Discovery of Novel c-di-GMP-Related Genes in Leptospira interrogans
Abstract
1. Introduction
2. Materials and Methods
2.1. Potential Proteins Involved in c-di-GMP Signaling in L. interrogans: Bioinformatic Analysis and Structural Prediction Models
2.2. Functional and Structural Characterization of Identified Proteins: An Analytical Approach
2.3. Multiple Amino Acid Sequence Alignment Using Three-Dimensional Structure Predictions
2.4. Pangenoma Analysis of Leptospira interrogans Isolates
2.5. Presence of the Genes Found in L. interrogans Serovar Copenhageni strain Fiocruz L1-130 in Other Leptospira Genomes
2.6. Graphical, Structural, and Imaging Software Tools
3. Results
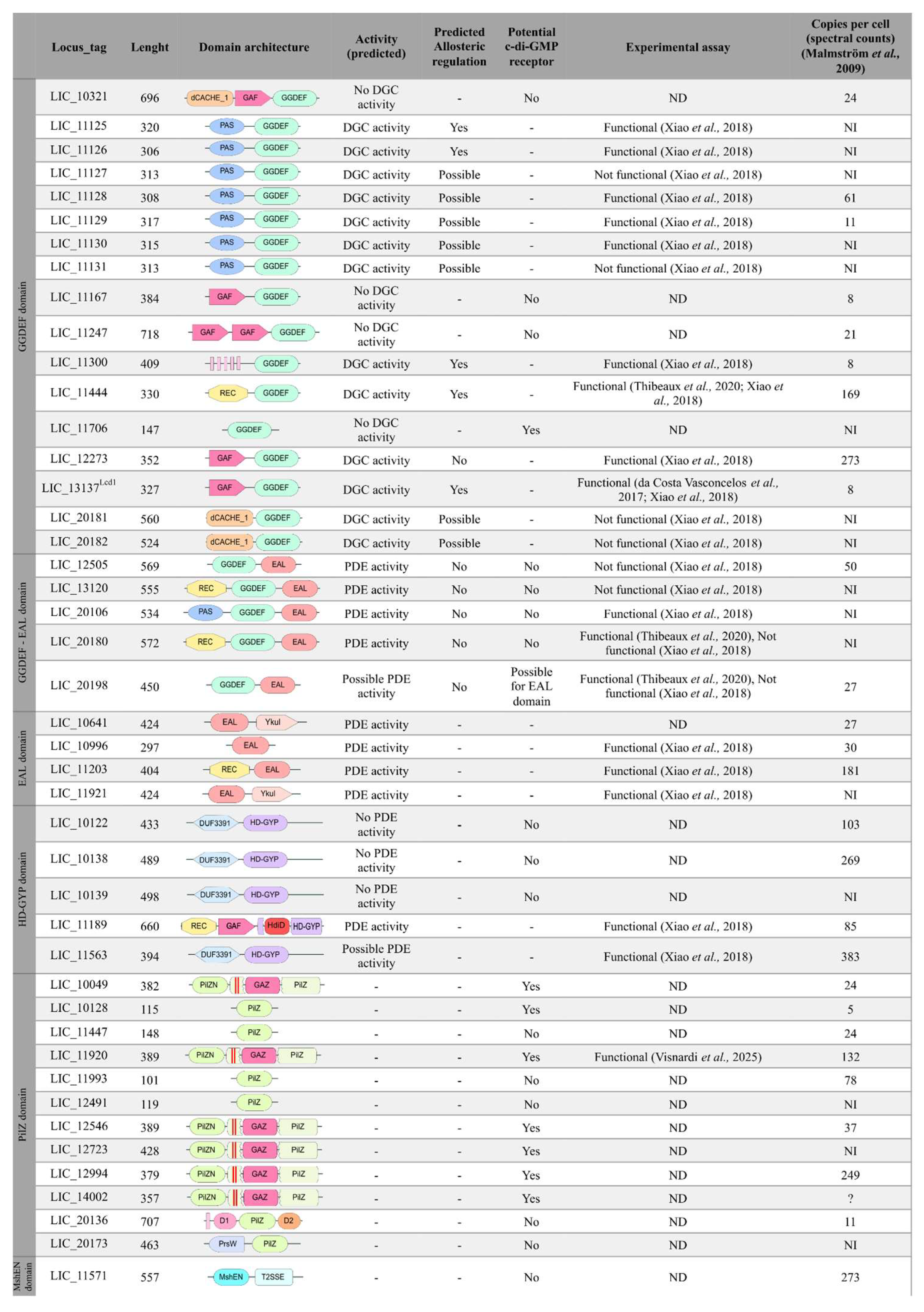
3.1. Identification of c-di-GMP-Related Genes in L. interrogans
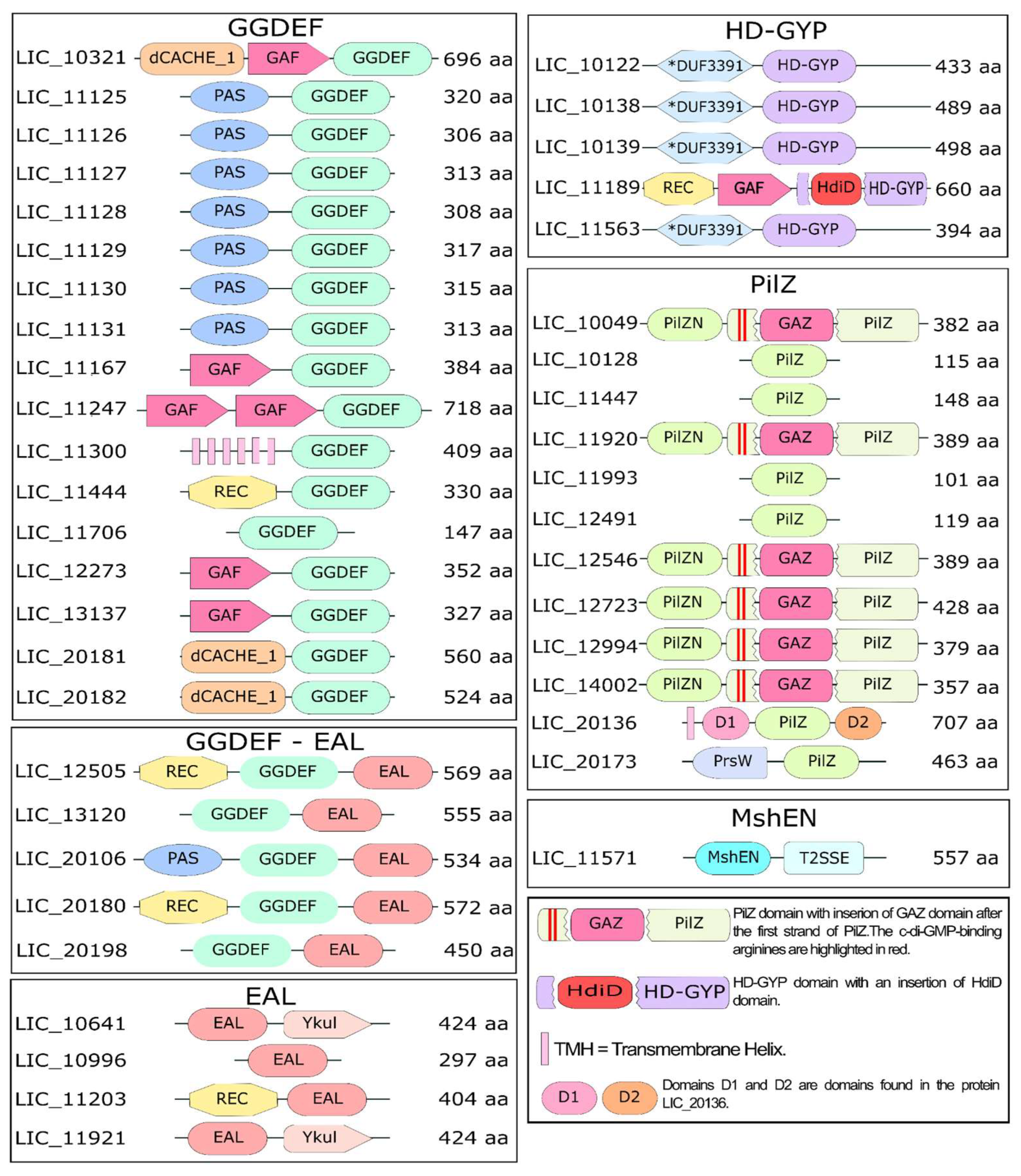
3.2. Diversity of Sensor Domains Present in c-di-GMP-Related Proteins
3.3. Diguanylate Cyclases
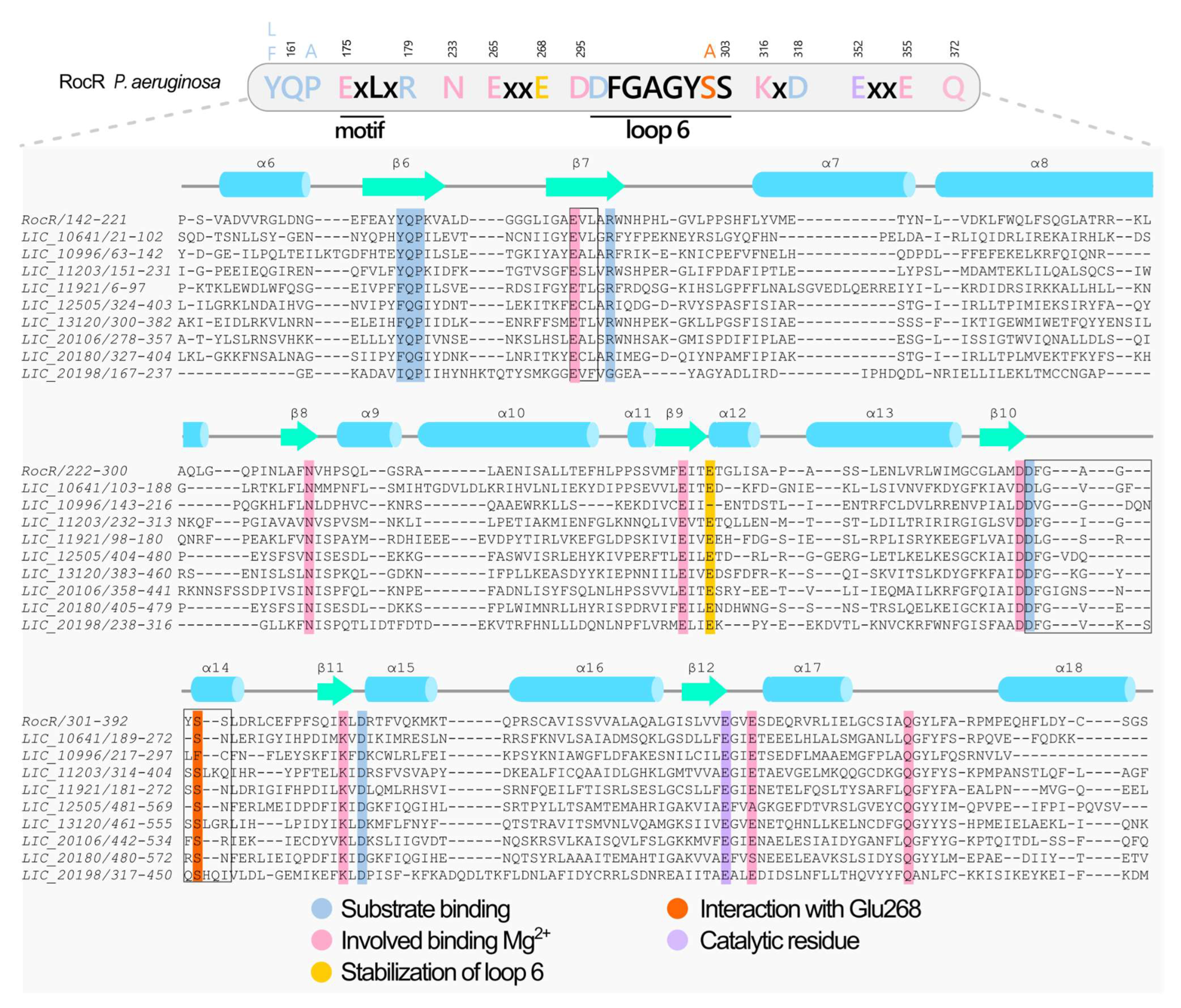
3.4. Proteins Containing EAL Domains
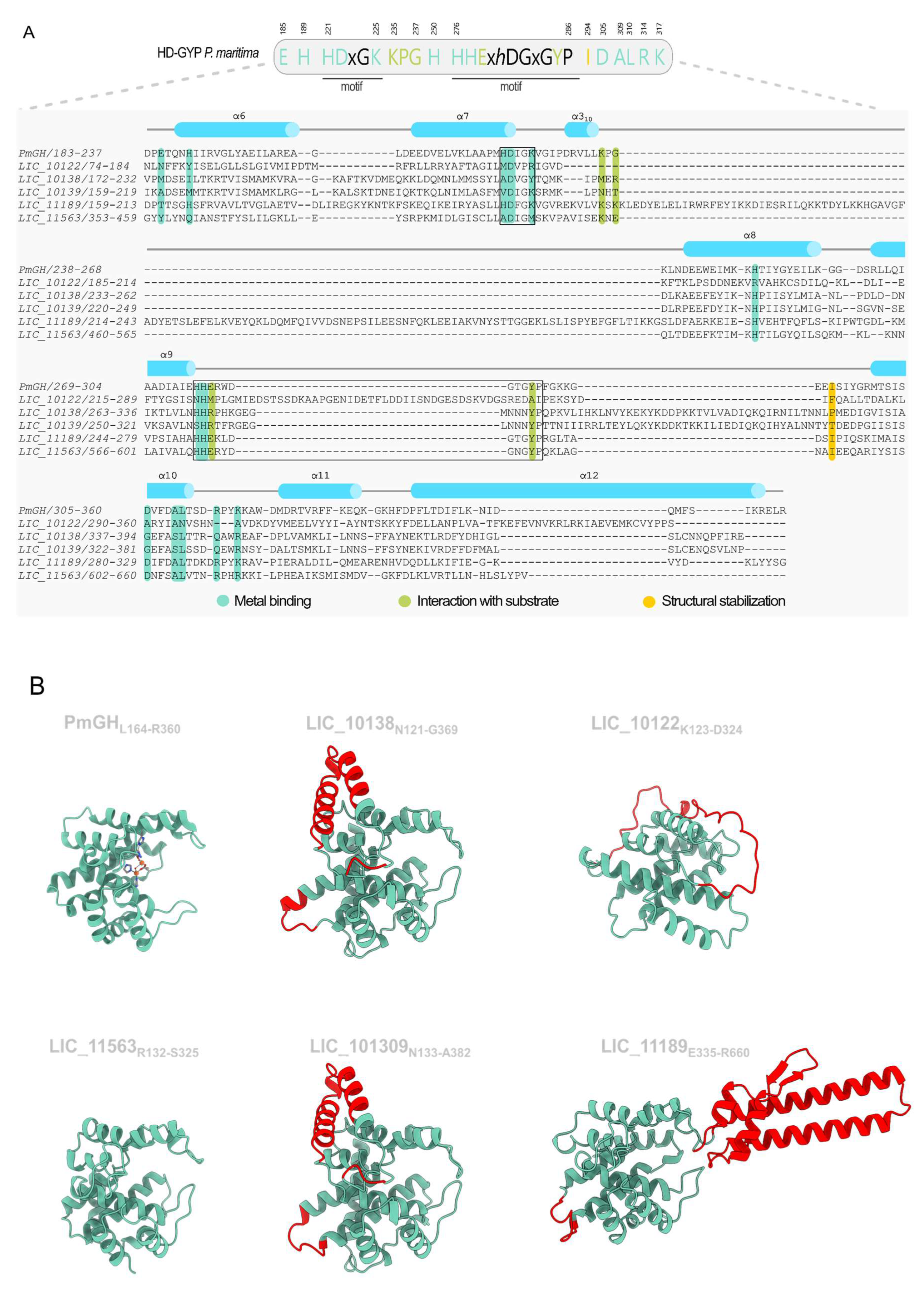
3.5. Proteins Containing HD-GYP Domains
3.6. The Insertion in the HD-GYP of LIC_11189 Is Widely Distributed in Response-Regulator HD-GYP Proteins
3.7. Potential Distant Members of DUF3391 Family
3.8. A Novel C-Terminal Domain in HD-GYP-Containing Proteins
3.9. Proteins Containing PilZ Domain
3.10. The LIC_20136 and LIC_20173 Represent Novel PilZ-like Families with Unique Domains Architecture
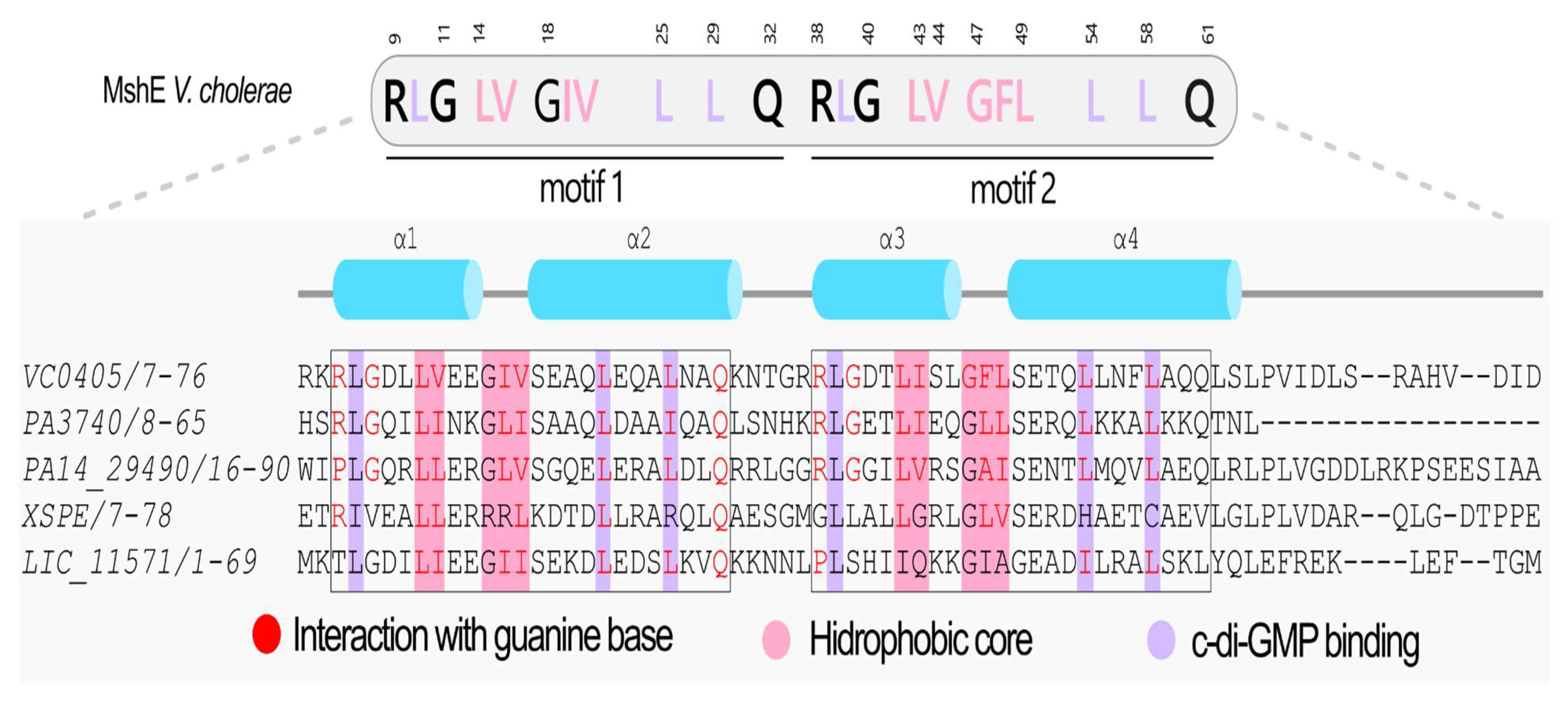
3.11. Proteins Containing the MshEN Domain in L. interrogans Serovar Copenhageni strain Fiocruz L1-130
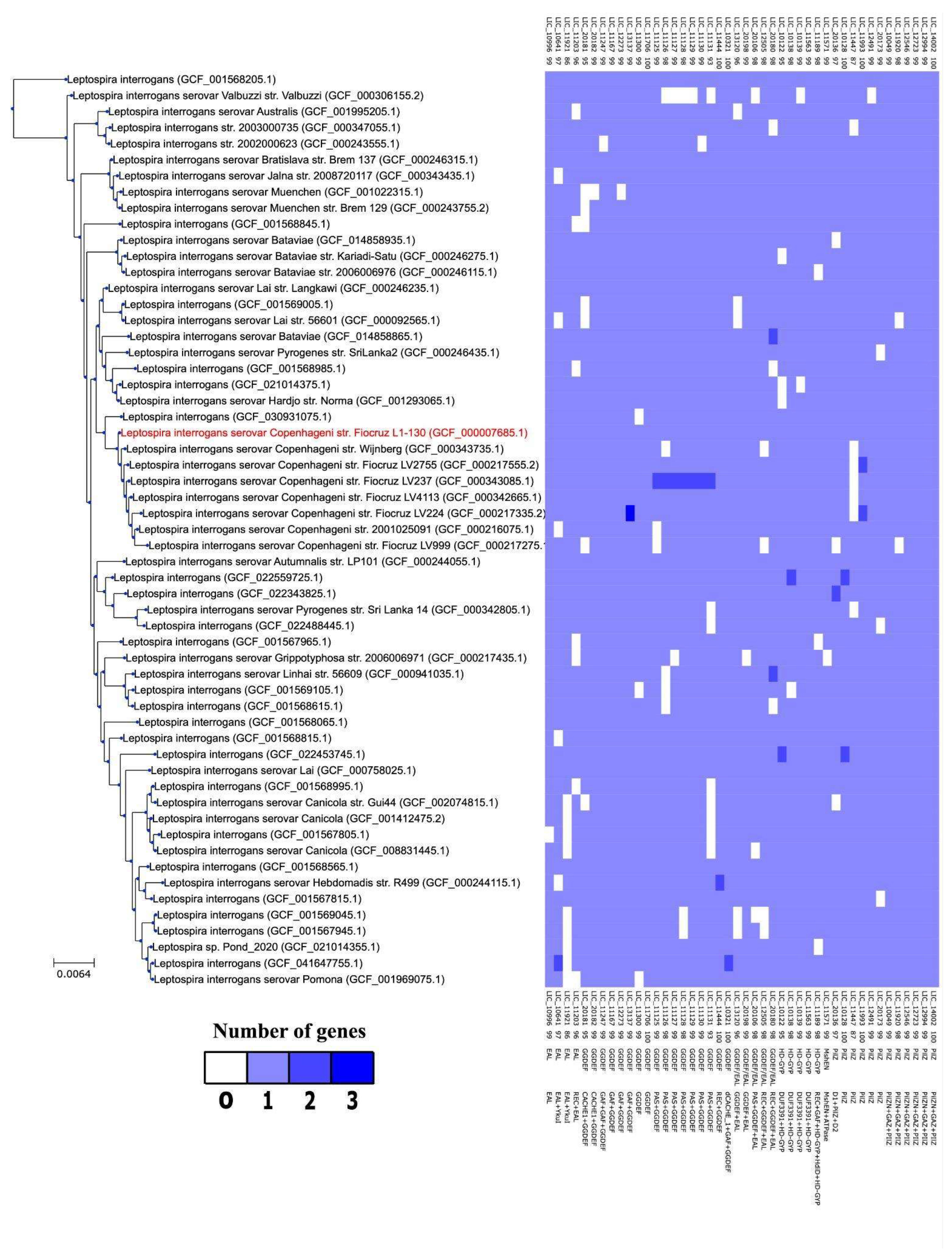
3.12. Distribution of c-di-GMP Genes Among L. interrogans Isolates
4. Discussion

5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Haake, D.A.; Levett, P.N. Leptospirosis in Humans. Curr. Top. Microbiol. Immunol. 2015, 387, 65–97. [Google Scholar] [CrossRef]
- Bonhomme, D.; Werts, C. Host and Species-Specificities of Pattern Recognition Receptors Upon Infection With. Front. Cell. Infect. Microbiol. 2022, 12, 932137. [Google Scholar] [CrossRef]
- Philip, N.; Bahtiar Affendy, N.; Ramli, S.N.A.; Arif, M.; Raja, P.; Nagandran, E.; Renganathan, P.; Taib, N.M.; Masri, S.N.; Yuhana, M.Y.; et al. Leptospira interrogans and Leptospira kirschneri Are the Dominant Leptospira Species Causing Human Leptospirosis in Central Malaysia. PLoS Negl. Trop. Dis. 2020, 14, e0008197. [Google Scholar] [CrossRef]
- Pětrošová, H.; Mikhael, A.; Culos, S.; Giraud-Gatineau, A.; Gomez, A.M.; Sherman, M.E.; Ernst, R.K.; Cameron, C.E.; Picardeau, M.; Goodlett, D.R. Lipid A Structural Diversity among Members of the Genus. Front. Microbiol. 2023, 14, 1181034. [Google Scholar] [CrossRef]
- Vincent, A.T.; Schiettekatte, O.; Goarant, C.; Neela, V.K.; Bernet, E.; Thibeaux, R.; Ismail, N.; Mohd Khalid, M.K.N.; Amran, F.; Masuzawa, T.; et al. Revisiting the Taxonomy and Evolution of Pathogenicity of the Genus Leptospira through the Prism of Genomics. PLoS Negl. Trop. Dis. 2019, 13, e0007270. [Google Scholar] [CrossRef]
- Ministry of Health—Brazil. Leptospirosis. Available online: https://www.gov.br/saude/pt-br/assuntos/saude-de-a-a-z/l/leptospirose/leptospirose (accessed on 10 October 2025).
- Rajapakse, S. Leptospirosis: Clinical Aspects. Clin. Med. 2022, 22, 14–17. [Google Scholar] [CrossRef] [PubMed]
- Bharti, A.R.; Nally, J.E.; Ricaldi, J.N.; Matthias, M.A.; Diaz, M.M.; Lovett, M.A.; Levett, P.N.; Gilman, R.H.; Willig, M.R.; Gotuzzo, E.; et al. Leptospirosis: A Zoonotic Disease of Global Importance. Lancet Infect. Dis. 2003, 3, 757–771. [Google Scholar] [CrossRef]
- Dupouey, J.; Faucher, B.; Edouard, S.; Richet, H.; Kodjo, A.; Drancourt, M.; Davoust, B. Human Leptospirosis: An Emerging Risk in Europe? Comp. Immunol. Microbiol. Infect. Dis. 2014, 37, 77–83. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.M.; Stull, J.W.; Moore, G.E. Potential Drivers for the Re-Emergence of Canine Leptospirosis in the United States and Canada. Trop. Med. Infect. Dis. 2022, 7, 377. [Google Scholar] [CrossRef]
- Davignon, G.; Pietrosemoli, N.; Benaroudj, N.; Soupé-Gilbert, M.-E.; Cagliero, J.; Turc, É.; Picardeau, M.; Guentas, L.; Goarant, C.; Thibeaux, R. Leptospira interrogans Biofilm Transcriptome Highlights Adaption to Starvation and General Stress While Maintaining Virulence. NPJ Biofilms Microbiomes 2024, 10, 95. [Google Scholar] [CrossRef] [PubMed]
- PAHO; World Health Organization (WHO); United Nations. PAHO Supports Emergency Response Following Flooding in Rio Grande Do Sul, Brazil. Available online: https://www.paho.org/en/news/3-7-2024-paho-supports-emergency-response-following-flooding-rio-grande-do-sul-brazil (accessed on 28 April 2025).
- OCHA; United Nations. Brazil: Floods in Rio Grande Do Sul—United Nations Situation Report, as of 20 September 2024. Available online: https://www.unocha.org/publications/report/brazil/brazil-floods-rio-grande-do-sul-united-nations-situation-report-20-september-2024 (accessed on 10 October 2025).
- Ziliotto, M.; Chies, J.A.B.; Ellwanger, J.H. Extreme Weather Events and Pathogen Pollution Fuel Infectious Diseases: The 2024 Flood-Related Leptospirosis Outbreak in Southern Brazil and Other Red Lights. Pollutants 2024, 4, 424–433. [Google Scholar] [CrossRef]
- Martins-Filho, P.R.; Croda, J.; de Souza Araújo, A.A.; Correia, D.; Quintans-Júnior, L.J. Catastrophic Floods in Rio Grande Do Sul, Brazil: The Need for Public Health Responses to Potential Infectious Disease Outbreaks. Rev. Soc. Bras. Med. Trop. 2024, 57, e006032024. [Google Scholar] [CrossRef]
- Thibeaux, R.; Soupé-Gilbert, M.-E.; Kainiu, M.; Girault, D.; Bierque, E.; Fernandes, J.; Bähre, H.; Douyère, A.; Eskenazi, N.; Vinh, J.; et al. The Zoonotic Pathogen Leptospira interrogans Mitigates Environmental Stress through Cyclic-Di-GMP-Controlled Biofilm Production. NPJ Biofilms Microbiomes 2020, 6, 24. [Google Scholar] [CrossRef]
- da Costa Vasconcelos, F.N.; Maciel, N.K.; Favaro, D.C.; de Oliveira, L.C.; Barbosa, A.S.; Salinas, R.K.; de Souza, R.F.; Farah, C.S.; Guzzo, C.R. Structural and Enzymatic Characterization of a cAMP-Dependent Diguanylate Cyclase from Pathogenic Leptospira Species. J. Mol. Biol. 2017, 429, 2337–2352. [Google Scholar] [CrossRef]
- Xiao, G.; Kong, L.; Che, R.; Yi, Y.; Zhang, Q.; Yan, J.; Lin, X. Identification and Characterization of c-Di-GMP Metabolic Enzymes of and c-Di-GMP Fluctuations After Thermal Shift and Infection. Front. Microbiol. 2018, 9, 764. [Google Scholar] [CrossRef]
- Vasconcelos, L.; Aburjaile, F.; Andrade, L.; Cancio, A.F.; Seyffert, N.; Aguiar, E.R.G.R.; Ristow, P. Genomic Insights into the c-Di-GMP Signaling and Biofilm Development in the Saprophytic Spirochete Leptospira biflexa. Arch. Microbiol. 2023, 205, 180. [Google Scholar] [CrossRef]
- Visnardi, A.B.; Ribeiro, R.A.; de Souza, A.S.; Churasacari Vinces, T.G.; Llontop, E.E.; de Almeida Ferrari, A.S.; França Henrique, P.A.; Valdivieso, D.; Sánchez-Limache, D.E.; Silva, G.R.; et al. Insertion of a Divergent GAF-like Domain Defines a Novel Family of YcgR Homologues That Bind c-Di-GMP in. ACS Omega 2025, 10, 3988–4006. [Google Scholar] [CrossRef] [PubMed]
- Cheang, Q.W.; Xin, L.; Chea, R.Y.F.; Liang, Z.-X. Emerging Paradigms for PilZ Domain-Mediated C-Di-GMP Signaling. Biochem. Soc. Trans. 2019, 47, 381–388. [Google Scholar] [CrossRef]
- Römling, U.; Galperin, M.Y.; Gomelsky, M. Cyclic Di-GMP: The First 25 Years of a Universal Bacterial Second Messenger. Microbiol. Mol. Biol. Rev. 2013, 77, 1–52. [Google Scholar] [CrossRef] [PubMed]
- Park, S.; Sauer, K. Controlling Biofilm Development Through Cyclic Di-GMP Signaling. Adv. Exp. Med. Biol. 2022, 1386, 69–94. [Google Scholar] [CrossRef] [PubMed]
- Blanco-Romero, E.; Garrido-Sanz, D.; Durán, D.; Rybtke, M.; Tolker-Nielsen, T.; Redondo-Nieto, M.; Rivilla, R.; Martín, M. Role of Extracellular Matrix Components in Biofilm Formation and Adaptation of F113 to the Rhizosphere Environment. Front. Microbiol. 2024, 15, 1341728. [Google Scholar] [CrossRef] [PubMed]
- Schirmer, T.; Jenal, U. Structural and Mechanistic Determinants of c-Di-GMP Signalling. Nat. Rev. Microbiol. 2009, 7, 724–735. [Google Scholar] [CrossRef]
- da Purificação, A.D.; de Azevedo, N.M.; de Araujo, G.G.; de Souza, R.F.; Guzzo, C.R. The World of Cyclic Dinucleotides in Bacterial Behavior. Molecules 2020, 25, 2462. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Yu, Z.; Zhu, L.; Li, Z.; Yin, W.; Shang, X.; Chou, S.-H.; Tan, Q.; He, J. The Multiple Regulatory Relationship Between RNA-Chaperone Hfq and the Second Messenger c-Di-GMP. Front. Microbiol. 2021, 12, 689619. [Google Scholar] [CrossRef]
- Valentini, M.; Filloux, A. Biofilms and Cyclic Di-GMP (c-Di-GMP) Signaling: Lessons from Pseudomonas aeruginosa and Other Bacteria. J. Biol. Chem. 2016, 291, 12547–12555. [Google Scholar] [CrossRef]
- UniProt Consortium. UniProt: The Universal Protein Knowledgebase in 2023. Nucleic Acids Res. 2023, 51, D523–D531. [Google Scholar] [CrossRef]
- Chou, S.-H.; Galperin, M.Y. Diversity of Cyclic Di-GMP-Binding Proteins and Mechanisms. J. Bacteriol. 2016, 198, 32–46. [Google Scholar] [CrossRef] [PubMed]
- Roelofs, K.G.; Jones, C.J.; Helman, S.R.; Shang, X.; Orr, M.W.; Goodson, J.R.; Galperin, M.Y.; Yildiz, F.H.; Lee, V.T. Systematic Identification of Cyclic-Di-GMP Binding Proteins in Vibrio cholerae Reveals a Novel Class of Cyclic-Di-GMP-Binding ATPases Associated with Type II Secretion Systems. PLoS Pathog. 2015, 11, e1005232. [Google Scholar] [CrossRef]
- Wang, Y.-C.; Chin, K.-H.; Tu, Z.-L.; He, J.; Jones, C.J.; Sanchez, D.Z.; Yildiz, F.H.; Galperin, M.Y.; Chou, S.-H. Nucleotide Binding by the Widespread High-Affinity Cyclic Di-GMP Receptor MshEN Domain. Nat. Commun. 2016, 7, 12481. [Google Scholar] [CrossRef]
- Amikam, D.; Galperin, M.Y. PilZ Domain Is Part of the Bacterial c-Di-GMP Binding Protein. Bioinformatics 2006, 22, 3–6. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic Local Alignment Search Tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Nascimento, A.L.T.O.; Verjovski-Almeida, S.; Van Sluys, M.A.; Monteiro-Vitorello, C.B.; Camargo, L.E.A.; Digiampietri, L.A.; Harstkeerl, R.A.; Ho, P.L.; Marques, M.V.; Oliveira, M.C.; et al. Genome Features of Leptospira interrogans serovar Copenhageni. Braz. J. Med. Biol. Res. 2004, 37, 459–477. [Google Scholar] [CrossRef]
- Abramson, J.; Adler, J.; Dunger, J.; Evans, R.; Green, T.; Pritzel, A.; Ronneberger, O.; Willmore, L.; Ballard, A.J.; Bambrick, J.; et al. Accurate Structure Prediction of Biomolecular Interactions with AlphaFold 3. Nature 2024, 630, 493–500. [Google Scholar] [CrossRef]
- van Kempen, M.; Kim, S.S.; Tumescheit, C.; Mirdita, M.; Lee, J.; Gilchrist, C.L.M.; Söding, J.; Steinegger, M. Fast and Accurate Protein Structure Search with Foldseek. Nat. Biotechnol. 2024, 42, 243–246. [Google Scholar] [CrossRef] [PubMed]
- Holm, L. Dali Server: Structural Unification of Protein Families. Nucleic Acids Res. 2022, 50, W210–W215. [Google Scholar] [CrossRef]
- Hildebrand, A.; Remmert, M.; Biegert, A.; Söding, J. Fast and Accurate Automatic Structure Prediction with HHpred. Proteins 2009, 77, 128–132. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Chitsaz, F.; Derbyshire, M.K.; Gonzales, N.R.; Gwadz, M.; Lu, S.; Marchler, G.H.; Song, J.S.; Thanki, N.; Yamashita, R.A.; et al. The Conserved Domain Database in 2023. Nucleic Acids Res. 2023, 51, D384–D388. [Google Scholar] [CrossRef]
- Blum, M.; Andreeva, A.; Florentino, L.C.; Chuguransky, S.R.; Grego, T.; Hobbs, E.; Pinto, B.L.; Orr, A.; Paysan-Lafosse, T.; Ponamareva, I.; et al. InterPro: The Protein Sequence Classification Resource in 2025. Nucleic Acids Res. 2025, 53, D444–D456. [Google Scholar] [CrossRef]
- Paysan-Lafosse, T.; Andreeva, A.; Blum, M.; Chuguransky, S.R.; Grego, T.; Pinto, B.L.; Salazar, G.A.; Bileschi, M.L.; Llinares-López, F.; Meng-Papaxanthos, L.; et al. The Pfam Protein Families Database: Embracing AI/ML. Nucleic Acids Res. 2025, 53, D523–D534. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.-S.; Chen, Y.-C.; Lu, C.-H.; Hwang, J.-K. Prediction of Protein Subcellular Localization. Proteins 2006, 64, 643–651. [Google Scholar] [CrossRef]
- Almagro Armenteros, J.J.; Tsirigos, K.D.; Sønderby, C.K.; Petersen, T.N.; Winther, O.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 5.0 Improves Signal Peptide Predictions Using Deep Neural Networks. Nat. Biotechnol. 2019, 37, 420–423. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.-X.; Pan, X.; Shen, H.-B. Signal-3L 3.0: Improving Signal Peptide Prediction through Combining Attention Deep Learning with Window-Based Scoring. J. Chem. Inf. Model. 2020, 60, 3679–3686. [Google Scholar] [CrossRef] [PubMed]
- Krogh, A.; Larsson, B.; von Heijne, G.; Sonnhammer, E.L. Predicting Transmembrane Protein Topology with a Hidden Markov Model: Application to Complete Genomes. J. Mol. Biol. 2001, 305, 567–580. [Google Scholar] [CrossRef]
- Hallgren, J.; Tsirigos, K.D.; Pedersen, M.D.; Almagro Armenteros, J.J.; Marcatili, P.; Nielsen, H.; Krogh, A.; Winther, O. DeepTMHMM Predicts Alpha and Beta Transmembrane Proteins Using Deep Neural Networks. bioRxiv 2022. [Google Scholar] [CrossRef]
- Waterhouse, A.M.; Procter, J.B.; Martin, D.M.A.; Clamp, M.; Barton, G.J. Jalview Version 2—A Multiple Sequence Alignment Editor and Analysis Workbench. Bioinformatics 2009, 25, 1189–1191. [Google Scholar] [CrossRef]
- Chan, C.; Paul, R.; Samoray, D.; Amiot, N.C.; Giese, B.; Jenal, U.; Schirmer, T. Structural Basis of Activity and Allosteric Control of Diguanylate Cyclase. Proc. Natl. Acad. Sci. USA 2004, 101, 17084–17089. [Google Scholar] [CrossRef]
- Marks, M.E.; Castro-Rojas, C.M.; Teiling, C.; Du, L.; Kapatral, V.; Walunas, T.L.; Crosson, S. The Genetic Basis of Laboratory Adaptation in Caulobacter crescentus. J. Bacteriol. 2010, 192, 3678–3688. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.W.; Kotaka, M.; Vonrhein, C.; Bricogne, G.; Rao, F.; Chuah, M.L.C.; Svergun, D.; Schneider, G.; Liang, Z.-X.; Lescar, J. Structural Insights into the Regulatory Mechanism of the Response Regulator RocR from Pseudomonas aeruginosa in Cyclic Di-GMP Signaling. J. Bacteriol. 2012, 194, 4837–4846. [Google Scholar] [CrossRef]
- Stover, C.K.; Pham, X.Q.; Erwin, A.L.; Mizoguchi, S.D.; Warrener, P.; Hickey, M.J.; Brinkman, F.S.; Hufnagle, W.O.; Kowalik, D.J.; Lagrou, M.; et al. Complete Genome Sequence of Pseudomonas aeruginosa PAO1, an Opportunistic Pathogen. Nature 2000, 406, 959–964. [Google Scholar] [CrossRef]
- Bellini, D.; Caly, D.L.; McCarthy, Y.; Bumann, M.; An, S.-Q.; Dow, J.M.; Ryan, R.P.; Walsh, M.A. Crystal Structure of an HD-GYP Domain Cyclic-Di-GMP Phosphodiesterase Reveals an Enzyme with a Novel Trinuclear Catalytic Iron Centre. Mol. Microbiol. 2014, 91, 26–38. [Google Scholar] [CrossRef]
- Reysenbach, A.-L.; Hamamura, N.; Podar, M.; Griffiths, E.; Ferreira, S.; Hochstein, R.; Heidelberg, J.; Johnson, J.; Mead, D.; Pohorille, A.; et al. Complete and Draft Genome Sequences of Six Members of the Aquificales. J. Bacteriol. 2009, 191, 1992–1993. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Yuan, Z.; Gu, L. Structural Basis for the Regulation of Chemotaxis by MapZ in the Presence of c-Di-GMP. Acta Crystallogr. D Struct. Biol. 2017, 73, 683–691. [Google Scholar] [CrossRef]
- Heidelberg, J.F.; Eisen, J.A.; Nelson, W.C.; Clayton, R.A.; Gwinn, M.L.; Dodson, R.J.; Haft, D.H.; Hickey, E.K.; Peterson, J.D.; Umayam, L.; et al. DNA Sequence of Both Chromosomes of the Cholera Pathogen Vibrio cholerae. Nature 2000, 406, 477–483. [Google Scholar] [CrossRef]
- Goldfarb, T.; Kodali, V.K.; Pujar, S.; Brover, V.; Robbertse, B.; Farrell, C.M.; Oh, D.-H.; Astashyn, A.; Ermolaeva, O.; Haddad, D.; et al. NCBI RefSeq: Reference Sequence Standards through 25 Years of Curation and Annotation. Nucleic Acids Res. 2024, 53, D243–D257. [Google Scholar] [CrossRef]
- Parks, D.H.; Chuvochina, M.; Rinke, C.; Mussig, A.J.; Chaumeil, P.-A.; Hugenholtz, P. GTDB: An Ongoing Census of Bacterial and Archaeal Diversity through a Phylogenetically Consistent, Rank Normalized and Complete Genome-Based Taxonomy. Nucleic Acids Res. 2021, 50, D785–D794. [Google Scholar] [CrossRef]
- Chklovski, A.; Parks, D.H.; Woodcroft, B.J.; Tyson, G.W. CheckM2: A Rapid, Scalable and Accurate Tool for Assessing Microbial Genome Quality Using Machine Learning. Nat. Methods 2023, 20, 1203–1212. [Google Scholar] [CrossRef] [PubMed]
- GitHub—Leepbioinfo/Rotifer. Available online: https://github.com/leepbioinfo/rotifer (accessed on 25 October 2025).
- Perrin, A.; Rocha, E.P.C. PanACoTA: A Modular Tool for Massive Microbial Comparative Genomics. NAR Genom. Bioinform. 2021, 3, lqaa106. [Google Scholar] [CrossRef] [PubMed]
- Jain, C.; Rodriguez-R, L.M.; Phillippy, A.M.; Konstantinidis, K.T.; Aluru, S. High Throughput ANI Analysis of 90K Prokaryotic Genomes Reveals Clear Species Boundaries. Nat. Commun. 2018, 9, 5114. [Google Scholar] [CrossRef]
- Talevich, E.; Invergo, B.M.; Cock, P.J.A.; Chapman, B.A. Bio.Phylo: A Unified Toolkit for Processing, Analyzing and Visualizing Phylogenetic Trees in Biopython. BMC Bioinform. 2012, 13, 209. [Google Scholar] [CrossRef]
- Ren, S.-X.; Fu, G.; Jiang, X.-G.; Zeng, R.; Miao, Y.-G.; Xu, H.; Zhang, Y.-X.; Xiong, H.; Lu, G.; Lu, L.-F.; et al. Unique Physiological and Pathogenic Features of Leptospira interrogans Revealed by Whole-Genome Sequencing. Nature 2003, 422, 888–893. [Google Scholar] [CrossRef]
- Bulach, D.M.; Zuerner, R.L.; Wilson, P.; Seemann, T.; McGrath, A.; Cullen, P.A.; Davis, J.; Johnson, M.; Kuczek, E.; Alt, D.P.; et al. Genome Reduction in Leptospira borgpetersenii Reflects Limited Transmission Potential. Proc. Natl. Acad. Sci. USA 2006, 103, 14560–14565. [Google Scholar] [CrossRef] [PubMed]
- Senavirathna, I.; Jayasundara, D.; Warnasekara, J.; Agampodi, S.; Putz, E.J.; Nally, J.E.; Bayles, D.O.; Chaurasia, R.; Vinetz, J.M. Genomic Analysis of Human-Infecting Leptospira borgpetersenii Isolates in Sri Lanka: Expanded PF07598 Gene Family Repertoire, Less Overall Genome Reduction than Bovine Isolates. bioRxiv 2024. [Google Scholar] [CrossRef]
- Putz, E.J.; Sivasankaran, S.K.; Fernandes, L.G.V.; Brunelle, B.; Lippolis, J.D.; Alt, D.P.; Bayles, D.O.; Hornsby, R.L.; Nally, J.E. Distinct Transcriptional Profiles of Leptospira borgpetersenii serovar Hardjo strains JB197 and HB203 Cultured at Different Temperatures. PLoS Negl. Trop. Dis. 2021, 15, e0009320. [Google Scholar] [CrossRef]
- Picardeau, M.; Bulach, D.M.; Bouchier, C.; Zuerner, R.L.; Zidane, N.; Wilson, P.J.; Creno, S.; Kuczek, E.S.; Bommezzadri, S.; Davis, J.C.; et al. Genome Sequence of the Saprophyte Leptospira biflexa Provides Insights into the Evolution of Leptospira and the Pathogenesis of Leptospirosis. PLoS ONE 2008, 3, e1607. [Google Scholar] [CrossRef]
- Shim, S.R.; Kim, S.-J.; Lee, J.; Rücker, G. Network Meta-Analysis: Application and Practice Using R Software. Epidemiol. Health 2019, 41, e2019013. [Google Scholar] [CrossRef]
- Morgan, M.; Ramos, M. BiocManager: Access the Bioconductor Project Package Repository. Version: 1.30.27. 2025. Available online: https://cran.r-project.org/web/packages/BiocManager/index.html (accessed on 1 December 2025).
- Gu, Z.; Eils, R.; Schlesner, M. Complex Heatmaps Reveal Patterns and Correlations in Multidimensional Genomic Data. Bioinformatics 2016, 32, 2847–2849. [Google Scholar] [CrossRef]
- Paradis, E.; Schliep, K. Ape 5.0: An Environment for Modern Phylogenetics and Evolutionary Analyses in R. Bioinformatics 2019, 35, 526–528. [Google Scholar] [CrossRef]
- Inkscape Project. Inkscape. Available online: https://inkscape.org/ (accessed on 13 October 2025).
- Schrödinger, L.L.C. The PyMOL Molecular Graphics System. Available online: https://www.pymol.org/ (accessed on 13 October 2025).
- Meng, E.C.; Goddard, T.D.; Pettersen, E.F.; Couch, G.S.; Pearson, Z.J.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Tools for Structure Building and Analysis. Protein Sci. 2023, 32, e4792. [Google Scholar] [CrossRef] [PubMed]
- Krasteva, P.V.; Giglio, K.M.; Sondermann, H. Sensing the Messenger: The Diverse Ways That Bacteria Signal through c-Di-GMP: The Ins and Outs of c-Di-GMP Signaling. Protein Sci. 2012, 21, 929–948. [Google Scholar] [CrossRef] [PubMed]
- Nie, H.; Xiao, Y.; He, J.; Liu, H.; Nie, L.; Chen, W.; Huang, Q. Phenotypic-Genotypic Analysis of GGDEF/EAL/HD-GYP Domain-Encoding Genes in Pseudomonas putida. Environ. Microbiol. Rep. 2020, 12, 38–48. [Google Scholar] [CrossRef]
- Yang, C.-Y.; Chin, K.-H.; Chuah, M.L.-C.; Liang, Z.-X.; Wang, A.H.-J.; Chou, S.-H. The Structure and Inhibition of a GGDEF Diguanylate Cyclase Complexed with (c-Di-GMP)(2) at the Active Site. Acta Crystallogr. D Biol. Crystallogr. 2011, 67, 997–1008. [Google Scholar] [CrossRef]
- Römling, U.; Liang, Z.-X.; Dow, J.M. Progress in Understanding the Molecular Basis Underlying Functional Diversification of Cyclic Dinucleotide Turnover Proteins. J. Bacteriol. 2017, 199, e00790-16. [Google Scholar] [CrossRef]
- Hallberg, Z.F.; Wang, X.C.; Wright, T.A.; Nan, B.; Ad, O.; Yeo, J.; Hammond, M.C. Hybrid Promiscuous (Hypr) GGDEF Enzymes Produce Cyclic AMP-GMP (3′, 3′-cGAMP). Proc. Natl. Acad. Sci. USA 2016, 113, 1790–1795. [Google Scholar] [CrossRef] [PubMed]
- Hallberg, Z.F.; Chan, C.H.; Wright, T.A.; Kranzusch, P.J.; Doxzen, K.W.; Park, J.J.; Bond, D.R.; Hammond, M.C. Structure and Mechanism of a Hypr GGDEF Enzyme That Activates cGAMP Signaling to Control Extracellular Metal Respiration. eLife 2019, 8, e43959. [Google Scholar] [CrossRef]
- Rangarajan, A.A.; Waters, C.M. Double Take: A Dual-Functional Hypr GGDEF Synthesizes Both Cyclic Di-GMP and Cyclic GMP-AMP to Control Predation in Bdellovibrio bacteriovorus. PLoS Genet. 2022, 18, e1010263. [Google Scholar] [CrossRef]
- Rao, F.; See, R.Y.; Zhang, D.; Toh, D.C.; Ji, Q.; Liang, Z.-X. YybT Is a Signaling Protein That Contains a Cyclic Dinucleotide Phosphodiesterase Domain and a GGDEF Domain with ATPase Activity. J. Biol. Chem. 2011, 286, 29441. [Google Scholar] [CrossRef]
- Schirmer, T. C-Di-GMP Synthesis: Structural Aspects of Evolution, Catalysis and Regulation. J. Mol. Biol. 2016, 428, 3683–3701. [Google Scholar] [CrossRef]
- Wassmann, P.; Chan, C.; Paul, R.; Beck, A.; Heerklotz, H.; Jenal, U.; Schirmer, T. Structure of BeF3− -Modified Response Regulator PleD: Implications for Diguanylate Cyclase Activation, Catalysis, and Feedback Inhibition. Structure 2007, 15, 915–927. [Google Scholar] [CrossRef] [PubMed]
- Galperin, M.Y.; Chou, S.-H. Sequence Conservation, Domain Architectures, and Phylogenetic Distribution of the HD-GYP Type c-Di-GMP Phosphodiesterases. J. Bacteriol. 2022, 204, e0056121. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, A.J.; Ryjenkov, D.A.; Gomelsky, M. The Ubiquitous Protein Domain EAL Is a Cyclic Diguanylate-Specific Phosphodiesterase: Enzymatically Active and Inactive EAL Domains. J. Bacteriol. 2005, 187, 4774–4781. [Google Scholar] [CrossRef]
- Tischler, A.D.; Camilli, A. Cyclic Diguanylate (c-Di-GMP) Regulates Vibrio cholerae Biofilm Formation. Mol. Microbiol. 2004, 53, 857–869. [Google Scholar] [CrossRef]
- Römling, U.; Gomelsky, M.; Galperin, M.Y. C-Di-GMP: The Dawning of a Novel Bacterial Signalling System. Mol. Microbiol. 2005, 57, 629–639. [Google Scholar] [CrossRef]
- Heo, K.; Lee, J.-W.; Jang, Y.; Kwon, S.; Lee, J.; Seok, C.; Ha, N.-C.; Seok, Y.-J. A pGpG-Specific Phosphodiesterase Regulates Cyclic Di-GMP Signaling in Vibrio cholerae. J. Biol. Chem. 2022, 298, 101626. [Google Scholar] [CrossRef]
- Rao, F.; Qi, Y.; Chong, H.S.; Kotaka, M.; Li, B.; Li, J.; Lescar, J.; Tang, K.; Liang, Z.-X. The Functional Role of a Conserved Loop in EAL Domain-Based Cyclic Di-GMP-Specific Phosphodiesterase. J. Bacteriol. 2009, 191, 4722–4731. [Google Scholar] [CrossRef] [PubMed]
- Römling, U. Rationalizing the Evolution of EAL Domain-Based Cyclic Di-GMP-Specific Phosphodiesterases. J. Bacteriol. 2009, 191, 4697–4700. [Google Scholar] [CrossRef]
- Tchigvintsev, A.; Xu, X.; Singer, A.; Chang, C.; Brown, G.; Proudfoot, M.; Cui, H.; Flick, R.; Anderson, W.F.; Joachimiak, A.; et al. Structural Insight into the Mechanism of c-Di-GMP Hydrolysis by EAL Domain Phosphodiesterases. J. Mol. Biol. 2010, 402, 524–538. [Google Scholar] [CrossRef] [PubMed]
- Barends, T.R.M.; Hartmann, E.; Griese, J.J.; Beitlich, T.; Kirienko, N.V.; Ryjenkov, D.A.; Reinstein, J.; Shoeman, R.L.; Gomelsky, M.; Schlichting, I. Structure and Mechanism of a Bacterial Light-Regulated Cyclic Nucleotide Phosphodiesterase. Nature 2009, 459, 1015–1018. [Google Scholar] [CrossRef]
- Tarutina, M.; Ryjenkov, D.A.; Gomelsky, M. An Unorthodox Bacteriophytochrome from Rhodobacter sphaeroides Involved in Turnover of the Second Messenger c-Di-GMP. J. Biol. Chem. 2006, 281, 34751–34758. [Google Scholar] [CrossRef] [PubMed]
- Johnson, J.G.; Murphy, C.N.; Sippy, J.; Johnson, T.J.; Clegg, S. Type 3 Fimbriae and Biofilm Formation Are Regulated by the Transcriptional Regulators MrkHI in Klebsiella pneumoniae. J. Bacteriol. 2011, 193, 3453–3460. [Google Scholar] [CrossRef]
- Sultan, S.Z.; Pitzer, J.E.; Boquoi, T.; Hobbs, G.; Miller, M.R.; Motaleb, M.A. Analysis of the HD-GYP Domain Cyclic Dimeric GMP Phosphodiesterase Reveals a Role in Motility and the Enzootic Life Cycle of Borrelia burgdorferi. Infect. Immun. 2011, 79, 3273–3283. [Google Scholar] [CrossRef]
- Cutruzzolà, F.; Paiardini, A.; Scribani Rossi, C.; Spizzichino, S.; Paone, A.; Giardina, G.; Rinaldo, S. A Conserved Scaffold with Heterogeneous Metal Ion Binding Site: The Multifaceted Example of HD-GYP Proteins. Coord. Chem. Rev. 2022, 450, 214228. [Google Scholar] [CrossRef]
- Ryan, R.P.; McCarthy, Y.; Andrade, M.; Farah, C.S.; Armitage, J.P.; Dow, J.M. Cell-Cell Signal-Dependent Dynamic Interactions between HD-GYP and GGDEF Domain Proteins Mediate Virulence in Xanthomonas campestris. Proc. Natl. Acad. Sci. USA 2010, 107, 5989–5994. [Google Scholar] [CrossRef]
- Sun, S.; Pandelia, M.-E. HD-[HD-GYP] Phosphodiesterases: Activities and Evolutionary Diversification within the HD-GYP Family. Biochemistry 2020, 59, 2340–2350. [Google Scholar] [CrossRef]
- Gong, H.; Gao, Y.; Zhou, X.; Xiao, Y.; Wang, W.; Tang, Y.; Zhou, S.; Zhang, Y.; Ji, W.; Yu, L.; et al. Cryo-EM Structure of Trimeric Mycobacterium smegmatis Succinate Dehydrogenase with a Membrane-Anchor SdhF. Nat. Commun. 2020, 11, 4245. [Google Scholar] [CrossRef]
- He, D.; You, L.; Wu, X.; Shi, J.; Wen, A.; Yan, Z.; Mu, W.; Fang, C.; Feng, Y.; Zhang, Y. Pseudomonas aeruginosa SutA Wedges RNAP Lobe Domain Open to Facilitate Promoter DNA Unwinding. Nat. Commun. 2022, 13, 4204. [Google Scholar] [CrossRef]
- Zhang, X.; Liang, Y.; Zheng, K.; Wang, Z.; Dong, Y.; Liu, Y.; Ren, L.; Wang, H.; Han, Y.; McMinn, A.; et al. Characterization and Genomic Analysis of Phage vB_ValR_NF, Representing a New Viral Family Prevalent in the Ulva prolifera Blooms. Front. Microbiol. 2023, 14, 1161265. [Google Scholar] [CrossRef]
- Macián, M.C.; Ludwig, W.; Schleifer, K.H.; Garay, E.; Pujalte, M.J. Thalassomonas viridans Gen. Nov., Sp. Nov., a Novel Marine Gamma-Proteobacterium. Int. J. Syst. Evol. Microbiol. 2001, 51, 1283–1289. [Google Scholar] [CrossRef] [PubMed]
- Ward, L.M.; Bertran, E.; Johnston, D.T. Genomic Sequence Analysis of Dissulfurirhabdus thermomarina SH388 and Proposed Reassignment to Dissulfurirhabdaceae Fam. Nov. Microb. Genom. 2020, 6, e000390. [Google Scholar] [CrossRef] [PubMed]
- Laskowski, R.A.; Jabłońska, J.; Pravda, L.; Vařeková, R.S.; Thornton, J.M. PDBsum: Structural Summaries of PDB Entries. Protein Sci. 2018, 27, 129–134. [Google Scholar] [CrossRef]
- Galperin, M.Y.; Chou, S.-H. Structural Conservation and Diversity of PilZ-Related Domains. J. Bacteriol. 2020, 202, e00664-19. [Google Scholar] [CrossRef]
- Sheng, S.; Xin, L.; Yam, J.K.H.; Salido, M.M.; Khong, N.Z.J.; Liu, Q.; Chea, R.A.; Li, H.Y.; Yang, L.; Liang, Z.-X.; et al. The MapZ-Mediated Methylation of Chemoreceptors Contributes to Pathogenicity of Pseudomonas aeruginosa. Front. Microbiol. 2019, 10, 67. [Google Scholar] [CrossRef]
- Chen, Y.; Tsai, B.; Li, N.; Gao, N. Structural Remodeling of Ribosome Associated Hsp40-Hsp70 Chaperones during Co-Translational Folding. Nat. Commun. 2022, 13, 3410. [Google Scholar] [CrossRef] [PubMed]
- Koplin, A.; Preissler, S.; Ilina, Y.; Koch, M.; Scior, A.; Erhardt, M.; Deuerling, E. A Dual Function for Chaperones SSB-RAC and the NAC Nascent Polypeptide-Associated Complex on Ribosomes. J. Cell Biol. 2010, 189, 57–68. [Google Scholar] [CrossRef]
- Peisker, K.; Braun, D.; Wölfle, T.; Hentschel, J.; Fünfschilling, U.; Fischer, G.; Sickmann, A.; Rospert, S. Ribosome-Associated Complex Binds to Ribosomes in Close Proximity of Rpl31 at the Exit of the Polypeptide Tunnel in Yeast. Mol. Biol. Cell 2008, 19, 5279–5288. [Google Scholar] [CrossRef]
- Pichlo, C.; Juetten, L.; Wojtalla, F.; Schacherl, M.; Diaz, D.; Baumann, U. Molecular Determinants of the Mechanism and Substrate Specificity of Clostridium difficile Proline-Proline Endopeptidase-1. J. Biol. Chem. 2019, 294, 11525–11535. [Google Scholar] [CrossRef]
- Pannifer, A.D.; Wong, T.Y.; Schwarzenbacher, R.; Renatus, M.; Petosa, C.; Bienkowska, J.; Lacy, D.B.; Collier, R.J.; Park, S.; Leppla, S.H.; et al. Crystal Structure of the Anthrax Lethal Factor. Nature 2001, 414, 229–233. [Google Scholar] [CrossRef]
- Visschedyk, D.; Rochon, A.; Tempel, W.; Dimov, S.; Park, H.-W.; Merrill, A.R. Certhrax Toxin, an Anthrax-Related ADP-Ribosyltransferase from Bacillus cereus. J. Biol. Chem. 2012, 287, 41089–41102. [Google Scholar] [CrossRef]
- Scheithauer, L.; Thiem, S.; Schmelz, S.; Dellmann, A.; Büssow, K.; Brouwer, R.M.H.J.; Ünal, C.M.; Blankenfeldt, W.; Steinert, M. Zinc Metalloprotease ProA of Legionella pneumophila Increases Alveolar Septal Thickness in Human Lung Tissue Explants by Collagen IV Degradation. Cell. Microbiol. 2021, 23, e13313. [Google Scholar] [CrossRef]
- Shen, Y.; Zhukovskaya, N.L.; Guo, Q.; Florián, J.; Tang, W.-J. Calcium-Independent Calmodulin Binding and Two-Metal-Ion Catalytic Mechanism of Anthrax Edema Factor. EMBO J. 2005, 24, 929–941. [Google Scholar] [CrossRef] [PubMed]
- Pei, J.; Mitchell, D.A.; Dixon, J.E.; Grishin, N.V. Expansion of Type II CAAX Proteases Reveals Evolutionary Origin of γ-Secretase Subunit APH-1. J. Mol. Biol. 2011, 410, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Ellermeier, C.D.; Losick, R. Evidence for a Novel Protease Governing Regulated Intramembrane Proteolysis and Resistance to Antimicrobial Peptides in Bacillus subtilis. Genes Dev. 2006, 20, 1911–1922. [Google Scholar] [CrossRef]
- Dalbey, R.E.; Kuhn, A.; Zhu, L.; Kiefer, D. The Membrane Insertase YidC. Biochim. Biophys. Acta 2014, 1843, 1489–1496. [Google Scholar] [CrossRef]
- Jones, C.J.; Utada, A.; Davis, K.R.; Thongsomboon, W.; Zamorano Sanchez, D.; Banakar, V.; Cegelski, L.; Wong, G.C.L.; Yildiz, F.H. C-Di-GMP Regulates Motile to Sessile Transition by Modulating MshA Pili Biogenesis and near-Surface Motility Behavior in Vibrio cholerae. PLoS Pathog. 2015, 11, e1005068. [Google Scholar] [CrossRef]
- Düvel, J.; Bense, S.; Möller, S.; Bertinetti, D.; Schwede, F.; Morr, M.; Eckweiler, D.; Genieser, H.-G.; Jänsch, L.; Herberg, F.W.; et al. Application of Synthetic Peptide Arrays to Uncover Cyclic Di-GMP Binding Motifs. J. Bacteriol. 2015, 198, 138–146. [Google Scholar] [CrossRef]
- Chen, Y.; Shiue, S.-J.; Huang, C.-W.; Chang, J.-L.; Chien, Y.-L.; Hu, N.-T.; Chan, N.-L. Structure and Function of the XpsE N-Terminal Domain, an Essential Component of the Xanthomonas campestris Type II Secretion System. J. Biol. Chem. 2005, 280, 42356–42363. [Google Scholar] [CrossRef]
- Huerta-Cepas, J.; Serra, F.; Bork, P. ETE 3: Reconstruction, Analysis, and Visualization of Phylogenomic Data. Mol. Biol. Evol. 2016, 33, 1635–1638. [Google Scholar] [CrossRef] [PubMed]
- Christen, M.; Christen, B.; Folcher, M.; Schauerte, A.; Jenal, U. Identification and Characterization of a Cyclic Di-GMP-Specific Phosphodiesterase and Its Allosteric Control by GTP. J. Biol. Chem. 2005, 280, 30829–30837. [Google Scholar] [CrossRef] [PubMed]
- Jenal, U.; Reinders, A.; Lori, C. Cyclic Di-GMP: Second Messenger Extraordinaire. Nat. Rev. Microbiol. 2017, 15, 271–284. [Google Scholar] [CrossRef]
- Martín-Rodríguez, A.J.; Higdon, S.M.; Thorell, K.; Tellgren-Roth, C.; Sjöling, Å.; Galperin, M.Y.; Krell, T.; Römling, U. Comparative Genomics of Cyclic Di-GMP Metabolism and Chemosensory Pathways in Shewanella algae Strains: Novel Bacterial Sensory Domains and Functional Insights into Lifestyle Regulation. mSystems 2022, 7, e0151821. [Google Scholar] [CrossRef]
- Galperin, M.Y. A Census of Membrane-Bound and Intracellular Signal Transduction Proteins in Bacteria: Bacterial IQ, Extroverts and Introverts. BMC Microbiol. 2005, 5, 35. [Google Scholar] [CrossRef] [PubMed]
- Galperin, M.Y.; Higdon, R.; Kolker, E. Interplay of Heritage and Habitat in the Distribution of Bacterial Signal Transduction Systems. Mol. Biosyst. 2010, 6, 721–728. [Google Scholar] [CrossRef]
- Guzzo, C.R.; Salinas, R.K.; Andrade, M.O.; Farah, C.S. PILZ Protein Structure and Interactions with PILB and the FIMX EAL Domain: Implications for Control of Type IV Pilus Biogenesis. J. Mol. Biol. 2009, 393, 848–866. [Google Scholar] [CrossRef]
- Khan, F.; Jeong, G.-J.; Tabassum, N.; Kim, Y.-M. Functional Diversity of c-Di-GMP Receptors in Prokaryotic and Eukaryotic Systems. Cell Commun. Signal. 2023, 21, 259. [Google Scholar] [CrossRef]
- Holland, L.M.; O’Donnell, S.T.; Ryjenkov, D.A.; Gomelsky, L.; Slater, S.R.; Fey, P.D.; Gomelsky, M.; O’Gara, J.P. A Staphylococcal GGDEF Domain Protein Regulates Biofilm Formation Independently of Cyclic Dimeric GMP. J. Bacteriol. 2008, 190, 5178–5189. [Google Scholar] [CrossRef]
- Li, M.-L.; Jiao, J.; Zhang, B.; Shi, W.-T.; Yu, W.-H.; Tian, C.-F. Global Transcriptional Repression of Diguanylate Cyclases by MucR1 Is Essential for Sinorhizobium-Soybean Symbiosis. mBio 2021, 12, e0119221. [Google Scholar] [CrossRef]
- Bobrov, A.G.; Kirillina, O.; Ryjenkov, D.A.; Waters, C.M.; Price, P.A.; Fetherston, J.D.; Mack, D.; Goldman, W.E.; Gomelsky, M.; Perry, R.D. Systematic Analysis of Cyclic Di-GMP Signalling Enzymes and Their Role in Biofilm Formation and Virulence in Yersinia pestis. Mol. Microbiol. 2011, 79, 533–551. [Google Scholar] [CrossRef] [PubMed]
- Bordeleau, E.; Fortier, L.-C.; Malouin, F.; Burrus, V. C-Di-GMP Turn-over in Clostridium difficile Is Controlled by a Plethora of Diguanylate Cyclases and Phosphodiesterases. PLoS Genet. 2011, 7, e1002039. [Google Scholar] [CrossRef] [PubMed]
- Aravind, L.; Koonin, E.V. The HD Domain Defines a New Superfamily of Metal-Dependent Phosphohydrolases. Trends Biochem. Sci. 1998, 23, 469–472. [Google Scholar] [CrossRef]
- Lovering, A.L.; Capeness, M.J.; Lambert, C.; Hobley, L.; Sockett, R.E. The Structure of an Unconventional HD-GYP Protein from Bdellovibrio Reveals the Roles of Conserved Residues in This Class of Cyclic-Di-GMP Phosphodiesterases. mBio 2011, 2, e00163-11. [Google Scholar] [CrossRef]
- McKee, R.W.; Kariisa, A.; Mudrak, B.; Whitaker, C.; Tamayo, R. A Systematic Analysis of the in Vitro and in Vivo Functions of the HD-GYP Domain Proteins of Vibrio cholerae. BMC Microbiol. 2014, 14, 272. [Google Scholar] [CrossRef] [PubMed]
- Stelitano, V.; Giardina, G.; Paiardini, A.; Castiglione, N.; Cutruzzolà, F.; Rinaldo, S. C-Di-GMP Hydrolysis by Pseudomonas Aeruginosa HD-GYP Phosphodiesterases: Analysis of the Reaction Mechanism and Novel Roles for pGpG. PLoS ONE 2013, 8, e74920. [Google Scholar] [CrossRef]
- Wang, X.; Hao, X.; Yang, Y.; Jia, S.; Chen, Y.; Yang, W.; Luo, Y.; Xie, Z.; Gu, Y.; Wu, Y.; et al. A Phosphodiesterase CpdB in Yersinia pseudotuberculosis Degrades CDNs to Inhibit Innate Immune Response. Vet. Microbiol. 2024, 297, 110194. [Google Scholar] [CrossRef]
- Remminghorst, U.; Rehm, B.H.A. Alg44, a Unique Protein Required for Alginate Biosynthesis in Pseudomonas Aeruginosa. FEBS Lett. 2006, 580, 3883–3888. [Google Scholar] [CrossRef] [PubMed]
- Moradali, M.F.; Donati, I.; Sims, I.; Ghods, S.; Rehm, B.H.A. Alginate Polymerization and Modification Are Linked in Pseudomonas aeruginosa. mBio 2015, 6, e00453-15. [Google Scholar] [CrossRef] [PubMed]
- Hou, Y.-J.; Yang, W.-S.; Hong, Y.; Zhang, Y.; Wang, D.-C.; Li, D.-F. Structural Insights into the Mechanism of c-Di-GMP-Bound YcgR Regulating Flagellar Motility in Escherichia coli. J. Biol. Chem. 2020, 295, 808–821. [Google Scholar] [CrossRef] [PubMed]
- Han, Q.; Wang, S.-F.; Qian, X.-X.; Guo, L.; Shi, Y.-F.; He, R.; Yuan, J.-H.; Hou, Y.-J.; Li, D.-F. Flagellar Brake Protein YcgR Interacts with Motor Proteins MotA and FliG to Regulate the Flagellar Rotation Speed and Direction. Front. Microbiol. 2023, 14, 1159974. [Google Scholar] [CrossRef]
- Chen, Y.; Tachiyama, S.; Li, Y.; Feng, X.; Zhao, H.; Wu, Y.; Guo, Y.; Lara-Tejero, M.; Hua, C.; Liu, J.; et al. Tetrameric PilZ Protein Stabilizes Stator Ring in Complex Flagellar Motor and Is Required for Motility in Campylobacter jejuni. Proc. Natl. Acad. Sci. USA 2025, 122, e2412594121. [Google Scholar] [CrossRef]
- Wilksch, J.J.; Yang, J.; Clements, A.; Gabbe, J.L.; Short, K.R.; Cao, H.; Cavaliere, R.; James, C.E.; Whitchurch, C.B.; Schembri, M.A.; et al. MrkH, a Novel c-Di-GMP-Dependent Transcriptional Activator, Controls Klebsiella pneumoniae Biofilm Formation by Regulating Type 3 Fimbriae Expression. PLoS Pathog. 2011, 7, e1002204. [Google Scholar] [CrossRef]
- Schumacher, M.A.; Zeng, W. Structures of the Activator of K. pneumonia Biofilm Formation, MrkH, Indicates PilZ Domains Involved in c-Di-GMP and DNA Binding. Proc. Natl. Acad. Sci. USA 2016, 113, 10067–10072. [Google Scholar] [CrossRef]
- Devkota, S.R.; Kwon, E.; Ha, S.C.; Chang, H.W.; Kim, D.Y. Structural Insights into the Regulation of Bacillus subtilis SigW Activity by Anti-Sigma RsiW. PLoS ONE 2017, 12, e0174284. [Google Scholar] [CrossRef]
- Heinrich, J.; Hein, K.; Wiegert, T. Two Proteolytic Modules Are Involved in Regulated Intramembrane Proteolysis of Bacillus subtilis RsiW. Mol. Microbiol. 2009, 74, 1412–1426. [Google Scholar] [CrossRef]
- Pannullo, A.G.; Ellermeier, C.D. Activation of the Extracytoplasmic Function σ Factor σV in Clostridioides difficile Requires Regulated Intramembrane Proteolysis of the Anti-σ Factor RsiV. mSphere 2022, 7, e0009222. [Google Scholar] [CrossRef] [PubMed]
- Helmann, J.D. Bacillus subtilis Extracytoplasmic Function (ECF) Sigma Factors and Defense of the Cell Envelope. Curr. Opin. Microbiol. 2016, 30, 122–132. [Google Scholar] [CrossRef]
- de Bruijn, F.J. (Ed.) Stress and Environmental Regulation of Gene Expression and Adaptation in Bacteria, 1st ed.; John Wiley & Sons: Hoboken, NJ, USA, 2016; ISBN 978-1-119-00488-2. [Google Scholar]
- Wang, Y.; Hay, I.D.; Rehman, Z.U.; Rehm, B.H.A. Membrane-Anchored MucR Mediates Nitrate-Dependent Regulation of Alginate Production in Pseudomonas aeruginosa. Appl. Microbiol. Biotechnol. 2015, 99, 7253–7265. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Heine, S.; Entian, M.; Sauer, K.; Frankenberg-Dinkel, N. NO-Induced Biofilm Dispersion in Pseudomonas aeruginosa Is Mediated by an MHYT Domain-Coupled Phosphodiesterase. J. Bacteriol. 2013, 195, 3531–3542. [Google Scholar] [CrossRef]
- Bharati, B.K.; Sharma, I.M.; Kasetty, S.; Kumar, M.; Mukherjee, R.; Chatterji, D. A Full-Length Bifunctional Protein Involved in c-Di-GMP Turnover Is Required for Long-Term Survival under Nutrient Starvation in Mycobacterium smegmatis. Microbiology 2012, 158, 1415–1427. [Google Scholar] [CrossRef]
- Trimble, M.J.; McCarter, L.L. Bis-(3′-5′)-Cyclic Dimeric GMP-Linked Quorum Sensing Controls Swarming in Vibrio parahaemolyticus. Proc. Natl. Acad. Sci. USA 2011, 108, 18079–18084. [Google Scholar] [CrossRef] [PubMed]
- Feirer, N.; Xu, J.; Allen, K.D.; Koestler, B.J.; Bruger, E.L.; Waters, C.M.; White, R.H.; Fuqua, C. A Pterin-Dependent Signaling Pathway Regulates a Dual-Function Diguanylate Cyclase-Phosphodiesterase Controlling Surface Attachment in Agrobacterium tumefaciens. MBio 2015, 6, e00156. [Google Scholar] [CrossRef]
- Bridges, A.A.; Prentice, J.A.; Fei, C.; Wingreen, N.S.; Bassler, B.L. Quantitative Input-Output Dynamics of a c-Di-GMP Signal Transduction Cascade in Vibrio cholerae. PLoS Biol. 2022, 20, e3001585. [Google Scholar] [CrossRef]
- Viruega-Góngora, V.I.; Acatitla-Jácome, I.S.; Zamorano-Sánchez, D.; Reyes-Carmona, S.R.; Xiqui-Vázquez, M.L.; Baca, B.E.; Ramírez-Mata, A. The GGDEF-EAL Protein CdgB from Azospirillum Baldaniorum Sp245, Is a Dual Function Enzyme with Potential Polar Localization. PLoS ONE 2022, 17, e0278036. [Google Scholar] [CrossRef]
- Guzzo, C.R.; Dunger, G.; Salinas, R.K.; Farah, C.S. Structure of the PilZ-FimXEAL-c-Di-GMP Complex Responsible for the Regulation of Bacterial Type IV Pilus Biogenesis. J. Mol. Biol. 2013, 425, 2174–2197. [Google Scholar] [CrossRef] [PubMed]
- Malmström, J.; Beck, M.; Schmidt, A.; Lange, V.; Deutsch, E.W.; Aebersold, R. Proteome-Wide Cellular Protein Concentrations of the Human Pathogen Leptospira interrogans. Nature 2009, 460, 762–765. [Google Scholar] [CrossRef] [PubMed]




| Locus | Genomes | Percentage |
|---|---|---|
| LIC_10049 | 364 | 99.73% |
| LIC_10128 | 365 | 100% |
| LIC_10139 | 362 | 99.18% |
| LIC_10321 | 365 | 100% |
| LIC_10996 | 364 | 99.73% |
| LIC_11125 | 362 | 99.18% |
| LIC_11127 | 362 | 99.18% |
| LIC_11129 | 362 | 99.18% |
| LIC_11130 | 363 | 99.45% |
| LIC_11167 | 364 | 99.73% |
| LIC_11247 | 362 | 99.18% |
| LIC_12273 | 364 | 99.73% |
| LIC_11300 | 362 | 99.18% |
| LIC_11444 | 365 | 100% |
| LIC_11563 | 363 | 99.45% |
| LIC_11571 | 364 | 99.73% |
| LIC_11706 | 365 | 100% |
| LIC_11993 | 365 | 100% |
| LIC_12994 | 364 | 99.73% |
| LIC_13137 | 363 | 99.45% |
| LIC_12491 | 363 | 99.45% |
| LIC_12723 | 363 | 99.45% |
| LIC_12546 | 364 | 99.73% |
| LIC_14002 | 365 | 100% |
| LIC_20173 | 362 | 99.18% |
| LIC_20198 | 364 | 99.73% |
| LIC_20182 | 363 | 99.45% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Ferrari, A.S.d.A.; Merighi, D.G.S.; Visnardi, A.B.; Silva, G.R.; Gonçalves, C.A.B.; Sanchez-Limache, D.E.; Ogusku, B.S.C.; Souza, A.S.d.; de Souza, R.F.; Guzzo, C.R. Discovery of Novel c-di-GMP-Related Genes in Leptospira interrogans. Pathogens 2026, 15, 151. https://doi.org/10.3390/pathogens15020151
Ferrari ASdA, Merighi DGS, Visnardi AB, Silva GR, Gonçalves CAB, Sanchez-Limache DE, Ogusku BSC, Souza ASd, de Souza RF, Guzzo CR. Discovery of Novel c-di-GMP-Related Genes in Leptospira interrogans. Pathogens. 2026; 15(2):151. https://doi.org/10.3390/pathogens15020151
Chicago/Turabian StyleFerrari, Anielle Salviano de Almeida, Davi Gabriel Salustiano Merighi, Aline Biazola Visnardi, Gabriela Roberto Silva, Cauê Augusto Boneto Gonçalves, Daniel Enrique Sanchez-Limache, Bruna Sayuri Cardoso Ogusku, Anacleto Silva de Souza, Robson Francisco de Souza, and Cristiane Rodrigues Guzzo. 2026. "Discovery of Novel c-di-GMP-Related Genes in Leptospira interrogans" Pathogens 15, no. 2: 151. https://doi.org/10.3390/pathogens15020151
APA StyleFerrari, A. S. d. A., Merighi, D. G. S., Visnardi, A. B., Silva, G. R., Gonçalves, C. A. B., Sanchez-Limache, D. E., Ogusku, B. S. C., Souza, A. S. d., de Souza, R. F., & Guzzo, C. R. (2026). Discovery of Novel c-di-GMP-Related Genes in Leptospira interrogans. Pathogens, 15(2), 151. https://doi.org/10.3390/pathogens15020151








