Porphyromonas gingivalis Virulence Factors and Clinical Significance in Periodontal Disease and Coronary Artery Diseases
Abstract
1. Introduction
2. Porphyromonas gingivalis (P. gingivalis)
2.1. Virulence Factors
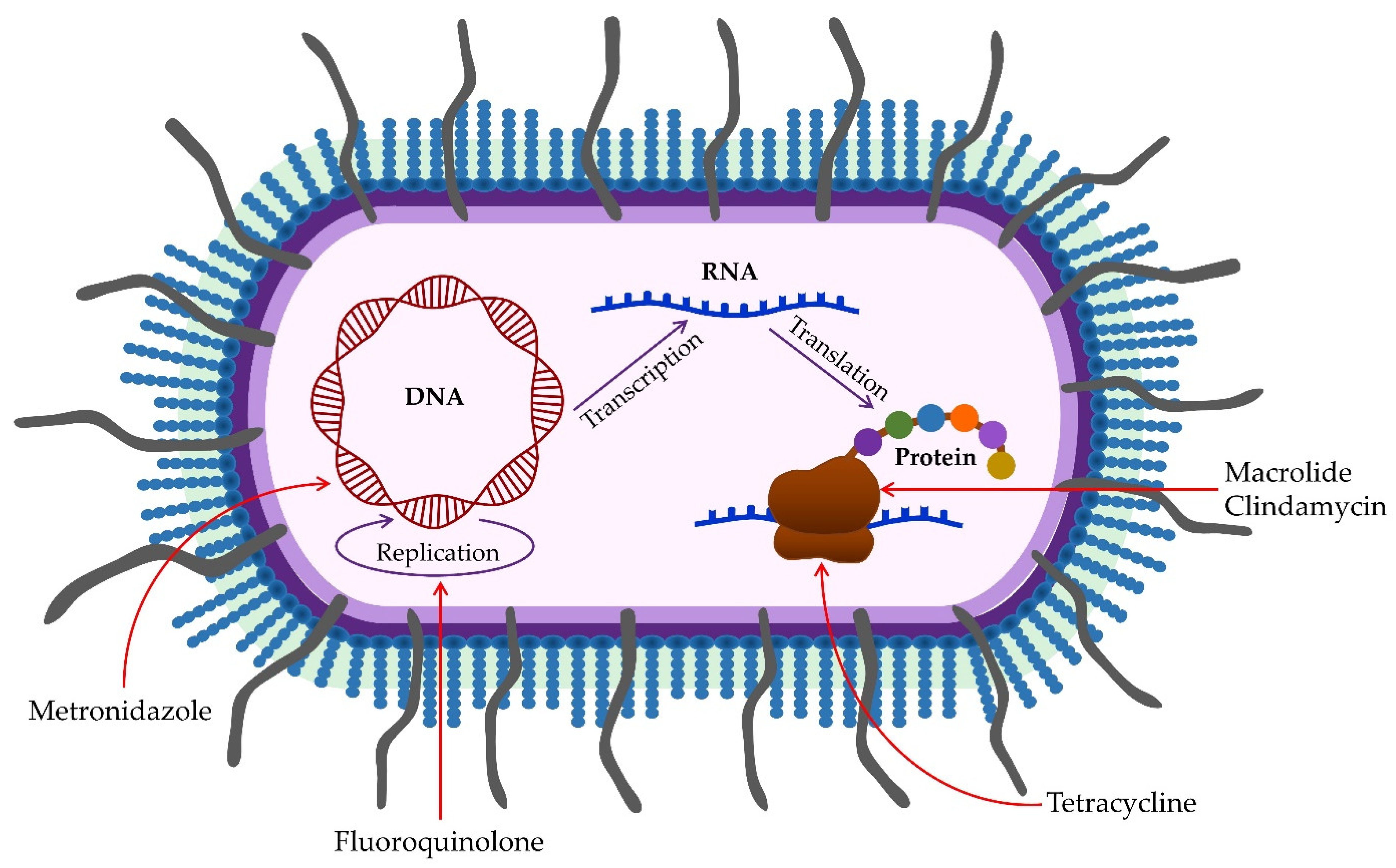
2.2. Biofilm
3. Clinical Importance of Porphyromonas gingivalis in Periodontal Disease
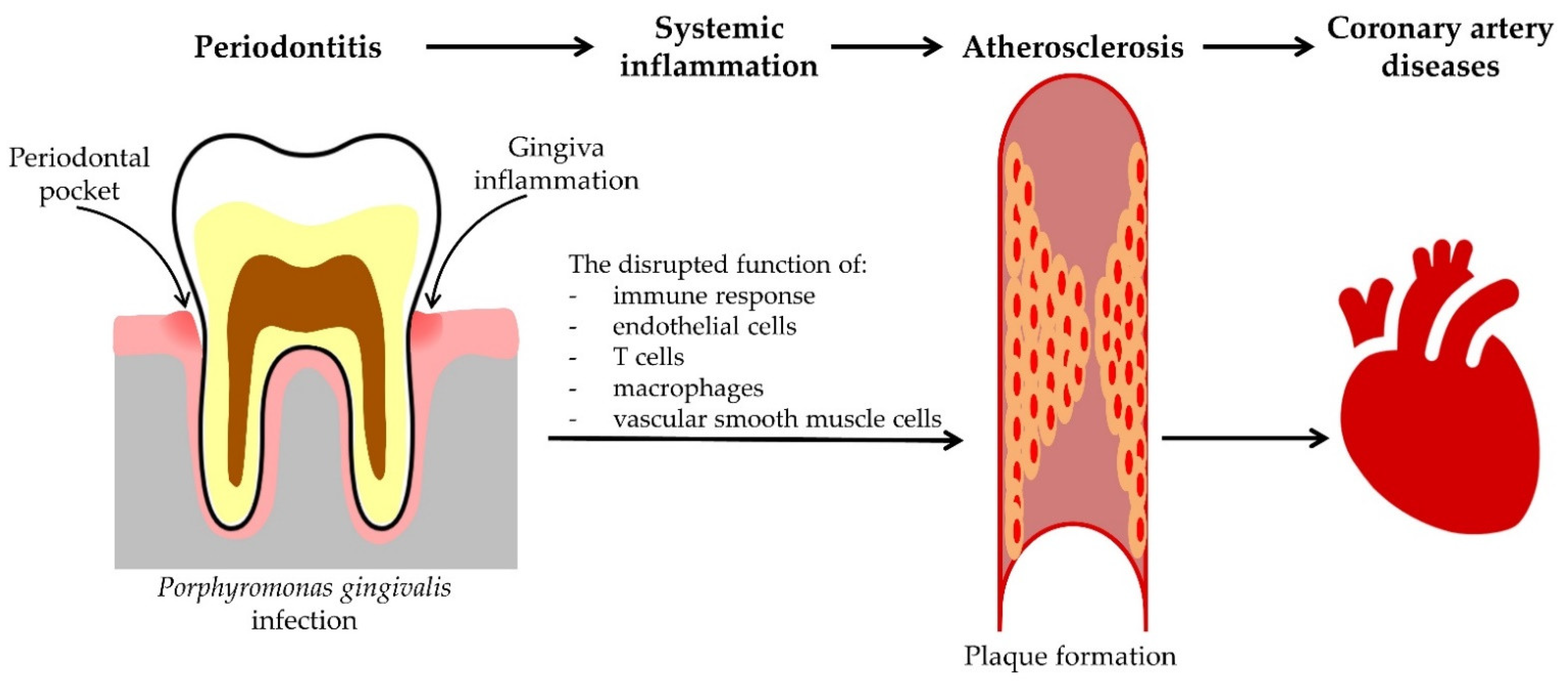
4. Clinical Implication of Periodontal Infections Caused by P. gingivalis on the Development of Coronary Artery Diseases
Atherosclerosis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Slots, J. Periodontitis: Facts, fallacies and the future. Periodontol. 2000 2017, 75, 7–23. [Google Scholar] [CrossRef]
- Xu, W.; Zhou, W.; Wang, H.; Liang, S. Roles of Porphyromonas gingivalis and its virulence factors in periodontitis. Adv. Protein Chem. Struct. Biol. 2020, 120, 45–84. [Google Scholar] [CrossRef] [PubMed]
- Könönen, E.; Gursoy, M.; Gursoy, U.K. Periodontitis: A Multifaceted Disease of Tooth-Supporting Tissues. J. Clin. Med. 2019, 8, 1135. [Google Scholar] [CrossRef] [PubMed]
- Nagano, K.; Hasegawa, Y. Periodontal Pathogens; Humana: New York, NY, USA, 2021; ISBN 9781071609385. [Google Scholar]
- Kinane, D.F.; Stathopoulou, P.G.; Papapanou, P.N. Periodontal diseases. Nat. Rev. Dis. Primers 2017, 3, 17038. [Google Scholar] [CrossRef] [PubMed]
- Preshaw, P.M.; Bissett, S.M. Periodontitis and diabetes. Br. Dent. J. 2019, 227, 577–584. [Google Scholar] [CrossRef] [PubMed]
- Bui, F.Q.; Almeida-da-Silva, C.L.C.; Huynh, B.; Trinh, A.; Liu, J.; Woodward, J.; Asadi, H.; Ojcius, D.M. Association between periodontal pathogens and systemic disease. Biomed. J. 2019, 42, 27–35. [Google Scholar] [CrossRef] [PubMed]
- Kwon, T.H.; Lamster, I.B.; Levin, L. Current Concepts in the Management of Periodontitis. Int. Dent. J. 2021, 71, 462–476. [Google Scholar] [CrossRef] [PubMed]
- Hajishengallis, G.; Lamont, R.J. The polymicrobial synergy and dysbiosis model of periodontal disease pathogenesis. In The Human Microbiota and Chronic Disease; Wiley: Hoboken, NJ, USA, 2016; pp. 227–242. [Google Scholar] [CrossRef]
- Mohanty, R.; Asopa, S.; Joseph, M.D.; Singh, B.; Rajguru, J.; Saidath, K.; Sharma, U. Red complex: Polymicrobial conglomerate in oral flora: A review. J. Fam. Med. Prim. Care 2019, 8, 3480. [Google Scholar] [CrossRef]
- Bourgeois, D.; Inquimbert, C.; Ottolenghi, L.; Carrouel, F. Periodontal Pathogens as Risk Factors of Cardiovascular Diseases, Diabetes, Rheumatoid Arthritis, Cancer, and Chronic Obstructive Pulmonary Disease-Is There Cause for Consideration? Microorganisms 2019, 7, 424. [Google Scholar] [CrossRef] [PubMed]
- Leite, F.R.M.; Nascimento, G.G.; Scheutz, F.; López, R. Effect of Smoking on Periodontitis: A Systematic Review and Meta-regression. Am. J. Prev. Med. 2018, 54, 831–841. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.Z.; Yuan, Y.H.; Liu, H.H.; Li, S.S.; Zhang, B.W.; Chen, W.; An, Z.J.; Chen, S.Y.; Wu, Y.Z.; Han, B.; et al. Epidemiologic relationship between periodontitis and type 2 diabetes mellitus. BMC Oral Health 2020, 20, 204. [Google Scholar] [CrossRef] [PubMed]
- Van der Velden, U. What exactly distinguishes aggressive from chronic periodontitis: Is it mainly a difference in the degree of bacterial invasiveness? Periodontol. 2000 2017, 75, 24–44. [Google Scholar] [CrossRef] [PubMed]
- Bodet, C.; Chandad, F.; Grenier, D. Pathogenic potential of Porphyromonas gingivalis, Treponema denticola and Tannerella forsythia, the red bacterial complex associated with periodontitis. Pathol. Biol. 2007, 55, 154–162. [Google Scholar] [CrossRef] [PubMed]
- Mysak, J.; Podzimek, S.; Sommerova, P.; Lyuya-Mi, Y.; Bartova, J.; Janatova, T.; Prochazkova, J.; Duskova, J. Porphyromonas gingivalis: Major periodontopathic pathogen overview. J. Immunol. Res. 2014, 2014, 476068. [Google Scholar] [CrossRef] [PubMed]
- Socransky, S.S.; Haffajee, A.D.; Cugini, M.A.; Smith, C.; Kent, R.L. Microbial complexes in subgingival plaque. J. Clin. Periodontol. 1998, 25, 134–144. [Google Scholar] [CrossRef]
- Malakar, A.K.; Choudhury, D.; Halder, B.; Paul, P.; Uddin, A.; Chakraborty, S. A review on coronary artery disease, its risk factors, and therapeutics. J. Cell. Physiol. 2019, 234, 16812–16823. [Google Scholar] [CrossRef]
- Sanz, M.; Marco del Castillo, A.; Jepsen, S.; Gonzalez-Juanatey, J.R.; D’Aiuto, F.; Bouchard, P.; Chapple, I.; Dietrich, T.; Gotsman, I.; Graziani, F.; et al. Periodontitis and cardiovascular diseases: Consensus report. J. Clin. Periodontol. 2020, 47, 268–288. [Google Scholar] [CrossRef]
- Zardawi, F.; Gul, S.; Abdulkareem, A.; Sha, A.; Yates, J. Association Between Periodontal Disease and Atherosclerotic Cardiovascular Diseases: Revisited. Front. Cardiovasc. Med. 2021, 7, 625579. [Google Scholar] [CrossRef]
- How, K.Y.; Song, K.P.; Chan, K.G. Porphyromonas gingivalis: An Overview of Periodontopathic Pathogen below the Gum Line. Front. Microbiol. 2016, 7, 53. [Google Scholar] [CrossRef]
- Eisenstein, M. Homing in on an oral link to inflammatory disease. Nature 2021. [Google Scholar] [CrossRef]
- Frédéric, L.J.; Michel, B.; Selena, T. Oral Microbes, Biofilms and Their Role in Periodontal and Peri-Implant Diseases. Materials 2018, 11, 1802. [Google Scholar] [CrossRef] [PubMed]
- Martínez-García, M.; Hernández-Lemus, E. Periodontal Inflammation and Systemic Diseases: An Overview. Front. Physiol. 2021, 12, 709438. [Google Scholar] [CrossRef] [PubMed]
- Zheng, S.; Yu, S.; Fan, X.; Zhang, Y.; Sun, Y.; Lin, L.; Wang, H.; Pan, Y.; Li, C. Porphyromonas gingivalis survival skills: Immune evasion. J. Periodontal Res. 2021, 56, 1007–1018. [Google Scholar] [CrossRef] [PubMed]
- Kuboniwa, M.; Amano, A.; Hashino, E.; Yamamoto, Y.; Inaba, H.; Hamada, N.; Nakayama, K.; Tribble, G.D.; Lamont, R.J.; Shizukuishi, S. Distinct roles of long/short fimbriae and gingipains in homotypic biofilm development by Porphyromonas gingivalis. BMC Microbiol. 2009, 9, 105. [Google Scholar] [CrossRef] [PubMed]
- Ng, H.M.; Slakeski, N.; Butler, C.A.; Veith, P.D.; Chen, Y.Y.; Liu, S.W.; Hoffmann, B.; Dashper, S.G.; Reynolds, E.C. The Role of Treponema denticola Motility in Synergistic Biofilm Formation with Porphyromonas gingivalis. Front. Cell. Infect. Microbiol. 2019, 9, 432. [Google Scholar] [CrossRef]
- Lu, B.; McBride, B.C. Stress response of Porphyromonas gingivalis. Oral Microbiol. Immunol. 1994, 9, 166–173. [Google Scholar] [CrossRef]
- Bostanci, N.; Belibasakis, G.N. Porphyromonas gingivalis: An invasive and evasive opportunistic oral pathogen. FEMS Microbiol. Lett. 2012, 333, 1–9. [Google Scholar] [CrossRef]
- Stobernack, T.; du Teil Espina, M.; Mulder, L.M.; Palma Medina, L.M.; Piebenga, D.R.; Gabarrini, G. A Secreted Bacterial Peptidylarginine Deiminase Can Neutralize Human Innate Immune Defenses. MBio 2018, 9, e01704-18. [Google Scholar] [CrossRef]
- Hajishengalis, G.; Krauss, J.L.; Jotwani, R.; Lambris, J.D. Differential capacity for complement receptor-mediated immune evasion by Porphyromonas gingivalis depending on the type of innate leukocyte George. Mol. Oral Microbiol. 2017, 32, 154–165. [Google Scholar] [CrossRef]
- Wang, M.; Shakhatreh, M.-A.K.; James, D.; Liang, S.; Nishiyama, S.; Yoshimura, F.; Demuth, D.R.; Hajishengallis, G. Fimbrial Proteins of Porphyromonas gingivalis Mediate In Vivo Virulence and Exploit TLR2 and Complement Receptor 3 to Persist in Macrophages. J. Immunol. 2007, 179, 2349–2358. [Google Scholar] [CrossRef]
- Coats, S.R.; Jones, J.W.; Do, C.T.; Braham, P.H.; Bainbridge, B.W.; To, T.T.; Goodlett, D.R.; Ernst, R.K.; Darveau, R.P. Human Toll-like receptor 4 responses to P. gingivalis are regulated by lipid A 1- and 4’- phosphatase activities. Cell Microbiol 2009, 11, 1587–1599. [Google Scholar] [CrossRef]
- Hajishengallis, G.; Shakhatreh, M.-A.K.; Wang, M.; Liang, S. Complement Receptor 3 Blockade Promotes IL-12-Mediated Clearance of Porphyromonas gingivalis and Negates Its Virulence In Vivo. J. Immunol. 2007, 179, 2359–2367. [Google Scholar] [CrossRef]
- Potempa, J.; Pike, R.N. Corruption of innate immunity by bacterial proteases. J. Innate Immun. 2009, 1, 70–87. [Google Scholar] [CrossRef] [PubMed]
- Hajishengallis, G.; Wang, M.; Liang, S.; Triantafilou, M.; Triantafilou, K. Pathogen induction of CXCR4/TLR2 cross-talk impairs host defense function. Proc. Natl. Acad. Sci. USA 2008, 105, 13532–13537. [Google Scholar] [CrossRef] [PubMed]
- Dominy, S.S.; Lynch, C.; Ermini, F.; Benedyk, M.; Marczyk, A.; Konradi, A.; Nguyen, M.; Haditsch, U.; Raha, D.; Griffin, C.; et al. Porphyromonas gingivalis in Alzheimer’s disease brains: Evidence for disease causation and treatment with small-molecule inhibitors. Sci. Adv. 2019, 5, aau3333. [Google Scholar] [CrossRef] [PubMed]
- Maekawa, T.; Krauss, J.L.; Abe, T.; Jotwani, R.; Triantafilou, M.; Triantafilou, K.; Hashim, A.; Hoch, S.; Curtis, M.A.; Nussbaum, G.; et al. Porphyromonas gingivalis manipulates complement and TLR signaling to uncouple bacterial clearance from inflammation and promote dysbiosis. Cell Host Microbe 2014, 15, 768–778. [Google Scholar] [CrossRef] [PubMed]
- Hasegawa, Y.; Nagano, K. Porphyromonas gingivalis FimA and Mfa1 fimbriae: Current insights on localization, function, biogenesis, and genotype. Jpn. Dent. Sci. Rev. 2021, 57, 190–200. [Google Scholar] [CrossRef] [PubMed]
- Jia, L.; Han, N.; Du, J.; Guo, L.; Luo, Z.; Liu, Y. Pathogenesis of important virulence factors of Porphyromonas gingivalis via toll-like receptors. Front. Cell. Infect. Microbiol. 2019, 9, 262. [Google Scholar] [CrossRef]
- Amano, A. Host-parasite interactions in periodontitis: Microbial pathogenicity and innate immunity. Periodontol. 2000 2010, 54, 9–14. [Google Scholar] [CrossRef]
- Ogawa, T.; Mukai, T.; Yasuda, K.; Shimauchi, H.; Toda, Y.; Hamada, S. Distribution and immunochemical specificities of fimbriae of Porphyromonas gingivalis and related bacterial species. Oral Microbiol. Immunol. 1991, 6, 332–340. [Google Scholar] [CrossRef]
- Amano, A. Molecular Interaction of Porphyromonas gingivalis with Host Cells: Implication for the Microbial Pathogenesis of Periodontal Disease. J. Periodontol. 2003, 74, 90–96. [Google Scholar] [CrossRef] [PubMed]
- Lamont, R.J.; Jenkinson, H.F. Subgingival colonization by Porphyromonas gingivalis. Oral Microbiol. Immunol. 2000, 15, 341–349. [Google Scholar] [CrossRef]
- Sojar, H.T.; Genco, R.J. Identification of glyceraldehyde-3-phosphate dehydrogenase of epithelial cells as a second molecule that binds to Porphyromonas gingivalis fimbriae. FEMS Immunol. Med. Microbiol. 2005, 45, 25–30. [Google Scholar] [CrossRef] [PubMed]
- Lamont, R.J.; Hsiao, G.W.; Gil, S. Identification of a molecule of Porphyromonas gingivalis that binds to Streptococcus. Microb. Pathog. 1994, 17, 355–360. [Google Scholar] [CrossRef] [PubMed]
- Hamada, N.; Sojar, H.T.; Cho, M.I.L.; Genco, R.J. Isolation and characterization of a minor fimbria from Porphyromonas gingivalis. Infect. Immun. 1996, 64, 4788–4794. [Google Scholar] [CrossRef] [PubMed]
- Smalley, J.W.; Olczak, T. Heme acquisition mechanisms of Porphyromonas gingivalis—Strategies used in a polymicrobial community in a heme-limited host environment. Mol. Oral Microbiol. 2017, 32, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Liu, D.; Liu, S.; Zhang, S.; Pan, Y. The Role of Porphyromonas gingivalis Outer Membrane Vesicles in Periodontal Disease and Related Systemic Diseases. Front. Cell. Infect. Microbiol. 2021, 10, 585917. [Google Scholar] [CrossRef]
- Potempa, J.; Pike, R.; Travis, J. Titration and Mapping of the Active Site of Cysteine Proteinases from Porphyromonas gingivalis (Gingipains) Using Peptidyl Chloromethanes. Biol. Chem. 1997, 378, 223–230. [Google Scholar] [CrossRef] [PubMed]
- Smalley, J.W.; Silver, J.; Marsh, P.J.; Birss, A.J. The periodontopathogen Porphyromonas gingivalis binds iron protoporphyrin IX in the μ-oxo dimeric form: An oxidative buffer and possible pathogenic mechanism. Biochem. J. 1998, 331, 601–685. [Google Scholar] [CrossRef]
- Amano, A.; Kuboniwa, M.; Kataoka, K.; Tazaki, K.; Inoshita, E.; Nagata, H.; Tamagawa, H.; Shizukuishi, S. Binding of hemoglobin by Porphyromonas gingivalis. FEMS Microbiol. Lett. 1995, 134, 63–67. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Mckee, A.S.; Mcdermid, A.S.; Baskerville, A.; Dowsett, A.B.; Ellwood, D.C.; Marshlt, P.D. Effect of hemin on the physiology and virulence of Bacteroides gingivalis W50. Infect. Immun. 1986, 52, 349–355. [Google Scholar] [CrossRef] [PubMed]
- Marsh, P.D.; McDermid, A.S.; McKee, A.S.; Baskerville, A. The effect of growth rate and haemin on the virulence and proteolytic activity of Porphyromonas gingivalis W50. Microbiology 1994, 140, 861–865. [Google Scholar] [CrossRef][Green Version]
- Smalley, J.W.; Birss, A.J.; Mckee, A.S.; Marsh, P.D. Haemin-restriction influences haemin-binding, haemagglutination and protease activity of cells and extracellular membrane vesicles of Porphyromonas gingivalis W50. FEMS Microbiol. Lett. 1991, 69, 63–67. [Google Scholar] [CrossRef]
- Al-Qutub, M.N.; Braham, P.H.; Karimi-Naser, L.M.; Liu, X.; Genco, C.A.; Darveau, R.P. Hemin-dependent modulation of the lipid A structure of Porphyromonas gingivalis lipopolysaccharide. Infect. Immun. 2006, 74, 4474–4485. [Google Scholar] [CrossRef] [PubMed]
- Cutler, C.W.; Eke, P.I.; Genco, C.A.; Van Dyke, T.E.; Arnold, R.R. Hemin-induced modifications of the antigenicity and hemin-binding capacity of Porphyromonas gingivalis lipopolysaccharide. Infect. Immun. 1996, 64, 2282–2287. [Google Scholar] [CrossRef]
- Smalley, J.W.; Birss, A.J.; Mckee, A.S.; Marsh, P.D. Changes in the affinity of haemin-binding by Porphyromonas gingivalis w50 under different environmental conditions. Microb. Ecol. Health Dis. 1994, 7, 9–15. [Google Scholar] [CrossRef]
- Smalley, J.W.; Birss, A.J.; Silver, J. The periodontal pathogen Porphyromonas gingivalis harnesses the chemistry of the μ-oxo bishaem of iron protoporphyrin IX to protect against hydrogen peroxide. FEMS Microbiol. Lett. 2000, 183, 159–164. [Google Scholar] [CrossRef]
- Smalley, J.W.; Birss, A.J.; Szmigielski, B.; Potempa, J. Mechanism of methaemoglobin breakdown by the lysine-specific gingipain of the periodontal pathogen Porphyromonas gingivalis. Biol. Chem. 2008, 389, 1235–1238. [Google Scholar] [CrossRef] [PubMed]
- Smalley, J.W.; Birss, A.J.; Szmigielski, B.; Potempa, J. Sequential action of R- and K-specific gingipains of Porphyromonas gingivalis in the generation of the heam-containing pigment from oxyhaemoglobin. Arch. Biochem. Biophys. 2007, 465, 44–49. [Google Scholar] [CrossRef] [PubMed]
- Smalley, J.W.; Birss, A.J.; Withnall, R.; Silver, J. Interactions of Porphyromonas gingivalis with oxyhaemoglobin and deoxyhaemoglobin. Biochem. J. 2002, 362, 239–245. [Google Scholar] [CrossRef] [PubMed]
- Potempa, J.; Sroka, A.; Imamura, T.; Travis, J. Gingipains, the Major Cysteine Proteinases and Virulence Factors of Porphyromonas gingivalis: Structure, Function and Assembly of Multidomain Protein Complexes. Curr. Protein Pept. Sci. 2003, 4, 397–407. [Google Scholar] [CrossRef] [PubMed]
- Laine, M.L.; Appelmelk, B.J.; Van Winkelhoff, A.J. Prevalence and distribution of six capsular serotypes of Porphyromonas gingivalis in periodontitis patients. J. Dent. Res. 1997, 76, 1840–1844. [Google Scholar] [CrossRef] [PubMed]
- Vernal, R.; León, R.; Silva, A.; Van Winkelhoff, A.J.; Garcia-Sanz, J.A.; Sanz, M. Differential cytokine expression by human dendritic cells in response to different Porphyromonas gingivalis capsular serotypes. J. Clin. Periodontol. 2009, 36, 823–829. [Google Scholar] [CrossRef]
- Rosen, G.; Sela, M.N. Coaggregation of Porphyromonas gingivalis and Fusobacterium nucleatum PK 1594 is mediated by capsular polysaccharide and lipopolysaccharide. FEMS Microbiol. Lett. 2006, 256, 304–310. [Google Scholar] [CrossRef]
- Igboin, C.O.; Tordoff, K.P.; Moeschberger, M.L.; Griffen, A.L.; Leys, E.J. Porphyromonas gingivalis-host interactions in a drosophila melanogaster model. Infect. Immun. 2011, 79, 449–458. [Google Scholar] [CrossRef] [PubMed]
- Dierickx, K.; Pauwels, M.; Laine, M.L.; Van Eldere, J.; Cassiman, J.-J.; Van Winkelhoff, A.J.; Van Steenberghe, D.; Quirynen, M. Adhesion of Porphyromonas gingivalis. Serotypes to Pocket Epithelium. J. Periodontol. 2003, 74, 844–848. [Google Scholar] [CrossRef]
- D’Empaire, G.; Baer, M.T.; Gibson, F.C. The K1 serotype capsular polysaccharide of Porphyromonas gingivalis elicits chemokine production from murine macrophages that facilitates cell migration. Infect. Immun. 2006, 74, 6236–6243. [Google Scholar] [CrossRef]
- Singh, A.; Wyant, T.; Anaya-Bergman, C.; Aduse-Opoku, J.; Brunner, J.; Laine, M.L.; Curtis, M.A.; Lewis, J.P. The capsule of Porphyromonas gingivalis leads to a reduction in the host inflammatory response, evasion of phagocytosis, and increase in Virulence. Infect. Immun. 2011, 79, 4533–4542. [Google Scholar] [CrossRef]
- Brunner, J.; Scheres, N.; El Idrissi, N.B.; Deng, D.M.; Laine, M.L.; Van Winkelhoff, A.J.; Crielaard, W. The capsule of Porphyromonas gingivalis reduces the immune response of human gingival fibroblasts. BMC Microbiol. 2010, 10, 5. [Google Scholar] [CrossRef]
- Cecil, J.D.; Sirisaengtaksin, N.; O’Brien-Simpson, N.M.; Krachler, A.M. Outer Membrane Vesicle-Host Cell Interactions. Protein Secret. Bact. 2019, 7, 201–214. [Google Scholar] [CrossRef]
- Ahmadi Badi, S.; Bruno, S.P.; Moshiri, A.; Tarashi, S.; Siadat, S.D.; Masotti, A. Small RNAs in Outer Membrane Vesicles and Their Function in Host-Microbe Interactions. Front. Microbiol. 2020, 11, 1209. [Google Scholar] [CrossRef] [PubMed]
- Bonnington, K.E.; Kuehn, M.J. Protein selection and export via outer membrane vesicles. Biochim. Biophys. Acta 2014, 1843, 1612–1619. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Cruz, C.; Delgado, L.; López-Iglesias, C.; Mercade, E. Outer-inner membrane vesicles naturally secreted by gram-negative pathogenic bacteria. PLoS ONE 2015, 10, e0116896. [Google Scholar] [CrossRef] [PubMed]
- Mantri, C.K.; Chen, C.H.; Dong, X.; Goodwin, J.S.; Pratap, S.; Paromov, V.; Xie, H. Fimbriae-mediated outer membrane vesicle production and invasion of Porphyromonas gingivalis. Microbiologyopen 2015, 4, 53–65. [Google Scholar] [CrossRef] [PubMed]
- Xie, H. Biogenesis and function of Porphyromonas gingivalis outer membrane vesicles. Futur. Microbiol 2015, 10, 1517–1527. [Google Scholar] [CrossRef] [PubMed]
- Gui, M.J.; Dashper, S.G.; Slakeski, N.; Chen, Y.Y.; Reynolds, E.C. Spheres of influence: Porphyromonas gingivalis outer membrane vesicles. Mol. Oral Microbiol. 2016, 31, 365–378. [Google Scholar] [CrossRef] [PubMed]
- Chew, C.L.; Conos, S.A.; Unal, B.; Tergaonkar, V. Noncoding RNAs: Master Regulators of Inflammatory Signaling. Trends Mol. Med. 2018, 24, 66–84. [Google Scholar] [CrossRef]
- Elhenawy, W.; Debelyy, M.O.; Feldman, M.F. Preferential packing of acidic glycosidases and proteases into Bacteroides outer membrane vesicles. MBio 2014, 5, e00909-14. [Google Scholar] [CrossRef]
- Ho, M.H.; Chen, C.H.; Goodwin, J.S.; Wang, B.Y.; Xie, H. Functional advantages of Porphyromonas gingivalis vesicles. PLoS ONE 2015, 10, e0123448. [Google Scholar] [CrossRef]
- Choi, J.W.; Kim, S.C.; Hong, S.H.; Lee, H.J. Secretable Small RNAs via Outer Membrane Vesicles in Periodontal Pathogens. J. Dent. Res. 2017, 96, 458–466. [Google Scholar] [CrossRef]
- Fleetwood, A.J.; Lee, M.K.S.; Singleton, W.; Achuthan, A.; Lee, M.C.; O’Brien-Simpson, N.M.; Cook, A.D.; Murphy, A.J.; Dashper, S.G.; Reynolds, E.C.; et al. Metabolic remodeling, inflammasome activation, and pyroptosis in macrophages stimulated by Porphyromonas gingivalis and its outer membrane vesicles. Front. Cell. Infect. Microbiol. 2017, 7, 351. [Google Scholar] [CrossRef] [PubMed]
- Medzhitov, R. Toll-like receptors and innate immunity. Immunology 2001, 1, 135–145. [Google Scholar] [CrossRef] [PubMed]
- Schromm, A.B.; Brandenburg, K.; Loppnow, H.; Moran, A.P.; Koch, M.H.J.; Rietschel, E.T.; Seydel, U. Biological activities of lipopolysaccharides are determined by the shape of their lipid A portion. Eur. J. Biochem. 2000, 267, 2008–2013. [Google Scholar] [CrossRef] [PubMed]
- Curtis, M.A.; Percival, R.S.; Devine, D.; Darveau, R.P.; Coats, S.R.; Rangarajan, M.; Tarelli, E.; Marsh, P.D. Temperature-dependent modulation of Porphyromonas gingivalis lipid A structure and interaction with the innate host defenses. Infect. Immun. 2011, 79, 1187–1193. [Google Scholar] [CrossRef]
- Herath, T.D.K.; Wang, Y.; Seneviratne, C.J.; Darveau, R.P.; Wang, C.Y.; Jin, L. The expression and regulation of matrix metalloproteinase-3 is critically modulated by Porphyromonas gingivalis lipopolysaccharide with heterogeneous lipid A structures in human gingival fibroblasts. BMC Microbiol. 2013, 13, 73. [Google Scholar] [CrossRef]
- Papadopoulos, G.; Weinberg, E.O.; Massari, P.; Gibson, F.C.; Wetzler, L.M.; Morgan, E.F.; Genco, C.A. Macrophage-Specific TLR2 Signaling Mediates Pathogen-Induced TNF-Dependent Inflammatory Oral Bone Loss. J. Immunol. 2013, 190, 1148–1157. [Google Scholar] [CrossRef]
- Wang, P.L.; Ohura, K. Porphyromonas gingivalis lipopolysaccharide signaling in gingival fibroblasts—CD14 and toll-like receptors. Crit. Rev. Oral Biol. Med. 2002, 13, 132–142. [Google Scholar] [CrossRef]
- Ding, P.H.; Wang, C.Y.; Darveau, R.P.; Jin, L. Porphyromonas gingivalis LPS stimulates the expression of LPS-binding protein in human oral keratinocytes in vitro. Innate Immun. 2013, 19, 66–75. [Google Scholar] [CrossRef]
- De Diego, I.; Veillard, F.; Sztukowska, M.N.; Guevara, T.; Potempa, B.; Pomowski, A.; Huntington, J.A.; Potempa, J.; Gomis-Rüth, F.X. Structure and mechanism of cysteine peptidase gingipain K (Kgp), a major virulence factor of Porphyromonas gingivalis. J. Biol. Chem. 2014, 289, 32291–32302. [Google Scholar] [CrossRef]
- Karin Kristoffersen, A.; Solli, S.J.; Duy Nguyen, T.; Enersen, M. Association of the rgpB gingipain genotype to the major fimbriae ( fimA ) genotype in clinical isolates of the periodontal pathogen Porphyromonas gingivalis. J. Oral Microbiol. 2015, 7, 29124. [Google Scholar] [CrossRef]
- Tokuda, M.; Karunakaran, T.; Duncan, M.; Hamada, N.; Kuramitsu, H. Role of Arg-gingipain A in virulence of Porphyromonas gingivalis. Infect. Immun. 1998, 66, 1159–1166. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, H.; Sasaki, N.; Yamaga, S.; Kuboniwa, M.; Matsusaki, M.; Amano, A. Porphyromonas gingivalis induces penetration of lipopolysaccharide and peptidoglycan through the gingival epithelium via degradation of junctional adhesion molecule. PLoS Pathog. Pathog. 2019, 15, e1008124. [Google Scholar] [CrossRef] [PubMed]
- Carlisle, M.D.; Srikantha, R.N.; Brogden, K.A. Degradation of human α- And β-defensins by culture supernatants of Porphyromonas gingivalis strain 381. J. Innate Immun. 2009, 1, 118–122. [Google Scholar] [CrossRef] [PubMed]
- Maisetta, G.; Brancatisano, F.L.; Esin, S.; Campa, M.; Batoni, G. Gingipains produced by Porphyromonas gingivalis ATCC49417 degrade human-β-defensin 3 and affect peptide’s antibacterial activity in vitro. Peptides 2011, 32, 1073–1077. [Google Scholar] [CrossRef] [PubMed]
- Wilensky, A.; Tzach-Nahman, R.; Potempa, J.; Shapira, L.; Nussbaum, G. Porphyromonas gingivalis gingipains selectively reduce CD14 expression, leading to macrophage hyporesponsiveness to bacterial infection. J. Innate Immun. 2015, 7, 127–135. [Google Scholar] [CrossRef]
- Xu, X.; Tong, T.; Yang, X.; Pan, Y.; Lin, L.; Li, C. Differences in survival, virulence and biofilm formation between sialidase-deficient and W83 wild-type Porphyromonas gingivalis strains under stressful environmental conditions. BMC Microbiol. 2017, 17, 178. [Google Scholar] [CrossRef]
- Kreve, S.; Reis, A.C.D. Bacterial adhesion to biomaterials: What regulates this attachment? A review. Jpn. Dent. Sci. Rev. 2021, 57, 85–96. [Google Scholar] [CrossRef]
- Heidler, T.V.; Ernits, K.; Ziolkowska, A.; Claesson, R.; Persson, K. Porphyromonas gingivalis fimbrial protein Mfa5 contains a von Willebrand factor domain and an intramolecular isopeptide. Commun. Biol. 2021, 4, 106. [Google Scholar] [CrossRef]
- Meghil, M.M.; Ghaly, M.; Cutler, C.W. A Tale of Two Fimbriae: How Invasion of Dendritic Cells by Porphyromonas gingivalis Disrupts DC Maturation and Depolarizes the T-Cell-Mediated Immune Response. Pathogens 2022, 11, 328. [Google Scholar] [CrossRef]
- Chopra, A.; Bhat, S.G.; Sivaraman, K. Porphyromonas gingivalis adopts intricate and unique molecular mechanisms to survive and persist within the host: A critical update. J. Oral Microbiol. 2020, 12, 1801090. [Google Scholar] [CrossRef]
- Guzmán-Soto, I.; McTiernan, C.; Gonzalez-Gomez, M.; Ross, A.; Gupta, K.; Suuronen, E.J.; Mah, T.F.; Griffith, M.; Alarcon, E.I. Mimicking biofilm formation and development: Recent progress in in vitro and in vivo biofilm models. iScience 2021, 24, 102443. [Google Scholar] [CrossRef] [PubMed]
- Chathoth, K.; Fostier, L.; Martin, B.; Baysse, C.; Mahé, F. A Multi-Skilled Mathematical Model of Bacterial Attachment in Initiation of Biofilms. Microorganisms 2022, 10, 686. [Google Scholar] [CrossRef] [PubMed]
- Aliko, A.; Kamińska, M.; Bergum, B.; Gawron, K.; Benedyk, M.; Lamont, R.J.; Malicki, S.; Delaleu, N.; Potempa, J.; Mydel, P. Impact of Porphyromonas gingivalis Peptidylarginine Deiminase on Bacterial Biofilm Formation, Epithelial Cell Invasion, and Epithelial Cell Transcriptional Landscape. Sci. Rep. 2018, 8, 2–10. [Google Scholar] [CrossRef]
- Kalimuthu, S.; Cheung, B.P.K.; Yau, J.Y.Y.; Shanmugam, K.; Solomon, A.P.; Neelakantan, P. A novel small molecule, 1,3-di-m-tolyl-urea, inhibits and disrupts multispecies oral biofilms. Microorganisms 2020, 8, 1261. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Li, X.; Cheng, T.; Sun, H.; Jin, L. Eradication of Porphyromonas gingivalis Persisters Through Colloidal Bismuth Subcitrate Synergistically Combined with Metronidazole. Front. Microbiol. 2021, 12, 748121. [Google Scholar] [CrossRef] [PubMed]
- Cugini, C.; Shanmugam, M.; Landge, N.; Ramasubbu, N. The Role of Exopolysaccharides in Oral Biofilms. J. Dent. Res. 2019, 98, 739–745. [Google Scholar] [CrossRef] [PubMed]
- Davies, J.R.; Kad, T.; Neilands, J.; Kinnby, B.; Prgomet, Z.; Bengtsson, T.; Khalaf, H.; Svensäter, G. Polymicrobial synergy stimulates Porphyromonas gingivalis survival and gingipain expression in a multi-species subgingival community. BMC Oral Health 2021, 21, 639. [Google Scholar] [CrossRef]
- Plančak, D.; Musić, L.; Puhar, I. Quorum sensing of periodontal pathogens. Acta Stomatol. Croat. 2015, 49, 234–241. [Google Scholar] [CrossRef]
- Lamont, R.J.; Koo, H.; Hajishengallis, G. The oral microbiota: Dynamic communities and host interactions. Nat. Rev. Microbiol. 2018, 16, 745–759. [Google Scholar] [CrossRef]
- Zhu, B.; Macleod, L.C.; Newsome, E.; Liu, J.; Xu, P. Aggregatibacter actinomycetemcomitans mediates protection of Porphyromonas gingivalis from Streptococcus sanguinis hydrogen peroxide production in multi-species biofilms. Sci. Rep. 2019, 9, 4944. [Google Scholar] [CrossRef]
- Miller, D.P.; Hutcherson, J.A.; Wang, Y.; Nowakowska, Z.M.; Potempa, J.; Yoder-Himes, D.R.; Scott, D.A.; Whiteley, M.; Lamont, R.J. Genes contributing to Porphyromonas gingivalis fitness in abscess and epithelial cell colonization environments. Front. Cell. Infect. Microbiol. 2017, 7, 378. [Google Scholar] [CrossRef] [PubMed]
- Muhammad, M.H.; Idris, A.L.; Fan, X.; Guo, Y.; Yu, Y.; Jin, X.; Qiu, J.; Guan, X.; Huang, T. Beyond Risk: Bacterial Biofilms and Their Regulating Approaches. Front. Microbiol. 2020, 11, 928. [Google Scholar] [CrossRef] [PubMed]
- Rumbaugh, K.P.; Sauer, K. Biofilm dispersion. Nat. Rev. Microbiol. 2020, 18, 571. [Google Scholar] [CrossRef] [PubMed]
- Khatoon, Z.; McTiernan, C.D.; Suuronen, E.J.; Mah, T.F.; Alarcon, E.I. Bacterial biofilm formation on implantable devices and approaches to its treatment and prevention. Heliyon 2018, 4, e01067. [Google Scholar] [CrossRef]
- Talapko, J.; Matijević, T.; Juzbašić, M.; Antolović-Požgain, A.; Škrlec, I. Antibacterial Activity of Silver and Its Application in Dentistry, Cardiology and Dermatology. Microorganisms 2020, 8, 1400. [Google Scholar] [CrossRef] [PubMed]
- Conrads, G.; Klomp, T.; Deng, D.; Wenzler, J.S.; Braun, A.; Abdelbary, M.M.H. The Antimicrobial Susceptibility of Porphyromonas gingivalis: Genetic Repertoire, Global Phenotype, and Review of the Literature. Antibiotics 2021, 10, 1438. [Google Scholar] [CrossRef] [PubMed]
- Bi, Y.; Xia, G.; Shi, C.; Wan, J.; Liu, L.; Chen, Y.; Wu, Y.; Zhang, W.; Zhou, M.; He, H.; et al. Therapeutic strategies against bacterial biofilms. Fundam. Res. 2021, 1, 193–212. [Google Scholar] [CrossRef]
- Rams, T.E.; Sautter, J.D.; van Winkelhoff, A.J. Comparative in vitro resistance of human periodontal bacterial pathogens to tinidazole and four other antibiotics. Antibiotics 2020, 9, 68. [Google Scholar] [CrossRef] [PubMed]
- Talapko, J.; Škrlec, I. The Principles, Mechanisms, and Benefits of Unconventional Agents in the Treatment of Biofilm Infection. Pharmaceuticals 2020, 13, 299. [Google Scholar] [CrossRef]
- Zhou, N.; Zou, F.; Cheng, X.; Huang, Y.; Zou, H.; Niu, Q.; Qiu, Y.; Shan, F.; Luo, A.; Teng, W.; et al. Porphyromonas gingivalis induces periodontitis, causes immune imbalance, and promotes rheumatoid arthritis. J. Leukoc. Biol. 2021, 110, 461–473. [Google Scholar] [CrossRef]
- Andrian, E.; Grenier, D.; Rouabhia, M. Porphyromonas gingivalis-epithelial cell interactions in periodontitis. J. Dent. Res. 2006, 85, 392–403. [Google Scholar] [CrossRef] [PubMed]
- Sandros, J.; Papapanou, P.N.; Dahlen, G. Porphyromonas gingivalis invades oral epithelial cells in vitro. J. Periodont. Res. 1993, 28, 219–226. [Google Scholar] [CrossRef]
- Lamont, R.J.; Chan, A.; Belton, C.M.; Izutsu, K.T.; Vasel, D.; Weinberg, A. Porphyromonas gingivalis invasion of gingival epithelial cells. Infect. Immun. 1995, 63, 3878–3885. [Google Scholar] [CrossRef] [PubMed]
- Groeger, S.E.; Meyle, J. Epithelial barrier and oral bacterial infection. Periodontol. 2000 2015, 69, 46–67. [Google Scholar] [CrossRef]
- Andrian, E.; Grenier, D.; Rouabhia, M. In vitro models of tissue penetration and destruction by Porphyromonas gingivalis. Infect. Immun. 2004, 72, 4689–4698. [Google Scholar] [CrossRef] [PubMed]
- Nakao, R.; Takashiba, S.; Kosono, S.; Yoshida, M.; Watanabe, H.; Ohnishi, M.; Senpuku, H. Effect of Porphyromonas gingivalis outer membrane vesicles on gingipain-mediated detachment of cultured oral epithelial cells and immune responses. Microbes Infect. 2014, 16, 6–16. [Google Scholar] [CrossRef] [PubMed]
- Abe-Yutori, M.; Chikazawa, T.; Shibasaki, K.; Murakami, S. Decreased expression of E-cadherin by Porphyromonas gingivalis-lipopolysaccharide attenuates epithelial barrier function. J. Periodontal Res. 2017, 52, 42–50. [Google Scholar] [CrossRef]
- Chen, W.; Alshaikh, A.; Kim, S.; Kim, J.; Chun, C.; Mehrazarin, S.; Lee, J.; Lux, R.; Kim, R.H.; Shin, K.H.; et al. Porphyromonas gingivalis Impairs Oral Epithelial Barrier through Targeting GRHL2. J. Dent. Res. 2019, 98, 1150–1158. [Google Scholar] [CrossRef] [PubMed]
- Eskan, M.A.; Hajishengallis, G.; Kinane, D.F. Differential activation of human gingival epithelial cells and monocytes by Porphyromonas gingivalis fimbriae. Infect. Immun. 2007, 75, 892–898. [Google Scholar] [CrossRef] [PubMed]
- Yilmaz, Ö.; Watanabe, K.; Lamont, R.J. Involvement of integrins in fimbriae-mediated binding and invasion by Porphyromonas gingivalis. Cell. Microbiol. 2002, 4, 305–314. [Google Scholar] [CrossRef]
- Axe, E.L.; Walker, S.A.; Manifava, M.; Chandra, P.; Roderick, H.L.; Habermann, A.; Griffiths, G.; Ktistakis, N.T. Autophagosome formation from membrane compartments enriched in phosphatidylinositol 3-phosphate and dynamically connected to the endoplasmic reticulum. J. Cell Biol. 2008, 182, 685–701. [Google Scholar] [CrossRef] [PubMed]
- Leea, K.; Robertsb, J.A.S.; Choic, C.H.; Atanasova, K.R.; Yilmaz, O. Porphyromonas gingivalis traffics into endoplasmic reticulum-richautophagosomes for successful survival in human gingival epithelial cells. Virulence 2018, 9, 845–859. [Google Scholar] [CrossRef]
- Park, M.H.; Jeong, S.Y.; Na, H.S.; Chung, J. Porphyromonas gingivalis induces autophagy in THP-1-derived macrophages. Mol. Oral Microbiol. 2017, 32, 48–59. [Google Scholar] [CrossRef]
- El-Awady, A.R.; Miles, B.; Scisci, E.; Kurago, Z.B.; Palani, C.D.; Arce, R.M.; Waller, J.L.; Genco, C.A.; Slocum, C.; Manning, M.; et al. Porphyromonas gingivalis Evasion of Autophagy and Intracellular Killing by Human Myeloid Dendritic Cells Involves DC-SIGN-TLR2 Crosstalk. PLoS Pathog. 2015, 11, e1004647. [Google Scholar] [CrossRef] [PubMed]
- Dorn, B.R.; Dunn, J.; Progulske-Fox, A. Porphyromonas gingivalis traffics to autophagosomes in human coronary artery endothelial cells. Infect. Immun. 2001, 69, 5698–5708. [Google Scholar] [CrossRef]
- Hirasawa, M.; Kurita-Ochiai, T. Porphyromonas gingivalis induces apoptosis and autophagy via ER stress in human umbilical vein endothelial cells. Mediat. Inflamm. 2018, 2018, 1967506. [Google Scholar] [CrossRef]
- Gonzales, J.R.; Groeger, S.; Johansson, A.; Meyle, J. T helper cells from aggressive periodontitis patients produce higher levels of interleukin-1 beta and interleukin-6 in interaction with Porphyromonas gingivalis. Clin. Oral Investig. 2014, 18, 1835–1843. [Google Scholar] [CrossRef] [PubMed]
- Vernal, R.; Díaz-Zúñiga, J.; Melgar-Rodríguez, S.; Pujol, M.; Diaz-Guerra, E.; Silva, A.; Sanz, M.; Garcia-Sanz, J.A. Activation of RANKL-induced osteoclasts and memory T lymphocytes by Porphyromonas gingivalis is serotype dependant. J. Clin. Periodontol. 2014, 41, 451–459. [Google Scholar] [CrossRef] [PubMed]
- Salminen, A.; Gursoy, U.K.; Paju, S.; Hyvärinen, K.; Mäntylä, P.; Könönen, E.; Nieminen, M.S.; Sorsa, T.; Sinisalo, J.; Pussinen, P.J. Salivary biomarkers of bacterial burden, inflammatory response, and tissue destruction in periodontitis. J. Clin. Periodontol. 2014, 41, 442–450. [Google Scholar] [CrossRef] [PubMed]
- Trombone, A.P.; Claudino, M.; Colavite, P.; De Assis, G.F.; Avila-Campos, M.J.; Silva, J.S.; Campanelli, A.P.; Ibañez, O.M.; De Franco, M.; Garlet, G.P. Periodontitis and arthritis interaction in mice involves a shared hyper-inflammatory genotype and functional immunological interferences. Genes Immun. 2010, 11, 479–489. [Google Scholar] [CrossRef] [PubMed]
- Ricklin, D.; Hajishengallis, G.; Yang, K.; Lambris, J.D. Complement: A key system for immune surveillance and homeostasis. Nat. Immunol 2010, 11, 785–797. [Google Scholar] [CrossRef] [PubMed]
- Hajishengallis, G.; Harokopakis, E. Porphyromonas gingivalis interactions with complement receptor 3 (CR3): Innate immunity or immune evasion? Front. Biosci. 2007, 12, 4547–4557. [Google Scholar] [CrossRef] [PubMed]
- Hajishengallis, G.; Lambris, J.D. Complement and dysbiosis in periodontal disease. Immunobiology 2012, 217, 1111–1116. [Google Scholar] [CrossRef] [PubMed]
- Huynh, A.H.S.; Veith, P.D.; Mcgregor, N.R.; Adams, G.G.; Chen, D.; Reynolds, E.C.; Ngo, L.H.; Darby, I.B. Gingival crevicular fluid proteomes in health, gingivitis and chronic periodontitis. J. Periodontal Res. 2015, 50, 637–649. [Google Scholar] [CrossRef] [PubMed]
- Hajishengallis, G.; Liang, S.; Payne, M.A.; Hashim, A.; Eskan, M.A.; Mcintosh, M.L.; Alsam, A.; Kirkwood, K.L.; Lambris, J.D.; Darveau, R.P.; et al. A Low-Abundance Biofilm Species Orchestrates Inflammatory Periodontal Disease through the Commensal Microbiota and the Complement Pathway. Cell Host Microbe 2011, 10, 497–506. [Google Scholar] [CrossRef] [PubMed]
- Olsen, I.; Lambris, J.D.; Hajishengallis, G. Porphyromonas gingivalis disturbs host–commensal homeostasis by changing complement function. J. Oral Microbiol. 2017, 9, 1340085. [Google Scholar] [CrossRef] [PubMed]
- Bielecka, E.; Scavenius, C.; Kantyka, T.; Jusko, M.; Mizgalska, D.; Szmigielski, B.; Potempa, B.; Enghild, J.J.; Prossnitz, E.R.; Blom, A.M.; et al. Peptidyl arginine deiminase from Porphyromonas gingivalis abolishes anaphylatoxin C5a activity. J. Biol. Chem. 2014, 289, 32481–32487. [Google Scholar] [CrossRef] [PubMed]
- Makkawi, H.; Hoch, S.; Burns, E.; Hosur, K.; Hajishengallis, G.; Kirschning, C.J.; Nussbaum, G. Porphyromonas gingivalis stimulates TLR2-PI3K signaling to escape immune clearance and induce bone resorption independently of MyD88. Front. Cell. Infect. Microbiol. 2017, 7, 359. [Google Scholar] [CrossRef] [PubMed]
- Bryzek, D.; Ciaston, I.; Dobosz, E.; Gasiorek, A.; Makarska, A.; Sarna, M.; Eickid, S.; Puklo, M.; Lech, M.; Potempa, B.; et al. Triggering NETosis via protease-activated receptor (PAR)-2 signaling as a mechanism of hijacking neutrophils function for pathogen benefits. PLoS Pathog. 2019, 15, e1007773. [Google Scholar] [CrossRef] [PubMed]
- Azzouz, L.; Cherry, A.; Riedl, M.; Khan, M.; Pluthero, F.G.; Kahr, W.H.A.; Palaniyar, N.; Licht, C. Relative antibacterial functions of complement and NETs: NETs trap and complement effectively kills bacteria. Mol. Immunol. 2018, 97, 71–81. [Google Scholar] [CrossRef] [PubMed]
- Radwan-Oczko, M.; Jaworski, A.; Duś, I.; Plonek, T.; Szulc, M.; Kustrzycki, W. Porphyromonas gingivalis in periodontal pockets and heart valves. Virulence 2014, 5, 575–580. [Google Scholar] [CrossRef] [PubMed]
- Czerniuk, M.R.; Surma, S.; Romańczyk, M.; Nowak, J.M.; Wojtowicz, A.; Filipiak, K.J. Unexpected Relationships: Periodontal Diseases: Atherosclerosis–Plaque Destabilization? From the Teeth to a Coronary Event. Biology 2022, 11, 272. [Google Scholar] [CrossRef] [PubMed]
- Bale, B.F.; Doneen, A.L.; Vigerust, D.J. High-Risk periodontal pathogens contribute to the pathogenesis of atherosclerosis. Postgrad. Med. J. 2017, 93, 215–220. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Xie, M.; Huang, X.; Chen, G.; Yin, Y.; Lu, X.; Feng, G.; Yu, R.; Chen, L. The Effects of Porphyromonas gingivalis on Atherosclerosis-Related Cells. Front. Immunol. 2021, 12, 766560. [Google Scholar] [CrossRef] [PubMed]
- Horliana, A.C.R.T.; Chambrone, L.; Foz, A.M.; Artese, H.P.C.; De Rabelo, M.S.; Pannuti, C.M.; Romito, G.A. Dissemination of periodontal pathogens in the bloodstream after periodontal procedures: A systematic review. PLoS ONE 2014, 9, 21–23. [Google Scholar] [CrossRef] [PubMed]
- Del Giudice, C.; Vaia, E.; Liccardo, D.; Marzano, F.; Valletta, A.; Spagnuolo, G.; Ferrara, N.; Rengo, C.; Cannavo, A.; Rengo, G. Infective endocarditis: A focus on oral microbiota. Microorganisms 2021, 9, 1218. [Google Scholar] [CrossRef]
- Miyauchi, S.; Tokuyama, T.; Shintani, T.; Uotani, Y.; Miyamoto, S.; Ikeuchi, Y.; Okamura, S.; Okubo, Y.; Miyauchi, M.; Nakano, Y. Porphyromonas gingivalis and left atrial appendage spontaneous echo contrast in atrial fibrillation ablation candidates. Heart Vessels 2021, 36, 1721–1729. [Google Scholar] [CrossRef] [PubMed]
- Shiheido, Y.; Maejima, Y.; Suzuki, J.-I.; Aoyama, N.; Kaneko, M.; Watanabe, R.; Sakamaki, Y.; Wakayama, K.; Ikeda, Y.; Akazawa, H.; et al. Porphyromonas gingivalis, a periodontal pathogen, enhances myocardial vulnerability, thereby promoting post-infarct cardiac rupture. J. Mol. Cell. Cardiol. 2016, 99, 123–137. [Google Scholar] [CrossRef]
- Matsuo, I.; Ohnuki, Y.; Suita, K.; Ishikawa, M.; Mototani, Y.; Ito, A.; Hayakawa, Y.; Nariyama, M.; Morii, A.; Kiyomoto, K.; et al. Effects of chronic Porphylomonas gingivalis lipopolysaccharide infusion on cardiac dysfunction in mice. J. Oral Biosci. 2021, 63, 394–400. [Google Scholar] [CrossRef]
- Li, J.; Deng, J.; Shang, S.; Liu, G.; Song, W.; Sun, P.; Jiang, W.; Pan, K. Effect of Porphyromonas gingivalis lipopolysaccharide on calcification of human umbilical artery smooth muscle cells co-cultured with human periodontal ligament cells. Exp. Ther. Med. 2021, 21, 655. [Google Scholar] [CrossRef]
- Sampath, C.; Okoro, E.U.; Gipson, M.J.; Chukkapalli, S.S.; Farmer-Dixon, C.M.; Gangula, P.R. Porphyromonas gingivalis infection alters Nrf2-Phase II enzymes and nitric oxide in primary human aortic endothelial cells. J. Periodontol. 2021, 92, 54–65. [Google Scholar] [CrossRef]
- Xie, M.; Tang, Q.; Yu, S.; Sun, J.; Mei, F.; Zhao, J.; Chen, L. Porphyromonas gingivalis disrupts vascular endothelial homeostasis in a TLR-NF-κB axis dependent manner. Int. J. Oral Sci. 2020, 12, 28. [Google Scholar] [CrossRef] [PubMed]
- Borén, J.; John Chapman, M.; Krauss, R.M.; Packard, C.J.; Bentzon, J.F.; Binder, C.J.; Daemen, M.J.; Demer, L.L.; Hegele, R.A.; Nicholls, S.J.; et al. Low-density lipoproteins cause atherosclerotic cardiovascular disease: Pathophysiological, genetic, and therapeutic insights: A consensus statement from the European Atherosclerosis Society Consensus Panel. Eur. Heart J. 2020, 41, 2313–2330. [Google Scholar] [CrossRef] [PubMed]
- Lönn, J.; Ljunggren, S.; Klarström-Engström, K.; Demirel, I.; Bengtsson, T.; Karlsson, H. Lipoprotein modifications by gingipains of Porphyromonas gingivalis. J. Periodontal Res. 2018, 53, 403–413. [Google Scholar] [CrossRef] [PubMed]
- Groeger, S.; Jarzina, F.; Domann, E.; Meyle, J. Porphyromonas gingivalis activates NFΚB and MAPK pathways in human oral epithelial cells. BMC Immunol. 2017, 18, 1. [Google Scholar] [CrossRef]
- Hajishengallis, G.; Sojar, H.; Genco, R.J.; DeNardin, E. Intracellular signaling and cytokine induction upon interactions of Porphyromonas gingivalis fimbriae with pattern-recognition receptors. Immunol. Investig. 2004, 33, 157–172. [Google Scholar] [CrossRef]
- Huynh, J.; Scholz, G.M.; Aw, J.; Kwa, M.Q.; Achuthan, A.; Hamilton, J.A.; Reynolds, E.C. IRF6 Regulates the Expression of IL-36γ by Human Oral Epithelial Cells in Response to Porphyromonas gingivalis. J. Immunol. 2016, 196, 2230–2238. [Google Scholar] [CrossRef]
- Sandros, J.; Karlsson, C.; Lappin, D.F.; Madianos, P.M.; Kinane, D.F.; Papapanou, P.M. Cytokine responses of oral epithelial cells to Porphyromonas gingivalis infection. J. Dent. Res. 2000, 79, 1808–1814. [Google Scholar] [CrossRef]
- Kim, H.J.; Cha, G.S.; Kim, H.J.; Kwon, E.Y.; Lee, J.Y.; Choi, J.; Joo, J.Y. Porphyromonas gingivalis accelerates atherosclerosis through oxidation of high-density lipoprotein. J. Periodontal Implant Sci. 2018, 48, 60–68. [Google Scholar] [CrossRef]
- Gibson, M.S.; Domingues, N.; Vieira, O.V. Lipid and non-lipid factors affecting macrophage dysfunction and inflammation in atherosclerosis. Front. Physiol. 2018, 9, 654. [Google Scholar] [CrossRef]
- Lorkowski, S.W.; Smith, J.D. HDL Is Not Dead Yet. Biomedicines 2022, 10, 128. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Hao, T.; Liu, Y.; Huang, J.; Wu, W.; Wu, J.; Sun, W. Th17/Treg balance and indoleamine 2,3 dioxygenase activity in periodontitis-associated atherosclerotic patients. J. Int. Med. Res. 2022, 50, 3000605221080877. [Google Scholar] [CrossRef] [PubMed]
- Metcalfe, S.; Anselmi, N.; Escobar, A.; Visser, M.B.; Kay, J.G. Innate Phagocyte Polarization in the Oral Cavity. Front. Immunol. 2022, 12, 768479. [Google Scholar] [CrossRef] [PubMed]
- Guerrini, V.; Gennaro, M.L. Foam cells: One size doesn’t fit all. Trends Immunol. 2019, 40, 1163–1179. [Google Scholar] [CrossRef]
- Kozarov, E.; Progulske-Fox, A. Atherosclerosis microbiome: Upcoming target for vaccine and drug development. Vessel Plus 2020, 4, 10. [Google Scholar] [CrossRef]
- Schenkein, H.A.; Papapanou, P.N.; Genco, R.; Sanz, M. Mechanisms underlying the association between periodontitis and atherosclerotic disease. Periodontol. 2000 2020, 83, 90–106. [Google Scholar] [CrossRef]
- Charoensaensuk, V.; Chen, Y.; Lin, Y.; Ou, K.; Yang, L.; Lu, D. Porphyromonas gingivalis Induces Proinflammatory Cytokine Expression Leading to Apoptotic Death through the Oxidative Stress/NF-κB Pathway in Brain Endothelial Cells. Cells 2021, 10, 3033. [Google Scholar] [CrossRef]

| Virulence Factors | Function | Ref |
|---|---|---|
| Fimbriae | Fimbriae promote biofilm formation, bacterial motility, adhesion, and invasion of host cells | [2,39,40] |
| Hemolysin | Serves to supply heme | [48] |
| Hemagglutinin | Serves to supply heme | [48] |
| Capsule | Encapsulation is associated with increased resistance to phagocytosis | [2,21] |
| Outer membrane vesicles (OMVs) | They are involved in the adaptation of bacteria to stress, nutrient metabolism, and communication with other periodontogens, and also host cells | [49] |
| Lipopolysaccharides (LPS) | Have the ability to cause an inflammatory reaction | [2] |
| Gingipains | Make up 85% of total proteolytic activity | [2,29,50] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aleksijević, L.H.; Aleksijević, M.; Škrlec, I.; Šram, M.; Šram, M.; Talapko, J. Porphyromonas gingivalis Virulence Factors and Clinical Significance in Periodontal Disease and Coronary Artery Diseases. Pathogens 2022, 11, 1173. https://doi.org/10.3390/pathogens11101173
Aleksijević LH, Aleksijević M, Škrlec I, Šram M, Šram M, Talapko J. Porphyromonas gingivalis Virulence Factors and Clinical Significance in Periodontal Disease and Coronary Artery Diseases. Pathogens. 2022; 11(10):1173. https://doi.org/10.3390/pathogens11101173
Chicago/Turabian StyleAleksijević, Lorena Horvat, Marko Aleksijević, Ivana Škrlec, Marko Šram, Miroslav Šram, and Jasminka Talapko. 2022. "Porphyromonas gingivalis Virulence Factors and Clinical Significance in Periodontal Disease and Coronary Artery Diseases" Pathogens 11, no. 10: 1173. https://doi.org/10.3390/pathogens11101173
APA StyleAleksijević, L. H., Aleksijević, M., Škrlec, I., Šram, M., Šram, M., & Talapko, J. (2022). Porphyromonas gingivalis Virulence Factors and Clinical Significance in Periodontal Disease and Coronary Artery Diseases. Pathogens, 11(10), 1173. https://doi.org/10.3390/pathogens11101173








