vIRA Inhibition of Antiviral Necroptosis and RIPK3 Binding Are Separable Events
Abstract
1. Introduction
2. Materials and Methods
2.1. Cells and Reagents
2.2. Plasmids, Transfections, Immunoblotting, Antibodies, and Immunoprecipitation
2.3. Immunofluorescence
2.4. Generation of RHIM-Swap MCMVs
2.5. Viruses and Infections
2.6. Cell Viability Assays
2.7. FACS Assay
3. Results
3.1. 1-277 M45 RHIM-Swap Constructs Bind RIPK3 but Do Not Protect from TNFα-Induced Necroptosis
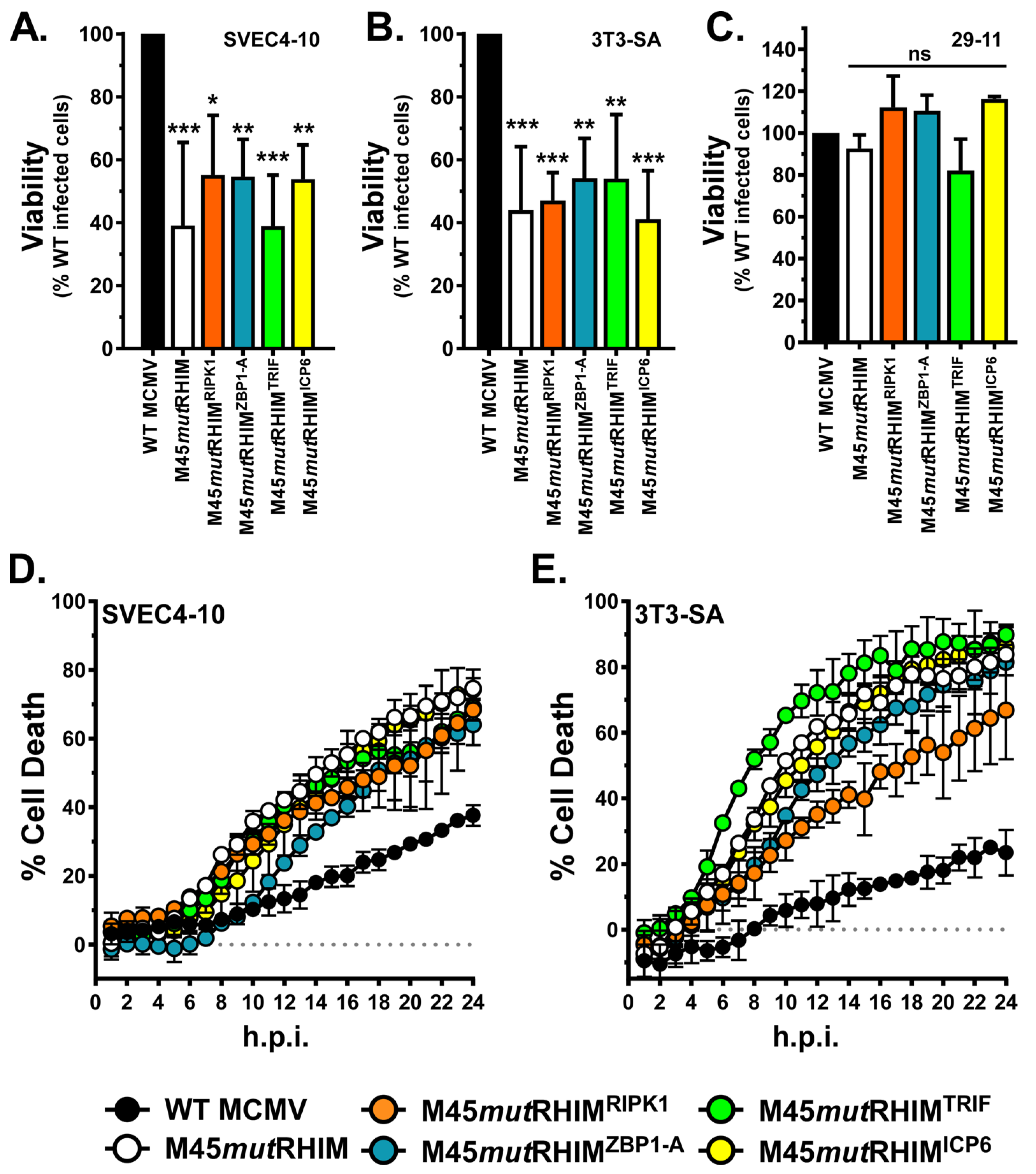
3.2. M45 RHIM-Swap Recombinant MCMV Replication Is Restricted in Cell Lines Permissive to Necroptosis
3.3. M45 RHIM-Swap MCMVs Do Not Inhibit Virus-Induced ZBP1-Dependent Necroptosis
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sridharan, H.; Upton, J.W. Programmed Necrosis in Microbial Pathogenesis. Trends Microbiol. 2014, 22, 199–207. [Google Scholar] [CrossRef]
- Zhou, W.; Yuan, J. Necroptosis in Health and Diseases. Semin. Cell Dev. Biol. 2014, 35, 14–23. [Google Scholar] [CrossRef]
- Mocarski, E.S.; Guo, H.; Kaiser, W.J. Necroptosis: The Trojan Horse in Cell Autonomous Antiviral Host Defense. Virology 2015, 479–480, 160–166. [Google Scholar] [CrossRef]
- Cho, Y.S.; Challa, S.; Moquin, D.; Genga, R.; Ray, T.D.; Guildford, M.; Chan, F.K.-M. Phosphorylation-Driven Assembly of the RIP1-RIP3 Complex Regulates Programmed Necrosis and Virus-Induced Inflammation. Cell 2009, 137, 1112–1123. [Google Scholar] [CrossRef]
- Sun, X.; Lee, J.; Navas, T.; Baldwin, D.T.; Stewart, T.A.; Dixit, V.M. RIP3, a Novel Apoptosis-Inducing Kinase. J. Biol. Chem. 1999, 274, 16871–16875. [Google Scholar] [CrossRef]
- Cai, Z.; Liu, Z.-G. Execution of RIPK3-Regulated Necrosis. Mol. Cell Oncol. 2014, 1, e960759. [Google Scholar] [CrossRef]
- Cai, Z.; Jitkaew, S.; Zhao, J.; Chiang, H.-C.; Choksi, S.; Liu, J.; Ward, Y.; Wu, L.-G.; Liu, Z.-G. Plasma Membrane Translocation of Trimerized MLKL Protein Is Required for TNF-Induced Necroptosis. Nat. Cell Biol. 2014, 16, 55–65. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Wang, H.; Wang, Z.; He, S.; Chen, S.; Liao, D.; Wang, L.; Yan, J.; Liu, W.; Lei, X.; et al. Mixed Lineage Kinase Domain-like Protein Mediates Necrosis Signaling Downstream of RIP3 Kinase. Cell 2012, 148, 213–227. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Jitkaew, S.; Cai, Z.; Choksi, S.; Li, Q.; Luo, J.; Liu, Z.-G. Mixed Lineage Kinase Domain-like Is a Key Receptor Interacting Protein 3 Downstream Component of TNF-Induced Necrosis. Proc. Natl. Acad. Sci. USA 2012, 109, 5322–5327. [Google Scholar] [CrossRef]
- Chen, X.; Li, W.; Ren, J.; Huang, D.; He, W.-T.; Song, Y.; Yang, C.; Li, W.; Zheng, X.; Chen, P.; et al. Translocation of Mixed Lineage Kinase Domain-like Protein to Plasma Membrane Leads to Necrotic Cell Death. Cell Res. 2014, 24, 105–121. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Huang, Z.; Ren, J.; Zhang, Z.; He, P.; Li, Y.; Ma, J.; Chen, W.; Zhang, Y.; Zhou, X.; et al. Mlkl Knockout Mice Demonstrate the Indispensable Role of Mlkl in Necroptosis. Cell Res. 2013, 23, 994–1006. [Google Scholar] [CrossRef]
- Wang, H.; Sun, L.; Su, L.; Rizo, J.; Liu, L.; Wang, L.-F.; Wang, F.-S.; Wang, X. Mixed Lineage Kinase Domain-like Protein MLKL Causes Necrotic Membrane Disruption upon Phosphorylation by RIP3. Mol. Cell 2014, 54, 133–146. [Google Scholar] [CrossRef]
- Kaiser, W.J.; Offermann, M.K. Apoptosis Induced by the Toll-like Receptor Adaptor TRIF Is Dependent on Its Receptor Interacting Protein Homotypic Interaction Motif. J. Immunol. 2005, 174, 4942–4952. [Google Scholar] [CrossRef]
- Kaiser, W.J.; Sridharan, H.; Huang, C.; Mandal, P.; Upton, J.W.; Gough, P.J.; Sehon, C.A.; Marquis, R.W.; Bertin, J.; Mocarski, E.S. Toll-like Receptor 3-Mediated Necrosis via TRIF, RIP3, and MLKL. J. Biol. Chem. 2013, 288, 31268–31279. [Google Scholar] [CrossRef] [PubMed]
- Upton, J.W.; Shubina, M.; Balachandran, S. RIPK3-Driven Cell Death during Virus Infections. Immunol. Rev. 2017, 277, 90–101. [Google Scholar] [CrossRef] [PubMed]
- Steain, M.; Baker, M.O.D.G.; Pham, C.L.L.; Shanmugam, N.; Gambin, Y.; Sierecki, E.; McSharry, B.P.; Avdic, S.; Slobedman, B.; Sunde, M.; et al. Varicella Zoster Virus Encodes a Viral Decoy RHIM to Inhibit Cell Death. PLoS Pathog. 2020, 16, e1008473. [Google Scholar] [CrossRef] [PubMed]
- Kaiser, W.J.; Upton, J.W.; Mocarski, E.S. Receptor-Interacting Protein Homotypic Interaction Motif-Dependent Control of NF-Kappa B Activation via the DNA-Dependent Activator of IFN Regulatory Factors. J. Immunol. 2008, 181, 6427–6434. [Google Scholar] [CrossRef]
- Rebsamen, M.; Heinz, L.X.; Meylan, E.; Michallet, M.-C.; Schroder, K.; Hofmann, K.; Vazquez, J.; Benedict, C.A.; Tschopp, J. DAI/ZBP1 Recruits RIP1 and RIP3 through RIP Homotypic Interaction Motifs to Activate NF-kappaB. EMBO Rep. 2009, 10, 916–922. [Google Scholar] [CrossRef]
- Upton, J.W.; Kaiser, W.J.; Mocarski, E.S. Cytomegalovirus M45 Cell Death Suppression Requires Receptor-Interacting Protein (RIP) Homotypic Interaction Motif (RHIM)-Dependent Interaction with RIP1. J. Biol. Chem. 2008, 283, 16966–16970. [Google Scholar] [CrossRef]
- Guo, H.; Kaiser, W.J.; Mocarski, E.S. Manipulation of Apoptosis and Necroptosis Signaling by Herpesviruses. Med. Microbiol. Immunol. 2015, 204, 439–448. [Google Scholar] [CrossRef]
- Guo, H.; Gilley, R.P.; Fisher, A.; Lane, R.; Landsteiner, V.J.; Ragan, K.B.; Dovey, C.M.; Carette, J.E.; Upton, J.W.; Mocarski, E.S.; et al. Species-Independent Contribution of ZBP1/DAI/DLM-1-Triggered Necroptosis in Host Defense against HSV1. Cell Death Dis. 2018, 9, 816. [Google Scholar] [CrossRef]
- Brune, W.; Ménard, C.; Heesemann, J.; Koszinowski, U.H. A Ribonucleotide Reductase Homolog of Cytomegalovirus and Endothelial Cell Tropism. Science 2001, 291, 303–305. [Google Scholar] [CrossRef]
- Lembo, D.; Brune, W. Tinkering with a Viral Ribonucleotide Reductase. Trends Biochem. Sci. 2009, 34, 25–32. [Google Scholar] [CrossRef]
- Wang, X.; Li, Y.; Liu, S.; Yu, X.; Li, L.; Shi, C.; He, W.; Li, J.; Xu, L.; Hu, Z.; et al. Direct Activation of RIP3/MLKL-Dependent Necrosis by Herpes Simplex Virus 1 (HSV-1) Protein ICP6 Triggers Host Antiviral Defense. Proc. Natl. Acad. Sci. USA 2014, 111, 15438–15443. [Google Scholar] [CrossRef]
- Huang, Z.; Wu, S.-Q.; Liang, Y.; Zhou, X.; Chen, W.; Li, L.; Wu, J.; Zhuang, Q.; Chen, C.; Li, J.; et al. RIP1/RIP3 Binding to HSV-1 ICP6 Initiates Necroptosis to Restrict Virus Propagation in Mice. Cell Host Microbe 2015, 17, 229–242. [Google Scholar] [CrossRef]
- Sun, X.; Yin, J.; Starovasnik, M.A.; Fairbrother, W.J.; Dixit, V.M. Identification of a Novel Homotypic Interaction Motif Required for the Phosphorylation of Receptor-Interacting Protein (RIP) by RIP3. J. Biol. Chem. 2002, 277, 9505–9511. [Google Scholar] [CrossRef] [PubMed]
- Mompeán, M.; Li, W.; Li, J.; Laage, S.; Siemer, A.B.; Bozkurt, G.; Wu, H.; McDermott, A.E. The Structure of the Necrosome RIPK1-RIPK3 Core, a Human Hetero-Amyloid Signaling Complex. Cell 2018, 173, 1244–1253.e10. [Google Scholar] [CrossRef] [PubMed]
- Pham, C.L.; Shanmugam, N.; Strange, M.; O’Carroll, A.; Brown, J.W.; Sierecki, E.; Gambin, Y.; Steain, M.; Sunde, M. Viral M45 and Necroptosis-Associated Proteins Form Heteromeric Amyloid Assemblies. EMBO Rep. 2019, 20, e46518. [Google Scholar] [CrossRef]
- Hu, H.; Wu, X.; Wu, G.; Nan, N.; Zhang, J.; Zhu, X.; Zhang, Y.; Shu, Z.; Liu, J.; Liu, X.; et al. RIP3-Mediated Necroptosis Is Regulated by Inter-Filament Assembly of RIP Homotypic Interaction Motif. Cell Death Differ. 2021, 28, 251–266. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Ma, Y.; Zhao, K.; Zhang, J.; Sun, Y.; Li, Y.; Dong, X.; Hu, H.; Liu, J.; Wang, J.; et al. The Structure of a Minimum Amyloid Fibril Core Formed by Necroptosis-Mediating RHIM of Human RIPK3. Proc. Natl. Acad. Sci. USA 2021, 118, e2022933118. [Google Scholar] [CrossRef]
- Wu, X.-L.; Hu, H.; Dong, X.-Q.; Zhang, J.; Wang, J.; Schwieters, C.D.; Liu, J.; Wu, G.-X.; Li, B.; Lin, J.-Y.; et al. The Amyloid Structure of Mouse RIPK3 (Receptor Interacting Protein Kinase 3) in Cell Necroptosis. Nat. Commun. 2021, 12, 1627. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Wu, X.-L.; Zhang, J.; Li, B.; Wang, H.-Y.; Wang, J.; Lu, J.-X. The Structure of Mouse RIPK1 RHIM-Containing Domain as a Homo-Amyloid and in RIPK1/RIPK3 Complex. Nat. Commun. 2024, 15, 6975. [Google Scholar] [CrossRef] [PubMed]
- Upton, J.W.; Kaiser, W.J.; Mocarski, E.S. Virus Inhibition of RIP3-Dependent Necrosis. Cell Host Microbe 2010, 7, 302–313. [Google Scholar] [CrossRef]
- Sridharan, H.; Ragan, K.B.; Guo, H.; Gilley, R.P.; Landsteiner, V.J.; Kaiser, W.J.; Upton, J.W. Murine Cytomegalovirus IE3-Dependent Transcription Is Required for DAI/ZBP1-Mediated Necroptosis. EMBO Rep. 2017, 18, 1429–1441. [Google Scholar] [CrossRef]
- Tandon, R.; Mocarski, E.S. Control of Cytoplasmic Maturation Events by Cytomegalovirus Tegument Protein Pp150. J. Virol. 2008, 82, 9433–9444. [Google Scholar] [CrossRef]
- Redwood, A.J.; Messerle, M.; Harvey, N.L.; Hardy, C.M.; Koszinowski, U.H.; Lawson, M.A.; Shellam, G.R. Use of a Murine Cytomegalovirus K181-Derived Bacterial Artificial Chromosome as a Vaccine Vector for Immunocontraception. J. Virol. 2005, 79, 2998–3008. [Google Scholar] [CrossRef]
- Lembo, D.; Donalisio, M.; Hofer, A.; Cornaglia, M.; Brune, W.; Koszinowski, U.; Thelander, L.; Landolfo, S. The Ribonucleotide Reductase R1 Homolog of Murine Cytomegalovirus Is Not a Functional Enzyme Subunit but Is Required for Pathogenesis. J. Virol. 2004, 78, 4278–4288. [Google Scholar] [CrossRef]
- Welsby, I.; Hutin, D.; Gueydan, C.; Kruys, V.; Rongvaux, A.; Leo, O. PARP12, an Interferon-Stimulated Gene Involved in the Control of Protein Translation and Inflammation. J. Biol. Chem. 2014, 289, 26642–26657. [Google Scholar] [CrossRef]
- Upton, J.W.; Kaiser, W.J.; Mocarski, E.S. DAI/ZBP1/DLM-1 Complexes with RIP3 to Mediate Virus-Induced Programmed Necrosis That Is Targeted by Murine Cytomegalovirus vIRA. Cell Host Microbe 2012, 11, 290–297. [Google Scholar] [CrossRef] [PubMed]



| Primer Name | Sequence | |
|---|---|---|
| pEGFP/M45 | HS38 | TAGGGCACGCTGTGCCCCCCACCCACCCCGA |
| pEGFP/M45 | HS39 | AGCATCGCCTCTTGCTGGAGTCCCTCCTACACTGACCGA |
| RIPK1 RHIM | HS40 | GGTGGGGGGCACAGCGTGCCCTACGTCAGGACTATATTCAATAGTTCTGGTATTCAGATTGGAAACCACAATTATATG |
| RIPK1 RHIM | HS41 | AGGAGGGACTCCAGCAAGAGGCGATGCTCATATAATTGTGGTTTCCAATCTGAATACCAGAACTATTGAATATAGT |
| RIPK3 RHIM | HS42 | GGTGGGGGGCACAGCGTGCCCTACGTCAGGGTCTTCAACAACTGTTCGAAGTGCAGATTGGGAACTACAACTCCTTG |
| RIPK3 RHIM | HS43 | AGGAGGGACTCCAGCAAGAGGCGATGCTCAAGGAGTTGTAGTTCCCAATCTGCACTTCAGAACAGTTGTTGAAGAC |
| TRIF RHIM | HS44 | GGTGGGGGGCACAGCGTGCCCTACGTCAGGATTATTCACCATGCCCAGATGGTTCAGCTGGGTGTCAACAATCACATG |
| TRIF RHIM | HS45 | AGGAGGGACTCCAGCAAGAGGCGATGCTCATGTGATTGTTGACACCCAGCTGAACCATCTGGGCATGGTGAATAAT |
| ZBP1-A RHIM | HS46 | GGTGGGGGGCACAGCGTGCCCTACGTCAGGTCCATTGCCAATTCAAACGCCATCCAGATTGGTCACGGGAATGTCATA |
| ZBP1-A RHIM | HS47 | AGGAGGGACTCCAGCAAGAGGCGATGCTTATGACATTCCCGTGACCAATCTGGATGGCGTTTGAATTGGCAATGGA |
| PARP12 RHIM | HS48 | GGTGGGGGGCACAGCGTGCCCTACGTCAGGGCCCAGGCAGCCACCTTGAAGTTCCAGGCTGGAAAACACAACTATGAG |
| PARP12 RHIM | HS49 | AGGAGGGACTCCAGCAAGAGGCGATGCTCTCATAGTTGTGTTTTCCAGCCTGGAACTTCAAGGTGGCTGCCTGGGC |
| ICP6 RHIM | HS50 | GGTGGGGGGCACAGCGTGCCCTACGTCAGGCGCATCAGCGATAGCAACTTTGTCCAATGTGGTTCCAACTGCACCATG |
| ICP6 RHIM | HS51 | AGGAGGGACTCCAGCAAGAGGCGATGCTCATGGTGCAGTTGGAACCACATTGGACAAAGTTGCTATCGCTGATGCG |
| ZBP1-B RHIM | HS66 | GGTGGGGGGCACAGCGTGCCCTACGTCAGGGACAAGTCCTTGCTCCAACAAGTGCAGCTTGGCCACCACAACGAGATG |
| ZBP1-B RHIM | HS67 | AGGAGGGACTCCAGCAAGAGGCGATGCTCATCTCGTTGTGGTGGCCAAGCTGCACTTGTTGGAGCAAGGACTTGTC |
| Primer Name | Sequence | |
|---|---|---|
| Sequencing M45 RHIM locus | HS36 | CCCAAAGTGTACTCCGACCC |
| Sequencing M45 RHIM locus | HS37 | GCTTCTTGGCTTGAGGTGC |
| RHIM amplification | ASP01C | CAGCTCTCCGTGGTT |
| RHIM amplification | ASP01B | AGCGGCAGGTGAGGA |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Ragan, K.B.; Sridharan, H.; Stark, A.S.; Ilami, K.; Fisher, A.D.; Brahms, O.N.; Kaiser, W.J.; Upton, J.W. vIRA Inhibition of Antiviral Necroptosis and RIPK3 Binding Are Separable Events. Pathogens 2026, 15, 79. https://doi.org/10.3390/pathogens15010079
Ragan KB, Sridharan H, Stark AS, Ilami K, Fisher AD, Brahms ON, Kaiser WJ, Upton JW. vIRA Inhibition of Antiviral Necroptosis and RIPK3 Binding Are Separable Events. Pathogens. 2026; 15(1):79. https://doi.org/10.3390/pathogens15010079
Chicago/Turabian StyleRagan, Katherine B., Haripriya Sridharan, Aaron S. Stark, Kaela Ilami, Amanda D. Fisher, Olivia N. Brahms, William J. Kaiser, and Jason W. Upton. 2026. "vIRA Inhibition of Antiviral Necroptosis and RIPK3 Binding Are Separable Events" Pathogens 15, no. 1: 79. https://doi.org/10.3390/pathogens15010079
APA StyleRagan, K. B., Sridharan, H., Stark, A. S., Ilami, K., Fisher, A. D., Brahms, O. N., Kaiser, W. J., & Upton, J. W. (2026). vIRA Inhibition of Antiviral Necroptosis and RIPK3 Binding Are Separable Events. Pathogens, 15(1), 79. https://doi.org/10.3390/pathogens15010079






