High Prevalence of Clostridioides difficile Ribotype 176 in the University Hospital in Kosice
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design and Patients
2.2. Cultivation and Identification
2.3. Isolation of Genomic DNA
2.4. Detection of Toxin Genes
2.5. PCR Ribotyping
2.6. Antimicrobial Susceptibility Testing
2.7. Statistical Analysis
3. Results
3.1. Characteristics of Patients and C.difficile Isolates
3.2. Toxins of C. difficile Isolates
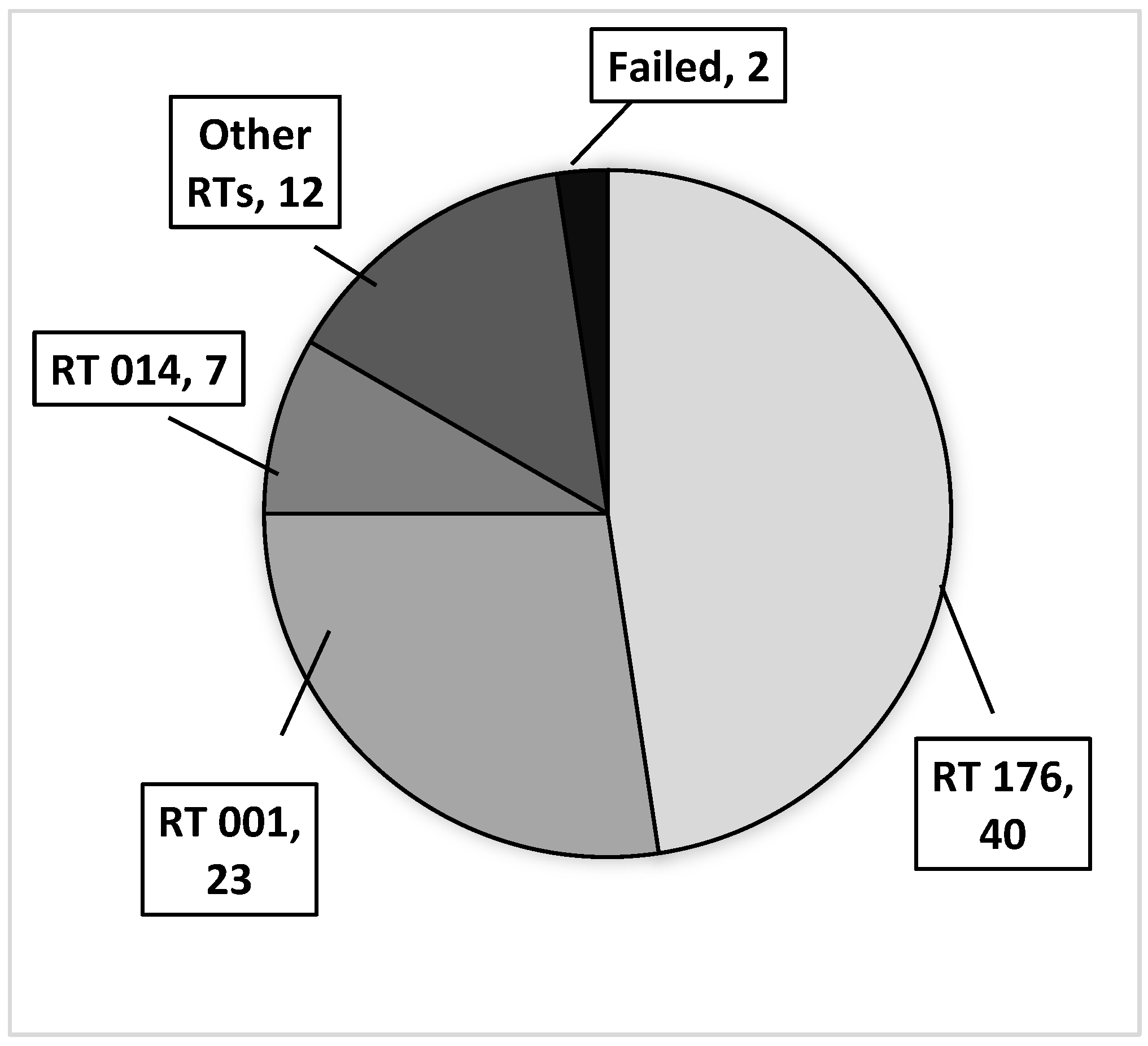
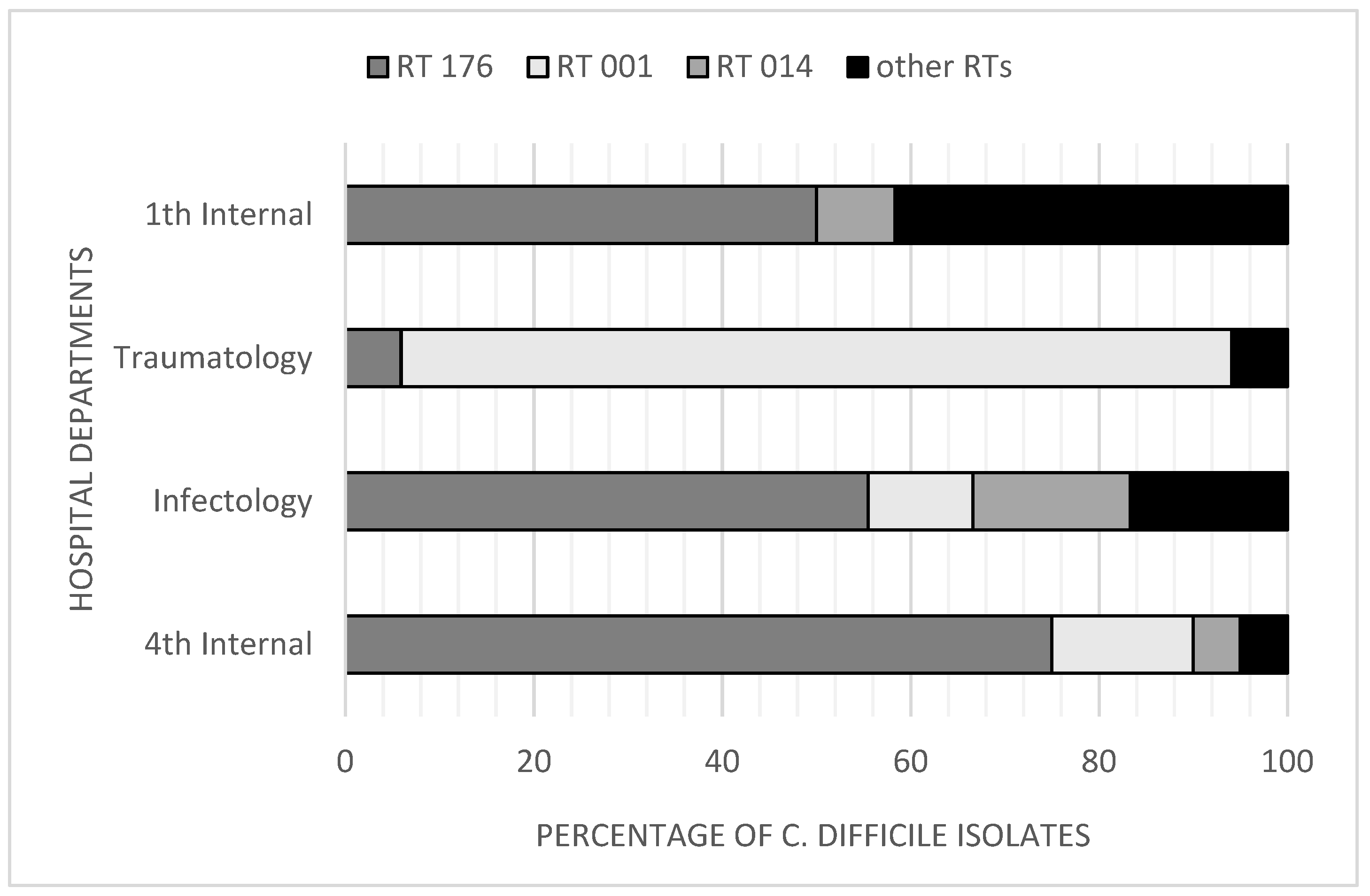
3.3. Ribotypes of C. difficile Isolates
3.4. Antimicrobial Resistance of C. difficile Isolates
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Fan, Y.; Pedersen, O. Gut microbiota in human metabolic health and disease. Nat. Rev. Microbiol. 2021, 19, 55–71. [Google Scholar] [CrossRef] [PubMed]
- Thaiss, C.A.; Itav, S.; Rothschild, D.; Meijer, M.T.; Levy, M.; Moresi, C.; Dohnalová, L.; Braverman, S.; Rozin, S.; Malitsky, S.; et al. Persistent microbiome alterations modulate the rate of post-dieting weight regain. Nature 2016, 540, 544–551. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Zhou, J.; Wang, L. Role and Mechanism of Gut Microbiota in Human Disease. Front. Cell. Infect. Microbiol. 2021, 11, 625913. [Google Scholar] [CrossRef] [PubMed]
- Natividad, J.M.M.; Verdu, E.F. Modulation of intestinal barrier by intestinal microbiota: Pathological and therapeutic implications. Pharmacol. Res. 2013, 69, 42–51. [Google Scholar] [CrossRef] [PubMed]
- Den Besten, G.; van Eunen, K.; Groen, A.K.; Venema, K.; Reijngoud, D.-J.; Bakker, B.M. The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J. Lipid Res. 2013, 54, 2325–2340. [Google Scholar] [CrossRef] [PubMed]
- Bäumler, A.J.; Sperandio, V. Interactions between the microbiota and pathogenic bacteria in the gut. Nature 2016, 535, 85–93. [Google Scholar] [CrossRef]
- Gensollen, T.; Iyer, S.S.; Kasper, D.L.; Blumberg, R.S. How colonization by microbiota in early life shapes the immune system. Science 2016, 352, 539–544. [Google Scholar] [CrossRef] [PubMed]
- Kho, Z.Y.; Lal, S.K. The Human Gut Microbiome—A Potential Controller of Wellness and Disease. Front. Microbiol. 2018, 9, 1835. [Google Scholar] [CrossRef] [PubMed]
- Carding, S.; Verbeke, K.; Vipond, D.T.; Corfe, B.M.; Owen, L.J. Dysbiosis of the gut microbiota in disease. Microb. Ecol. Health Dis. 2015, 26, 26191. [Google Scholar] [CrossRef]
- Czepiel, J.; Dróżdż, M.; Pituch, H.; Kuijper, E.J.; Perucki, W.; Mielimonka, A.; Goldman, S.; Wultańska, D.; Garlicki, A.; Biesiada, G. Clostridium difficile infection: Review. Eur. J. Clin. Microbiol. Infect. Dis. 2019, 38, 1211–1221. [Google Scholar] [CrossRef] [PubMed]
- Aktories, K.; Papatheodorou, P.; Schwan, C. Binary Clostridium difficile toxin (CDT) - A virulence factor disturbing the cytoskeleton. Anaerobe 2018, 53, 21–29. [Google Scholar] [CrossRef]
- Smits, W.K.; Lyras, D.; Lacy, D.B. Clostridium difficile infection. Nat. Rev. Dis. Primer. 2016, 2, 16020. [Google Scholar] [CrossRef] [PubMed]
- Sorg, J.A.; Sonenshein, A.L. Bile Salts and Glycine as Cogerminants for Clostridium difficile Spores. J. Bacteriol. 2008, 190, 2505–2512. [Google Scholar] [CrossRef] [PubMed]
- Theriot, C.M.; Koenigsknecht, M.J.; Carlson, P.E., Jr.; Hatton, G.E.; Nelson, A.M.; Li, B.; Huffnagle, G.B.; Li, J.Z.; Young, V.B. Antibiotic-induced shifts in the mouse gut microbiome and metabolome increase susceptibility to Clostridium difficile infection. Nat. Commun. 2014, 5, 3114. [Google Scholar] [CrossRef]
- Elliott, B.; Androga, G.O.; Knight, D.R.; Riley, T.V. Clostridium difficile infection: Evolution, phylogeny and molecular epidemiology. Infect. Genet. Evol. 2017, 49, 1–11. [Google Scholar] [CrossRef]
- A Davies, K.; Ashwin, H.; Longshaw, C.M.; A Burns, D.; Davis, G.L.; Wilcox, M.H.; on behalf of the EUCLID study group. Diversity of Clostridium difficile PCR ribotypes in Europe: Results from the European, multicentre, prospective, biannual, point-prevalence study of Clostridium difficile infection in hospitalised patients with diarrhoea (EUCLID), 2012 and 2013. Eurosurveillance 2016, 21, 30294. [Google Scholar] [CrossRef]
- Couturier, J.; Davies, K.; Gateau, C.; Barbut, F. Ribotypes and New Virulent Strains Across Europe. Adv. Exp. Med. Biol. 2018, 1050, 45–58. [Google Scholar] [CrossRef] [PubMed]
- Persson, S.; Torpdahl, M.; Olsen, K. New multiplex PCR method for the detection of Clostridium difficile toxin A (tcdA) and toxin B (tcdB) and the binary toxin (cdtA/cdtB) genes applied to a Danish strain collection. Clin. Microbiol. Infect. 2008, 14, 1057–1064. [Google Scholar] [CrossRef]
- Fawley, W.N.; Knetsch, W.; MacCannell, D.R.; Harmanus, C.; Du, T.; Mulvey, M.R.; Paulick, A.; Anderson, L.; Kuijper, E.; Wilcox, M.H. Development and Validation of an Internationally-Standardized, High-Resolution Capillary Gel-Based Electrophoresis PCR-Ribotyping Protocol for Clostridium difficile. PLos ONE 2015, 10, e0118150. [Google Scholar] [CrossRef] [PubMed]
- Bidet, P.; Lalande, V.; Salauze, B.; Burghoffer, B.; Avesani, V.; Delmée, M.; Rossier, A.; Barbut, F.; Petit, J.-C. Comparison of PCR-Ribotyping, Arbitrarily Primed PCR, and Pulsed-Field Gel Electrophoresis for Typing Clostridium difficile. J. Clin. Microbiol. 2000, 38, 2484–2487. [Google Scholar] [CrossRef]
- WEBRIBO. Available online: https://webribo.ages.at/ (accessed on 27 June 2022).
- Jarmo, O.; Veli-Jukka, A.; Eero, M. Treatment of Clostridioides (Clostridium) difficile infection. Ann. Med. 2019, 52, 12–20. [Google Scholar] [CrossRef] [PubMed]
- van Rossen, T.M.; Ooijevaar, R.E.; Vandenbroucke-Grauls, C.M.; Dekkers, O.M.; Kuijper, E.J.; Keller, J.J.; van Prehn, J. Prognostic factors for severe and recurrent Clostridioides difficile infection: A systematic review. Clin. Microbiol. Infect. 2021, 28, 321–331. [Google Scholar] [CrossRef] [PubMed]
- López-De-Andrés, A.; Esteban-Vasallo, M.D.; de Miguel-Díez, J.; Hernández-Barrera, V.; de Miguel-Yanes, J.M.; Méndez-Bailón, M.; Jiménez-García, R. Incidence and in-hospital outcomes of Clostridium difficile infection among type 2 diabetes patients in Spain. Int. J. Clin. Pr. 2018, 72, e13251. [Google Scholar] [CrossRef] [PubMed]
- Maslennikov, R.; Ivashkin, V.; Ufimtseva, A.; Poluektova, E.; Ulyanin, A. Clostridioides difficile co-infection in patients with COVID-19. Futur. Microbiol. 2022, 17, 653–663. [Google Scholar] [CrossRef]
- Mutai, W.C.; Mureithi, M.; Anzala, O.; Kullin, B.; Ofwete, R.; Kyany’a, C.; Odoyo, E.; Musila, L.; Revathi, G. Assessment of independent comorbidities and comorbidity measures in predicting healthcare facility-onset Clostridioides difficile infection in Kenya. PLoS Glob. Public Health 2022, 2, e0000090. [Google Scholar] [CrossRef]
- Spigaglia, P. Recent advances in the understanding of antibiotic resistance in Clostridium difficile infection. Ther. Adv. Infect. Dis. 2015, 3, 23–42. [Google Scholar] [CrossRef] [PubMed]
- Binyamin, D.; Nitzan, O.; Azrad, M.; Hamo, Z.; Koren, O.; Peretz, A. The microbial diversity following antibiotic treatment of Clostridioides difficile infection. BMC Gastroenterol. 2021, 21, 166. [Google Scholar] [CrossRef] [PubMed]
- Czepiel, J.; Krutova, M.; Mizrahi, A.; Khanafer, N.; Enoch, D.; Patyi, M.; Deptuła, A.; Agodi, A.; Nuvials, X.; Pituch, H.; et al. Mortality Following Clostridioides difficile Infection in Europe: A Retrospective Multicenter Case-Control Study. Antibiotics 2021, 10, 299. [Google Scholar] [CrossRef]
- Kim, B.; Kim, J.; Pai, H. Association between Antibiotic Consumption and Incidence of Clostridioides difficile Infection in a Hospital. J. Korean Med Sci. 2020, 35, e407. [Google Scholar] [CrossRef] [PubMed]
- Chalmers, J.D.; Akram, A.R.; Singanayagam, A.; Wilcox, M.H.; Hill, A.T. Risk factors for Clostridium difficile infection in hospitalized patients with community-acquired pneumonia. J. Infect. 2016, 73, 45–53. [Google Scholar] [CrossRef]
- Zycinska, K.; Chmielewska, M.; Lenartowicz, B.; Hadzik-Blaszczyk, M.; Cieplak, M.; Kur, Z.; Krupa, R.; Wardyn, K.A. Antibiotic Treatment of Hospitalized Patients with Pneumonia Complicated by Clostridium difficile Infection. Adv. Exp. Med. Biol. 2016, 952, 59–64. [Google Scholar] [CrossRef]
- Perić, A.; Rančić, N.; Dragojević-Simić, V.; Milenković, B.; Ljubenović, N.; Rakonjac, B.; Begović-Kuprešanin, V.; Šuljagić, V. Association between Antibiotic Use and Hospital-Onset Clostridioides difficile Infection in University Tertiary Hospital in Serbia, 2011–2021: An Ecological Analysis. Antibiotics 2022, 11, 1178. [Google Scholar] [CrossRef] [PubMed]
- Appaneal, H.J.; Caffrey, A.R.; Laplante, K.L. What Is the Role for Metronidazole in the Treatment of Clostridium difficile Infection? Results From a National Cohort Study of Veterans with Initial Mild Disease. Clin. Infect. Dis. 2018, 69, 1288–1295. [Google Scholar] [CrossRef]
- van Prehn, J.; Reigadas, E.; Vogelzang, E.H.; Bouza, E.; Hristea, A.; Guery, B.; Krutova, M.; Norén, T.; Allerberger, F.; Coia, J.E.; et al. European Society of Clinical Microbiology and Infectious Diseases: 2021 update on the treatment guidance document for Clostridioides difficile infection in adults. Clin. Microbiol. Infect. 2021, 27 (Suppl. S2), S1–S21. [Google Scholar] [CrossRef] [PubMed]
- Juul, F.E.; Garborg, K.; Bretthauer, M.; Skudal, H.; Øines, M.N.; Wiig, H.; Rose, Ø.; Seip, B.; Lamont, J.T.; Midtvedt, T.; et al. Fecal Microbiota Transplantation for Primary Clostridium difficile Infection. N. Engl. J. Med. 2018, 378, 2535–2536. [Google Scholar] [CrossRef]
- Haifer, C.; Paramsothy, S.; Borody, T.J.; Clancy, A.; Leong, R.W.; Kaakoush, N.O. Long-Term Bacterial and Fungal Dynamics following Oral Lyophilized Fecal Microbiota Transplantation in Clostridioides difficile Infection. Msystems 2021, 6, e00905-20. [Google Scholar] [CrossRef] [PubMed]
- Smith, A. Outbreak of Clostridium difficile infection in an English hospital linked to hypertoxin-producing strains in Canada and the US. Eurosurveillance 2005, 10, E050630.2. [Google Scholar] [CrossRef] [PubMed]
- Krutova, M.; Nyc, O.; Matejkova, J.; Allerberger, F.; Wilcox, M.; Kuijper, E. Molecular characterisation of Czech Clostridium difficile isolates collected in 2013–2015. Int. J. Med. Microbiol. 2016, 306, 479–485. [Google Scholar] [CrossRef] [PubMed]
- Sundram, F.; Guyot, A.; Carboo, I.; Green, S.; Lilaonitkul, M.; Scourfield, A. Clostridium difficile ribotypes 027 and 106: Clinical outcomes and risk factors. J. Hosp. Infect. 2009, 72, 111–118. [Google Scholar] [CrossRef]
- Krehelova, M.; Nyč, O.; Sinajová, E.; Krutova, M. The predominance and clustering of Clostridioides (Clostridium) difficile PCR ribotype 001 isolates in three hospitals in Eastern Slovakia, 2017. Folia Microbiol. 2018, 64, 49–54. [Google Scholar] [CrossRef] [PubMed]
- Novakova, E.; Kotlebova, N.; Gryndlerova, A.; Novak, M.; Vladarova, M.; Wilcox, M.; Kuijper, E.; Krutova, M. An Outbreak of Clostridium (Clostridioides) difficile Infections within an Acute and Long-Term Care Wards due to Moxifloxacin-Resistant PCR Ribotype 176 Genotyped as PCR Ribotype 027 by a Commercial Assay. J. Clin. Med. 2020, 9, 3738. [Google Scholar] [CrossRef] [PubMed]
- Novakova, E.; Stefkovicova, M.; Kopilec, M.G.; Novak, M.; Kotlebova, N.; Kuijper, E.; Krutova, M. The emergence of Clostridium difficile ribotypes 027 and 176 with a predominance of the Clostridium difficile ribotype 001 recognized in Slovakia following the European standardized Clostridium difficile infection surveillance of 2016. Int. J. Infect. Dis. 2020, 90, 111–115. [Google Scholar] [CrossRef]
- Krutova, M.; Capek, V.; Nycova, E.; Vojackova, S.; Balejova, M.; Geigerova, L.; Tejkalova, R.; Havlinova, L.; Vagnerova, I.; Cermak, P.; et al. The association of a reduced susceptibility to moxifloxacin in causative Clostridium (Clostridioides) difficile strain with the clinical outcome of patients. Antimicrob. Resist. Infect. Control. 2020, 9, 98. [Google Scholar] [CrossRef] [PubMed]
- Beran, V.; Kuijper, E.J.; Harmanus, C.; Sanders, I.M.; van Dorp, S.M.; Knetsch, C.W.; Janeckova, J.; Seidelova, A.; Barekova, L.; Tvrdik, J.; et al. Molecular typing and antimicrobial susceptibility testing to six antimicrobials of Clostridium difficile isolates from three Czech hospitals in Eastern Bohemia in 2011–2012. Folia Microbiol. 2017, 62, 445–451. [Google Scholar] [CrossRef] [PubMed]
- Kracík, M.; Dolinová, I.; Žemličková, H. Ribotyping of Clostridioides difficile in the Liberec Regional Hospital: A tertiary health care facility. Folia Microbiol. 2022, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Tóth, J.; Urbán, E.; Osztie, H.; Benczik, M.; Indra, A.; Nagy, E.; Allerberger, F. Distribution of PCR ribotypes among recent Clostridium difficile isolates collected in two districts of Hungary using capillary gel electrophoresis and review of changes in the circulating ribotypes over time. J. Med. Microbiol. 2016, 65, 1158–1163. [Google Scholar] [CrossRef] [PubMed]
- Aptekorz, M.; Szczegielniak, A.; Wiechuła, B.; Harmanus, C.; Kuijper, E.; Martirosian, G. Occurrence of Clostridium difficile ribotype 027 in hospitals of Silesia, Poland. Anaerobe 2017, 45, 106–113. [Google Scholar] [CrossRef]
- Vaverková, K.; Kracík, M.; Ryšková, L.; Paterová, P.; Kukla, R.; Hobzová, L.; Špánek, R.; Žemličková, H. Effect of Restriction of Fluoroquinolone Antibiotics on Clostridioides difficile Infections in the University Hospital Hradec Králové. Antibiotics 2021, 10, 519. [Google Scholar] [CrossRef]
- Rabold, D.; Espelage, W.; Abu Sin, M.; Eckmanns, T.; Schneeberg, A.; Neubauer, H.; Möbius, N.; Hille, K.; Wieler, L.H.; Seyboldt, C.; et al. The zoonotic potential of Clostridium difficile from small companion animals and their owners. PLoS ONE 2018, 13, e0193411. [Google Scholar] [CrossRef]
- Qin, J.; Dai, Y.; Ma, X.; Wang, Y.; Gao, Q.; Lu, H.; Li, T.; Meng, H.; Liu, Q.; Li, M. Nosocomial transmission of Clostridium difficile Genotype ST81 in a General Teaching Hospital in China traced by whole genome sequencing. Sci. Rep. 2017, 7, 9627. [Google Scholar] [CrossRef]
- Kabała, M.; Gofron, Z.; Aptekorz, M.; Sacha, K.; Harmanus, C.; Kuijper, E.; Martirosian, G. Clostridioides difficile Ribotype 027 (RT027) Outbreak Investigation Due to the Emergence of Rifampicin Resistance Using Multilocus Variable-Number Tandem Repeat Analysis (MLVA). Infect. Drug Resist. 2021, 14, 3247–3254. [Google Scholar] [CrossRef]
- Freeman, J.; Vernon, J.; Pilling, S.; Morris, K.; Nicholson, S.; Shearman, S.; Longshaw, C.; Wilcox, M.H.; Pan-European Longitudinal Surveillance of Antibiotic Resistance among Prevalent Clostridium difficile Ribotypes Study Group. The ClosER study: Results from a three-year pan-European longitudinal surveillance of antibiotic resistance among prevalent Clostridium difficile ribotypes, 2011–2014. Clin. Microbiol. Infect. 2017, 24, 724–731. [Google Scholar] [CrossRef] [PubMed]
- Sholeh, M.; Krutova, M.; Forouzesh, M.; Mironov, S.; Sadeghifard, N.; Molaeipour, L.; Maleki, A.; Kouhsari, E. Antimicrobial resistance in Clostridioides (Clostridium) difficile derived from humans: A systematic review and meta-analysis. Antimicrob. Resist. Infect. Control. 2020, 9, 158. [Google Scholar] [CrossRef] [PubMed]
| Clinical Characteristics | Overall (n = 84) No (%) | Age Over 65 (n = 70) No (%) | p-Value |
|---|---|---|---|
| Hospital-acquired CDI | 75 (89.3) | 62 (88.6) | |
| Death | 17 (20.2) | 16 (22.9) | |
| Previous use of ATBs * | 68 (80.9) | 57 (81.4) | n.s. |
| Previous use of PPIs * | 42 (50) | 34 (48.6) | |
| Comorbities | |||
| Hypertension | 56 (66.6) | 49 (70) | |
| Diabetes mellitus | 19 (22.6) | 17 (24.3) | |
| Malignancy | 13 (15.4) | 12 (17.1) | n.s. |
| COVID-19 | 9 (10.7) | 7 (10) |
| Antibiotics * | Overall (n = 84) No (%) | Age Over 65 (n = 70) No (%) | p-Value |
|---|---|---|---|
| Cephalosporins | 41 (48.8) | 38 (54.3) | 0.0029 ** |
| Fluoroquinolones | 23 (27.3) | 22 (31.4) | |
| Clindamycin | 18 (21.4) | 17 (24.3) | |
| Trimethoprim/Sulphametoxazole | 11 (13.1) | 9 (12.9) | |
| Macrolides | 10 (11.9) | 7 (10) | |
| Carbapenems | 8 (9.5) | 5 (7.1) | |
| Aminoglycosides | 7 (8.3) | 6 (8.6) | |
| Metronidazole | 6 (7.1) | 6 (8.6) | n.s. |
| Amoxicillin/Clavulanic Acid | 6 (7.1) | 4 (5.7) | |
| Number of ATB classes * | |||
| 1 ATB | 19 (22.6) | 17 (24.3) | |
| 2 and more ATBs | 49 (58.3) | 40 (57.1) | n.s. |
| No ATBs | 14 (16.7) | 11 (15.7) | |
| Unknown | 2 (2.4) | 2 (2.9) |
| Ribotype | No of Isolates | Toxin Gene Profile | ||
|---|---|---|---|---|
| tcdA+ tcdB+ cdtA+ cdtB+ | tcdA+ tcdB+ cdtA− cdtB− | tcdA− tcdB− cdtA− cdtB− | ||
| RT 176 | 40 | 40 | ||
| RT 001 | 23 | 23 | ||
| RT 014 | 7 | 7 | ||
| RT 078 | 2 | 2 | ||
| RT 020 | 2 | 2 | ||
| RT 027 | 1 | 1 | ||
| RT 003 | 1 | 1 | ||
| RT 010 | 1 | 1 | ||
| New | 5 | 2 | 1 | 2 |
| Failed | 2 | 1 | 1 | |
| Total | 84 | 46 | 35 | 3 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Curova, K.; Novotny, M.; Ambro, L.; Kamlarova, A.; Lovayova, V.; Hrabovsky, V.; Siegfried, L.; Jarcuska, P.; Jarcuska, P.; Toporova, A. High Prevalence of Clostridioides difficile Ribotype 176 in the University Hospital in Kosice. Pathogens 2023, 12, 430. https://doi.org/10.3390/pathogens12030430
Curova K, Novotny M, Ambro L, Kamlarova A, Lovayova V, Hrabovsky V, Siegfried L, Jarcuska P, Jarcuska P, Toporova A. High Prevalence of Clostridioides difficile Ribotype 176 in the University Hospital in Kosice. Pathogens. 2023; 12(3):430. https://doi.org/10.3390/pathogens12030430
Chicago/Turabian StyleCurova, Katarina, Martin Novotny, Lubos Ambro, Anna Kamlarova, Viera Lovayova, Vladimir Hrabovsky, Leonard Siegfried, Pavol Jarcuska, Peter Jarcuska, and Annamaria Toporova. 2023. "High Prevalence of Clostridioides difficile Ribotype 176 in the University Hospital in Kosice" Pathogens 12, no. 3: 430. https://doi.org/10.3390/pathogens12030430
APA StyleCurova, K., Novotny, M., Ambro, L., Kamlarova, A., Lovayova, V., Hrabovsky, V., Siegfried, L., Jarcuska, P., Jarcuska, P., & Toporova, A. (2023). High Prevalence of Clostridioides difficile Ribotype 176 in the University Hospital in Kosice. Pathogens, 12(3), 430. https://doi.org/10.3390/pathogens12030430








