Integrating MALDI-TOF Mass Spectrometry and Machine Learning for Rapid and Clinically Relevant Differentiation of MRSA and MSSA
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Isolates and Culture Conditions
2.2. Antimicrobial Susceptibility Testing (AST)
2.3. MALDI-TOF MS Sample Preparation
2.4. MALDI-TOF MS Data Acquisition
2.5. Instrument Calibration and Quality Control
2.6. Spectral Preprocessing and Peak Detection
2.7. Statistical Analysis of Discriminatory Peaks
2.8. Machine Learning Classification and Model Evaluation
3. Results
3.1. Study Population and MALDI-TOF MS Data Overview
3.2. Global Spectral Variability Between MRSA and MSSA Isolates
3.3. Identification of Discriminatory MALDI-TOF MS Peaks
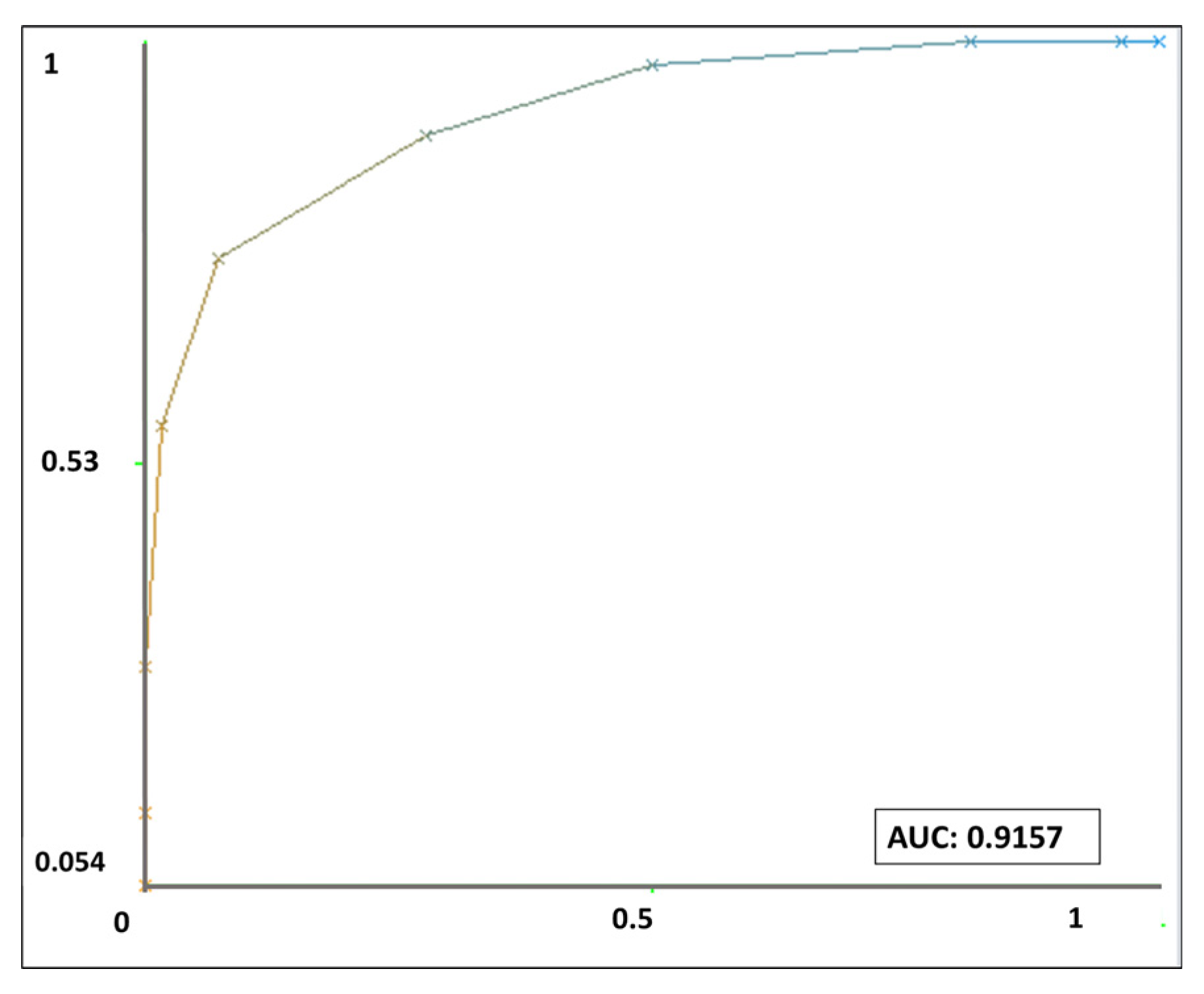
3.4. Machine Learning Classification Performance
3.5. Class-Specific Performance and Confusion Matrix Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| MALDI-TOF MS | Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry |
| MRSA | Methicillin-Resistant Staphylococcus aureus |
| MSSA | Methicillin-Susceptible Staphylococcus aureus |
| AST | Antimicrobial Susceptibility Testing |
| PCA | Principal Component Analysis |
| RF | Random Forest |
| ROC-AUC | Receiver Operating Characteristic Area Under the Curve |
| FDR | False Discovery Rate |
| EUCAST | European Committee on Antimicrobial Susceptibility Testing |
References
- Ojima-Kato, T.; Yamamoto, N.; Iijima, Y.; Tamura, H. Assessing the performance of novel software Strain Solution on automated discrimination of Escherichia coli serotypes and their mixtures using matrix-assisted laser desorption ionization-time of flight mass spectrometry. J. Microbiol. Methods 2015, 119, 233–238. [Google Scholar] [CrossRef] [PubMed]
- Santos, P.; Alho, I.; Ribeiro, E. MALDI-TOF MS Biomarkers for Methicillin-Resistant Staphylococcus aureus Detection: A Systematic Review. Metabolites 2025, 15, 540. [Google Scholar] [CrossRef] [PubMed]
- Florio, W.; Baldeschi, L.; Rizzato, C.; Tavanti, A.; Ghelardi, E.; Lupetti, A. Detection of Antibiotic-Resistance by MALDI-TOF Mass Spectrometry: An Expanding Area. Front. Cell. Infect. Microbiol. 2020, 10, 572909. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.M.; Kim, I.; Chung, S.H.; Chung, Y.; Han, M.; Kim, J.S. Rapid Discrimination of Methicillin-Resistant Staphylococcus aureus by MALDI-TOF MS. Pathogens 2019, 8, 214. [Google Scholar] [CrossRef]
- Alksne, L.; Makarova, S.; Avsejenko, J.; Cibrovska, A.; Trofimova, J.; Valciņa, O. Determination of methicillin-resistant Staphylococcus aureus and Staphylococcus epidermidis by MALDI-TOF MS in clinical isolates from Latvia. Clin. Mass Spectrom. 2020, 16, 33–39. [Google Scholar] [CrossRef]
- Gao, W.; Li, B.; Ling, L.; Zhang, L.; Yu, S. MALDI-TOF MS method for differentiation of methicillin-sensitive and methicillin-resistant Staphylococcus aureus using (E)-Propyl α-cyano-4-Hydroxyl cinnamylate. Talanta 2022, 244, 123405. [Google Scholar] [CrossRef]
- Yu, J.; Tien, N.; Liu, Y.C.; Cho, D.Y.; Chen, J.W.; Tsai, Y.T.; Huang, Y.C.; Chao, H.J.; Chen, C.J. Rapid Identification of Methicillin-Resistant Staphylococcus aureus Using MALDI-TOF MS and Machine Learning from over 20,000 Clinical Isolates. Microbiol. Spectr. 2022, 10, e0048322. [Google Scholar] [CrossRef]
- Weis, C.V.; Jutzeler, C.R.; Borgwardt, K. Machine learning for microbial identification and antimicrobial susceptibility testing on MALDI-TOF mass spectra: A systematic review. Clin. Microbiol. Infect. 2020, 26, 1310–1317. [Google Scholar] [CrossRef]
- Jeon, K.; Kim, J.-M.; Rho, K.; Jung, S.H.; Park, H.S.; Kim, J.-S. Performance of a Machine Learning-Based Methicillin Resistance of Staphylococcus aureus Identification System Using MALDI-TOF MS and Comparison of the Accuracy according to SCCmec Types. Microorganisms 2022, 10, 1903. [Google Scholar] [CrossRef]
- Santiago, L.S.; Guerrero-López, A.; Sevilla-Salcedo, C.; Rodríguez-Temporal, D.; Rodríguez-Sánchez, B.; Gómez-Verdejo, V. Machine Learning applied to MALDI-TOF data in a clinical setting: A systematic review. bioRxiv 2025. [Google Scholar] [CrossRef]
- Vela-Bernal, S.; Bea-Serrano, C.; de Gracia-Leon, A.I.; Belmonte-Domingo, A.; Pinto-Pla, C.; Ferrer-Ribera, A.; de Castro-Oliver, A.; Torres, I.; Colomina, J.; Oltra-Sempere, M.R. Impact of MALDI-TOF implementation and antimicrobial stewardship programs on the optimization of antibiotic therapy in Staphylococcus aureus bloodstream infections. Enfermedades Infecc. Microbiol. Clin. 2025, 43, 674–681. [Google Scholar] [CrossRef]
- Ceylan, A.N.; Sümbül, B.; Doymaz, M.Z. Screening of mecC Gene in Methicillin Resistant Staphylococcus aureus Isolates. Bezmiâlem Sci. 2022, 10, 226–230. [Google Scholar] [CrossRef]
- Gibb, S.; Strimmer, K. MALDIquant: A versatile R package for the analysis of mass spectrometry data. Bioinformatics 2012, 28, 2270–2271. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.Y.; Chung, C.R.; Wang, Z.; Li, S.; Chu, B.Y.; Horng, J.T.; Lu, J.J.; Lee, T.Y. A large-scale investigation and identification of methicillin-resistant Staphylococcus aureus based on peaks binning of matrix-assisted laser desorption ionization-time of flight MS spectra. Brief. Bioinform. 2021, 22, bbaa138. [Google Scholar] [CrossRef]
- Złoch, M.; Pomastowski, P.; Maślak, E.; Monedeiro, F.; Buszewski, B. Study on Molecular Profiles of Staphylococcus aureus Strains: Spectrometric Approach. Molecules 2020, 25, 4894. [Google Scholar] [CrossRef]
- Rhoads, D.D.; Wang, H.; Karichu, J.; Richter, S.S. The presence of a single MALDI-TOF mass spectral peak predicts methicillin resistance in staphylococci. Diagn. Microbiol. Infect. Dis. 2016, 86, 257–261. [Google Scholar] [CrossRef]
- Du, Z.; Yang, R.; Guo, Z.; Song, Y.; Wang, J. Identification of Staphylococcus aureus and determination of its methicillin resistance by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Anal. Chem. 2002, 74, 5487–5491. [Google Scholar] [CrossRef]
- Paskova, V.; Chudejova, K.; Sramkova, A.; Kraftova, L.; Jakubu, V.; Petinaki, E.A.; Zemlickova, H.; Neradova, K.; Papagiannitsis, C.C.; Hrabak, J. Insufficient repeatability and reproducibility of MALDI-TOF MS-based identification of MRSA. Folia Microbiol. 2020, 65, 895–900. [Google Scholar] [CrossRef]
- Hu, Y.; Huang, Y.; Lizou, Y.; Li, J.; Zhang, R. Evaluation of Staphylococcus aureus Subtyping Module for Methicillin-Resistant Staphylococcus aureus Detection Based on Matrix-Assisted Laser Desorption Ionization Time-of-Flight Mass Spectrometry. Front. Microbiol. 2019, 10, 2504. [Google Scholar] [CrossRef]
- Josten, M.; Dischinger, J.; Szekat, C.; Reif, M.; Al-Sabti, N.; Sahl, H.G.; Parcina, M.; Bekeredjian-Ding, I.; Bierbaum, G. Identification of agr-positive methicillin-resistant Staphylococcus aureus harbouring the class A mec complex by MALDI-TOF mass spectrometry. Int. J. Med. Microbiol. 2014, 304, 1018–1023. [Google Scholar] [CrossRef]
- Ng, W. Annotation of ribosomal protein mass peaks in MALDI-TOF mass spectra of bacterial species and their phylogenetic significance. Arch. Clin. Biomed. Res. 2021, 5, 862–904. [Google Scholar] [CrossRef]
- Kong, P.H.; Chiang, C.H.; Lin, T.C.; Kuo, S.C.; Li, C.F.; Hsiung, C.A.; Shiue, Y.L.; Chiou, H.Y.; Wu, L.C.; Tsou, H.H. Discrimination of Methicillin-resistant Staphylococcus aureus by MALDI-TOF Mass Spectrometry with Machine Learning Techniques in Patients with Staphylococcus aureus Bacteremia. Pathogens 2022, 11, 586. [Google Scholar] [CrossRef]
- Liu, X.; Su, T.; Hsu, Y.S.; Yu, H.; Yang, H.S.; Jiang, L.; Zhao, Z. Rapid identification and discrimination of methicillin-resistant Staphylococcus aureus strains via matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Rapid Commun. Mass Spectrom. 2021, 35, e8972. [Google Scholar] [CrossRef]
- Rybak, M.J.; Le, J.; Lodise, T.P.; Levine, D.P.; Bradley, J.S.; Liu, C.; Mueller, B.A.; Pai, M.P.; Wong-Beringer, A.; Rotschafer, J.C.; et al. Therapeutic Monitoring of Vancomycin for Serious Methicillin-resistant Staphylococcus aureus Infections: A Revised Consensus Guideline and Review by the American Society of Health-system Pharmacists, the Infectious Diseases Society of America, the Pediatric Infectious Diseases Society, and the Society of Infectious Diseases Pharmacists. Clin. Infect. Dis. 2020, 71, 1361–1364. [Google Scholar] [CrossRef]
- Barlam, T.F.; Cosgrove, S.E.; Abbo, L.M.; MacDougall, C.; Schuetz, A.N.; Septimus, E.J.; Srinivasan, A.; Dellit, T.H.; Falck-Ytter, Y.T.; Fishman, N.O.; et al. Implementing an Antibiotic Stewardship Program: Guidelines by the Infectious Diseases Society of America and the Society for Healthcare Epidemiology of America. Clin. Infect. Dis. 2016, 62, e51–e77. [Google Scholar] [CrossRef]



| Class | Precision | Recall (TP Rate) | F-Measure | FP Rate | ROC-AUC |
|---|---|---|---|---|---|
| MSSA | 0.768 | 0.981 | 0.862 | 0.432 | 0.916 |
| MRSA | 0.955 | 0.568 | 0.712 | 0.019 | 0.916 |
| Weighted average | 0.844 | 0.813 | 0.801 | 0.264 | 0.916 |
| Actual Class | Predicted MSSA | Predicted MRSA | Total |
|---|---|---|---|
| MSSA | 53 | 1 | 54 |
| MRSA | 16 | 21 | 37 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Gülmez, A.; Ceylan, A.N.; Kömeç, S.; Öncel, B.; Sağlam, Y. Integrating MALDI-TOF Mass Spectrometry and Machine Learning for Rapid and Clinically Relevant Differentiation of MRSA and MSSA. Pathogens 2026, 15, 191. https://doi.org/10.3390/pathogens15020191
Gülmez A, Ceylan AN, Kömeç S, Öncel B, Sağlam Y. Integrating MALDI-TOF Mass Spectrometry and Machine Learning for Rapid and Clinically Relevant Differentiation of MRSA and MSSA. Pathogens. 2026; 15(2):191. https://doi.org/10.3390/pathogens15020191
Chicago/Turabian StyleGülmez, Abdurrahman, Ayşe Nur Ceylan, Selda Kömeç, Beyza Öncel, and Yasin Sağlam. 2026. "Integrating MALDI-TOF Mass Spectrometry and Machine Learning for Rapid and Clinically Relevant Differentiation of MRSA and MSSA" Pathogens 15, no. 2: 191. https://doi.org/10.3390/pathogens15020191
APA StyleGülmez, A., Ceylan, A. N., Kömeç, S., Öncel, B., & Sağlam, Y. (2026). Integrating MALDI-TOF Mass Spectrometry and Machine Learning for Rapid and Clinically Relevant Differentiation of MRSA and MSSA. Pathogens, 15(2), 191. https://doi.org/10.3390/pathogens15020191





