Performance of Loop-Mediated Isothermal Amplification Technique in Milk Samples for the Diagnosis of Bovine Tuberculosis in Dairy Cattle Using a Bayesian Approach
Abstract
:1. Introduction
2. Results
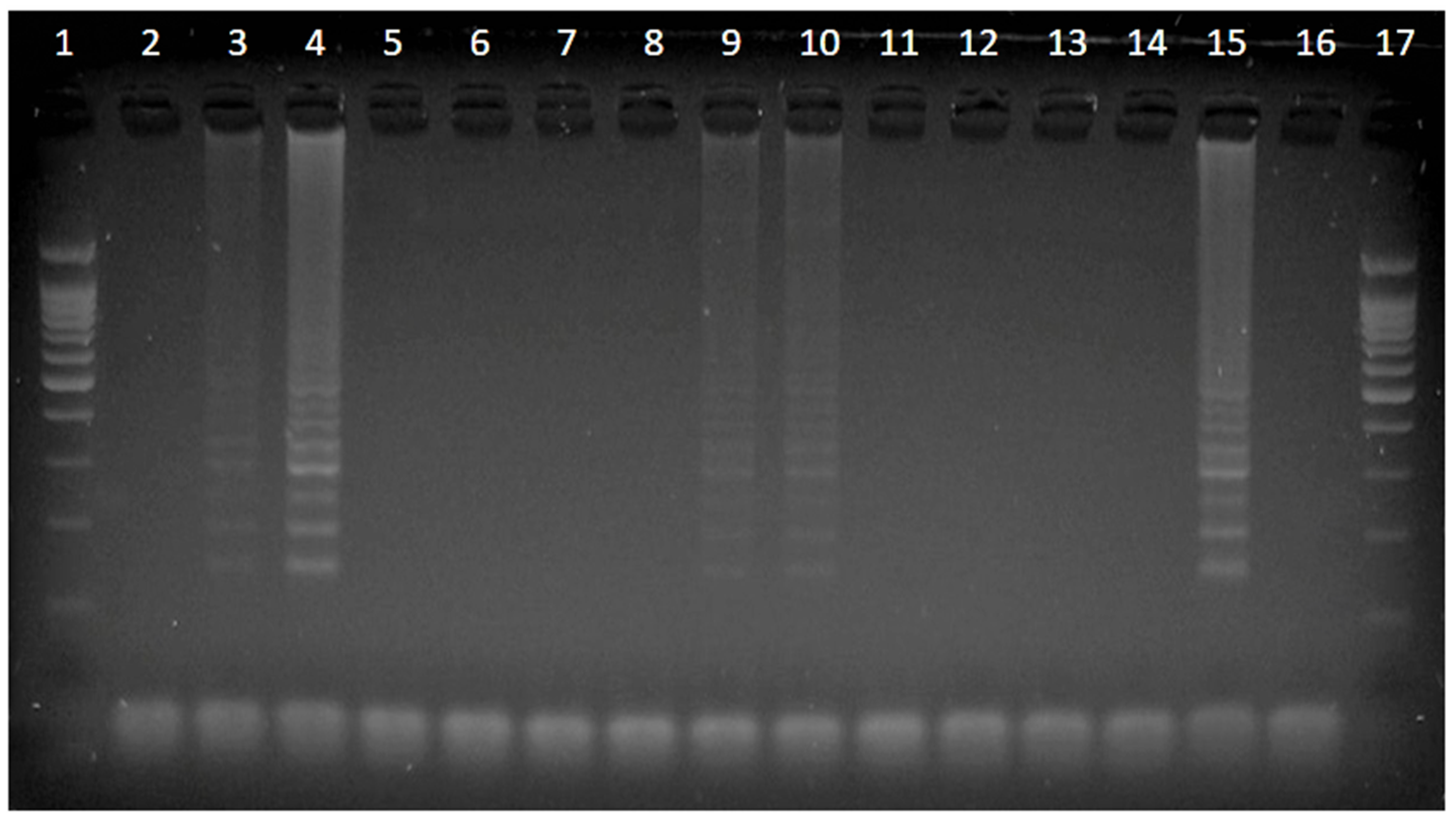
2.1. bTB Detection Using SIT and LAMP Tests
2.2. Test Performance of SIT and LAMP Tests
3. Discussion
4. Materials and Methods
4.1. Study Population
4.2. SIT Test
4.3. Milk Sample Collection
4.4. Genomic DNA Extraction from Milk
4.5. LAMP Test
4.6. Sensitivity and Specificity Estimations
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ayele, W.Y.; Neill, S.D.; Zinsstag, J.; Weiss, M.G.; Pavlik, I. Bovine Tuberculosis: An Old Disease but a New Threat to Africa. Int. J. Tuberc. Lung Dis. 2004, 8, 924–937. [Google Scholar]
- WHO. Global Tuberculosis Report 2018; WHO Document Services: Geneva, Switzerland, 2018. [Google Scholar]
- Waters, W.R.; Palmer, M.V.; Buddle, B.M.; Vordermeier, H.M. Bovine Tuberculosis Vaccine Research: Historical Perspectives and Recent Advances. Vaccine 2012, 30, 2611–2622. [Google Scholar] [CrossRef]
- De la Rua-Domenech, R.; Goodchild, A.T.; Vordermeier, H.M.; Hewinson, R.G.; Christiansen, K.H.; Clifton-Hadley, R.S. Ante Mortem Diagnosis of Tuberculosis in Cattle: A Review of the Tuberculin Tests, Gamma-Interferon Assay and Other Ancillary Diagnostic Techniques. Res. Vet. Sci. 2006, 81, 190–210. [Google Scholar] [CrossRef]
- Singhla, T.; Boonyayatra, S.; Chulakasian, S.; Lukkana, M.; Alvarez, J.; Sreevatsan, S.; Wells, S.J. Determination of the Sensitivity and Specificity of Bovine Tuberculosis Screening Tests in Dairy Herds in Thailand Using a Bayesian Approach. BMC Vet. Res. 2019, 15, 149. [Google Scholar] [CrossRef]
- Grabau, J.C.; DiFerdinando, G.T., Jr.; Novick, L.F. False Positive Tuberculosis Skin Test Results. Public Health Rep. 1995, 110, 703–706. [Google Scholar]
- Rahman, M.M.; Noor, M.; Islam, K.M.; Uddin, M.B.; Hossain, F.M.; Zinnah, M.A.; Al Mamun, M.; Islam, M.R.; Eo, S.K.; Ashour, H.M. Molecular Diagnosis of Bovine Tuberculosis in Bovine and Human Samples: Implications for Zoonosis. Future Microbiol. 2015, 10, 527–535. [Google Scholar] [CrossRef]
- OIE. Bovine Tuberculosis; World Organisation for Animal Health: Paris, France, 2014; Chapter 2.4.7. [Google Scholar]
- Song, L.; Li, J.; Hou, S.; Li, X.; Chen, S. Establishment of Loop-Mediated Isothermal Amplification (LAMP) for Rapid Detection of Brucella spp. and Application to Milk and Blood Samples. J. Microbiol. Methods 2012, 90, 292–297. [Google Scholar] [CrossRef]
- Suwancharoen, D.; Limlertvatee, S.; Chetiyawan, P.; Tongpan, P.; Sangkaew, N.; Sawaddee, Y.; Inthakan, K.; Wiratsudakul, A. A Nationwide Survey of Pathogenic Leptospires in Urine of Cattle and Buffaloes by Loop-Mediated Isothermal Amplification (LAMP) Method in Thailand, 2011–2013. J. Vet. Med. Sci. 2016, 78, 1495–1500. [Google Scholar] [CrossRef] [Green Version]
- Kapalamula, T.F.; Thapa, J.; Akapelwa, M.L.; Hayashida, K.; Gordon, S.V.; Ombe, B.M.H.; Munyeme, M.; Solo, E.S.; Bwalya, P.; Nyenje, M.E.; et al. Development of a loop-mediated isothermal amplification (LAMP) method for specific detection of Mycobacterium bovis. PLoS Negl. Trop. Dis. 2021, 15, e0008996. [Google Scholar] [CrossRef]
- Appelt, S.; Aly, S.S.; Tonooka, K.; Glenn, K.; Xue, Z.; Lehenbauer, T.W.; Marco, M.L. Development and Comparison of Loop-Mediated Isothermal Amplification and Quantitative Polymerase Chain Reaction Assays for the Detection of Mycoplasma bovis in Milk. J. Dairy Sci. 2019, 102, 1985–1996. [Google Scholar] [CrossRef]
- Branscum, A.J.; Gardner, I.A.; Johnson, W.O. Estimation of Diagnostic-Test Sensitivity and Specificity Through Bayesian Modeling. Prev. Vet. Med. 2005, 68, 145–163. [Google Scholar] [CrossRef]
- Rahman, A.K.; Saegerman, C.; Berkvens, D.; Fretin, D.; Gani, M.O.; Ershaduzzaman, M.; Ahmed, M.U.; Emmanuel, A. Bayesian Estimation of True Prevalence, Sensitivity and Specificity of Indirect ELISA, Rose Bengal Test and Slow Agglutination Test for the Diagnosis of Brucellosis in Sheep and Goats in Bangladesh. Prev. Vet. Med. 2013, 110, 242–252. [Google Scholar] [CrossRef]
- Singhla, T.; Tankaew, P.; Sthitmatee, N. Validation of a Novel ELISA for the Diagnosis of Hemorrhagic Septicemia in Dairy Cattle From Thailand Using a Bayesian Approach. Vet. Sci. 2020, 7, 163. [Google Scholar] [CrossRef]
- Alvarez, J.; Perez, A.; Bezos, J.; Marqués, S.; Grau, A.; Saez, J.L.; Mínguez, O.; de Juan, L.; Domínguez, L. Evaluation of the Sensitivity and Specificity of Bovine Tuberculosis Diagnostic Tests in Naturally Infected Cattle Herds Using a Bayesian Approach. Vet. Microbiol. 2012, 155, 38–43. [Google Scholar] [CrossRef]
- Srivastava, K.; Chauhan, D.S.; Gupta, P.; Singh, H.B.; Sharma, V.D.; Yadav, V.S.; Katoch, V.M. Isolation of Mycobacterium bovis and M. tuberculosis from Cattle of Some Farms in North India—Possible Relevance in Human Health. Indian J. Med. Res. 2008, 128, 26–31. [Google Scholar]
- Zarden, C.F.; Marassi, C.D.; Figueiredo, E.E.; Lilenbaum, W. Mycobacterium bovis Detection From Milk of Negative Skin Test Cows. Vet. Rec. 2013, 172, 130. [Google Scholar] [CrossRef] [Green Version]
- Wood, P.R.; Corner, L.A.; Rothel, J.S.; Baldock, C.; Jones, S.L.; Cousins, D.B.; McCormick, B.S.; Francis, B.R.; Creeper, J.; Tweddle, N.E. Field Comparison of the Interferon-Gamma Assay and the Intradermal Tuberculin Test for the Diagnosis of Bovine Tuberculosis. Aust. Vet. J. 1991, 68, 286–290. [Google Scholar] [CrossRef]
- Norby, B.; Bartlett, P.C.; Fitzgerald, S.D.; Granger, L.M.; Bruning-Fann, C.S.; Whipple, D.L.; Payeur, J.B. The Sensitivity of Gross Necropsy, Caudal Fold and Comparative Cervical Tests for the Diagnosis of Bovine Tuberculosis. J. Vet. Diagn. Investig. 2004, 16, 126–131. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Zhang, G.H.; Yang, L.; Huang, R.; Zhang, Y.; Jia, K.; Yuan, W.; Li, S.J. Development of a Loop-Mediated Isothermal Amplification Assay for the Detection of Mycobacterium bovis. Vet. J. 2011, 187, 393–396. [Google Scholar] [CrossRef]
- Zhang, H.; Wang, Z.; Cao, X.; Wang, Z.; Sheng, J.; Wang, Y.; Zhang, J.; Li, Z.; Gu, X.; Chen, C. Loop-Mediated Isothermal Amplification Assay Targeting the mpb70 Gene for Rapid Differential Detection of Mycobacterium bovis. Arch. Microbiol. 2016, 198, 905–911. [Google Scholar] [CrossRef]
- Moon, Y.J.; Lee, S.Y.; Oh, S.W. A Review of Isothermal Amplification Methods and Food-Origin Inhibitors Against Detecting Food-borne Pathogens. Foods 2022, 11, 322. [Google Scholar] [CrossRef]
- Monaghan, M.L.; Doherty, M.L.; Collins, J.D.; Kazda, J.F.; Quinn, P.J. The tuberculin test. Vet. Microbiol. 1994, 40, 111–124. [Google Scholar] [CrossRef]
- Thom, M.L.; Morgan, J.H.; Hope, J.C.; Villarreal-Ramos, B.; Martin, M.; Howard, C.J. The effect of repeated tuberculin skin testing of cattle on immune responses and disease following experimental infection with Mycobacterium bovis. Vet. Immunol. Immunopathol. 2004, 102, 399–412. [Google Scholar] [CrossRef]
- Coad, M.; Clifford, D.; Rhodes, S.G.; Hewinson, R.G.; Vordermeier, H.M.; Whelan, A.O. Repeat tuberculin skin testing leads to desensitisation in naturally infected tuberculous cattle which is associated with elevated interleukin-10 and decreased interleukin-1 beta responses. Vet. Res. 2010, 41, 14. [Google Scholar] [CrossRef] [Green Version]
- Ministry of Agricultural and Cooperatives. Diagnostic Test of Bovine Tuberculosis; Royal Gazette: Bangkok, Thailand, 2004.
- Oliver, S.P.; Gonzalez, R.N.; Hogan, J.S.; Jayarao, B.M.; Owens, W.E. Microbiological Procedures for the Diagnosis of Bovine Udder Infection and Determination of Milk Quality, 4th ed.; National Mastitis Council: Verona, WI, USA, 2004. [Google Scholar]
- Hong, M.; Zha, L.; Fu, W.; Zou, M.; Li, W.; Xu, D. A Modified Visual Loop-Mediated Isothermal Amplification Method for Diagnosis and Differentiation of Main Pathogens From Mycobacterium tuberculosis Complex. World J. Microbiol. Biotechnol. 2012, 28, 523–531. [Google Scholar] [CrossRef]
- Landis, J.R.; Koch, G.G. The Measurement of Observer Agreement for Categorical Data. Biometrics 1977, 33, 159–174. [Google Scholar] [CrossRef] [Green Version]
- Whipple, D.L.; Palmer, M.V.; Slaughter, R.E.; Jones, S.L. Comparison of Purified Protein Derivatives and Effect of Skin Testing on Results of a Commercial Gamma Interferon Assay for Diagnosis of Tuberculosis in Cattle. J. Vet. Diagn. Investig. 2001, 13, 117–122. [Google Scholar] [CrossRef] [Green Version]
- Farnham, M.W.; Norby, B.; Goldsmith, T.J.; Wells, S.J. Meta-Analysis of Field Studies on Bovine Tuberculosis Skin Tests in United States Cattle Herds. Prev. Vet. Med. 2012, 103, 234–242. [Google Scholar] [CrossRef]
- Plummer, M.; Stukalov, A.; Denwood, M. Rjags: Bayesian Graphical Models Using MCMC. 2019. Available online: https://cran.r-project.org/web/packages/rjags/rjags.pdf (accessed on 19 August 2021).
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2021. [Google Scholar]
- Su, Y.S.; Yajima, M.R. 2jags: Using R to Run. J. Am. Geriatr. Soc. 2021. Available online: https://cran.r-project.org/web/packages/R2jags/R2jags.pdf (accessed on 19 August 2021).
- Gelman, A.; Rubin, D.B. Inference From Iterative Simulation Using Multiple Sequences. Stat. Sci. 1992, 7, 457–511. [Google Scholar] [CrossRef]
| Test Results | SIT (Standard) a | SIT (Severe) b | |||
|---|---|---|---|---|---|
| Positive | Negative | Positive | Negative | ||
| LAMP c | positive | 0 | 38 | 6 | 32 |
| negative | 2 | 163 | 9 | 156 | |
| Diagnostic Tests | Parameters | Median (%) | 95% PPI a (%) |
|---|---|---|---|
| SIT (standard) a | Sensitivity | 63.5 | 42.1–81.9 |
| Specificity | 99.1 | 97.1–99.9 | |
| LAMP b | Sensitivity | 67.2 | 40.5–88.4 |
| Specificity | 82.0 | 76.1–87.1 | |
| Disease prevalence | 3.7 | 1.4–7.8 |
| Diagnostic Tests | Parameters | Median (%) | 95% PPI a (%) |
|---|---|---|---|
| SIT (severe) a | Sensitivity | 76.1 | 55.7–90.9 |
| Specificity | 96 | 92.6–98.5 | |
| LAMP b | Sensitivity | 68.8 | 44.8–88.9 |
| Specificity | 84.1 | 78.2–89.2 | |
| Disease prevalence | 6.7 | 3.2–12.1 |
| Primer | DNA Sequence (5′-3′) | Length | Target |
|---|---|---|---|
| F3 | CCGGGTGAGGATCCTGAC | 18 bp | esat6 |
| B3 | GACTGGTCGAGCTTCAGC | 18 bp | esat6 |
| FIP | GAAAGCACCGCGACGGTGTCTTTTCAGACGGATGACCGATTTGG | 44 bp | esat6 |
| BIP | CGAGGTGTTGGAAGACACGCCTTTTGAACGCCCACACGCCTT | 42 bp | esat6 |
| Diagnostic Tests | Parameters | Mode | 95% CI a |
|---|---|---|---|
| SIT test (standard) b | Sensitivity | 71.0 | >53.2 |
| Specificity | 98.6 | >89.2 | |
| LAMP test c | Sensitivity | 75.0 | >50.0 |
| Specificity | 95.0 | >50.0 | |
| Disease prevalence | 10.0 | <20.0 |
| Diagnostic Tests | Parameters | Mode | 95% CI a |
|---|---|---|---|
| SIT test (standard) b | Sensitivity | 81.0 | >63.0 |
| Specificity | 95.6 | >89.2 | |
| LAMP test c | Sensitivity | 75.0 | >50.0 |
| Specificity | 95.0 | >50.0 | |
| Disease prevalence | 10.0 | <20.0 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Singhla, T.; Pikulkaew, S.; Boonyayatra, S. Performance of Loop-Mediated Isothermal Amplification Technique in Milk Samples for the Diagnosis of Bovine Tuberculosis in Dairy Cattle Using a Bayesian Approach. Pathogens 2022, 11, 573. https://doi.org/10.3390/pathogens11050573
Singhla T, Pikulkaew S, Boonyayatra S. Performance of Loop-Mediated Isothermal Amplification Technique in Milk Samples for the Diagnosis of Bovine Tuberculosis in Dairy Cattle Using a Bayesian Approach. Pathogens. 2022; 11(5):573. https://doi.org/10.3390/pathogens11050573
Chicago/Turabian StyleSinghla, Tawatchai, Surachai Pikulkaew, and Sukolrat Boonyayatra. 2022. "Performance of Loop-Mediated Isothermal Amplification Technique in Milk Samples for the Diagnosis of Bovine Tuberculosis in Dairy Cattle Using a Bayesian Approach" Pathogens 11, no. 5: 573. https://doi.org/10.3390/pathogens11050573
APA StyleSinghla, T., Pikulkaew, S., & Boonyayatra, S. (2022). Performance of Loop-Mediated Isothermal Amplification Technique in Milk Samples for the Diagnosis of Bovine Tuberculosis in Dairy Cattle Using a Bayesian Approach. Pathogens, 11(5), 573. https://doi.org/10.3390/pathogens11050573







