Prevalence, Risk Factors, and Molecular Detection of Campylobacter in Farmed Cattle of Selected Districts in Bangladesh
Abstract
1. Introduction
2. Results
2.1. Dairy Farm Management Descriptive Statistics
2.2. Prevalence of Campylobacter spp.
2.2.1. Farm-Level Prevalence
2.2.2. Sample-Level Prevalence
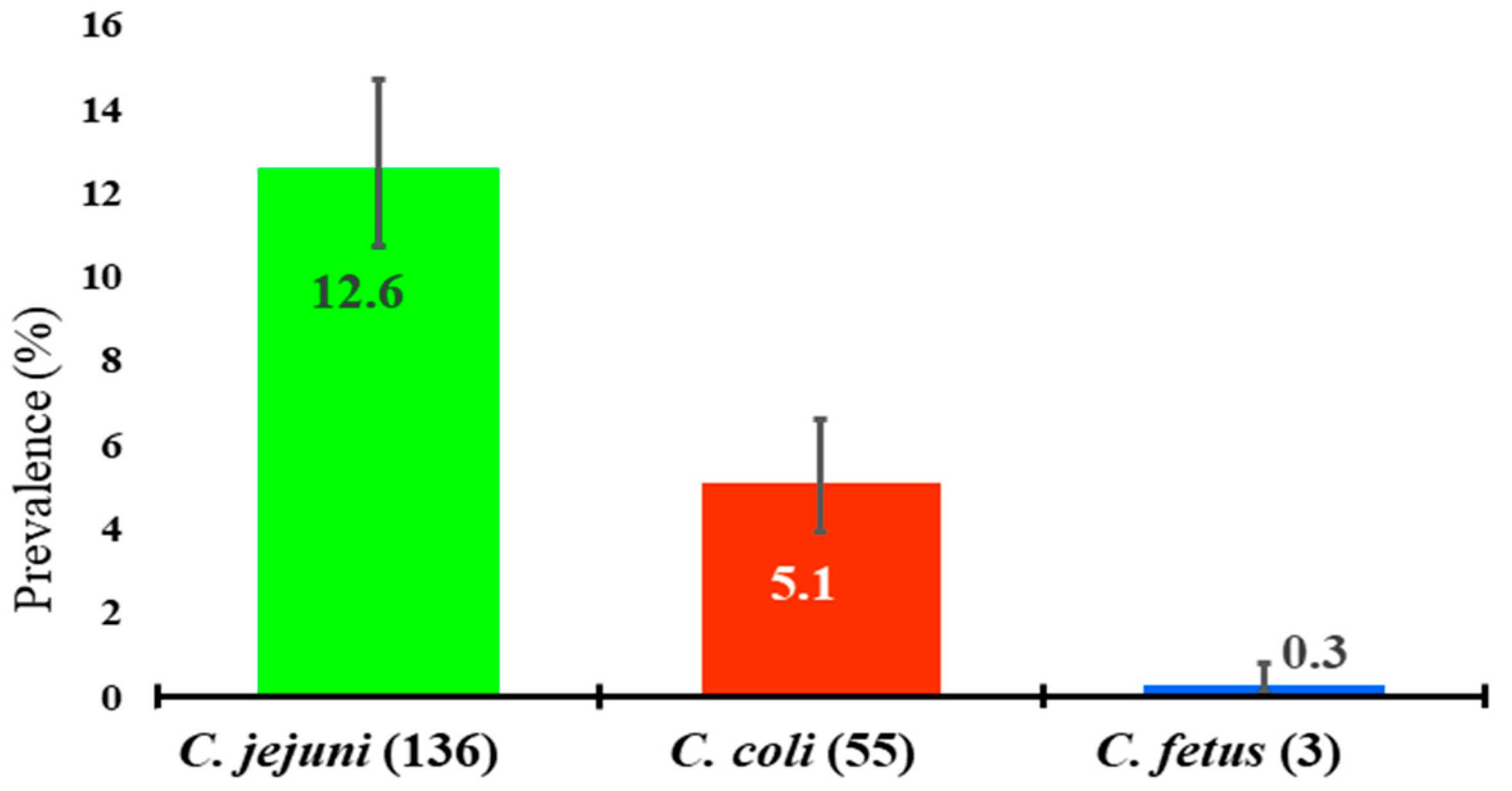
2.3. Molecular Detection of Campylobacter spp.
2.4. Evaluation of Risk Factors
2.4.1. Univariate Analysis
2.4.2. Multivariate Analysis
3. Discussion
4. Materials and Methods
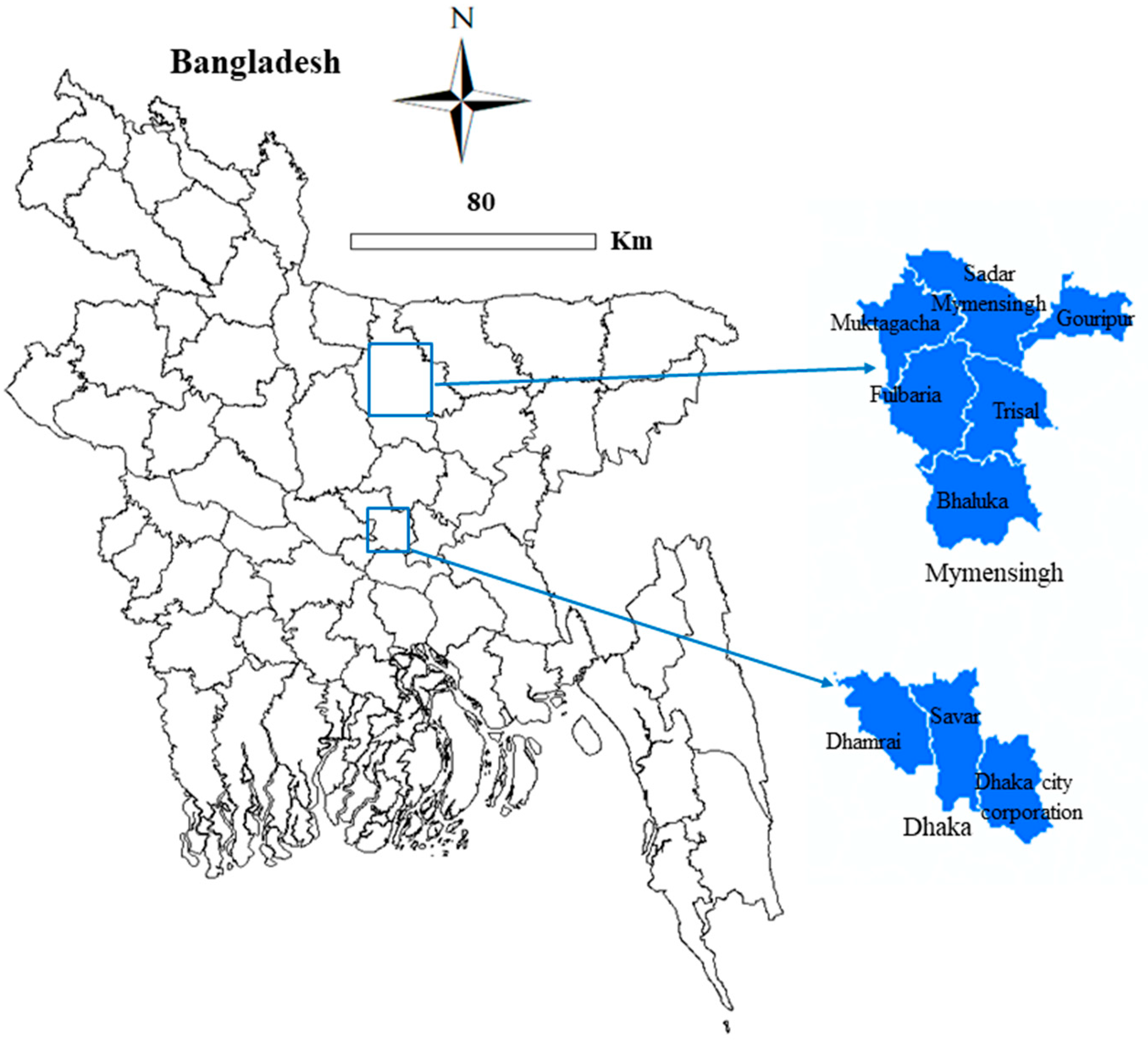
4.1. Study Location, Design, and Survey Farms
4.2. Face-to-Face Data Collection in Field Survey
4.3. Sample Size and Sampling Procedure
4.3.1. Sample Size Calculation
4.3.2. Sample Collection from Animals
4.4. Laboratory Evaluation
4.4.1. Culture and Biochemical Tests
4.4.2. Molecular Detection through PCR
4.4.3. Sequencing of 16S rRNA Gene
4.5. Statistical Evaluation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kirk, M.D.; Pires, S.M.; Black, R.E.; Caipo, M.; Crump, J.A.; Devleesschauwer, B.; Döpfer, D.; Fazil, A.; Fischer-Walker, C.L.; Hald, T. World Health Organization estimates of the global and regional disease burden of 22 foodborne bacterial, protozoal, and viral diseases, 2010: A data synthesis. PLoS Med. 2015, 12, e1001921. [Google Scholar] [CrossRef]
- Tack, D.M.; Marder, E.P.; Griffin, P.M.; Cieslak, P.R.; Dunn, J.; Hurd, S.; Scallan, E.; Lathrop, S.; Muse, A.; Ryan, P. Preliminary incidence and trends of infections with pathogens transmitted commonly through food—Foodborne Diseases Active Surveillance Network, 10 US Sites, 2015–2018. Morb. Mortal. Wkly. Rep. 2019, 68, 369. [Google Scholar] [CrossRef]
- CDC. Antibiotic Resistance Threats in the United States; U.S. Department of Health and Human Services, Ed.; CDC: Atlanta, GA, USA, 2019. [Google Scholar]
- Mäesaar, M.; Tedersoo, T.; Meremäe, K.; Roasto, M. The source attribution analysis revealed the prevalent role of poultry over cattle and wild birds in human campylobacteriosis cases in the Baltic States. PLoS ONE 2020, 15, e0235841. [Google Scholar] [CrossRef] [PubMed]
- Sahin, O.; Yaeger, M.; Wu, Z.; Zhang, Q. Campylobacter-associated diseases in animals. Annu. Rev. Anim. Biosci. 2017, 5, 21–42. [Google Scholar] [CrossRef]
- Manyi-Loh, C.E.; Mamphweli, S.N.; Meyer, E.L.; Makaka, G.; Simon, M.; Okoh, A.I. An overview of the control of bacterial pathogens in cattle manure. Int. J. Environ. Res. Public Health 2016, 13, 843. [Google Scholar] [CrossRef]
- Ogden, I.D.; Dallas, J.F.; MacRae, M.; Rotariu, O.; Reay, K.W.; Leitch, M.; Thomson, A.P.; Sheppard, S.K.; Maiden, M.; Forbes, K.J. Campylobacter excreted into the environment by animal sources: Prevalence, concentration shed, and host association. Foodborne Pathog. Dis. 2009, 6, 1161–1170. [Google Scholar] [CrossRef]
- Quinn, J. Clinical strategies for serious infection: A North American perspective. Diagn. Microbiol. Infect. Dis. 1998, 31, 389–395. [Google Scholar] [CrossRef]
- Friedman, C.R.J.; Neiman, H.C.; Wegener, R.V. Tauxe. Epidemiology of Campylobacter Jejuni Infections in the United States and Other Industrialised Nation; ASM Press: Washington, DC, USA, 2000. [Google Scholar]
- The European Union summary report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks in 2013. EFSA J. 2015, 13, 3991. [CrossRef]
- Gillespie, I.A.; O’Brien, S.J.; Frost, J.A.; Adak, G.K.; Horby, P.; Swan, A.V.; Painter, M.J.; Neal, K.R.; Collaborators, C.S.S.S. A case-case comparison of Campylobacter coli and Campylobacter jejuni infection: A tool for generating hypotheses. Emerg. Infect. Dis. 2011, 8, 937–942. [Google Scholar] [CrossRef]
- Bullman, S.; Corcoran, D.; O’Leary, J.; O’Hare, D.; Lucey, B.; Sleator, R.D. Emerging dynamics of human campylobacteriosis in Southern Ireland. FEMS Immunol. Med. Mic. 2011, 63, 248–253. [Google Scholar] [CrossRef]
- Wassenaar, T.M.; Newell, D.G. Genotyping of Campylobacter spp. Appl. Environ. Microbiol. 2000, 66, 1–9. [Google Scholar] [CrossRef]
- Patton, C.; Wachsmuth, I.; Evins, G.; Kiehlbauch, J.; Plikaytis, B.; Troup, N.; Tompkins, L.; Lior, H. Evaluation of 10 methods to distinguish epidemic-associated Campylobacter strains. J. Clin. Microbiol. 1991, 29, 680–688. [Google Scholar] [CrossRef] [PubMed]
- Al Amri, A.; Senok, A.C.; Ismaeel, A.Y.; Al-Mahmeed, A.E.; Botta, G.A. Multiplex PCR for direct identification of Campylobacter spp. in human and chicken stools. J. Med. Microbiol. 2007, 56, 1350–1355. [Google Scholar] [CrossRef]
- Kabir, S.M.L.; Kikuchi, K.; Asakura, M.; Shiramaru, S.; Tsuruoka, N.; Goto, A.; Hinenoya, A.; Yamasaki, S. Evaluation of a cytolethal distending toxin (cdt) gene-based species-specific multiplex PCR assay for the identification of Campylobacter strains isolated from diarrheal patients in Japan. Jpn. J. Infect. Dis. 2011, 64, 19–27. [Google Scholar]
- Asakura, M.; Samosornsuk, W.; Taguchi, M.; Kobayashi, K.; Misawa, N.; Kusumoto, M.; Nishimura, K.; Matsuhisa, A.; Yamasaki, S. Comparative analysis of cytolethal distending toxin (cdt) genes among Campylobacter jejuni, C. coli and C. fetus strains. Microb. Pathog. 2007, 42, 174–183. [Google Scholar] [CrossRef] [PubMed]
- Samosornsuk, W.; Asakura, M.; Yoshida, E.; Taguchi, T.; Nishimura, K.; Eampokalap, B.; Phongsisay, V.; Chaicumpa, W.; Yamasaki, S. Evaluation of a cytolethal distending toxin (cdt) gene-based species-specific multiplex PCR assay for the identification of Campylobacter strains isolated from poultry in Thailand. Microbiol. Immunol. 2007, 51, 909–917. [Google Scholar] [CrossRef]
- Berthenet, E.; Thépault, A.; Chemaly, M.; Rivoal, K.; Ducournau, A.; Buissonnière, A.; Bénéjat, L.; Bessède, E.; Mégraud, F.; Sheppard, S.K.; et al. Source attribution of Campylobacter jejuni shows variable importance of chicken and ruminants reservoirs in non-invasive and invasive French clinical isolates. Sci. Rep. 2019, 9, 8098. [Google Scholar] [CrossRef]
- EFSA Panel on Biological Hazards (BIOHAZ). Scientific Opinion on Quantification of the risk posed by broiler meat to human campylobacteriosis in the EU. EFSA J. 2010, 8, 1437. [Google Scholar] [CrossRef]
- Kaakoush, N.O.; Castaño-Rodríguez, N.; Mitchell, H.M.; Man, S.M. Global epidemiology of Campylobacter infection. Clin. Microbiol. Rev. 2015, 28, 687–720. [Google Scholar] [CrossRef]
- Nichols, G.L.; Richardson, J.F.; Sheppard, S.K.; Lane, C.; Sarran, C. Campylobacter epidemiology: A descriptive study reviewing 1 million cases in England and Wales between 1989 and 2011. BMJ Open 2012, 2, e001179. [Google Scholar] [CrossRef] [PubMed]
- Sheppard, S.K.; Dallas, J.F.; Strachan, N.J.; MacRae, M.; McCarthy, N.D.; Wilson, D.J.; Gormley, F.J.; Falush, D.; Ogden, I.D.; Maiden, M.C. Campylobacter genotyping to determine the source of human infection. Clin. Infect. Dis. 2009, 48, 1072–1078. [Google Scholar] [CrossRef]
- Thépault, A.; Méric, G.; Rivoal, K.; Pascoe, B.; Mageiros, L.; Touzain, F.; Rose, V.; Béven, V.; Chemaly, M.; Sheppard, S.K. Genome-wide identification of host-segregating epidemiological markers for source attribution in Campylobacter jejuni. Appl. Environ. Microbiol. 2017, 83, e03085-16. [Google Scholar] [CrossRef] [PubMed]
- Hlashwayo, D.F.; Sigaúque, B.; Bila, C.G. Epidemiology and antimicrobial resistance of Campylobacter spp. in animals in Sub-Saharan Africa: A Systemic Review. Heliyon 2020, 6, e03537. [Google Scholar] [CrossRef] [PubMed]
- Hannon, S.J.; Allan, B.; Waldner, C.; Russell, M.L.; Potter, A.; Babiuk, L.A.; Townsend, H.G. Prevalence and risk factor investigation of Campylobacter species in beef cattle feces from seven large commercial feedlots in Alberta, Canada. Can. J. Vet. Res. 2009, 73, 275–282. [Google Scholar] [PubMed]
- Wesley, I.; Wells, S.; Harmon, K.; Green, A.; Schroeder-Tucker, L.; Glover, M.; Siddique, I. Fecal shedding of Campylobacter and Arcobacter spp. in dairy cattle. Appl. Environ. Microbiol. 2000, 66, 1994–2000. [Google Scholar] [CrossRef] [PubMed]
- Klein, D.; Alispahic, M.; Sofka, D.; Iwersen, M.; Drillich, M.; Hilbert, F. Prevalence and risk factors for shedding of thermophilic Campylobacter in calves with and without diarrhea in Austrian dairy herds. J. Dairy Sci. 2013, 96, 1203–1210. [Google Scholar] [CrossRef] [PubMed]
- Kashoma, I.P.; Kassem, I.I.; Kumar, A.; Kessy, B.M.; Gebreyes, W.; Kazwala, R.R.; Rajashekara, G. Antimicrobial resistance and genotypic diversity of Campylobacter isolated from pigs, dairy, and beef cattle in Tanzania. Front. Microbiol. 2015, 6, 1240. [Google Scholar] [CrossRef]
- Blaser, M.J.; Taylor, D.N.; Feldman, R.A. Epidemiology of Campylobacter jejuni infections. Epidemiol. Rev. 1983, 5, 157–176. [Google Scholar] [CrossRef]
- Stanley, K.N.; Wallace, J.S.; Currie, J.E.; Diggle, P.J.; Jones, K. The seasonal variation of thermophilic Campylobacters in beef cattle, dairy cattle and calves. J. Appl. Microbiol. 1998, 85, 472–480. [Google Scholar] [CrossRef]
- Stampi, S.; Varoli, O.; Zanetti, F.; De Luca, G. Arcobacter cryaerophilus and thermophilic Campylobacters in a sewage treatment plant in Italy: Two secondary treatments compared. Epidemiol. Infect. 1993, 110, 633–639. [Google Scholar] [CrossRef] [PubMed]
- Tauxe, R.V. Epidemiology of Campylobacter jejuni infections in the United States and other industrialized nations. In Campylobacter jejuni: Current Status; Nachamkin, I., Blaser, M.J., Tompkins, L.S., Eds.; American Society for Microbiology (ASM) Press: Washington, DC, USA, 1992; pp. 9–16. [Google Scholar]
- National Livestock Development Policy, Department of Livestock Services, Ministry of Fisheries and Livestock, Peoples Republic of Bangladesh. 2007. Available online: http://old.dls.gov.bd/files/Livestock_Policy_Final.pdf (accessed on 29 September 2020).
- Livestock Economy at a Glance 2019–2020. Department of Livestock Services, Ministry of Fisheries and Livestock, Government of the People’s Republic of Bangladesh. 2020. Available online: http://dls.portal.gov.bd/sites/default/files/files/dls.portal.gov.bd/page/ee5f4621_fa3a_40ac_8bd9_898fb8ee4700/2020-07-22-19-34-e4cd5ed65f45419ee038e00b8939c1a0.pdf (accessed on 29 December 2020).
- Mourkas, E.; Taylor, A.J.; Méric, G.; Bayliss, S.C.; Pascoe, B.; Mageiros, L.; Calland, J.K.; Hitchings, M.D.; Ridley, A.; Vidal, A.; et al. Agricultural intensification and the evolution of host specialism in the enteric pathogen Campylobacter jejuni. Proc. Natl. Acad. Sci. USA 2020, 117, 11018–11028. [Google Scholar] [CrossRef]
- Batut, B.; Knibbe, C.; Marais, G.; Daubin, V. Reductive genome evolution at both ends of the bacterial population size spectrum. Nat. Rev. Microbiol. 2014, 12, 841–850. [Google Scholar] [CrossRef] [PubMed]
- Ocejo, M.; Oporto, B.; Hurtado, A. Occurrence of Campylobacter jejuni and Campylobacter coli in cattle and sheep in Northern Spain and changes in antimicrobial resistance in two studies 10-years apart. Pathogens 2019, 8, 98. [Google Scholar] [CrossRef] [PubMed]
- Kabir, S.M.L.; Lubna, M.M.; Islam, M.; Haque, A.K.M.Z.; Neogi, S.B.; Yamasaki, S. Isolation, molecular identification and antimicrobial resistance patterns of Campylobacter species of dairy origin: First report from Bangladesh. Vet. Sci. Dev. 2018, 8, 7838. [Google Scholar] [CrossRef]
- Mohakud, N.K.; Patra, S.D.; Kumar, S.; Sahu, P.S.; Misra, N.; Shrivastava, A.K. Detection and molecular typing of campylobacter isolates from human and animal faeces in coastal belt of Odisha, India. Indian J. Med. Microbiol. 2019, 37, 345–350. [Google Scholar] [CrossRef]
- Thépault, A.; Poezevara, T.; Quesne, S.; Rose, V.; Chemaly, M.; Rivoal, K. Prevalence of thermophilic Campylobacter in cattle production at slaughterhouse level in France and link between C. jejuni bovine strains and campylobacteriosis. Front. Microbiol. 2018, 9, 471. [Google Scholar] [CrossRef]
- Hansson, I.; Engvall, E.O.; Ferrari, S.; Harbom, B.; Lahti, E. Detection of Campylobacter species in different types of samples from dairy farms. Vet. Rec. 2020, 186, 605. [Google Scholar] [CrossRef] [PubMed]
- Oporto, B.; Esteban, J.; Aduriz, G.; Juste, R.; Hurtado, A. Prevalence and strain diversity of thermophilic campylobacters in cattle, sheep and swine farms. J. Appl. Microbiol. 2007, 103, 977–984. [Google Scholar] [CrossRef]
- Ramonaitė, S.; Rokaitytė, A.; Tamulevičienė, E.; Malakauskas, A.; Alter, T.; Malakauskas, M. Prevalence, quantitative load and genetic diversity of Campylobacter spp. in dairy cattle herds in Lithuania. Acta. Vet. Scand. 2013, 55, 87. [Google Scholar] [CrossRef]
- Padungtod, P.; Kaneene, J.B. Campylobacter in food animals and humans in northern Thailand. J. Food Prot. 2005, 68, 2519–2526. [Google Scholar] [CrossRef] [PubMed]
- Boonmar, S.; Chanda, C.; Markvichitr, K.; Chaunchom, S.; Yingsakmongkon, S.; Yamamot, S.; Morita, Y. Prevalence of Campylobacter spp. in slaughtered cattle and buffaloes in Vientiane, Lao People’s Democratic Republic. J. Vet. Med. Sci. 2007, 69, 853–855. [Google Scholar] [CrossRef][Green Version]
- An, J.-U.; Ho, H.; Kim, J.; Kim, W.H.; Kim, J.; Lee, S.; Mun, S.H.; Guk, J.H.; Hong, S.; Cho, S. Dairy cattle, a potential reservoir of human campylobacteriosis: Epidemiological and molecular characterization of Campylobacter jejuni from cattle farms. Front. Microbiol. 2018, 9, 3136. [Google Scholar] [CrossRef]
- Membre, J.M.; Laroche, M.; Magras, C. Meta-analysis of Campylobacter spp. Survival Data within a Temperature Range of 0 to 42 °C. J. Food Prot. 2013, 76, 1726–1732. [Google Scholar] [CrossRef]
- Inglis, G.D.; McAllister, T.A.; Larney, F.J.; Topp, E. Prolonged survival of Campylobacter species in bovine manure compost. Appl. Environ. Microbiol. 2010, 76, 1110–1119. [Google Scholar] [CrossRef]
- Guan, T.Y.; Holley, R.A. Pathogen survival in swine manure environments and transmission of human enteric illness--a review. J Environ Qual. 2003, 32, 383–392. [Google Scholar] [CrossRef]
- Flink, C.N.K.F. A sjukdomsframkallande bakterier I opastoriserad mjolk(in Swedish), (presence of pathogenic bacteria in unpasteurized). Swed. Food Agency Rep. 2016, 12. Available online: https://www.livsmedelsverket.se/globalassets/publikationsdatabas/rapporter/2016/forekomst-av-sjukdomsframkallande-bakterier-i-opastoriserad-mjolk_rapport-12_2016.pdf (accessed on 29 December 2020).
- Heuvelink, A.E.; van Heerwaarden, C.; Zwartkruis-Nahuis, A.; Tilburg, J.J.; Bos, M.H.; Heilmann, F.G.; Hofhuis, A.; Hoekstra, T.; de Boer, E. Two outbreaks of campylobacteriosis associated with the consumption of raw cows’ milk. Int. J. Food Microbiol. 2009, 134, 70–74. [Google Scholar] [CrossRef] [PubMed]
- Gilpin, B.J.; Scholes, P.; Robson, B.; Savill, M.G. The transmission of thermotolerant Campylobacter spp. to people living or working on dairy farms in New Zealand. Zoonoses Public Health. 2008, 55, 352–360. [Google Scholar] [CrossRef] [PubMed]
- Davis, K.R.; Dunn, A.C.; Burnett, C.; McCullough, L.; Dimond, M.; Wagner, J.; Smith, L.; Carter, A.; Willardson, S.; Nakashima, A.K. Campylobacter jejuni infections associated with raw milk consumption—Utah, 2014. Morb. Mortal. Wkly. 2016, 65, 301–305. [Google Scholar] [CrossRef]
- Mungai, E.A.; Behravesh, C.B.; Gould, L.H. Increased outbreaks associated with nonpasteurized milk, United States, 2007–2012. Emerg. Infect. Dis. 2015, 21, 119–122. [Google Scholar] [CrossRef] [PubMed]
- Smith, K.E.; Stenzel, S.A.; Bender, J.B.; Wagstrom, E.; Soderlund, D.; Leano, F.T.; Taylor, C.M.; Belle-Isle, P.A.; Danila, R. Outbreaks of enteric infections caused by multiple pathogens associated with calves at a farm day camp. Pediatr. Infect. Dis. J. 2004, 23, 1098–1104. [Google Scholar] [CrossRef] [PubMed]
- Eberhart-Phillips, J.; Walker, N.; Garrett, N.; Bell, D.; Sinclair, D.; Rainger, W.; Bates, M. Campylobacteriosis in New Zealand: Results of a case-control study. J. Epidemiol. Community Health 1997, 51, 686–691. [Google Scholar] [CrossRef] [PubMed]
- Ruegg, P. Practical food safety interventions for dairy production. J. Dairy Sci. 2003, 86, E1–E9. [Google Scholar] [CrossRef]
- Vissers, M.; Driehuis, F. On-Farm Hygienic Milk Production. Milk Processing and Quality Management; Blackwell Publishing Ltd.: Hoboken, NJ, USA, 2009; pp. 1–22. [Google Scholar]
- Grove-White, D.; Leatherbarrow, A.; Cripps, P.; Diggle, P.; French, N. Temporal and farm-management-associated variation in the faecal-pat prevalence of Campylobacter jejuni in ruminants. Epidemiol. Infect. 2010, 138, 549–558. [Google Scholar] [CrossRef]
- California Department of Public Health. Environmental Investigation of a Campylobacter jejuni Outbreak in 2012 Associated with Claravale Farms Raw Whole Milk; Final Report; California Department of Public Health: Sacramento, CA, USA, 2013; Available online: https://www.cdph.ca.go/rogram/E/FDC/DPH%20Document%20Librar/D/oodSafetyProgra/nvInvReport/dbEIRCV2013.pdf. (accessed on 29 December 2020).
- Sommer, H.M.; Høg, B.B.; Larsen, L.S.; Sørensen, A.I.V.; Williams, N.; Merga, J.; Cerdà-Cuéllar, M.; Urdaneta, S.; Dolz, R.; Wieczorek, K. Analysis of farm specific risk factors for Campylobacter colonization of broilers in six European countries. Microb. Risk Anal. 2016, 2, 16–26. [Google Scholar] [CrossRef]
- Hussain, M.; Hoq, M.E. Impacts of climate change on coastal and marine fisheries resources in Bangladesh. In Sustainable Management of Fisheries Resources of the Bay of Bengal; Department of Fisheries: Dhaka, Bangladesh, 2010; pp. 53–73. [Google Scholar]
- Park, J.; Jung, W. The operators’ non-compliance behavior to conduct emergency operating procedures—Comparing with the work experience and the complexity of procedural steps. Reliab. Eng. Syst. Safe. 2003, 82, 115–131. [Google Scholar] [CrossRef]
- Newell, D.; Elvers, K.; Dopfer, D.; Hansson, I.; Jones, P.; James, S.; Gittins, J.; Stern, N.; Davies, R.; Connerton, I.; et al. Biosecurity-based interventions and strategies to reduce Campylobacter spp. on poultry farms. Appl. Environ. Microbiol. 2011, 77, 8605–8614. [Google Scholar] [CrossRef]
- Sibanda, N.; McKenna, A.; Richmond, A.; Ricke, S.C.; Callaway, T.; Stratakos, A.C.; Gundogdu, O.; Corcionivoschi, N. A review of the effect of management practices on Campylobacter prevalence in poultry farms. Front. Microbiol. 2018, 9, 2002. [Google Scholar] [CrossRef]
- Pires, A.F.A.; Patterson, L.; Kukielka, E.A.; Aminabadi, P.; Navarro-Gonzalez, N.; Jay-Russell, M.T. Prevalence and risk factors associated with Campylobacter spp. and Salmonella enterica in livestock raised on diversified small-scale farms in California. Epidemiol. Infect. 2019, 147, 1–9. [Google Scholar] [CrossRef]
- Gormley, F.J.; MacRae, M.; Forbes, K.J.; Ogden, I.D.; Dallas, J.F.; Strachan, N.J. Has retail chicken played a role in the decline of human campylobacteriosis? Appl. Environ. Microbiol. 2008, 74, 383–390. [Google Scholar] [CrossRef] [PubMed][Green Version]
- District-Wise Livestock Data Base; Department of Livestock Services, Ministry of Fisheries and Livestock, Government of the People’s Republic of Bangladesh, 2020. Available online: http://old.dls.gov.bd/ (accessed on 29 December 2020).
- Hamid, M.; Rahman, A.; Zaman, M.; Hossain, K. Cattle Genetic Resources and their Conservation in Bangladesh. Asian J. Anim. Sci. 2017, 11, 54–64. [Google Scholar] [CrossRef]
- Thrusfield, M. Veterinary Epidemiology, 3rd ed.; Wiley-Blackwell: Oxford, UK, 2009. [Google Scholar]
- Bolton, F.; Hutchinson, D.; Parker, G. Reassessment of selective agars and filtration techniques for isolation of Campylobacter species from faeces. Eur. J. Clin. Microbiol. 1988, 7, 155–160. [Google Scholar] [CrossRef]
- Nachamkin, I. Campylobacter and Arcobacter. In Clinical Microbiology Reviews; Murray, P.R., Baron, E.J., Jorgensen, J.H., Pfaller, M.A., Yolken, R.H., Eds.; American Society for Microbiology: Washington, DC, USA, 2003; pp. 902–914. [Google Scholar]
- Foster, G.; Holmes, B.; Steigerwalt, A.G.; Lawson, P.A.; Thorne, P.; Byrer, D.E.; Ross, H.M.; Xerry, J.; Thompson, P.M.; Collins, M.D. Campylobacter insulaenigrae sp. nov., isolated from marine mammals. Int. J. Syst. Evol. Microbiol. 2004, 54, 2369–2373. [Google Scholar] [CrossRef] [PubMed]
- Swai, E.; Hulsebosch, J.; Van der Heijden, W. Prevalence of genital campylobacteriosis and trichomonosis in crossbred breeding bulls kept on zero-grazed smallholder dairy farms in the Tanga region of Tanzania. J. S. Afr. Vet. Assoc. 2005, 76, 224–227. [Google Scholar] [CrossRef]
- Hoshino, K.; Yamasaki, S.; Mukhopadhyay, A.K.; Chakraborty, S.; Basu, A.; Bhattacharya, S.K.; Nair, G.B.; Shimada, T.; Takeda, Y. Development and evaluation of a multiplex PCR assay for rapid detection of toxigenic Vibrio cholerae O1 and O139. FEMS Immunol. Med. Microbiol. 1998, 20, 201–207. [Google Scholar] [CrossRef] [PubMed]
- Linton, D.; Lawson, A.J.; Owen, R.J.; Stanley, J. PCR detection, identification to species level, and fingerprinting of Campylobacter jejuni and Campylobacter coli direct from diarrheic samples. J. Clin. Microbiol. 1997, 35, 2568–2572. [Google Scholar] [CrossRef]
- Asakura, M.; Samosornsuk, W.; Hinenoya, A.; Misawa, N.; Nishimura, K.; Matsuhisa, A.; Yamasaki, S. Development of a cytolethal distending toxin (cdt) gene-based species-specific multiplex PCR assay for the detection and identification of Campylobacter jejuni, Campylobacter coli and Campylobacter fetus. FEMS Immunol. Med. Microbiol. 2008, 52, 260–266. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention (CDC). Epi Info™ 7. User Guide. Available online: https://www.cdc.gov/epiinfo/support/userguide.html (accessed on 29 December 2020).
| Variables | Category | Number of Positive Farms (%) | Odds Ratio | 95% Confidence Interval (CI) | p Value |
|---|---|---|---|---|---|
| Farm location (District) | Mymensingh (n = 50) | 28 (56) | 1 | 0.57 | |
| Dhaka (n = 40) | 20 (50) | 0.8 | 0.3–1.8 | ||
| Age of the farm | Up to five years (n = 34) | 13 (38.2) | 1 | 0.03 | |
| >5 years (n = 56) | 35 (62.5) | 2.7 | 1.1–6.5 | ||
| Animal shed | Newly constructed within a year (n = 24) | 8 (33.3) | 1 | 0.021 | |
| Old (more than one year) (n = 66) | 40 (60.6) | 3.1 | 1.1–8.2 | ||
| Farm (herd) size | Up to 20 cattle (n = 53) | 28 (52.8) | 1 | 0.90 | |
| >20 cattle (n = 37) | 20 (54.0) | 1.1 | 0.4–2.4 | ||
| Stocking density | More than 50 sq. ft./animal (n = 49) | 18 (36.7) | 1 | 0.63 | |
| Less than 50 sq. ft./animal (n = 41) | 23 (56.1) | 1.2 | 0.5–2.8 | ||
| Milking type | Machine milking (n = 5) | 2 (40) | 1 | 0.53 | |
| Hand milking (n = 85) | 46 (54.1) | 1.7 | 0.3–11.1 | ||
| Feed used | Readymade feed (n = 28) | 15 (53.6) | 1 | 0.975 | |
| Prepared by farmer (n = 62) | 33 (53.2) | 0.9 | 0.4–2.4 | ||
| Farmers’ training | Yes (n = 32) | 11 (34.4) | 1 | 0.0007 | |
| No (n = 58) | 37 (63.8) | 3.4 | 1.4–8.3 | ||
| Knowledge on risk perception of cattle access outside or freely roaming | Yes (n = 19) | 5 (26.3) | 1 | ||
| No (n = 71) | 43 (60.6) | 4.3 | 1.4–13.3 | 0.007 | |
| Cattle handler type | Family member (n = 28) | 16 (57.1) | 1 | 0.3–1.9 | 0.62 |
| Employee (n = 62) | 32 (51.6) | 0.8 | |||
| Prophylactic use of antibiotics | Yes (n = 58) | 30 (51.7) | 1 | 0.50–2.8 | 0.68 |
| No (n = 32) | 18 (56.2) | 1.2 | |||
| Animal health care provider | Registered veterinarian (n = 38) | 15 (39.5) | 1 | 1.1–6.3 | 0.02 |
| Non-vet (para professional/quack/ farmer himself) (n = 52) | 33 (63.5) | 2.7 | |||
| Floor condition | Dry (n = 78) | 37 (47.4) | 1 | ||
| Wet (n = 12) | 11 (91.7) | 12.2 | 1.5–90.0 | 0.004 | |
| Sunlight accessibility in the cattle shed | Yes (n = 86) | 45 (52.3) | 1 | ||
| No (n = 4) | 3 (75) | 2.7 | 0.3–27.3 | 0.37 |
| Variable | Positive | Prevalence (%) | 95% Confidence Interval | p Value |
|---|---|---|---|---|
| Number of herd/farms (N = 90) | 48 | 53.3 | 42.5–63.9 | - |
| District | ||||
| Mymensingh (n = 50) | 28 | 56 | 41.3–70 | 0.57 |
| Dhaka (n = 40) | 20 | 50 | 33.8–66.2 | |
| Sub-districts/city corporation area | ||||
| Sadar Mymensingh (n = 26) | 15 | 57.7 | 36.9–76.6 | 0.64 |
| Muktagacha (n = 6) | 2 | 33.3 | 4.3–77.7 | |
| Trisal (n = 6) | 5 | 83.3 | 35.9–99.6 | |
| Bhaluka (n = 4) | 2 | 50.0 | 6.8–93.2 | |
| Gouripur (n = 3) | 2 | 66.7 | 9.4–99.2 | |
| Fulbaria (n = 5) | 2 | 40.0 | 5.3–85.3 | |
| Savar (n = 14) | 7 | 50.0 | 23–77 | |
| Dhamrai (n = 2) | 2 | 100.0 | 15.8–100 | |
| Dhaka City Corporation (n = 24) | 11 | 45.8 | 25.5–67.1 | |
| Sample type | ||||
| Feces (n = 540) | 167 | 30.9 | 27–35 | 0.000 |
| Milk (n = 180) | 3 | 1.7 | 0.3–4.8 | |
| Feed (n = 90) | 0 | 0 | 0–4 | |
| Water (n = 90) | 0 | 0 | 0–4 | |
| Manure swab (n = 90) | 14 | 15.6 | 8.8–24.7 | |
| Hand-rinse water of animal attendants (n = 90) | 10 | 11.11 | 5.5–19.5 | |
| Overall (N = 1080) | 194 | 18 | 15.7–20.4 | |
| Animal category | ||||
| Calves (n = 180) | 51 | 28.3 | 21.9–35.5 | 0.0008 |
| Heifers (n = 180) | 42 | 23.3 | 17.4–30.2 | |
| Cows (n = 180) | 74 | 41.1 | 33.8–48.7 | |
| Total sample (N = 540) | 167 | 30.9 | 27–35 | |
| Season | ||||
| Pre-monsoon (March–May) (n = 300) | 87 | 29 | 23.9–34.5 | 0.47 |
| Monsoon (June–October) (n = 156) | 54 | 34.6 | 27.2–42.6 | |
| Winter (November–February) (n = 84) | 26 | 31 | 21.3–42 | |
| Variables | Category | Number of Positive Farms (%) | OR | 95% CI | p -Value |
|---|---|---|---|---|---|
| Cleaning and disinfection practices (floor cleaning, cleaning of manger, and drink regularly) | Good practices (n = 60) | 26 (43.3) | 1 | ||
| Poor/no practices (n = 30) | 22 (73.3) | 11.2 | 3.5–36.4 | 0 | |
| Worker boot disinfection | Yes (n = 12) | 6 (50) | 1 | ||
| No (n = 78) | 42 (53.8) | 1.2 | 0.3–3.9 | 0.8 | |
| Isolation of animal | Yes (n = 19) | 9 (47.4) | 1 | ||
| No (n = 71) | 39 (55) | 1.3 | 0.5–3.7 | 0.55 | |
| Access of other animals (poultry/goats/sheep/wild animals) in the farm | No (n = 58) | 26 (44.8) | 1 | ||
| Yes (n = 32) | 22 (68.7) | 2.7 | 1.1–6.7 | 0.03 | |
| Udder cleaning | With antiseptic (n = 14) | 8 (57.1) | 1 | ||
| With water (n = 76) | 40 (52.6) | 0.8 | 0.2–2.6 | 0.75 | |
| Manure storage | Solid (n = 32) | 15 (46.9) | 1 | ||
| Semi-solid(n = 58) | 33 (56.9) | 1.5 | 0.6–3.6 | 0.36 | |
| Animal roams outside of the farm | No (n = 21) | 1 (4.7) | 1 | ||
| Yes (n = 69) | 47 (68.1) | 42.7 | 5.4–339.0 | 0 | |
| Cattle feces use purpose | Fertilizer (n = 41) | 22 (53.7) | 1 | ||
| Aquaculture (n = 49) | 26 (53.1) | 0.9 | 0.4–2.2 | 0.95 | |
| History of diarrhea in the farmed cattle | No (n = 84) | 45 (53.6) | 1 | ||
| Yes (n = 6) | 3 (50) | 0.9 | 0.1–4.5 | 0.86 | |
| Interface (Share same premices with cattle) | No (n = 70) | 40 (57.1) | 1 | ||
| Yes (n = 20) | 8 (40) | 0.5 | 0.2–1.4 | 0.175 |
| Risk Factors | Category | AOR | 95% CI | SE | p Value |
|---|---|---|---|---|---|
| Age of the farm | 1–5 years | 1 | |||
| >5 years | 10.6 | 1.9–59.8 | 0.882 | 0.0007 | |
| Animal shed | Newly constructed | 1 | |||
| Old | 4.0 | 0.8–19.9 | 0.82 | 0.09 | |
| Training | Yes | 1 | |||
| No | 3.9 | 0.7–21.2 | 0.861 | 0.112 | |
| Knowledge | Yes | 1 | |||
| No | 3.5 | 0.4–28.5 | 1.06 | 0.23 | |
| Cleaning and disinfection practices | Good practices | 1 | |||
| No/minimum practices | 12.4 | 2.1–71.6 | 0.893 | 0.0048 | |
| Floor Condition | Dry | 1 | |||
| Wet | 2.0 | 0.1–56.3 | 1.69 | 0.67 | |
| Animals roaming outside | No | 1 | |||
| Yes | 44.0 | 3.6–537.10 | 1.27 | 0.003 | |
| Other animal (poultry/goat/sheep/wild animal) access | No | 1 | |||
| Yes | 3.1 | 0.6–16.1 | 0.84 | 0.178 | |
| Animal health service provider | Registered veterinarian | 1 | |||
| Quack/farmer himself | 3.1 | 0.6–16.3 | 0.84 | 0.174 |
| Primers | Sequence (5′-3′) | Target/Purpose | Amplicon Size (bp) | PCR Condition (30 cycle) | Reference | ||
|---|---|---|---|---|---|---|---|
| Denaturation | Annealing | Extension | |||||
| 16S9F 16S1540R | GAGTTTGATCCTGGCTC AAGGAGGTGATCCAGCC | 16S rRNA | 1530 | 94 °C, 30 s | 47 °C, 30 s | 72 °C, 90 s | [18] |
| HIP400F HIP1134R | GAAGAGGGTTTGGGTGGTG AGCTAGCTTCGCATAATAACTTG | hipO gene | 735 | 94 °C, 30 s | 55 °C, 30 s | 72 °C, 45 s | [77] |
| Cj-CdtAU2 Cj-CdtAR2 | AGGACTTGAACCTACTTTTC AGGTGGAGTAGTTAAAAACC | CjcdtA | 631 | 94 °C, 30 s | 53 °C, 30 s | 72 °C, 30 s | [78] |
| Cc-CdtAU1 Cc-CdtAR1 | ATTGCCAAGGCTAAAATCTC GATAAAGTCTCCAAAACTGC | CccdtA | 329 | ||||
| Cf-CdtAU1 Cf-CdtAR1 | AACGACAAATGTAAGCACTC TATTTATGCAAGTCGTGCGA | CfcdtA | 489 | ||||
| 16S520F 16S1199F 16S741R 16S1240R | GTGCCAGCAGCCGCGG GCAACGAGCGCAACCC GTATCTAATCCTGTTTGC CCATTGTAGCACGTGT | Sequence for Cj-, Cc- and Cf- 16S rRNA | NA | NA | NA | NA | [16] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hoque, N.; Islam, S.S.; Uddin, M.N.; Arif, M.; Haque, A.K.M.Z.; Neogi, S.B.; Hossain, M.M.; Yamasaki, S.; Kabir, S.M.L. Prevalence, Risk Factors, and Molecular Detection of Campylobacter in Farmed Cattle of Selected Districts in Bangladesh. Pathogens 2021, 10, 313. https://doi.org/10.3390/pathogens10030313
Hoque N, Islam SS, Uddin MN, Arif M, Haque AKMZ, Neogi SB, Hossain MM, Yamasaki S, Kabir SML. Prevalence, Risk Factors, and Molecular Detection of Campylobacter in Farmed Cattle of Selected Districts in Bangladesh. Pathogens. 2021; 10(3):313. https://doi.org/10.3390/pathogens10030313
Chicago/Turabian StyleHoque, Nazmul, SK Shaheenur Islam, Md. Nasir Uddin, Mohammad Arif, A. K. M. Ziaul Haque, Sucharit Basu Neogi, Md. Mehedi Hossain, Shinji Yamasaki, and S. M. Lutful Kabir. 2021. "Prevalence, Risk Factors, and Molecular Detection of Campylobacter in Farmed Cattle of Selected Districts in Bangladesh" Pathogens 10, no. 3: 313. https://doi.org/10.3390/pathogens10030313
APA StyleHoque, N., Islam, S. S., Uddin, M. N., Arif, M., Haque, A. K. M. Z., Neogi, S. B., Hossain, M. M., Yamasaki, S., & Kabir, S. M. L. (2021). Prevalence, Risk Factors, and Molecular Detection of Campylobacter in Farmed Cattle of Selected Districts in Bangladesh. Pathogens, 10(3), 313. https://doi.org/10.3390/pathogens10030313









