Transcriptional Profiling of a Cross-Protective Salmonella enterica serovar Typhimurium UK-1 dam Mutant Identifies a Set of Genes More Transcriptionally Active Compared to Wild-Type, and Stably Transcribed across Biologically Relevant Microenvironments
Abstract
:1. Introduction
2. Results
2.1. UK-1 dam Mutant Produces Higher Levels of Transcription in Direct Comparison to UK-1 wt Parent Strain
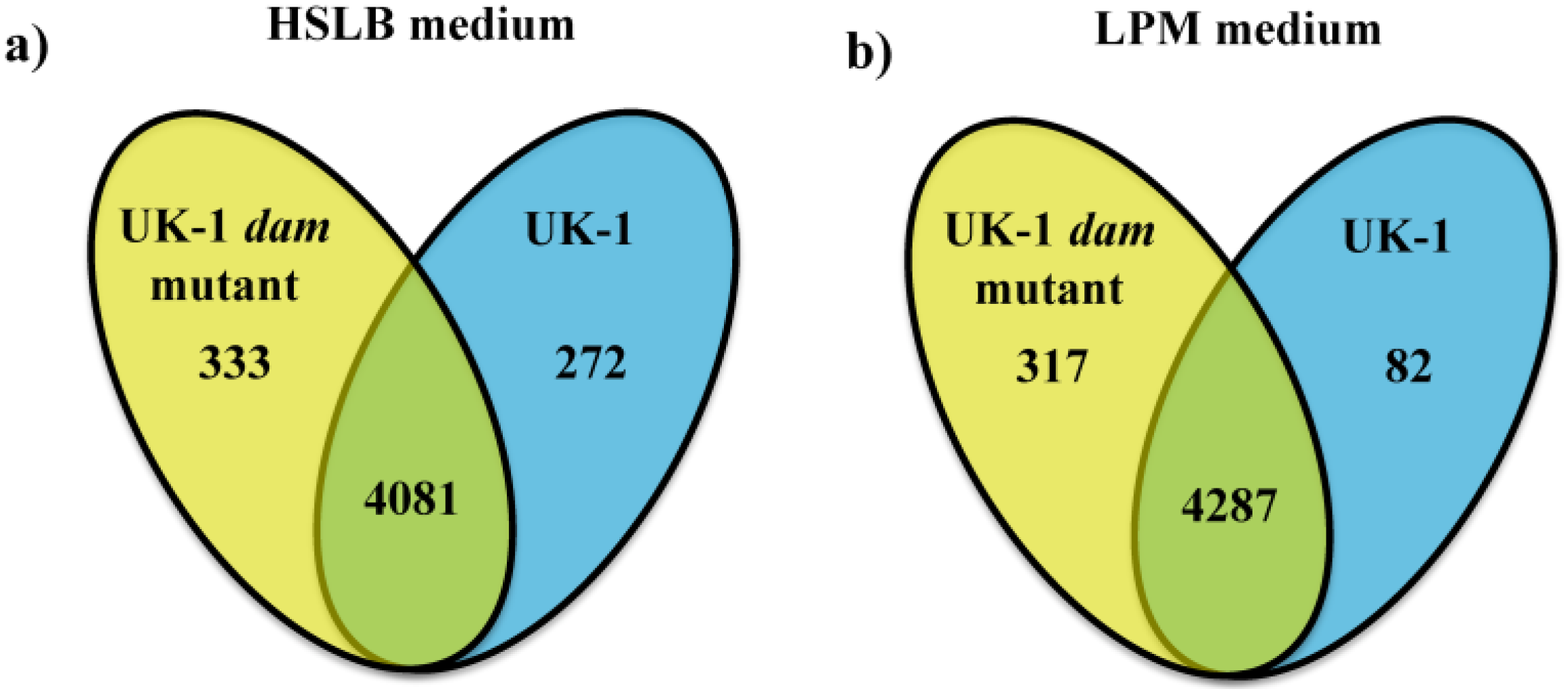

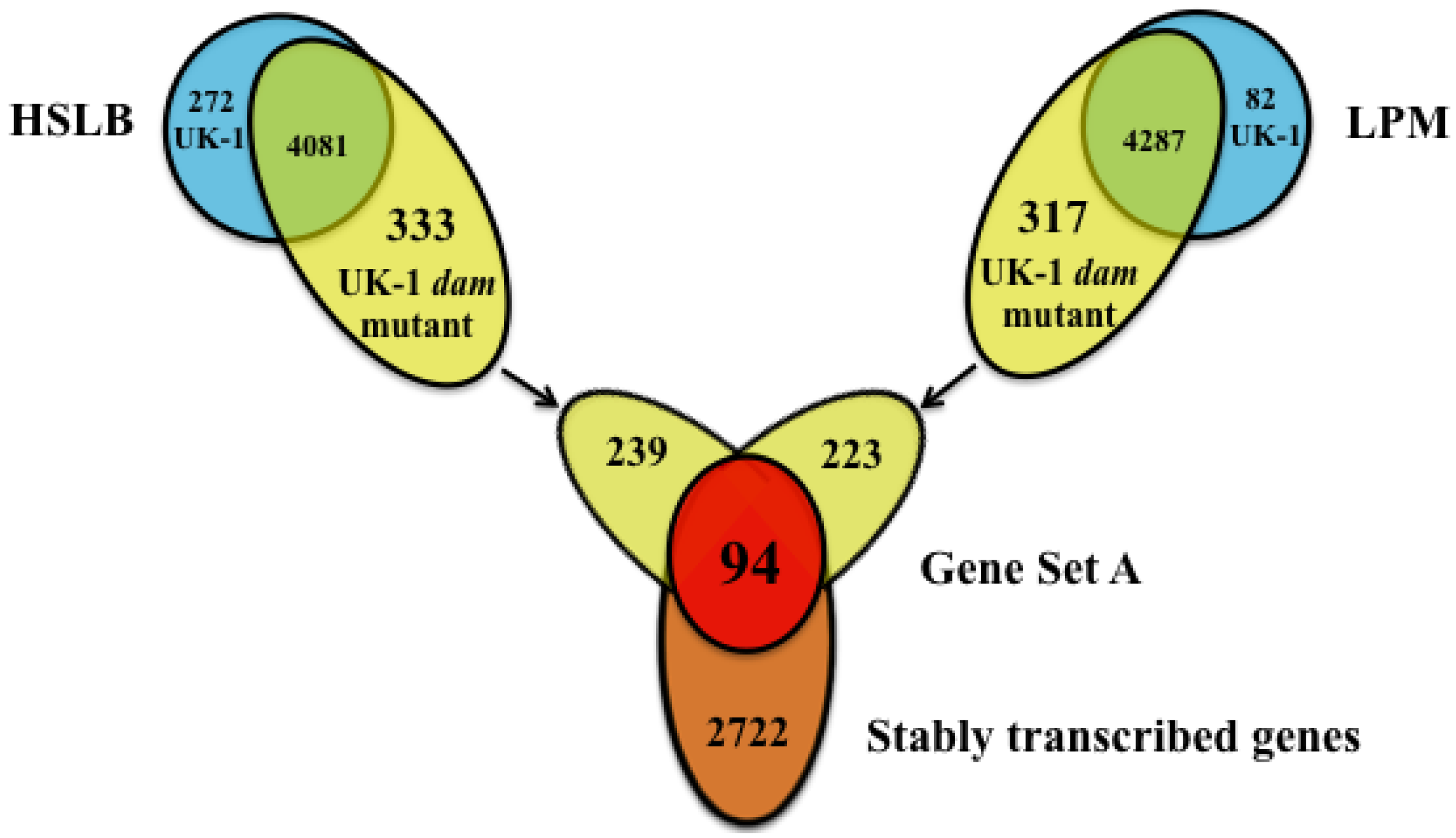
2.2. UK-1 dam Mutant and UK-1 wt Parent Strain Show Comparable Stable Transcription in Response to Growth in Differing Microenvironments
2.3. Identification of Genes More Highly Transcribed by the UK-1 dam Mutant and UK-1 wt Parent Strain Regardless of Culture Condition
| Functional Category | Gene ID | Fold Change of UK-1 dam Mutant a | Gene Product | |||
|---|---|---|---|---|---|---|
| vs. UK-1 in HSLB | vs. UK-1 in LPM | vs. Dublin in HSLB | vs. Newport in HSLB | |||
| Fimbriae | STMUK_3014 b | 350.44 | 93.28 | 350.55 | 859.27 | Hypothetical protein; std operon |
| STMUK_3015 | 267.40 | 119.79 | 239.21 | NP c | Putative outer membrane protein; std operon | |
| Bacteriophage | STMUK_1981 | 6.81 | 4.53 | 15.96 | NP | Hypothetical bacteriophage protein |
| STMUK_1982 | 6.90 | 10.08 | 17.47 | NP | Putative phage tail-like protein | |
| STMUK_1983 | 8.27 | 11.74 | 15.83 | NP | Hypothetical bacteriophage protein | |
| STMUK_1984 | 8.06 | 9.40 | 15.30 | NP | Similar to tail protein | |
| STMUK_1985 | 6.32 | 7.46 | 25.19 | NP | Similar to tail protein | |
| STMUK_1986 | 4.91 | 10.49 | 18.11 | NP | Similar to phage protein | |
| STMUK_1987 | 9.94 | 7.56 | 35.25 | NP | Putative bacteriophage protein | |
| STMUK_1988 | 8.74 | 8.21 | 21.68 | NP | Putative bacteriophage protein | |
| STMUK_1989 | 7.30 | 10.28 | 24.59 | NP | Putative bacteriophage protein | |
| STMUK_1990 | 7.13 | 13.77 | 29.52 | NP | Putative bacteriophage protein | |
| STMUK_1991 | 7.04 | 13.51 | 32.31 | NP | Hypothetical bacteriophage protein | |
| STMUK_1992 | 8.08 | 18.23 | 23.22 | NP | Putative tail tube protein | |
| STMUK_1993 | 7.41 | 22.23 | 80.27 | NP | Putative bacteriophage protein | |
| STMUK_1994 | 7.28 | 21.15 | 69.99 | NP | Hypothetical bacteriophage protein | |
| STMUK_1995 | 8.96 | 30.02 | 76.36 | NP | Hypothetical bacteriophage protein | |
| STMUK_1996 | 9.84 | 26.91 | 150.38 | NP | Hypothetical bacteriophage protein | |
| STMUK_1997 | 7.64 | 23.75 | 98.95 | NP | Hypothetical bacteriophage protein | |
| STMUK_1998 | 8.35 | 21.73 | 43.65 | NP | Hypothetical bacteriophage protein | |
| STMUK_1999 | 7.27 | 24.96 | 66.19 | NP | Hypothetical bacteriophage protein | |
| STMUK_2000 | 7.11 | 24.09 | 189.35 | NP | Putative bacteriophage protein | |
| STMUK_2001 | 7.97 | 25.17 | 311.71 | NP | Putative bacteriophage protein | |
| STMUK_2002 | 9.29 | 19.64 | 52.79 | NP | Putative bacteriophage protein | |
| STMUK_2003 | 6.66 | 22.35 | 68.68 | NP | Hypothetical bacteriophage protein | |
| STMUK_2004 | 7.86 | 20.07 | 30.54 | NP | Putative bacteriophage protein | |
| STMUK_2005 | 10.31 | 23.71 | 28.43 | NP | Hypothetical bacteriophage protein | |
| STMUK_2006 | 8.28 | 16.09 | 80.27 | NP | Hypothetical bacteriophage protein | |
| STMUK_2007 | 8.38 | 18.22 | 404.88 | 352.87 | Putative bacteriophage protein | |
| STMUK_2008 | 9.24 | 24.28 | NP | NP | Hypothetical bacteriophage protein | |
| STMUK_2009 | 3.68 | 5.56 | NP | NP | Hypothetical bacteriophage protein | |
| STMUK_2010 | 8.16 | 8.61 | NP | NP | Hypothetical bacteriophage protein | |
| STMUK_2011 | 7.18 | 7.15 | 9.98 | NP | Putative cytoplasmic protein | |
| STMUK_2012 | 7.72 | 8.67 | 5.83 | NP | Hypothetical bacteriophage protein | |
| STMUK_2013 | 12.83 | 9.73 | 7.24 | NP | Hypothetical bacteriophage protein | |
| STMUK_2014 | 10.00 | 16.93 | NP | NP | Putative bacteriophage protein | |
| STMUK_2015 | 10.12 | 21.50 | NP | NP | Hypothetical bacteriophage protein | |
| STMUK_2016 | 7.86 | 31.44 | 6.02 | NP | Hypothetical bacteriophage protein | |
| STMUK_2017 | 9.07 | 17.72 | NP | NP | Hypothetical bacteriophage protein | |
| STMUK_2018 | 9.06 | 13.96 | 4.03 | NP | Hypothetical bacteriophage protein | |
| STMUK_2019 | 7.35 | 12.62 | 4.26 | NP | Hypothetical bacteriophage protein | |
| STMUK_2021 | 13.14 | 26.39 | 3.18 | NP | Hypothetical bacteriophage protein | |
| STMUK_2022 | 11.69 | 20.75 | 3.46 | NP | Hypothetical bacteriophage protein | |
| STMUK_2023 | 12.09 | 24.14 | 2.81 | NP | Similar to phage protein | |
| STMUK_2024 | 8.37 | 12.71 | NP | NP | Hypothetical bacteriophage protein | |
| STMUK_2025 | 9.37 | 10.70 | NP | NP | Hypothetical bacteriophage protein | |
| STMUK_2026 | 10.27 | 9.84 | NP | NP | Hypothetical bacteriophage protein | |
| STMUK_2027 | 5.98 | 7.97 | 7.79 | NP | Putative bacteriophage protein | |
| STMUK_2028 | 7.20 | 12.42 | 5.09 | NP | Hypothetical bacteriophage protein | |
| STMUK_2029 | 4.88 | 4.19 | 2.62 | NP | Similar to phage integrase | |
| SOS regulon | STMUK_1276 | 3.92 | 5.28 | −1.18 | 2.93 | Nucleotide excision repair endonuclease |
| dinF | 4.50 | 4.68 | 2.42 | 2.04 | DNA-damage-inducible SOS response protein | |
| dinP | 4.76 | 5.07 | 2.67 | 2.13 | DNA polymerase IV | |
| lexA | 4.15 | 3.90 | 2.34 | 2.31 | LexA repressor | |
| polB | 2.66 | 2.73 | 2.17 | NP | DNA polymerase II | |
| recA | 4.86 | 4.86 | 3.42 | 3.18 | Recombinase A | |
| recN | 10.81 | 10.62 | 5.80 | 4.34 | Recombination and repair protein | |
| ruvA | 2.78 | 2.57 | 2.43 | 2.30 | Holliday junction DNA helicase RuvA | |
| ruvB | 2.28 | 2.11 | 2.49 | 2.04 | Holliday junction DNA helicase | |
| sulA | 6.17 | 11.44 | 2.92 | 3.29 | SOS cell division inhibitor | |
| umuC | 13.56 | 5.11 | NP | 4.46 | DNA polymerase V subunit | |
| umuD | 14.35 | 8.31 | NP | 4.90 | DNA polymerase V subunit | |
| uvrA | 2.60 | 2.16 | 2.47 | 2.08 | Excinuclease ABC subunit A | |
| ydjM | 2.30 | 2.02 | 1.31 | 1.12 | LexA-regulated gene | |
| yebE | 4.84 | 3.76 | 4.59 | 2.37 | Putative inner membrane protein | |
| yebF | 5.08 | 2.58 | 4.89 | 2.00 | Putative secreted Protein | |
| yebG | 11.08 | 15.76 | 7.74 | 6.03 | DNA damage-inducible protein | |
| Replication | STMUK_0980 | 2.45 | 2.33 | 1.12 | 3.013 | Putative replication regulatory protein |
| Metabolism | STMUK_2034 | 2.58 | 2.03 | 5.49 | 2.505 | Putative endoprotease |
| STMUK_4015 | 3.00 | 3.08 | 2.46 | 1.47 | Putative acetyl esterase | |
| Transport | ompS | 2.99 | 2.04 | 14.14 | 1.36 | Putative porin |
| setB | 4.17 | 3.01 | 1.14 | 2.70 | Proton efflux pump | |
| Translation | asnT | 6.53 | 6.00 | −1.18 | −3.06 | tRNA-Asn |
| cysT | 2.21 | 2.62 | 3.01 | 1.30 | tRNA-Cys | |
| glyW | 2.13 | 2.77 | 4.37 | 1.61 | tRNA-Gly | |
| leuW | 2.67 | 2.09 | 3.34 | 2.33 | tRNA-Leu | |
| leuZ | 3.40 | 3.17 | 1.58 | 1.51 | tRNA-Leu | |
| metW | 2.08 | 2.39 | 1.22 | 1.80 | tRNA-Met | |
| tyrT | 2.26 | 2.82 | −1.27 | 1.94 | tRNA-Tyr | |
| STMUK_3506 | 3.75 | 9.86 | 5.78 | NP | Putative ribonucleoprotein-related protein | |
| Virulence | STMUK_1011 | 16.94 | 3.11 | 13.26 | 13.73 | Attachment/invasion protein |
| Hypothetical Proteins | STMUK_0985 | 2.54 | 4.97 | 2.98 | 1.04 | Hypothetical protein |
| STMUK_0986 | 4.54 | 3.45 | 10.26 | 1.83 | Hypothetical protein | |
| STMUK_1493 | 2.50 | 4.11 | 3.12 | 2.788 | Putative outer membrane protein | |
| STMUK_1849 | 5.81 | 13.81 | 4.77 | 28.65 | Hypothetical protein | |
| STMUK_2239 | 9.29 | 21.77 | −1.26 | 15.76 | Putative inner membrane protein | |
| STMUK_2240 | 15.57 | 28.49 | 1.27 | 59.26 | Putative inner membrane protein | |
| STMUK_2655 | 4.77 | 4.67 | 3.39 | 1.76 | Hypothetical protein | |
| STMUK_2656 | 5.04 | 12.62 | 12.65 | 26.71 | Hypothetical protein | |
| rtcB | 2.61 | 2.35 | 1.41 | NP | Putative cytoplasmic protein | |
| yeeA | 4.79 | 3.61 | 3.40 | 3.42 | Putative inner membrane protein | |
| Plasmid | ccdA | 7.39 | 4.61 | NP | NP | Toxin addiction system: antidote |
| ccdB | 4.32 | 2.95 | NP | NP | Toxin addiction system: toxin | |
| rsdB | 7.40 | 3.17 | NP | NP | Resolvase | |
| Functional Category | Gene ID | Fold Change of UK-1 wt Strain a | Gene Product | |
|---|---|---|---|---|
| vs. UK-1 dam Mutant in HSLB Medium | vs. UK-1 dam Mutant in LPM Medium | |||
| Metabolism | gph | 4.43 | 2.41 | Phosphoglycolate phosphatase |
| rpe | 6.56 | 5.82 | Ribulose-phosphate 3-epimerase | |
| sdaB | 3.56 | 4.47 | L-serine dehydratase/L-threonine deaminase 2 | |
| yadI | 3.45 | 2.10 | Putative PTS enzyme | |
| Transport | sdaC | 4.30 | 4.19 | Putative serine transport protein |
| Transcription | yiaJ | 2.53 | 2.11 | Transcriptional repressor |
| Translation | trpS | 3.43 | 2.00 | Tryptophanyl-tRNA synthetase |
2.4. Analysis of Gene Set A in S. Dublin and S. Newport
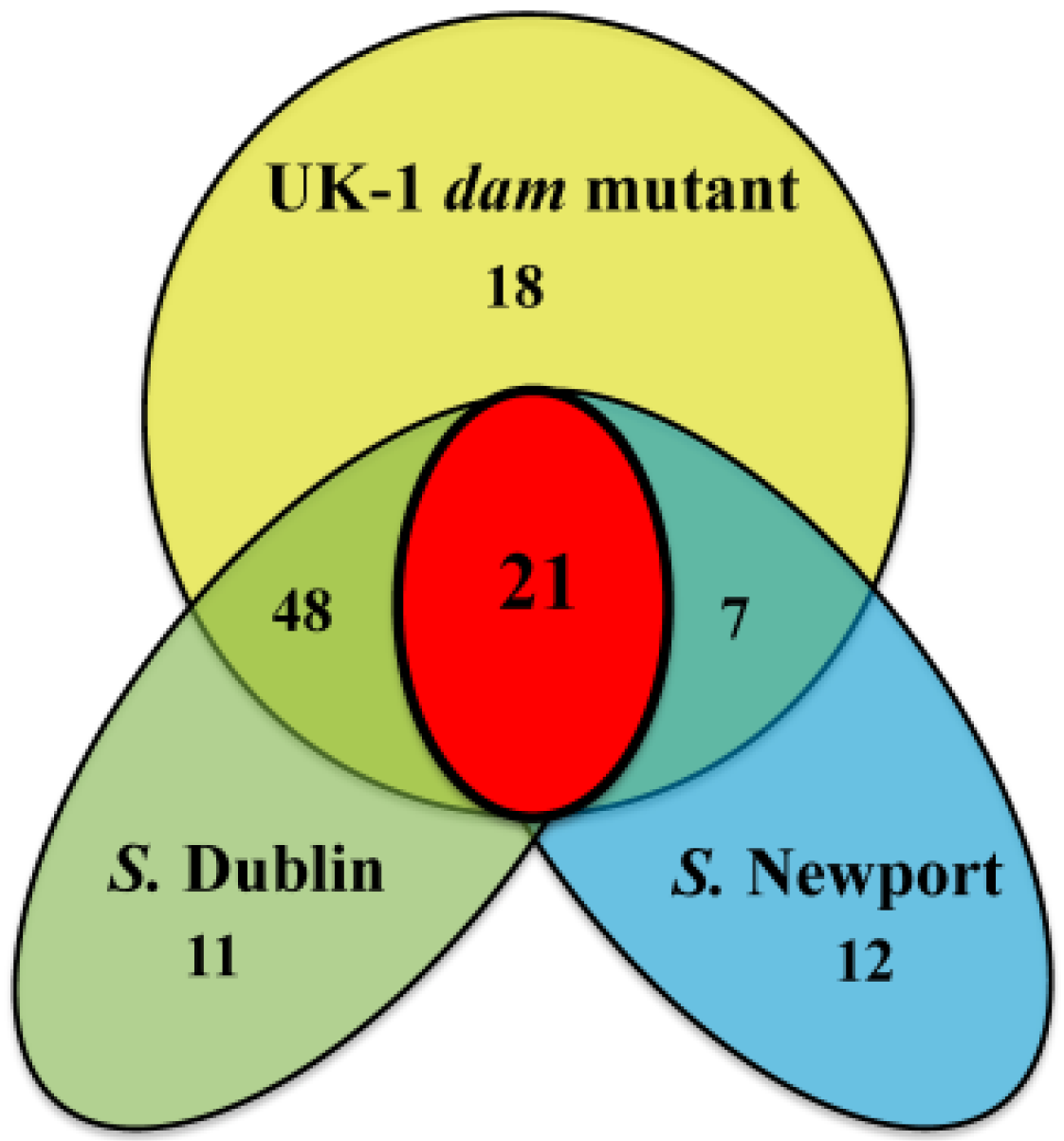
2.5. Validation of RNAseq Data with RT-PCR.
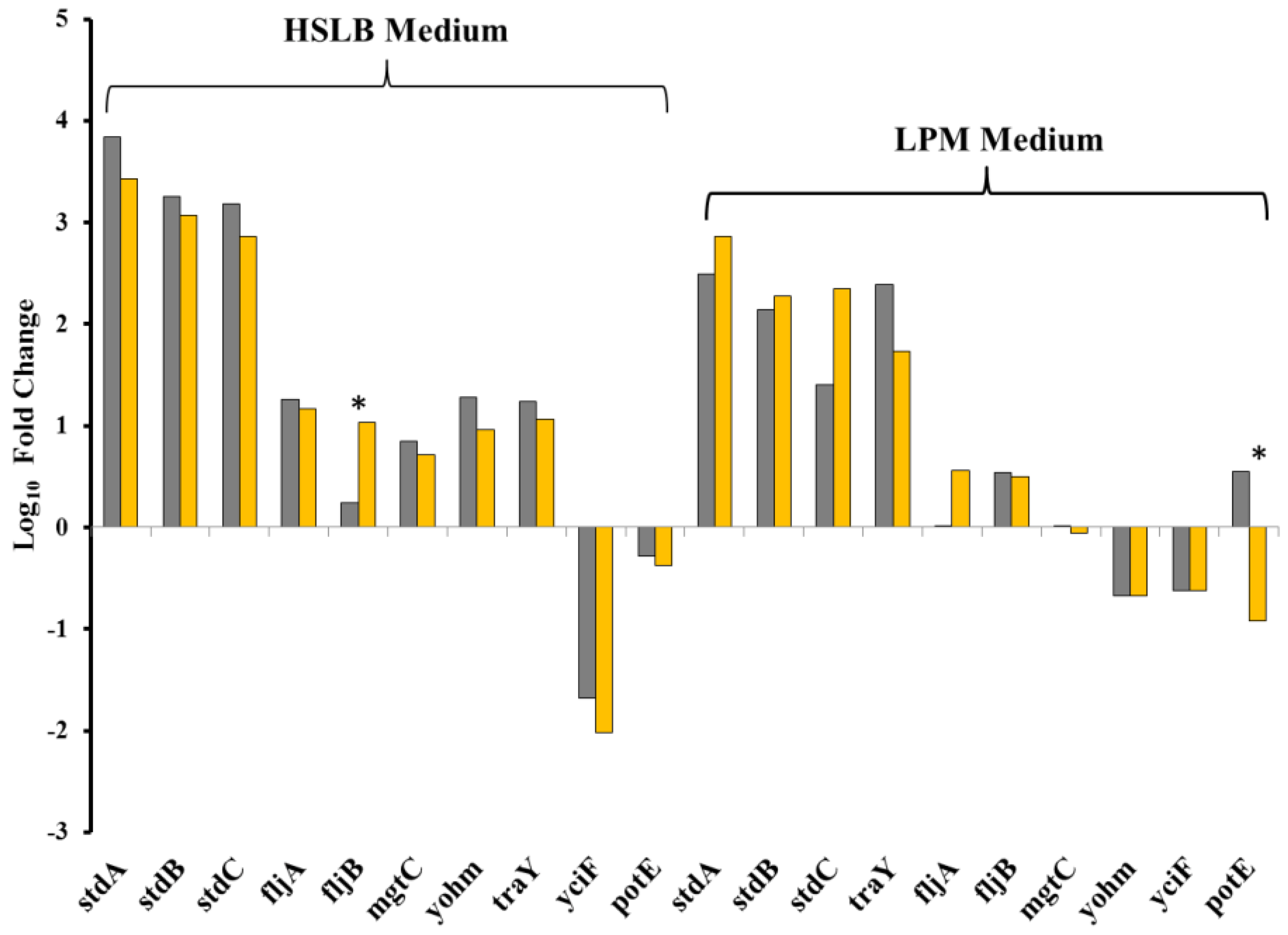

3. Discussion
4. Experimental Section
4.1. Bacterial Strains
| Strain | Clinical Significance | Source | PMID |
|---|---|---|---|
| Typhimurium dam102: Mud-Cm MT2313 | Avirulent | Heithoff 2001 Deuger 2003 | 11705984 |
| Typhimurium UK-1 | Highly virulent in mice, poultry, pigs, calves and horses | Curtiss 1991 Barrow 2001 Zhang 1999 | 21622747 |
| Newport 03-721 | Wild type isolated from an outbreak of salmonellosis in neonatal dairy calves | Mohler 2002 | 18329764 |
| Dublin MNO3-9704 | Wild type isolated from an outbreak of salmonellosis in neonatal dairy calves | Mohler 2006 | 16300866 |
4.2. Bacterial Culture Conditions
4.3. Transcriptome Sequencing and Analysis
4.4. Real-Time Reverse Transcriptase—PCR (RT-PCR)
| Gene ID | Forward Sequence 5ʹ-3ʹ | Reverse Sequence 5ʹ-3ʹ | Product Size (bp) |
|---|---|---|---|
| stdA | ACCATCACCAACTCACCCTGTGAT | GTGGTTGCATTGGCGGTATTCAGT | 104 |
| stdB | AGACGTACCTCAGCTCCGCTATTT | GCATTACTGTTCGCAATGCCGCTA | 133 |
| stdC | AGGACAGGGAAACACTGTTCTGGT | TCCGCTCAGCAGTCAGCTTTCTTT | 171 |
| fljA | CGTAAATGCGTGTCAGGTTGGTGT | TGATCCTGCTCACCCAGTCAAACA | 80 |
| fljB | ACCGTTTCACCGCGAACATCAAAG | TCAGTGGTCTGCGCAATGGAGATA | 82 |
| mgtC | ATTTACTGGCCGCTATGCTGTTGG | TAGTGCGTAATCCCGCCATACGTT | 81 |
| yohM | AGGTACGGTTAAACAGGCCGTCAT | TTCCACTGATTGTGCGGTGAATGC | 124 |
| yciF | ATTACGAAATCGCCAGCTACGGCA | TCGTCGAGGGTTTCTTTGAGCAGT | 94 |
| potE | AATAGCCTTCGTCAGCGGAGGATT | TTTGGTCTGGCGTTTGCACAGATG | 145 |
| traY | TGTGAGGAGGAGAAACGCAAGAGA | TTCCACTCCGATCTTTAGCCCTGA | 104 |
| gmk | TTGGCAGGGAGGCGTTT | GCGCGAAGTGCCGTAGTAAT | 61 |
| rpoD | ACATGGGTATTCAGGTAATGGAAGA | CGGTGCTGGTGGTATTTTCA | 74 |
5. Conclusions
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Flint, J.A.; Van Duynhoven, Y.T.; Angulo, F.J.; DeLong, S.M.; Braun, P.; Kirk, M.; Scallan, E.; Fitzgerald, M.; Adak, G.K.; Sockett, P.; et al. Estimating the burden of acute gastroenteritis, foodborne disease, and pathogens commonly transmitted by food: An international review. Clin. Infect. Dis. 2005, 41, 698–704. [Google Scholar] [CrossRef]
- Barrow, P.A.; Methner, U. Salmonella in Domestic Animals, 2nd ed.; CABI: Wallingford, UK, 2013. [Google Scholar]
- Anderson, R.J.; House, J.K.; Smith, B.P.; Kinde, H.; Walker, R.L.; Vande Steeg, B.J.; Breitmeyer, R.E. Epidemiologic and biological characteristics of salmonellosis in three dairy herds. J. Am. Vet. Med. Assoc. 2001, 219, 310–322. [Google Scholar] [CrossRef]
- Smith, B.P.; Da Roden, L.; Thurmond, M.C.; Dilling, G.W.; Konrad, H.; Pelton, J.A.; Picanso, J.P. Prevalence of salmonellae in cattle and in the environment on California dairies. J. Am. Vet. Med. Assoc. 1994, 205, 467–471. [Google Scholar]
- Gunn, J.S. Bacterial modification of LPS and resistance to antimicrobial peptides. J. Endotoxin Res. 2001, 7, 57–62. [Google Scholar] [CrossRef]
- Groisman, E.A. The pleiotropic two-component regulatory system PhoP-PhoQ. J. Bacteriol. 2001, 183, 1835–1842. [Google Scholar] [CrossRef]
- Bajaj, V.; Lucas, R.L.; Hwang, C.; Lee, C.A. Co-ordinate regulation of Salmonella typhimurium invasion genes by environmental and regulatory factors is mediated by control of hilA expression. Mol. Microbiol. 1996, 22, 703–714. [Google Scholar]
- Alpuche Aranda, C.M.; Swanson, J.A.; Loomis, W.P.; Miller, S.I. Salmonella typhimurium activates virulence gene transcription within acidified macrophage phagosomes. Proc. Natl. Acad. Sci. USA 1992, 89, 10079–10083. [Google Scholar] [CrossRef]
- Heider, L.C.; Meiring, R.W.; Hoet, A.E.; Gebreyes, W.A.; Funk, J.A.; Wittum, T.E. Evaluation of vaccination with a commercial subunit vaccine on shedding of Salmonella enterica in subclinically infected dairy cows. J. Am. Vet. Med. Assoc. 2008, 233, 466–469. [Google Scholar] [CrossRef]
- Hermesch, D.R.; Thomson, D.U.; Loneragan, G.H.; Renter, D.R.; White, B.J. Effects of a commercially available vaccine against Salmonella enterica serotype Newport on milk production, somatic cell count, and shedding of Salmonella organisms in female dairy cattle with no clinical signs of salmonellosis. Am. J. Vet. Res. 2008, 69, 1229–1234. [Google Scholar] [CrossRef]
- Alaniz, R.C.; Cummings, L.A.; Bergman, M.A.; Rassoulian-Barrett, S.L.; Cookson, B.T. Salmonella typhimurium coordinately regulates FliC location and reduces dendritic cell activation and antigen presentation to CD4+ T cells. J. Immunol. 2006, 177, 3983–3993. [Google Scholar] [CrossRef]
- Stewart, M.K.; Cummings, L.A.; Johnson, M.L.; Berezow, A.B.; Cookson, B.T. Regulation of phenotypic heterogeneity permits Salmonella evasion of the host caspase-1 inflammatory response. Proc. Natl. Acad. Sci. USA 2011, 108, 20742–20747. [Google Scholar]
- Heithoff, D.M.; Enioutina, E.Y.; Daynes, R.A.; Sinsheimer, R.L.; Low, D.A.; Mahan, M.J. Salmonella DNA adenine methylase mutants confer cross-protective immunity. Infect. Immun. 2001, 69, 6725–6730. [Google Scholar] [CrossRef]
- Dueger, E.L.; House, J.K.; Heithoff, D.M.; Mahan, M.J. Salmonella DNA adenine methylase mutants elicit early and late onset protective immune responses in calves. Vaccine 2003, 21, 3249–3258. [Google Scholar] [CrossRef]
- Dueger, E.L.; House, J.K.; Heithoff, D.M.; Mahan, M.J. Salmonella DNA adenine methylase mutants prevent colonization of newly hatched chickens by homologous and heterologous serovars. Int. J. Food Microbiol. 2003, 80, 153–159. [Google Scholar] [CrossRef]
- Dueger, E.L.; House, J.K.; Heithoff, D.M.; Mahan, M.J. Salmonella DNA adenine methylase mutants elicit protective immune responses to homologous and heterologous serovars in chickens. Infect. Immun. 2001, 69, 7950–7954. [Google Scholar] [CrossRef]
- Mohler, V.L.; Heithoff, D.M.; Mahan, M.J.; Walker, K.H.; Hornitzky, M.A.; McConnell, C.S.; Shum, L.W.C.; House, J.K. Cross-protective immunity in calves conferred by a DNA adenine methylase deficient Salmonella enterica serovar Typhimurium vaccine. Vaccine 2006, 24, 1339–1345. [Google Scholar] [CrossRef]
- Mohler, V.L.; Heithoff, D.M.; Mahan, M.J.; Walker, K.H.; Hornitzky, M.A.; Shum, L.W.C.; Makin, K.J.; House, J.K. Cross-protective immunity conferred by a DNA adenine methylase deficient Salmonella enterica serovar Typhimurium vaccine in calves challenged with Salmonella serovar Newport. Vaccine 2008, 26, 1751–1758. [Google Scholar] [CrossRef]
- Balbontin, R.; Rowley, G.; Pucciarelli, M.G.; Lopez-Garrido, J.; Wormstone, Y.; Lucchini, S.; Portillo, F.G.; Hinton, J.C.D.; Casadesus, J. DNA adenine methylation regulates virulence gene expression in Salmonella enterica serovar Typhimurium. J. Bacteriol. 2006, 188, 8160–8168. [Google Scholar] [CrossRef]
- Badie, G.; Heithoff, D.M.; Sinsheimer, R.L.; Mahan, M.J. Altered levels of Salmonella DNA adenine methylase are associated with defects in gene expression, motility, flagellar synthesis, and bile resistance in the pathogenic strain 14028 but not in the laboratory strain LT2. J. Bacteriol. 2007, 189, 1556–1564. [Google Scholar] [CrossRef]
- Heithoff, D.M.; Sinsheimer, R.L.; Low, D.A.; Mahan, M.J. An essential role for DNA adenine methylation in bacterial virulence. Science 1999, 284, 967–970. [Google Scholar] [CrossRef]
- Coombes, B.K.; Brown, N.F.; Valdez, Y.; Brumell, J.H.; Finlay, B.B. Expression and secretion of Salmonella pathogenicity island-2 virulence genes in response to acidification exhibit differential requirements of a functional type III secretion apparatus and SsaL. J. Biol. Chem. 2004, 279, 49804–49815. [Google Scholar]
- Ibarra, J.A.; Knodler, L.A.; Sturdevant, D.E.; Virtaneva, K.; Carmody, A.B.; Fischer, E.R.; Porcella, S.F.; Steele-Mortimer, O. Induction of Salmonella pathogenicity island 1 under different growth conditions can affect Salmonella-host cell interactions in vitro. Micobiology 2010, 156, 1120–1133. [Google Scholar]
- Kall, L.; Krogh, A.; Sonnhammer, E.L. A combined transmembrane topology and signal peptide prediction method. J. Mol. Biol 2004, 338, 1027–1036. [Google Scholar] [CrossRef]
- Lyngstadaas, A.; Lobner-Olesen, A.; Boye, E. Characterization of three genes in the dam-containing operon of Escherichia coli. Mol. Gen. Genet. 1995, 247, 546–554. [Google Scholar] [CrossRef]
- Lyngstadaas, A.; Lobner-Olesen, A.; Grelland, E.; Boye, E. The gene for 2-phosphoglycolate phosphatase (gph) in Escherichia coli is located in the same operon as dam and at least five other diverse genes. Biochim. Biophys. Acta 1999, 1472, 376–384. [Google Scholar] [CrossRef]
- Molina-Henares, A.J.; Krell, T.; Eugenia Guazzaroni, M.; Segura, A.; Ramos, J.L. Members of the IclR family of bacterial transcriptional regulators function as activators and/or repressors. FEMS Microbiol. Rev. 2006, 30, 157–186. [Google Scholar] [CrossRef]
- Townsend, S.M.; Kramer, N.E.; Edwards, R.; Baker, S.; Hamlin, N.; Simmonds, M.; Stevens, K.; Maloy, S.; Parkhill, J.; Dougan, G.; et al. Salmonella enterica serovar Typhi possesses a unique repertoire of fimbrial gene sequences. Infect. Immun. 2001, 69, 2894–2901. [Google Scholar] [CrossRef]
- Baumler, A.J.; Tsolis, R.M.; Bowe, F.A.; Kusters, J.G.; Hoffmann, S.; Heffron, F. The pef fimbrial operon of Salmonella typhimurium mediates adhesion to murine small intestine and is necessary for fluid accumulation in the infant mouse. Infect. Immun. 1996, 64, 61–68. [Google Scholar]
- Baumler, A.J.; Tsolis, R.M.; Heffron, F. Fimbrial adhesins of Salmonella typhimurium. Role in bacterial interactions with epithelial cells. Adv. Exp. Med. Biol. 1997, 412, 149–158. [Google Scholar] [CrossRef]
- Humphries, A.D.; Townsend, S.M.; Kingsley, R.A.; Nicholson, T.L.; Tsolis, R.M.; Baumler, A.J. Role of fimbriae as antigens and intestinal colonization factors of Salmonella serovars. FEMS Microbiol. Lett. 2001, 201, 121–125. [Google Scholar] [CrossRef]
- Humphries, A.; Deridder, S.; Baumler, A.J. Salmonella enterica serotype Typhimurium fimbrial proteins serve as antigens during infection of mice. Infect. Immun. 2005, 73, 5329–5338. [Google Scholar] [CrossRef]
- Khatri, N.S.; Maskey, P.; Poudel, S.; Jaiswal, V.K.; Karkey, A.; Koirala, S.; Shakya, N.; Agrawal, K.; Arjyal, A.; Basnyat, B.; et al. Gallbladder carriage of Salmonella paratyphi A may be an important factor in the increasing incidence of this infection in South Asia. Ann. Intern. Med. 2009, 150, 567–568. [Google Scholar] [CrossRef]
- Chessa, D.; Winter, M.G.; Nuccio, S.P.; Tukel, C.; Baumler, A.J. RosE represses Std fimbrial expression in Salmonella enterica serotype Typhimurium. Mol. Microbiol. 2008, 68, 573–587. [Google Scholar] [CrossRef]
- Mmolawa, P.T.; Schmieger, H.; Heuzenroeder, M.W. Bacteriophage ST64B, a genetic mosaic of genes from diverse sources isolated from Salmonella enterica serovar Typhimurium DT 64. J. Bacteriol. 2003, 185, 6481–6485. [Google Scholar] [CrossRef]
- Alonso, A.; Pucciarelli, M.G.; Figueroa-Bossi, N.; Garcia-del Portillo, F. Increased excision of the Salmonella prophage ST64B caused by a deficiency in Dam methylase. J. Bacteriol. 2005, 187, 7901–7911. [Google Scholar] [CrossRef]
- Gorski, A.; Miedzybrodzki, R.; Borysowski, J.; Dabrowska, K.; Wierzbicki, P.; Ohams, M.; Korczak-Kowalska, G.; Olszowska-Zaremba, N.; Lusiak-Szelachowska, M.; Klak, M.; et al. Phage as a modulator of immune responses: Practical implications for phage therapy. Adv. Virus Res. 2012, 83, 41–71. [Google Scholar] [CrossRef]
- Weber-Dabrowska, B.; Zimecki, M.; Mulczyk, M. Effective phage therapy is associated with normalization of cytokine production by blood cell cultures. Arch. Immunol. Ther. Exp. (Warsz) 2000, 48, 31–37. [Google Scholar]
- Shtrichman, R.; Heithoff, D.M.; Mahan, M.J.; Samuel, C.E. Tissue selectivity of interferon-stimulated gene expression in mice infected with Dam+ versus Dam− Salmonella enterica serovar Typhimurium strains. Infect. Immun. 2002, 70, 5579–5588. [Google Scholar] [CrossRef]
- Simon, R.; Heithoff, D.M.; Mahan, M.J.; Samuel, C.E. Comparison of tissue-selective proinflammatory gene induction in mice infected with wild-type, DNA adenine methylase-deficient, and flagellin-deficient Salmonella enterica. Infect. Immun. 2007, 75, 5627–5639. [Google Scholar] [CrossRef]
- Eriksson, F.; Tsagozis, P.; Lundberg, K.; Parsa, R.; Mangsbo, S.M.; Persson, M.A.; Harris, R.A.; Pisa, P. Tumor-specific bacteriophages induce tumor destruction through activation of tumor-associated macrophages. J. Immunol. 2009, 182, 3105–3111. [Google Scholar] [CrossRef]
- Luo, Y.Q.; Kong, Q.K.; Yang, J.; Golden, G.; Wanda, S.Y.; Jensen, R.V.; Ernst, P.B.; Curtiss, R. Complete Genome Sequence of the Universal Killer Salmonella enterica Serovar Typhimurium UK-1 (ATCC 68169). J. Bacteriol. 2011, 193, 4035–4036. [Google Scholar] [CrossRef]
- Kal, A.J.; van Zonneveld, A.J.; Benes, V.; van den Berg, M.; Koerkamp, M.G.; Albermann, K.; Strack, N.; Ruijter, J.M.; Richter, A.; Dujon, B.; et al. Dynamics of gene expression revealed by comparison of serial analysis of gene expression transcript profiles from yeast grown on two different carbon sources. Mol. Biol. Cell. 1999, 10, 1859–1872. [Google Scholar] [CrossRef]
- Botteldoorn, N.; Van Coillie, E.; Grijspeerdt, K.; Werbrouck, H.; Haesebrouck, F.; Donne, E.; D'Haese, E.; Heyndrickx, M.; Pasmans, F.; Herman, L. Real-time reverse transcription PCR for the quantification of the mntH expression of Salmonella enterica as a function of growth phase and phagosome-like conditions. J. Microbiol. Meth. 2006, 66, 125–135. [Google Scholar] [CrossRef]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Miller, C.B.; Pierlé, S.A.; Brayton, K.A.; Ochoa, J.N.; Shah, D.H.; Lahmers, K.K. Transcriptional Profiling of a Cross-Protective Salmonella enterica serovar Typhimurium UK-1 dam Mutant Identifies a Set of Genes More Transcriptionally Active Compared to Wild-Type, and Stably Transcribed across Biologically Relevant Microenvironments. Pathogens 2014, 3, 417-436. https://doi.org/10.3390/pathogens3020417
Miller CB, Pierlé SA, Brayton KA, Ochoa JN, Shah DH, Lahmers KK. Transcriptional Profiling of a Cross-Protective Salmonella enterica serovar Typhimurium UK-1 dam Mutant Identifies a Set of Genes More Transcriptionally Active Compared to Wild-Type, and Stably Transcribed across Biologically Relevant Microenvironments. Pathogens. 2014; 3(2):417-436. https://doi.org/10.3390/pathogens3020417
Chicago/Turabian StyleMiller, Claire B., Sebastian Aguilar Pierlé, Kelly A. Brayton, Jennine N. Ochoa, Devendra H. Shah, and Kevin K. Lahmers. 2014. "Transcriptional Profiling of a Cross-Protective Salmonella enterica serovar Typhimurium UK-1 dam Mutant Identifies a Set of Genes More Transcriptionally Active Compared to Wild-Type, and Stably Transcribed across Biologically Relevant Microenvironments" Pathogens 3, no. 2: 417-436. https://doi.org/10.3390/pathogens3020417
APA StyleMiller, C. B., Pierlé, S. A., Brayton, K. A., Ochoa, J. N., Shah, D. H., & Lahmers, K. K. (2014). Transcriptional Profiling of a Cross-Protective Salmonella enterica serovar Typhimurium UK-1 dam Mutant Identifies a Set of Genes More Transcriptionally Active Compared to Wild-Type, and Stably Transcribed across Biologically Relevant Microenvironments. Pathogens, 3(2), 417-436. https://doi.org/10.3390/pathogens3020417





