrossRef] [PubMed]
- Geoghegan, V.; Stainton, K.; Rainey, S.M.; Ant, T.H.; Dowle, A.A.; Larson, T.; Hester, S.; Charles, P.D.; Thomas, B.; Sinkins, S.P. Perturbed cholesterol and vesicular trafficking associated with dengue blocking in Wolbachia-infected Aedes aegypti cells. Nat. Commun. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.L.; Zhang, J.L.; Chen, W.; Xu, X.F.; Gao, N.; Fan, D.Y.; An, J. Roles of small GTPase Rac1 in the regulation of actin cytoskeleton during dengue virus infection. PLoS Negl. Trop. Dis. 2010, 4. [Google Scholar] [CrossRef] [PubMed]
- Shrivastava, N.; Sripada, S.; Kaur, J.; Shah, P.S.; Cecilia, D. Insights into the internalization and retrograde trafficking of dengue 2 virus in BHK-21 cells. PLoS ONE 2011, 6. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Garcia, M.D.; Meertens, L.; Chazal, M.; Hafirassou, M.L.; Dejarnac, O.; Zamborlini, A.; Despres, P.; Sauvonnet, N.; Arenzana-Seisdedos, F.; Jouvenet, N.; et al. Vaccine and wild-type strains of yellow fever virus engage distinct entry mechanisms and differentially stimulate antiviral immune responses. mBio 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Robinson, M.S. Forty years of clathrin-coated vesicles. Traffic 2015, 16, 1210–1238. [Google Scholar] [CrossRef] [PubMed]
- Mercer, J.; Schelhaas, M.; Helenius, A. Virus entry by endocytosis. Annu. Rev. Biochem. 2010, 79, 803–833. [Google Scholar] [CrossRef] [PubMed]
- McMahon, H.T.; Boucrot, E. Molecular mechanism and physiological functions of clathrin-mediated endocytosis. Nat. Rev. Mol. Cell Biol. 2011, 12, 517–533. [Google Scholar] [CrossRef] [PubMed]
- Krishnan, M.N.; Ng, A.; Sukumaran, B.; Gilfoy, F.D.; Uchil, P.D.; Sultana, H.; Brass, A.L.; Adametz, R.; Tsui, M.; Qian, F.; et al. RNA interference screen for human genes associated with West Nile virus infection. Nature 2008, 455, 242–245. [Google Scholar] [CrossRef] [PubMed]
- Sessions, O.M.; Barrows, N.J.; Souza-Neto, J.A.; Robinson, T.J.; Hershey, C.L.; Rodgers, M.A.; Ramirez, J.L.; Dimopoulos, G.; Yang, P.L.; Pearson, J.L.; et al. Discovery of insect and human dengue virus host factors. Nature 2009, 458, 1047–1050. [Google Scholar] [CrossRef] [PubMed]
- Le Sommer, C.; Barrows, N.J.; Bradrick, S.S.; Pearson, J.L.; Garcia-Blanco, M.A. G protein-coupled receptor kinase 2 promotes flaviviridae entry and replication. PLoS Negl. Trop. Dis. 2012, 6. [Google Scholar] [CrossRef] [PubMed]
- Bekerman, E.; Neveu, G.; Shulla, A.; Brannan, J.; Pu, S.Y.; Wang, S.; Xiao, F.; Barouch-Bentov, R.; Bakken, R.R.; Mateo, R.; et al. Anticancer kinase inhibitors impair intracellular viral trafficking and exert broad-spectrum antiviral effects. J. Clin. Investig. 2017, 127, 1338–1352. [Google Scholar] [CrossRef] [PubMed]
- Schoggins, J.W.; Wilson, S.J.; Panis, M.; Murphy, M.Y.; Jones, C.T.; Bieniasz, P.; Rice, C.M. A diverse range of gene products are effectors of the type I interferon antiviral response. Nature 2011, 472, 481–485. [Google Scholar] [CrossRef] [PubMed]
- Hackett, B.A.; Cherry, S. Flavivirus internalization is regulated by a size-dependent endocytic pathway. Proc. Natl. Acad. Sci. USA 2018, 115, 4246–4251. [Google Scholar] [CrossRef] [PubMed]
- Hackett, B.A.; Yasunaga, A.; Panda, D.; Tartell, M.A.; Hopkins, K.C.; Hensley, S.E.; Cherry, S. RNASEK is required for internalization of diverse acid-dependent viruses. Proc. Natl. Acad. Sci. USA 2015, 112, 7797–7802. [Google Scholar] [CrossRef] [PubMed]
- Perreira, J.M.; Aker, A.M.; Savidis, G.; Chin, C.R.; McDougall, W.M.; Portmann, J.M.; Meraner, P.; Smith, M.C.; Rahman, M.; Baker, R.E.; et al. RNASEK is a V-ATPase-associated factor required for endocytosis and the replication of rhinovirus, influenza A virus, and dengue virus. Cell Rep. 2015, 12, 850–863. [Google Scholar] [CrossRef] [PubMed]
- Harrison, S.C. Viral membrane fusion. Nat. Struct. Mol. Biol. 2008, 15, 690–698. [Google Scholar] [CrossRef] [PubMed]
- Hasan, S.S.; Sevvana, M.; Kuhn, R.J.; Rossmann, M.G. Structural biology of Zika virus and other flaviviruses. Nat. Struct. Mol. Biol. 2018, 25, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Smit, J.M.; Moesker, B.; Rodenhuis-Zybert, I.; Wilschut, J. Flavivirus cell entry and membrane fusion. Viruses 2011, 3, 160–171. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Zhao, X.; Na, R.; Li, L.; Warkentin, E.; Witt, J.; Lu, X.; Yu, Y.; Wei, Y.; Peng, G.; et al. The structure differences of Japanese encephalitis virus SA14 and SA14-14-2 E proteins elucidate the virulence attenuation mechanism. Protein Cell 2018. [Google Scholar] [CrossRef] [PubMed]
- Kanai, R.; Kar, K.; Anthony, K.; Gould, L.H.; Ledizet, M.; Fikrig, E.; Marasco, W.A.; Koski, R.A.; Modis, Y. Crystal structure of West Nile virus envelope glycoprotein reveals viral surface epitopes. J. Virol. 2006, 80, 11000–11008. [Google Scholar] [CrossRef] [PubMed]
- Nybakken, G.E.; Nelson, C.A.; Chen, B.R.; Diamond, M.S.; Fremont, D.H. Crystal structure of the West Nile virus envelope glycoprotein. J. Virol. 2006, 80, 11467–11474. [Google Scholar] [CrossRef] [PubMed]
- Luca, V.C.; Nelson, C.A.; Fremont, D.H. Structure of the St. Louis encephalitis virus postfusion envelope trimer. J. Virol. 2013, 87, 818–828. [Google Scholar] [CrossRef] [PubMed]
- Dai, L.; Song, J.; Lu, X.; Deng, Y.Q.; Musyoki, A.M.; Cheng, H.; Zhang, Y.; Yuan, Y.; Song, H.; Haywood, J.; et al. Structures of the Zika virus envelope protein and its complex with a flavivirus broadly protective antibody. Cell Host Microbe 2016, 19, 696–704. [Google Scholar] [CrossRef] [PubMed]
- Modis, Y.; Ogata, S.; Clements, D.; Harrison, S.C. A ligand-binding pocket in the dengue virus envelope glycoprotein. Proc. Natl. Acad. Sci. USA 2003, 100, 6986–6991. [Google Scholar] [CrossRef] [PubMed]
- Modis, Y.; Ogata, S.; Clements, D.; Harrison, S.C. Variable surface epitopes in the crystal structure of dengue virus type 3 envelope glycoprotein. J. Virol. 2005, 79, 1223–1231. [Google Scholar] [CrossRef] [PubMed]
- Cockburn, J.J.; Navarro Sanchez, M.E.; Goncalvez, A.P.; Zaitseva, E.; Stura, E.A.; Kikuti, C.M.; Duquerroy, S.; Dussart, P.; Chernomordik, L.V.; Lai, C.J.; et al. Structural insights into the neutralization mechanism of a higher primate antibody against dengue virus. EMBO J. 2012, 31, 767–779. [Google Scholar] [CrossRef] [PubMed]
- Stiasny, K.; Bressanelli, S.; Lepault, J.; Rey, F.A.; Heinz, F.X. Characterization of a membrane-associated trimeric low-pH-induced form of the class II viral fusion protein E from tick-borne encephalitis virus and its crystallization. J. Virol. 2004, 78, 3178–3183. [Google Scholar] [CrossRef] [PubMed]
- Kaufmann, B.; Rossmann, M.G. Molecular mechanisms involved in the early steps of flavivirus cell entry. Microbes Infect. 2011, 13, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Kaufmann, B.; Chipman, P.R.; Holdaway, H.A.; Johnson, S.; Fremont, D.H.; Kuhn, R.J.; Diamond, M.S.; Rossmann, M.G. Capturing a flavivirus pre-fusion intermediate. PLoS Pathog. 2009, 5. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.R.; Zou, G.; Hsieh, S.C.; Qing, M.; Tsai, W.Y.; Shi, P.Y.; Wang, W.K. The helical domains of the stem region of dengue virus envelope protein are involved in both virus assembly and entry. J. Virol. 2011, 85, 5159–5171. [Google Scholar] [CrossRef] [PubMed]
- Pangerl, K.; Heinz, F.X.; Stiasny, K. Mutational analysis of the zippering reaction during flavivirus membrane fusion. J. Virol. 2011, 85, 8495–8501. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, A.G.; Yang, P.L.; Harrison, S.C. Peptide inhibitors of dengue virus entry target a late-stage fusion intermediate. PLoS Pathog. 2010, 6. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, A.G.; Yang, P.L.; Harrison, S.C. Peptide inhibitors of flavivirus entry derived from the E protein stem. J. Virol. 2010, 84, 12549–12554. [Google Scholar] [CrossRef] [PubMed]
- Stiasny, K.; Kiermayr, S.; Bernhart, A.; Heinz, F.X. The membrane-proximal “stem” region increases the stability of the flavivirus E protein postfusion trimer and modulates its structure. J. Virol. 2013, 87, 9933–9938. [Google Scholar] [CrossRef] [PubMed]
- Rey, F.A.; Stiasny, K.; Vaney, M.C.; Dellarole, M.; Heinz, F.X. The bright and the dark side of human antibody responses to flaviviruses: Lessons for vaccine design. EMBO Rep. 2018, 19, 206–224. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Liu, H.; Zu, X.; Liu, Y.; Chen, L.; Zhu, X.; Zhang, L.; Zhou, Z.; Xiao, G.; Wang, W. The ubiquitin-proteasome system is essential for the productive entry of Japanese encephalitis virus. Virology 2016, 498, 116–127. [Google Scholar] [CrossRef] [PubMed]
- Nour, A.M.; Li, Y.; Wolenski, J.; Modis, Y. Viral membrane fusion and nucleocapsid delivery into the cytoplasm are distinct events in some flaviviruses. PLoS Pathog. 2013, 9. [Google Scholar] [CrossRef] [PubMed]
- Byk, L.A.; Iglesias, N.G.; De Maio, F.A.; Gebhard, L.G.; Rossi, M.; Gamarnik, A.V. Dengue virus genome uncoating requires ubiquitination. mBio 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Garcia, M.D.; Meertens, L.; Bonazzi, M.; Cossart, P.; Arenzana-Seisdedos, F.; Amara, A. Appraising the roles of CBLL1 and the ubiquitin/proteasome system for flavivirus entry and replication. J. Virol. 2011, 85, 2980–2989. [Google Scholar] [CrossRef] [PubMed]
- Hase, T.; Summers, P.L.; Ray, P. Entry and replication of Japanese encephalitis virus in cultured neurogenic cells. J. Virol. Methods 1990, 30, 205–214. [Google Scholar] [CrossRef]
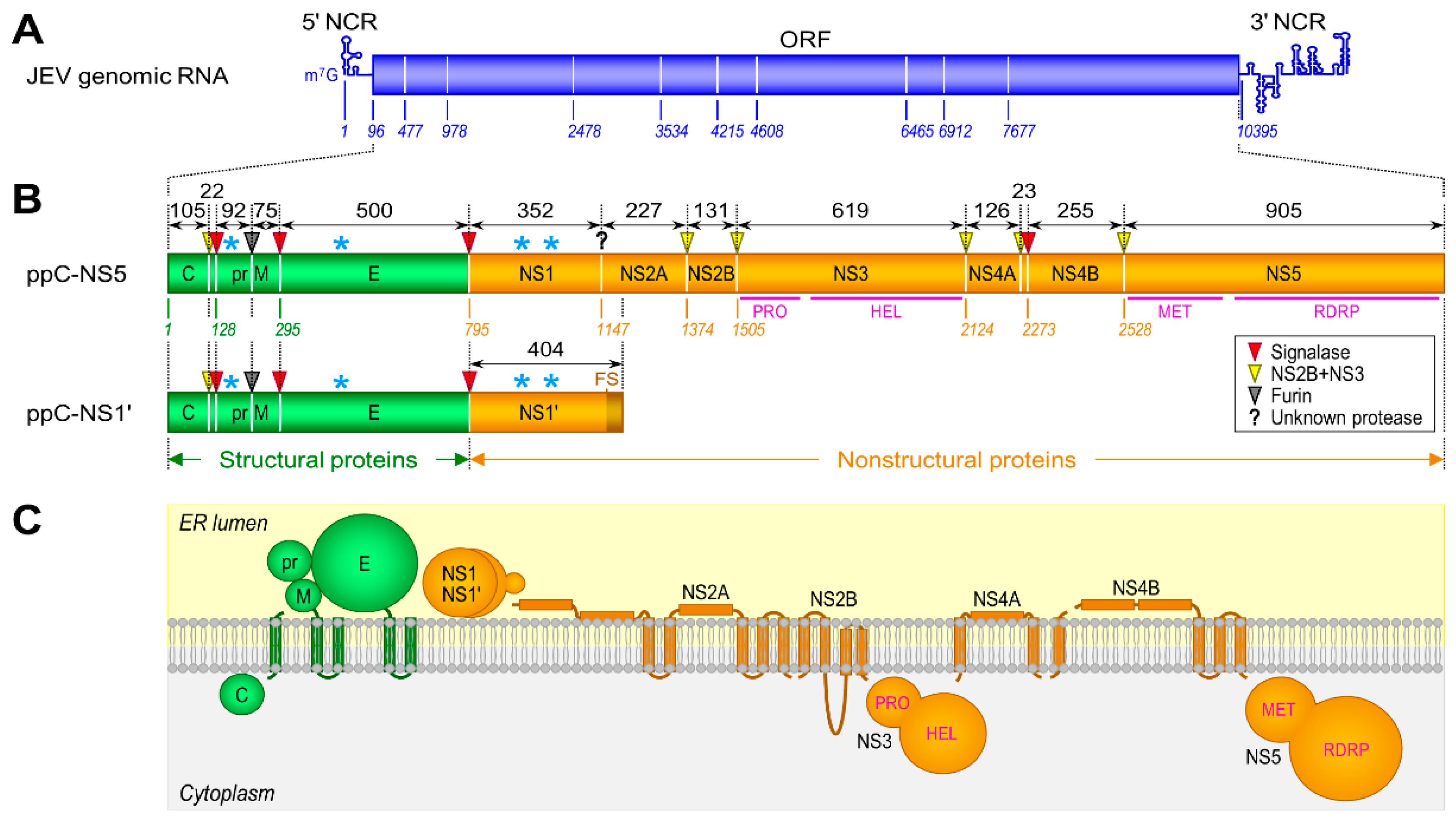
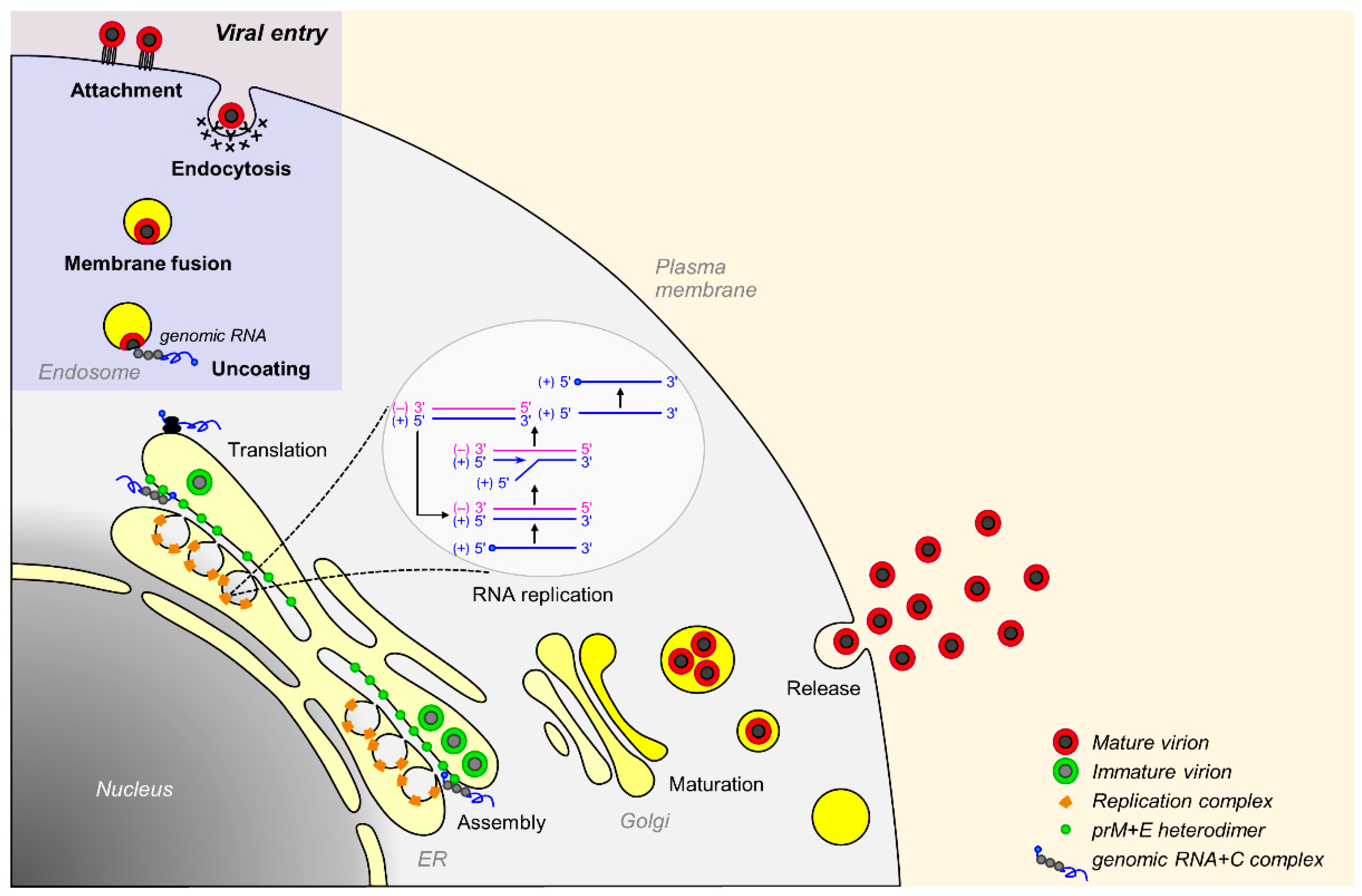

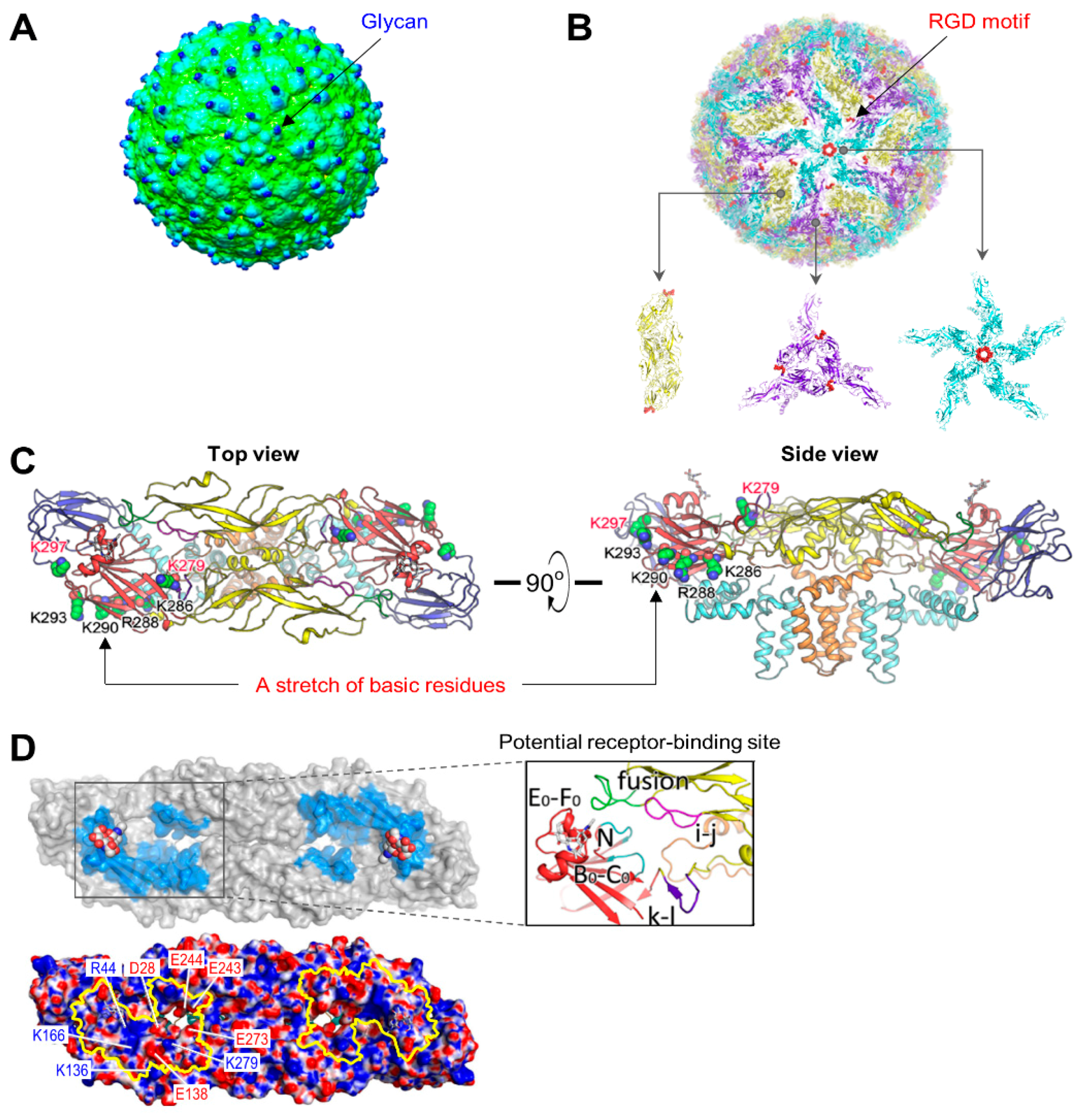
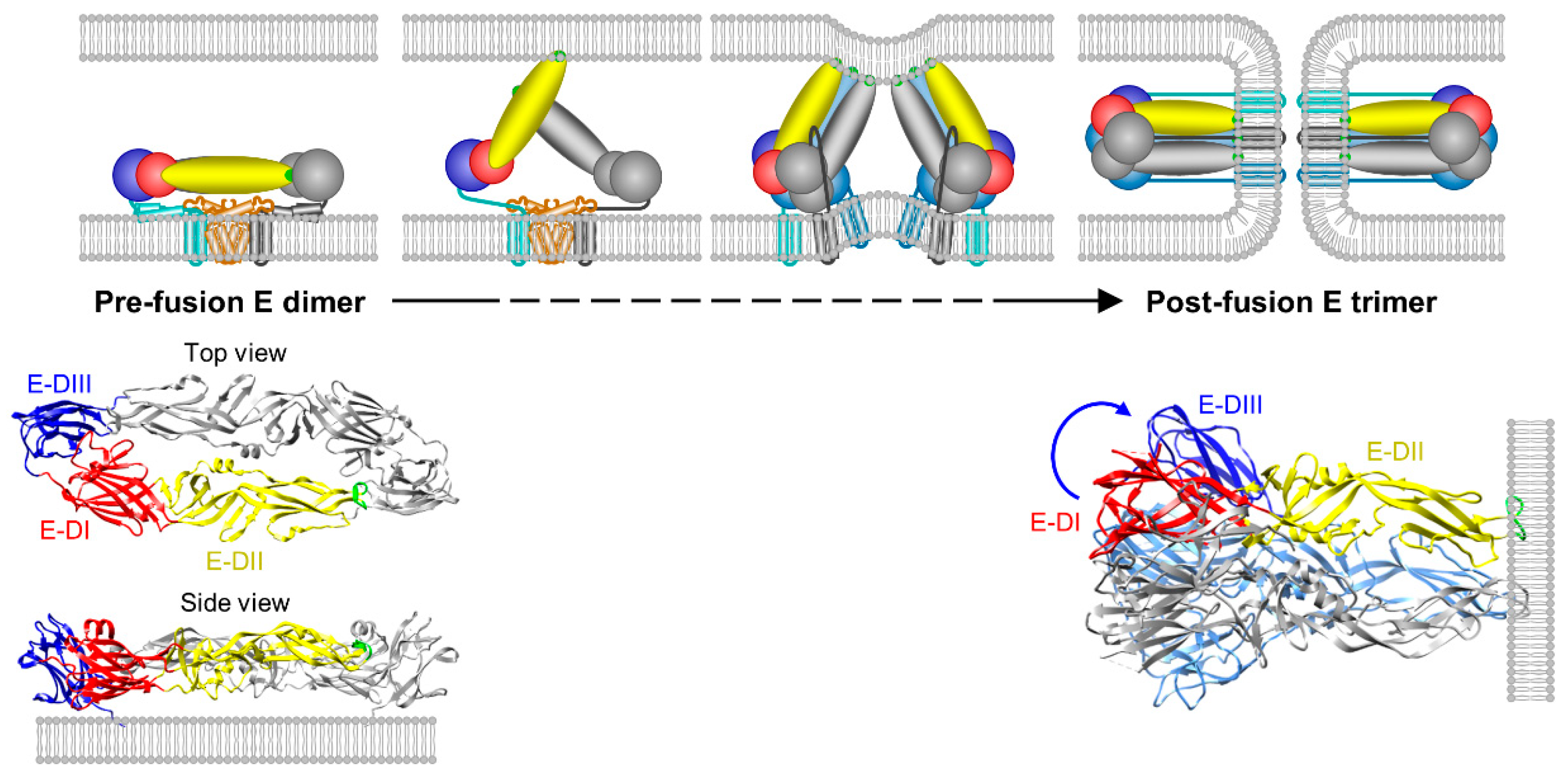
| Cellular Component | Type/Property | Host Cell | Virus |
|---|---|---|---|
| Heparan sulfate | Glycosaminoglycan | Mammal | JEV, WNV, MVEV, ZIKV, DENV, YFV, TBEV |
| DC-SIGN/L-SIGN | C-type lectin | Mammal | JEV, WNV, DENV |
| MR | C-type lectin | Mammal | JEV, DENV, TBEV |
| CLEC5A | C-type lectin | Mammal | JEV, DENV |
| LSECtin | C-type lectin | Mammal | JEV |
| mosGCTL-7, 1, and 3 | C-type lectin | Mosquito | JEV, WNV, DENV |
| αvβ3 | Integrin | Mammal | JEV, WNV |
| HSP70/HSC70/GRP78 | 70-kDa heat shock protein | Mammal, Mosquito | JEV, DENV |
| HSP90 | 90-kDa heat shock protein | Mammal | JEV, DENV |
| 37/67-kDa LR | High-affinity laminin receptor | Mammal | JEV |
| CD4 | Immunoglobulin superfamily | Mammal | JEV |
| CD14 | Pattern recognition receptor | Mammal | JEV |
| Vimentin | Type III intermediate filament | Mammal | JEV |
| LDLR | Low-density lipoprotein receptor | Mammal | JEV |
| 74-kDa protein | Not characterized | Mammal | JEV |
| 53-kDa protein | Not characterized | Mosquito | JEV |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yun, S.-I.; Lee, Y.-M. Early Events in Japanese Encephalitis Virus Infection: Viral Entry. Pathogens 2018, 7, 68. https://doi.org/10.3390/pathogens7030068
Yun S-I, Lee Y-M. Early Events in Japanese Encephalitis Virus Infection: Viral Entry. Pathogens. 2018; 7(3):68. https://doi.org/10.3390/pathogens7030068
Chicago/Turabian StyleYun, Sang-Im, and Young-Min Lee. 2018. "Early Events in Japanese Encephalitis Virus Infection: Viral Entry" Pathogens 7, no. 3: 68. https://doi.org/10.3390/pathogens7030068
APA StyleYun, S.-I., & Lee, Y.-M. (2018). Early Events in Japanese Encephalitis Virus Infection: Viral Entry. Pathogens, 7(3), 68. https://doi.org/10.3390/pathogens7030068






