Genotype-Specific HPV E6/E7 mRNA Triage Improves Risk Stratification and Reduces Referrals in DNA-Positive ASC-US/LSIL: A Real-World Cohort from Nordland, Norway
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design, Setting, and Population
2.2. Screening Tests and Triage Workflow
2.3. Definitions and Endpoints
2.4. Subgroup and Supportive Analyses
2.5. Referral Strategy Comparison (Simulation)
- (i)
- Refer all DNA-positive women to colposcopy, without additional stratification;
- (ii)
- mRNA triage, referring only to mRNA-positive women and managing mRNA-negative women through routine surveillance with repeat HPV DNA testing at 12 months, with colposcopy referral reserved for women with persistent HPV positivity.
2.6. Data Sources and Quality Control
2.7. Statistical Analysis
2.8. Ethics
3. Results
3.1. Study Population
3.2. Diagnostic Performance of mRNA Triage
- CIN2+: Among mRNA-positive women, 33.3% (26/78) had CIN2+, compared with 15.5% (15/97) among mRNA-negative women (relative risk [RR] = 2.16; 95% CI 1.23–3.78; χ2 = 7.70; p = 0.006). The corresponding test characteristics were: sensitivity 63.4%, specificity 61.2%, PPV 33.3%, and NPV 84.5% (Table 2A). Thus, 15 of 41 CIN2+ cases (36.6%) occurred in the mRNA-negative group and would not be detected at baseline triage, but would be expected to be identified at follow-up among women with persistent HPV positivity.
- CIN3+: For mRNA-positive women, the risk of CIN3+ was 14.1% (11/78) compared with 6.2% (6/97) among mRNA-negative women (RR = 2.27; 95% CI 0.88–5.87; χ2 = 3.07; p = 0.08). Sensitivity, specificity, PPV, and NPV were 64.7%, 57.6%, 14.1%, and 93.8%, respectively (Table 2B). Accordingly, 6 of 17 CIN3+ cases (35.3%) occurred in the mRNA-negative group and would not be detected at baseline triage, but would be expected to be identified at follow-up among women with persistent HPV positivity.
3.3. Genotype-Specific Risk Stratification
3.4. Co-Infections Detected by the 7-Type HPV mRNA Assay
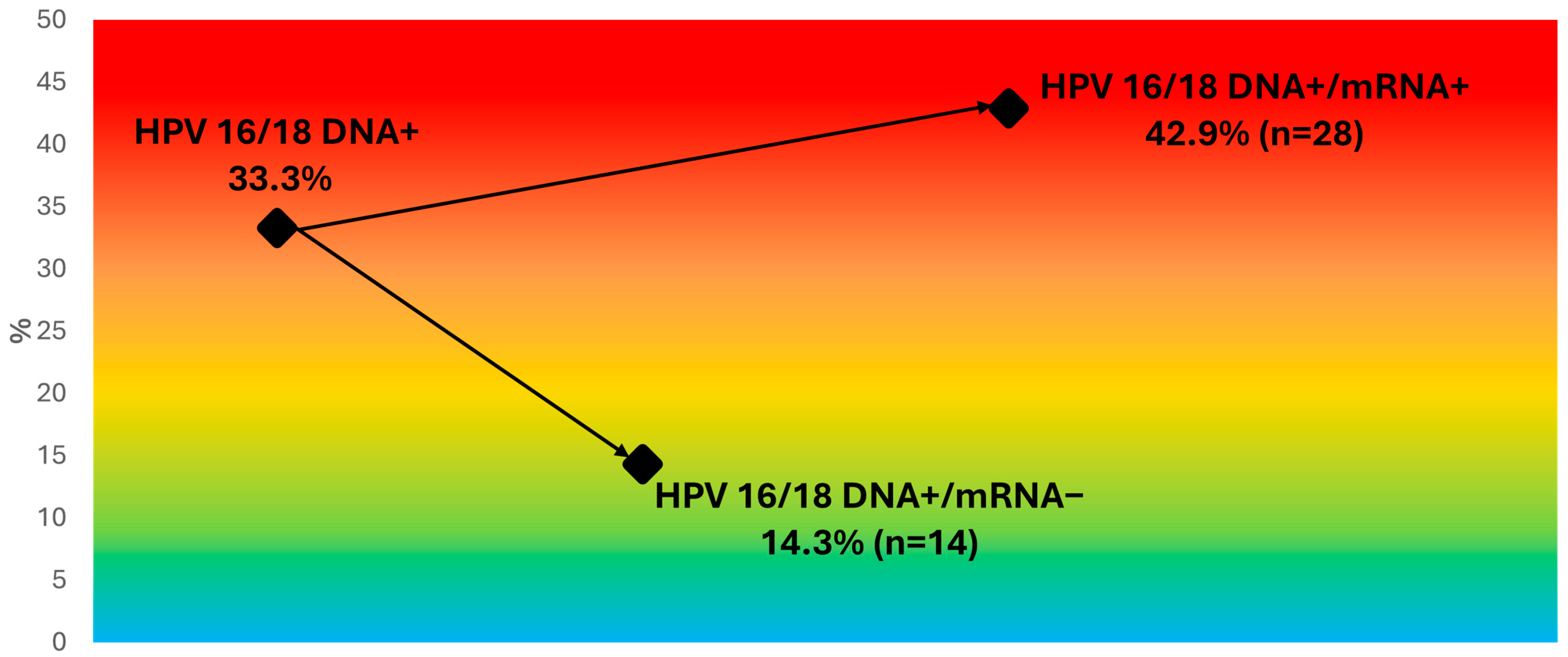
3.5. Age-Specific Comparison of HPV16/18 DNA and mRNA Detection, and Triage Within HPV16/18 DNA-Positive Women
3.6. Simulated Referral Strategies and Procedural Yield
3.7. Summary of Principal Findings
- (i)
- approximately 2-fold higher risk separation between test-positive and test-negative women for CIN2+ and CIN3+,
- (ii)
- the highest genotype-specific risks for HPV33, HPV18, HPV16, and HPV31, and
- (iii)
4. Discussion
4.1. Principal Findings
4.2. Context with Prior Evidence
4.3. Why Genotype-Specific mRNA Matters in a DNA-Positive, Low-Grade Group
4.4. Refining HPV16/18 Risk: DNA–mRNA Differentiation and Implications for Targeted Triage
4.5. Clinical and Programmatic Implications
- Substantial reduction in referrals: mRNA-based triage would reduce colposcopy referrals by ~55%, easing pressure on overstretched colposcopy services.
- Improved diagnostic efficiency: Fewer procedures are required per detected CIN2+/CIN3+ lesion, reflecting a more efficient allocation of diagnostic resources.
- Targeted evaluation of women at highest immediate risk: mRNA positivity identifies infections with transcriptionally active oncogene expression, allowing diagnostic work-ups to focus on women most likely to harbor CIN2+ or rapidly progressing lesions.
- Safe surveillance of low-risk individuals: mRNA-negative women—over half of the cohort—formed a low-risk group with a low CIN2+/CIN3+ yield, supporting surveillance rather than immediate referral. This is particularly important for younger women (<35), in whom CIN2 frequently regresses and overtreatment poses reproductive risks.
4.6. Strengths and Limitations
4.7. Future Directions
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ASC-US | Atypical squamous cells of undetermined significance |
| CI | Confidence interval |
| CIN | Cervical intraepithelial neoplasia |
| CIN2+ | Cervical intraepithelial neoplasia grade 2 or worse |
| CIN3+ | Cervical intraepithelial neoplasia grade 3 or worse |
| DNA | Deoxyribonucleic acid |
| HPV | Human papillomavirus |
| HSIL | High-grade squamous intraepithelial lesion |
| LBC | Liquid-based cytology |
| LSIL | Low-grade squamous intraepithelial lesion |
| mRNA | Messenger ribonucleic acid |
| NPV | Negative predictive value |
| PPV | Positive predictive value |
| REK | Regional Committee for Medical and Health Research Ethics (Norway) |
| RR | Relative risk |
References
- IARC Working Group on the Evaluation of Cancer-Preventive Interventions. Screen-and-Treat Approach and Women at Differential Risk. In Cervical Cancer Screening; IARC Handbooks of Cancer Prevention, No. 18; International Agency for Research on Cancer: Lyon, France, 2022. Available online: https://www.ncbi.nlm.nih.gov/books/NBK601994/ (accessed on 26 October 2025).
- Ronco, G.; Dillner, J.; Elfström, K.M.; Tunesi, S.; Snijders, P.J.; Arbyn, M.; Kitchener, H.; Segnan, N.; Gilham, C.; Giorgi-Rossi, P.; et al. Efficacy of HPV-based screening for prevention of invasive cervical cancer: Follow-up of four European randomised controlled trials. Lancet 2014, 383, 524–532. [Google Scholar] [CrossRef] [PubMed]
- Nygård, M.; Engesæter, B.; Castle, P.E.; Berland, J.M.; Eide, M.L.; Iversen, O.E.; Jonassen, C.M.; Christiansen, I.K.; Vintermyr, O.K.; Tropé, A. Randomized implementation of a primary human papillomavirus testing-based cervical cancer screening protocol for women 34 to 69 years in Norway. Cancer Epidemiol. Biomarkers Prev. 2022, 31, 1812–1822. [Google Scholar] [CrossRef] [PubMed]
- Hashim, D.; Engesæter, B.; Baadstrand Skare, G.; Castle, P.E.; Bjørge, T.; Tropé, A.; Nygård, M. Real-world data on cervical cancer risk stratification by cytology and HPV genotype to inform the management of HPV-positive women in routine cervical screening. Br. J. Cancer 2020, 122, 1715–1723. [Google Scholar] [CrossRef] [PubMed]
- Wentzensen, N.; Clarke, M.A.; Bremer, R.; Poitras, N.; Tokugawa, D.; Goldhoff, P.E.; Castle, P.E.; Schiffman, M.; Kingery, J.D.; Grewal, K.K.; et al. Clinical Evaluation of Human Papillomavirus Screening with p16/Ki-67 Dual Stain Triage in a Large Organized Cervical Cancer Screening Program. JAMA Intern. Med. 2019, 179, 881–888. [Google Scholar] [CrossRef]
- St-Martin, G.; Thamsborg, L.H.; Andersen, B.; Christensen, J.; Ejersbo, D.; Jochumsen, K.; Johansen, T.; Larsen, L.G.; Waldstrøm, M.; Lynge, E. Management of low-grade cervical cytology in young women: A cohort study from Denmark. Acta Oncol. 2021, 60, 444–451. [Google Scholar] [CrossRef]
- Okunade, K.S.; Adejimi, A.A.; John-Olabode, S.O.; Oshodi, Y.A.; Oluwole, A.A. An overview of HPV screening tests to improve access to cervical cancer screening amongst underserved populations: From development to implementation. Risk Manag. Healthc. Policy 2022, 15, 1823–1830. [Google Scholar] [CrossRef]
- Gravitt, P.E.; Winer, R.L. Natural history of HPV infection across the lifespan: Role of viral latency. Viruses 2017, 9, 267. [Google Scholar] [CrossRef]
- Arbyn, M.; Martin-Hirsch, P.; Buntinx, F.; Van Ranst, M.; Paraskevaidis, E.; Dillner, J. Triage of women with equivocal or low-grade cervical cytology results: A meta-analysis of the HPV test positivity rate. J. Cell. Mol. Med. 2009, 13, 648–659. [Google Scholar] [CrossRef]
- Folkehelseinstituttet (Norwegian Institute of Public Health). Flowchart for the Assessment of Liquid-Based Cervical Samples for Laboratories Performing Extended Genotyping; Version 5; Norwegian Institute of Public Health: Oslo, Norway, 2026. Available online: https://www.fhi.no/globalassets/livmorhalsprogrammet/algoritmerflytskjema/2026/utvidet-genotyping_oppdatert_05.01.2026.jpg (accessed on 23 January 2026).
- Bonde, J.; Bottari, F.; Iacobone, A.D.; Cocuzza, C.E.; Sandri, M.-T.; Bogliatto, F.; Khan, K.S.; Ejegod, D.M.; Gary, D.S.; Andrews, J.C. Human Papillomavirus Same Genotype Persistence and Risk: A Systematic Review. J. Low. Genit. Tract Dis. 2021, 25, 27–37. [Google Scholar] [CrossRef]
- Tjalma, W.A.; Fiander, A.; Reich, O.; Powell, N.; Nowakowski, A.M.; Kirschner, B.; Koiss, R.; O’Leary, J.; Joura, E.A.; Rosenlund, M.; et al. Differences in human papillomavirus type distribution in high-grade cervical intraepithelial neoplasia and invasive cervical cancer in Europe. Int. J. Cancer 2013, 132, 854–867. [Google Scholar] [CrossRef]
- Pal, A.; Kundu, R. Human papillomavirus E6 and E7: The cervical cancer hallmarks and targets for therapy. Front. Microbiol. 2020, 10, 3116. [Google Scholar] [CrossRef]
- Ratnam, S.; Coutlee, F.; Fontaine, D.; Bentley, J.; Escott, N.; Ghatage, P.; Gadag, V.; Holloway, G.; Bartellas, E.; Kum, N.; et al. Aptima HPV E6/E7 mRNA Test Is as Sensitive as Hybrid Capture 2 Assay but More Specific at Detecting Cervical Precancer and Cancer. J. Clin. Microbiol. 2011, 49, 557–564. [Google Scholar] [CrossRef]
- Westre, B.; Giske, A.; Guttormsen, H.; Sørbye, S.W.; Skjeldestad, F.E. Five-type HPV mRNA versus 14-type HPV DNA test: Test performance, over-diagnosis, and overtreatment in triage of women with minor cervical lesions. BMC Clin. Pathol. 2016, 16, 9. [Google Scholar] [CrossRef] [PubMed]
- Sørbye, S.; Falang, B.M.; Antonsen, M.; Mortensen, E. Genotype-specific HPV mRNA triage improves CIN2+ detection efficiency compared to cytology: A population-based study of HPV DNA-positive women. Pathogens 2025, 14, 749. [Google Scholar] [CrossRef] [PubMed]
- Lagheden, C.; Eklund, C.; Lamin, H.; Kleppe, S.N.; Lei, J.; Elfström, K.M.; Sundström, K.; Andrae, B.; Sparén, P.; Dillner, J. Nationwide Comprehensive Human Papillomavirus (HPV) Genotyping of Invasive Cervical Cancer. Br. J. Cancer 2018, 118, 1377–1381. [Google Scholar] [CrossRef] [PubMed]
- Vänskä, S.; Luostarinen, T.; Lagheden, C.; Eklund, C.; Nordqvist Kleppe, S.; Andrae, B.; Sparén, P.; Sundström, K.; Lehtinen, M.; Dillner, J. Differing Age-Specific Cervical Cancer Incidence between Different Types of Human Papillomavirus: Implications for Predicting the Impact of Elimination Programs. Am. J. Epidemiol. 2021, 190, 506–514. [Google Scholar] [CrossRef]
- Hammer, A.; Rositch, A.; Qeadan, F.; Gravitt, P.E.; Blaakaer, J. Age-Specific Prevalence of HPV16/18 Genotypes in Cervical Cancer: A Systematic Review and Meta-Analysis. Int. J. Cancer 2016, 138, 2795–2803. [Google Scholar] [CrossRef]
- Nayar, R.; Wilbur, D.C. The Bethesda System for Reporting Cervical Cytology: A historical perspective. Acta Cytol. 2017, 61, 359–372. [Google Scholar] [CrossRef]
- Origoni, M.; Cristoforoni, P.; Carminati, G.; Stefani, C.; Costa, S.; Sandri, M.T.; Mariani, L.; Preti, M. E6/E7 mRNA Testing for Human Papilloma Virus-Induced High-Grade Cervical Intraepithelial Disease (CIN2/CIN3): A Promising Perspective. Ecancermedicalscience 2015, 9, 533. [Google Scholar] [CrossRef]
- Derbie, A.; Mekonnen, D.; Biadglegne, F.; Abebe, T.; De Vos, D. HPV E6/E7 mRNA Test for the Detection of High-Grade Cervical Intraepithelial Neoplasia (CIN2+): A Systematic Review. Infect. Agents Cancer 2020, 15, 9. [Google Scholar] [CrossRef]
- Benevolo, M.; Vocaturo, A.; Caraceni, D.; French, D.; Rosini, S.; Zappacosta, R.; Terrenato, I.; Ciccocioppo, L.; Frega, A.; Giorgi Rossi, P. Sensitivity, Specificity, and Clinical Value of Human Papillomavirus (HPV) E6/E7 mRNA Assay as a Triage Test for Cervical Cytology and HPV DNA Test. J. Clin. Microbiol. 2011, 49, 2643–2650. [Google Scholar] [CrossRef]
- Alaghehbandan, R.; Fontaine, D.; Bentley, J.; Escott, N.; Ghatage, P.; Lear, A.; Coutlee, F.; Ratnam, S. Performance of ProEx C and PreTect HPV-Proofer E6/E7 mRNA Tests in Comparison with the Hybrid Capture 2 HPV DNA Test for Triaging ASCUS and LSIL Cytology. Diagn. Cytopathol. 2013, 41, 767–775. [Google Scholar] [CrossRef] [PubMed]
- Luttmer, R.; Berkhof, J.; Dijkstra, M.G.; van Kemenade, F.J.; Snijders, P.J.; Heideman, D.A.; Meijer, C.J. Comparing triage algorithms using HPV DNA genotyping, HPV E7 mRNA detection and cytology in HPV DNA-positive women. J. Clin. Virol. 2015, 67, 59–66. [Google Scholar] [CrossRef] [PubMed]
- Verdoodt, F.; Szarewski, A.; Halfon, P.; Cuschieri, K.; Arbyn, M. Triage of Women with Minor Abnormal Cervical Cytology: Meta-Analysis of the Accuracy of an Assay Targeting Messenger Ribonucleic Acid of 5 High-Risk Human Papillomavirus Types. Cancer Cytopathol. 2013, 121, 675–687. [Google Scholar] [CrossRef] [PubMed]
- Sørbye, S.W.; Falang, B.M.; Antonsen, M. Distribution of HPV Types in Tumor Tissue from Non-Vaccinated Women with Cervical Cancer in Norway. J. Mol. Pathol. 2023, 4, 166–177. [Google Scholar] [CrossRef]
- Reinholdt, K.; Juul, K.E.; Dehlendorff, C.; Munk, C.; Kjær, S.K.; Thomsen, L.T. Triage of Low-Grade Squamous Intraepithelial Lesions Using Human Papillomavirus Messenger Ribonucleic Acid Tests—A Prospective Population-Based Register Study. Acta Obstet. Gynecol. Scand. 2020, 99, 204–212. [Google Scholar] [CrossRef]
- Bruno, M.T.; Ferrara, M.; Fava, V.; Barrasso, G.; Panella, M.M. A Prospective Study of Women with ASC-US or LSIL Pap Smears at Baseline and HPV E6/E7 mRNA Positive: A 3-Year Follow-Up. Epidemiol. Infect. 2018, 146, 612–618. [Google Scholar] [CrossRef]
- Zappacosta, R.; Gatta, D.M.P.; Marinucci, P.; Capanna, S.; Lattanzio, G.; Caraceni, D.; Rosini, S. Role of E6/E7 mRNA Test in the Diagnostic Algorithm of HPV-Positive Patients Showing ASC-US and LSIL: Clinical and Economic Implications in a Publicly Financed Healthcare System. Expert Rev. Mol. Diagn. 2015, 15, 137–150. [Google Scholar] [CrossRef]
- Kafasi, A.; Pitiriga, V.C.; Spanakis, N.; Vlachos, N.; Thomakos, N.; Stournaras, S.; Tsakris, A.; Kaparos, G. Evaluation of the PreTect HPV-Proofer E6/E7 mRNA Assay for the Detection of Precancerous Cervical Lesions in the Greek Female Population. Pathogens 2025, 14, 853. [Google Scholar] [CrossRef]
- Dabeski, D.; Duvlis, S.; Basheska, N.; Antovska, V.; Stojovski, M.; Trajanova, M.; Dimitrov, G.; Dabeski, A.; Gureva-Gjorgievska, N. Comparison between HPV DNA Testing and HPV E6/E7 mRNA Testing in Women with Squamous Cell Abnormalities of the Uterine Cervix. Pril 2019, 40, 51–58. [Google Scholar] [CrossRef]
- Bruno, M.T.; Cassaro, N.; Vitale, S.G.; Guaita, A.; Boemi, S. Possible Role of Negative Human Papillomavirus E6/E7 mRNA as a Predictor of Regression of Cervical Intraepithelial Neoplasia 2 Lesions in HPV-Positive Women. Virol. J. 2022, 19, 95. [Google Scholar] [CrossRef]
- Giorgi Rossi, P.; Benevolo, M.; Vocaturo, A.; Caraceni, D.; Ciccocioppo, L.; Frega, A.; Terrenato, I.; Zappacosta, R.; French, D.; Rosini, S.; et al. Prognostic Value of HPV E6/E7 mRNA Assay in Women with Negative Colposcopy or CIN1 Histology Result: A Follow-Up Study. PLoS ONE 2013, 8, e57600. [Google Scholar] [CrossRef]
- Frega, A.; Pavone, M.; Sesti, F.; Leone, C.; Bianchi, P.; Cozza, G.; Colombrino, C.; Lukic, A.; Marziani, R.; De Sanctis, L.; et al. Sensitivity and Specificity Values of HPV DNA, p16/Ki-67 and HPV mRNA in Young Women with Atypical Squamous Cells of Undetermined Significance (ASC-US) or Low-Grade Squamous Intraepithelial Lesion (LSIL). Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 10672–10677. [Google Scholar] [CrossRef]
- Sørbye, S.W.; Falang, B.M.; Botha, M.H.; Snyman, L.C.; van der Merwe, H.; Visser, C.; Richter, K.; Dreyer, G. Enhancing Cervical Cancer Prevention in South African Women: Primary HPV mRNA Screening with Different Genotype Combinations. Cancers 2023, 15, 5453. [Google Scholar] [CrossRef]
- Flores, C.E.A.; Falang, B.M.; Gómez-Laguna, L.; Gutiérrez, G.G.; León, J.M.O.; Uribe, M.; Cruz, O.; Sørbye, S.W. Enhancing Cervical Cancer Screening with 7-Type HPV mRNA E6/E7 Testing on Self-Collected Samples: Multicentric Insights from Mexico. Cancers 2024, 16, 2485. [Google Scholar] [CrossRef]
- Tainio, K.; Athanasiou, A.; Tikkinen, K.A.O.; Aaltonen, R.; Cardenas Hernandes, J.; Glazer-Livson, S.; Jakobsson, M.; Joronen, K.; Kiviharju, M.; Louvanto, K.; et al. Clinical Course of Untreated Cervical Intraepithelial Neoplasia Grade 2 under Active Surveillance: Systematic Review and Meta-Analysis. BMJ 2018, 360, k499. [Google Scholar] [CrossRef]

| Characteristic | n/N (%) |
|---|---|
| Cytology | |
| ASC-US | 115/175 (65.7) |
| LSIL | 60/175 (34.3) |
| Histology outcome | |
| CIN2+ prevalence (overall) | 41/175 (23.4) |
| ASC-US subgroup | 26/115 (22.6) |
| LSIL subgroup | 15/60 (25.0) |
| CIN3+ prevalence (overall) | 17/175 (9.7) |
| ASC-US subgroup | 10/115 (8.7) |
| LSIL subgroup | 7/60 (11.7) |
| mRNA (7-type) positive | 78/175 (44.6) |
| ASC-US subgroup | 52/115 (45.2) |
| LSIL subgroup | 26/60 (43.3) |
| (A) | ||||
| mRNA Result | CIN2+ (n) | ≤CIN1 (n) | Total (n) | Predictive Value |
| Positive | 26 | 52 | 78 | PPV 33.3% |
| Negative | 15 | 82 | 97 | NPV 84.5% |
| Total | 41 | 134 | 175 | - |
| (B) | ||||
| mRNA Result | CIN3+ (n) | ≤CIN2 (n) | Total (n) | Predictive Value |
| Positive | 11 | 67 | 78 | PPV 14.1% |
| Negative | 6 | 91 | 97 | NPV 93.8% |
| Total | 17 | 160 | 175 | - |
| mRNA Genotype | N (Total Positive) | CIN2+ (n, PPV%) | CIN3+ (n, PPV%) |
|---|---|---|---|
| 16 | 22 | 9 (40.9%) | 4 (18.2%) |
| 18 | 6 | 3 (50.0%) | 1 (16.7%) |
| 31 | 20 | 7 (35.0%) | 2 (10.0%) |
| 33 | 6 | 4 (66.7%) | 3 (50.0%) |
| 45 | 5 | 1 (20.0%) | 1 (20.0%) |
| 52 | 11 | 1 (9.1%) | 0 (0.0%) |
| 58 | 8 | 1 (12.5%) | 0 (0.0%) |
| mRNA-negative | 97 | 15 (15.5%) | 6 (6.2%) |
| mRNA Genotypes Detected (Co-Infection Pattern) | N (%) |
|---|---|
| HPV16 + HPV58 | 2/78 (2.6) |
| HPV33 + HPV52 | 1/78 (1.3) |
| HPV16 + HPV52 | 1/78 (1.3) |
| HPV31 + HPV33 | 1/78 (1.3) |
| HPV18 + HPV31 | 1/78 (1.3) |
| HPV18 + HPV52 | 1/78 (1.3) |
| HPV18 + HPV45 + HPV58 | 1/78 (1.3) |
| HPV16 + HPV18 + HPV52 + HPV58 | 1/78 (1.3) |
| Age Group (Years) | HPV16/18 DNA n pos/Total (% Positive) | HPV16/18 mRNA n pos/Total (% Positive) | Relative Reduction in Positivity (mRNA vs. DNA) |
|---|---|---|---|
| <25 | 0/9 (0.0%) | 0/9 (0.0%) | — |
| 25–33 | 14/68 (20.6%) | 10/68 (14.7%) | −28.6% |
| 34–69 | 28/97 (28.9%) | 18/97 (18.6%) | −35.7% |
| >69 | 0/1 (0.0%) | 0/1 (0.0%) | — |
| Total | 42/175 (24.0%) | 28/175 (16.0%) | −33.3% |
| mRNA16/18 Result | CIN2+ (n) | ≤CIN1 (n) | Total (n) | Predictive Value |
|---|---|---|---|---|
| Positive | 12 | 16 | 28 | PPV 42.9% |
| Negative | 2 | 12 | 14 | NPV 85.7% |
| Total | 14 | 28 | 42 | - |
| Referral Strategy | Referrals (n, %) | CIN2+ Detected | Colposcopies per CIN2+ | CIN3+ Detected | Colposcopies per CIN3+ |
|---|---|---|---|---|---|
| Refer all HPV DNA-positive with ASC-US/LSIL to colposcopy | 175 (100%) | 41 | 4.27 | 17 | 10.3 |
| mRNA triage (refer only to mRNA-positive among HPV DNA-positive with ASC-US/LSIL) | 78 (44.6%) | 26 | 3.00 | 11 | 7.1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Al-Shibli, K.; Nguyen, D.T.; Mohammed, H.A.L.; Sørbye, S.W. Genotype-Specific HPV E6/E7 mRNA Triage Improves Risk Stratification and Reduces Referrals in DNA-Positive ASC-US/LSIL: A Real-World Cohort from Nordland, Norway. Pathogens 2026, 15, 178. https://doi.org/10.3390/pathogens15020178
Al-Shibli K, Nguyen DT, Mohammed HAL, Sørbye SW. Genotype-Specific HPV E6/E7 mRNA Triage Improves Risk Stratification and Reduces Referrals in DNA-Positive ASC-US/LSIL: A Real-World Cohort from Nordland, Norway. Pathogens. 2026; 15(2):178. https://doi.org/10.3390/pathogens15020178
Chicago/Turabian StyleAl-Shibli, Khalid, Dat Tan Nguyen, Hiba Abdul Latif Mohammed, and Sveinung Wergeland Sørbye. 2026. "Genotype-Specific HPV E6/E7 mRNA Triage Improves Risk Stratification and Reduces Referrals in DNA-Positive ASC-US/LSIL: A Real-World Cohort from Nordland, Norway" Pathogens 15, no. 2: 178. https://doi.org/10.3390/pathogens15020178
APA StyleAl-Shibli, K., Nguyen, D. T., Mohammed, H. A. L., & Sørbye, S. W. (2026). Genotype-Specific HPV E6/E7 mRNA Triage Improves Risk Stratification and Reduces Referrals in DNA-Positive ASC-US/LSIL: A Real-World Cohort from Nordland, Norway. Pathogens, 15(2), 178. https://doi.org/10.3390/pathogens15020178







