Host Diversity and Potential Transmission Pathways of SARS-CoV-2 at the Human-Animal Interface
Abstract
1. Introduction
2. Epidemiology of Human SARS-CoV-2 Infections
3. Transmission Pathways of SARS-CoV-2 in Humans
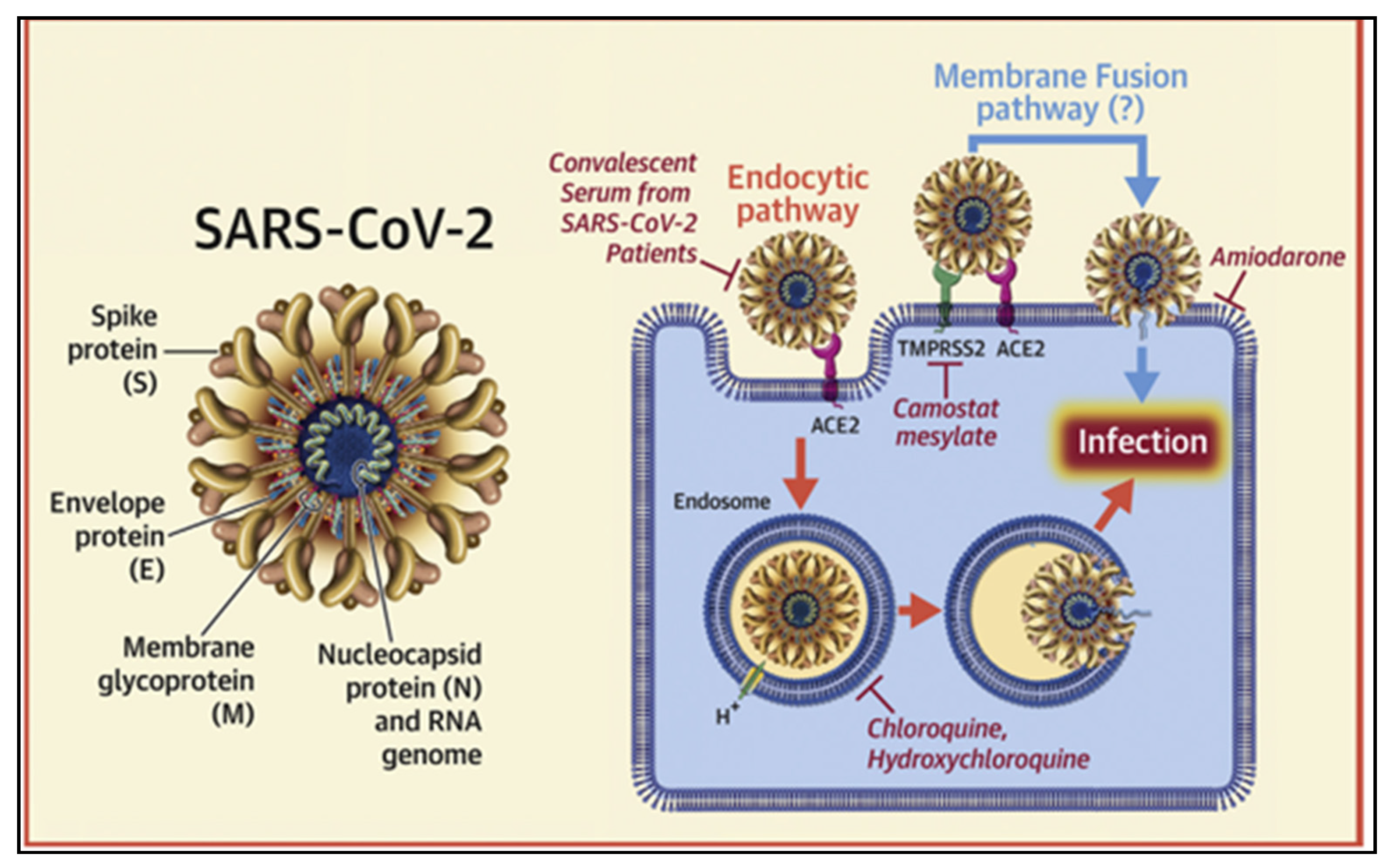
4. SARS-CoV-2 Spike Protein and ACE-2
5. Animal Host Diversity of SARS-CoV-2
5.1. Bats
5.2. Pangolins
5.3. Felids
5.4. Mustelids
5.5. Rodents
5.6. Eulipotyphlans
5.7. Tree Shrews
5.8. Lagomorphs
5.9. Canids
5.10. Non-Human Primates
5.11. Livestock
5.12. Aves
5.13. Reptilia
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mousavizadeh, L.; Ghasemi, S. Genotype and phenotype of COVID-19: Their roles in pathogenesis. J. Microbiol. Immunol. Infect. 2020, 10. [Google Scholar] [CrossRef] [PubMed]
- Ramadan, N.; Shaib, H. Middle East respiratory syndrome coronavirus (MERS-CoV): A review. Germs 2019, 9, 35–42. [Google Scholar] [CrossRef]
- Miłek, J.; Domanska-Blicharz, K. Coronaviruses in avian species—Review with focus on epidemiology and diagnosis in wild birds. J. Vet. Res. 2018, 62, 249–255. [Google Scholar] [CrossRef] [PubMed]
- Chu, D.K.W.; Leung, C.Y.H.; Gilbert, M.; Joyner, P.H.; Ng, E.M.; Tse, T.M.; Guan, Y.; Peiris, J.S.M.; Poon, L.L.M. Avian Coronavirus in Wild Aquatic Birds. J. Virol. 2011, 85, 12815–12820. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, R.; Dhama, K.; Sharun, K.; Yatoo, M.I.; Malik, Y.S.; Singh, R.; Michalak, I.; Sah, R.; Bonilla-Aldana, D.K.; Rodriguez-Morales, A.J. COVID-19: Animals, veterinary and zoonotic links. Vet. Q. 2020, 40, 169–182. [Google Scholar] [CrossRef]
- Corman, V.M.; Muth, D.; Niemeyer, D.; Drosten, C. Hosts and Sources of Endemic Human Coronaviruses. Adv. Virus Res. 2018, 100, 163–188. [Google Scholar] [CrossRef]
- Saif, L. Animal coronaviruses: What can they teach us about the severe acute respiratory syndrome? Rev. Sci. Tech. l’OIE 2004, 23, 643–660. [Google Scholar] [CrossRef]
- McIntosh, K.; Dees, J.H.; Becker, W.B.; Kapikian, A.Z.; Chanock, R.M. Recovery in tracheal organ cultures of novel viruses from patients with respiratory disease. Proc. Natl. Acad. Sci. USA 1967, 57, 933–940. [Google Scholar] [CrossRef]
- Hamre, D.; Procknow, J.J. A New Virus Isolated from the Human Respiratory Tract. Exp. Biol. Med. 1966, 121, 190–193. [Google Scholar] [CrossRef]
- Woo, P.C.Y.; Lau, S.K.P.; Chu, C.-M.; Chan, K.-H.; Tsoi, H.-W.; Huang, Y.; Wong, B.H.L.; Poon, R.W.S.; Cai, J.J.; Luk, W.-K.; et al. Characterization and Complete Genome Sequence of a Novel Coronavirus, Coronavirus HKU1, from Patients with Pneumonia. J. Virol. 2005, 79, 884–895. [Google Scholar] [CrossRef]
- Van der Hoek, L.; Pyrc, K.; Jebbink, M.F.; Vermeulen-Oost, W.; Berkhout, R.J.M.; Wolthers, K.C.; Dillen, P.M.E.W.-V.; Kaandorp, J.; Spaargaren, J.; Berkhout, B. Identification of a new human coronavirus. Nat. Med. 2004, 10, 368–373. [Google Scholar] [CrossRef]
- Cummings, D.A.; Radonovich, L.J.; Gorse, G.J.; Gaydos, C.A.; Bessesen, M.T.; Brown, A.C.; Gibert, C.L.; Hitchings, M.D.T.; Lessler, J.; Nyquist, A.-C.; et al. Risk Factors for Healthcare Personnel Infection With Endemic Coronaviruses (HKU1, OC43, NL63, 229E): Results from the Respiratory Protection Effectiveness Clinical Trial (ResPECT). Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Wang, L.-F.; Shi, Z.; Zhang, S.; Field, H.; Daszak, P.; Eaton, B.T. Review of Bats and SARS. Emerg. Infect. Dis. 2006, 12, 1834–1840. [Google Scholar] [CrossRef]
- World Health Organization (WHO). Summary Table of SARS Cases by Country, 1 November 2002–August 2003. Summary Table of SARS Cases by Country N-A; World Health Organisation (WHO): Geneva, Switzerland; Available online: https://covid19.who.int/ (accessed on 1 December 2020).
- World Health Organization (WHO). Middle East Respiratory Syndrome Coronavirus (MERS-CoV); WHO: Geneva, Switzerland, 2013. [Google Scholar]
- Lu, G.; Wang, Q.; Gao, G.F. Bat-to-human: Spike features determining ‘host jump’ of coronaviruses SARS-CoV, MERS-CoV, and beyond. Trends Microbiol. 2015, 23, 468–478. [Google Scholar] [CrossRef]
- Wang, C.; Horby, P.W.; Hayden, F.G.; Gao, G.F. A novel coronavirus outbreak of global health concern. Lancet 2020, 395, 470–473. [Google Scholar] [CrossRef]
- Wu, F.; Zhao, S.; Yu, B.; Chen, Y.-M.; Wang, W.; Song, Z.-G.; Hu, Y.; Tao, Z.-W.; Tian, J.-H.; Pei, Y.-Y.; et al. A new coronavirus associated with human respiratory disease in China. Nature 2020, 579, 265–269. [Google Scholar] [CrossRef] [PubMed]
- Ashour, H.M.; Elkhatib, W.F.; Rahman, M.; Elshabrawy, H.A. Insights into the Recent 2019 Novel Coronavirus (SARS-CoV-2) in Light of Past Human Coronavirus Outbreaks. Pathogens 2020, 9, 186. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Moneim, A.S.; Abdelwhab, E.M. Evidence for SARS-CoV-2 Infection of Animal Hosts. Pathogens 2020, 9, 529. [Google Scholar] [CrossRef] [PubMed]
- Gortázar, C.; de la Fuente, J. COVID-19 is likely to impact animal health. Prev. Vet. Med. 2020, 180, 105030. [Google Scholar] [CrossRef]
- Lai, A.; Bergna, A.; Acciarri, C.; Galli, M.; Zehender, G. Early phylogenetic estimate of the effective reproduction number of SARS-CoV-2. J. Med. Virol. 2020, 92, 675–679. [Google Scholar] [CrossRef]
- Nie, Q.; Li, X.; Chen, W.; Liu, D.; Chen, Y.; Li, H.; Li, D.; Tian, M.; Tan, W.; Zai, J. Phylogenetic and phylodynamic analyses of SARS-CoV-2. Virus Res. 2020, 287, 198098. [Google Scholar] [CrossRef]
- World Health Organization (WHO). Coronavirus Disease (COVID-19) Dashboard. 2020. Available online: https://covid19.who.int/ (accessed on 29 September 2020).
- Bobay, L.-M.; O’Donnell, A.C.; Ochman, H. Recombination events are concentrated in the spike protein region of Betacoronaviruses. PLoS Genet. 2020, 16, e1009272. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhao, W.; Mao, Y.; Chen, Y.; Wang, S.; Zhong, Y.; Su, T.; Gong, M.; Du, D.; Lu, X.; et al. Site-specific N-glycosylation Characterization of Recombinant SARS-CoV-2 Spike Proteins. Mol. Cell. Proteom. 2020. [Google Scholar] [CrossRef]
- Boni, M.F.; Lemey, P.; Jiang, X.; Lam, T.T.-Y.; Perry, B.W.; Castoe, T.A.; Rambaut, A.; Robertson, D.L. Evolutionary origins of the SARS-CoV-2 sarbecovirus lineage responsible for the COVID-19 pandemic. Nat. Microbiol. 2020, 5, 1–10. [Google Scholar] [CrossRef]
- Cao, B.; Wang, Y.; Wen, D.; Liu, W.; Wang, J.; Fan, G.; Ruan, L.; Song, B.; Cai, Y.; Wei, M.; et al. A Trial of Lopinavir–Ritonavir in Adults Hospitalized with Severe Covid-19. N. Engl. J. Med. 2020, 382, 1787–1799. [Google Scholar] [CrossRef] [PubMed]
- Singhal, T. A Review of Coronavirus Disease-2019 (COVID-19). Indian J. Pediatr. 2020, 87, 281–286. [Google Scholar] [CrossRef]
- Helmy, Y.A.; Fawzy, M.; Elaswad, A.; Sobieh, A.; Kenney, S.P.; Shehata, A.A. The COVID-19 Pandemic: A Comprehensive Review of Taxonomy, Genetics, Epidemiology, Diagnosis, Treatment, and Control. J. Clin. Med. 2020, 9, 1225. [Google Scholar] [CrossRef] [PubMed]
- Flores-Alanis, A.; Sandner-Miranda, L.; Delgado, G.; Cravioto, A.; Morales-Espinosa, R. The receptor binding domain of SARS-CoV-2 spike protein is the result of an ancestral recombination between the bat-CoV RaTG13 and the pangolin-CoV MP789. BMC Res. Notes 2020, 13, 1–6. [Google Scholar] [CrossRef]
- Lam, T.T.-Y.; Jia, N.; Zhang, Y.-W.; Shum, M.H.-H.; Jiang, J.-F.; Zhu, H.-C.; Tong, Y.-G.; Shi, Y.-X.; Ni, X.-B.; Liao, Y.-S.; et al. Identifying SARS-CoV-2-related coronaviruses in Malayan pangolins. Nature 2020, 583, 282–285. [Google Scholar] [CrossRef]
- Lau, S.K.P.; Woo, P.C.Y.; Yip, C.C.Y.; Fan, R.Y.Y.; Huang, Y.; Wang, M.; Guo, R.; Lam, C.S.F.; Tsang, A.K.L.; Lai, K.K.Y.; et al. Isolation and Characterization of a Novel Betacoronavirus Subgroup A Coronavirus, Rabbit Coronavirus HKU14, from Domestic Rabbits. J. Virol. 2012, 86, 5481–5496. [Google Scholar] [CrossRef]
- Shang, J.; Ye, G.; Shi, K.; Wan, Y.; Luo, C.; Aihara, H.; Geng, Q.; Auerbach, A.; Li, F. Structural basis of receptor recognition by SARS-CoV-2. Nat. Cell Biol. 2020, 581, 221–224. [Google Scholar] [CrossRef] [PubMed]
- Frutos, R.; Serra-Cobo, J.; Chen, T.; Devaux, C.A. COVID-19: Time to exonerate the pangolin from the transmission of SARS-CoV-2 to humans. Infect. Genet. Evol. 2020, 84, 104493. [Google Scholar] [CrossRef]
- Guan, Q.; Sadykov, M.; Mfarrej, S.; Hala, S.; Naeem, R.; Nugmanova, R.; Al-Omari, A.; Salih, S.; Al-Mutair, A.; Carr, M.J.; et al. A genetic barcode of SARS-CoV-2 for monitoring global distribution of different clades during the COVID-19 pandemic. Int. J. Infect. Dis. 2020, 100, 216–223. [Google Scholar] [CrossRef] [PubMed]
- Xiao, K.; Zhai, J.; Feng, Y.; Zhou, N.; Zhang, X.; Zou, J.-J.; Li, N.; Guo, Y.; Li, X.; Shen, X.; et al. Isolation of SARS-CoV-2-related coronavirus from Malayan pangolins. Nat. Cell Biol. 2020, 583, 286–289. [Google Scholar] [CrossRef]
- Konda, M.; Dodda, B.; Konala, V.M.; Naramala, S.; Adapa, S. Potential Zoonotic Origins of SARS-CoV-2 and Insights for Preventing Future Pandemics Through One Health Approach. Cureus 2020, 12, e8932. [Google Scholar] [CrossRef]
- Zhao, J.; Cui, W.; Tian, B.-P. The Potential Intermediate Hosts for SARS-CoV-2. Front. Microbiol. 2020, 11, 11. [Google Scholar] [CrossRef]
- Kiros, M.; Andualem, H.; Kiros, T.; Hailemichael, W.; Getu, S.; Geteneh, A.; Alemu, D.; Abegaz, W.E. COVID-19 pandemic: Current knowledge about the role of pets and other animals in disease transmission. Virol. J. 2020, 17, 1–8. [Google Scholar] [CrossRef]
- Pandey, K.; Acharya, A.; Mohan, M.; Ng, C.L.; Reid, S.P.; Byrareddy, S.N. Animal models for SARS-CoV-2 research: A comprehensive literature review. Transbound. Emerg. Dis. 2020, 13907. [Google Scholar] [CrossRef] [PubMed]
- Raj, V.S.; Farag, E.A.; Reusken, C.B.; Lamers, M.M.; Pas, S.D.; Voermans, J.; Smits, S.L.; Osterhaus, A.D.; Al-Mawlawi, N.; Al-Romaihi, H.E.; et al. Isolation of MERS Coronavirus from a Dromedary Camel, Qatar, 2014. Emerg. Infect. Dis. 2014, 20, 1339–1342. [Google Scholar] [CrossRef]
- El-Zowalaty, M.E.; Järhult, J.D. From SARS to COVID-19: A previously unknown SARS- related coronavirus (SARS-CoV-2) of pandemic potential infecting humans—Call for a One Health approach. One Health 2020, 9, 100124. [Google Scholar] [CrossRef]
- Tazerji, S.S.; Duarte, P.M.; Rahimi, P.; Shahabinejad, F.; Dhakal, S.; Malik, Y.S.; Shehata, A.A.; Lama, J.; Klein, J.; Safdar, M.; et al. Transmission of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) to animals: An updated review. J. Transl. Med. 2020, 18, 1–11. [Google Scholar] [CrossRef]
- McNamara, T.; Richt, J.A.; Glickman, L. A Critical Needs Assessment for Research in Companion Animals and Livestock Following the Pandemic of COVID-19 in Humans. Vector Borne Zoonotic Dis. 2020, 20, 393–405. [Google Scholar] [CrossRef]
- Munnink, B.B.O.; Sikkema, R.S.; Nieuwenhuijse, D.F.; Molenaar, R.J.; Munger, E.; Molenkamp, R.; van der Spek, A.; Tolsma, P.; Rietveld, A.; Brouwer, M.; et al. Transmission of SARS-CoV-2 on mink farms between humans and mink and back to humans. Science 2021, 371, 172–177. [Google Scholar] [CrossRef] [PubMed]
- Enserink, M. Coronavirus rips through Dutch mink farms, triggering culls. Science 2020, 368, 1169. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control (CDC). One Health Basics. Available online: https://www.cdc.gov/onehealth/basics/index.html (accessed on 28 January 2021).
- Phelan, A.L.; Katz, R.; Gostin, L.O. The Novel Coronavirus Originating in Wuhan, China. JAMA 2020, 323, 709. [Google Scholar] [CrossRef] [PubMed]
- Chowdhury, M.A.; Hossain, N.; Kashem, M.A.; Shahid, A.; Alam, A. Immune response in COVID-19: A review. J. Infect. Public Health 2020, 13, 1619–1629. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Pei, S.; Chen, B.; Song, Y.; Zhang, T.; Yang, W.; Shaman, J. Substantial undocumented infection facilitates the rapid dissemination of novel coronavirus (SARS-CoV-2). Science 2020, 368, 489–493. [Google Scholar] [CrossRef] [PubMed]
- Tay, M.Z.; Poh, C.M.; Rénia, L.; Macary, P.A.; Ng, L.F.P. The trinity of COVID-19: Immunity, inflammation and intervention. Nat. Rev. Immunol. 2020, 20, 363–374. [Google Scholar] [CrossRef]
- Shi, J.; Wen, Z.; Zhong, G.; Yang, H.; Wang, C.; Huang, B.; Liu, R.; He, X.; Shuai, L.; Sun, Z.; et al. Susceptibility of ferrets, cats, dogs, and other domesticated animals to SARS–coronavirus 2. Science 2020, 368, 1016–1020. [Google Scholar] [CrossRef]
- Sun, J.; He, W.-T.; Wang, L.; Lai, A.; Ji, X.; Zhai, X.; Li, G.; Suchard, M.A.; Tian, J.; Zhou, J.; et al. COVID-19: Epidemiology, Evolution, and Cross-Disciplinary Perspectives. Trends Mol. Med. 2020, 26, 483–495. [Google Scholar] [CrossRef]
- Oran, D.P.; Topol, E.J. Prevalence of Asymptomatic SARS-CoV-2 Infection. Ann. Intern. Med. 2020, 173, 362–367. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Zhang, L.; Du, H.; Zhang, J.; Li, Y.Y.; Qu, J.; Zhang, W.; Wang, Y.; Bao, S.; Li, Y.; et al. SARS-CoV-2 Infection in Children. N. Engl. J. Med. 2020, 382, 1663–1665. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.F.-W.; Yuan, S.; Kok, K.-H.; To, K.K.-W.; Chu, H.; Yang, J.; Xing, F.; Liu, J.; Yip, C.C.-Y.; Poon, R.W.-S.; et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: A study of a family cluster. Lancet 2020, 395, 514–523. [Google Scholar] [CrossRef]
- Gudbjartsson, D.F.; Helgason, A.; Jonsson, H.; Magnusson, O.T.; Melsted, P.; Norddahl, G.L.; Saemundsdottir, J.; Sigurdsson, A.; Sulem, P.; Agustsdottir, A.B.; et al. Spread of SARS-CoV-2 in the Icelandic Population. N. Engl. J. Med. 2020, 382, 2302–2315. [Google Scholar] [CrossRef]
- Burki, T. Mass testing for COVID-19. Lancet Microbe 2020, 1, e317. [Google Scholar] [CrossRef]
- Guan, W.J.; Ni, Z.Y.; Hu, Y.; Liang, W.H.; Ou, C.Q.; He, J.X.; Liu, L.; Shan, H.; Lei, C.L.; Hui, D.S.C.; et al. Clinical Characteristics of Coronavirus Disease 2019 in China. N. Engl. J. Med. 2020, 382, 1708–1720. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Guan, X.; Wu, P.; Wang, X.; Zhou, L.; Tong, Y.; Ren, R.; Leung, K.S.M.; Lau, E.H.Y.; Wong, J.Y.; et al. Early transmission dynamics in Wuhan, China, of Novel Coronavirus–infected pneumonia. N. Engl. J. Med. 2020, 382, 1199–1207. [Google Scholar] [CrossRef]
- Lauer, S.A.; Grantz, K.H.; Bi, Q.; Jones, F.K.; Zheng, Q.; Meredith, H.R.; Azman, A.S.; Reich, N.G.; Lessler, J. The Incubation Period of Coronavirus Disease 2019 (COVID-19) From Publicly Reported Confirmed Cases: Estimation and Application. Ann. Intern. Med. 2020, 172, 577–582. [Google Scholar] [CrossRef] [PubMed]
- Bunders, M.J.; Altfeld, M. Implications of Sex Differences in Immunity for SARS-CoV-2 Pathogenesis and Design of Therapeutic Interventions. Immunity 2020, 53, 487–495. [Google Scholar] [CrossRef]
- Turan, O.; Hakim, A.; Dashraath, P.; Jeslyn, W.J.L.; Wright, A.; Abdul-Kadir, R. Clinical characteristics, prognostic factors, and maternal and neonatal outcomes of SARS-CoV-2 infection among hospitalized pregnant women: A systematic review. Int. J. Gynecol. Obstet. 2020, 151, 7–16. [Google Scholar] [CrossRef]
- Arias-Reyes, C.; Zubieta-De-Urioste, N.; Poma-Machicao, L.; Aliaga-Raduan, F.; Carvajal-Rodriguez, F.; Dutschmann, M.; Schneider-Gasser, E.M.; Zubieta-Calleja, G.; Soliz, J. Does the pathogenesis of SARS-CoV-2 virus decrease at high-altitude? Respir. Physiol. Neurobiol. 2020, 277, 103443. [Google Scholar] [CrossRef]
- Segovia-Juarez, J.; Castagnetto, J.M.; Gonzales, G.F. High altitude reduces infection rate of COVID-19 but not case-fatality rate. Respir. Physiol. Neurobiol. 2020, 281, 103494. [Google Scholar] [CrossRef] [PubMed]
- Chennakesavulu, K.; Reddy, G.R. The effect of latitude and PM2.5 on spreading of SARS-CoV-2 in tropical and temperate zone countries. Environ. Pollut. 2020, 266, 115176. [Google Scholar] [CrossRef] [PubMed]
- Hooper, M.W.; Nápoles, A.M.; Pérez-Stable, E.J. COVID-19 and Racial/Ethnic Disparities. JAMA 2020, 323, 2466. [Google Scholar] [CrossRef] [PubMed]
- McClure, E.S.; Vasudevan, P.; Bailey, Z.; Patel, S.; Robinson, W.R. Racial Capitalism Within Public Health—How Occupational Settings Drive COVID-19 Disparities. Am. J. Epidemiol. 2020, 189, 1244–1253. [Google Scholar] [CrossRef] [PubMed]
- Stokes, E.K.; Zambrano, L.D.; Anderson, K.N.; Marder, E.P.; Raz, K.M.; Felix, S.E.B.; Tie, Y.; Fullerton, K.E. Coronavirus Disease 2019 Case Surveillance—United States, January 22–May 30, 2020. MMWR. Morb. Mortal. Wkly. Rep. 2020, 69, 759–765. [Google Scholar] [CrossRef]
- Killerby, M.E.; Link-Gelles, R.; Haight, S.C.; Schrodt, C.A.; England, L.; Gomes, D.J.; Shamout, M.; Pettrone, K.; O’Laughlin, K.; Kimball, A.; et al. Characteristics Associated with Hospitalization Among Patients with COVID-19—Metropolitan Atlanta, Georgia, March–April 2020. MMWR. Morb. Mortal. Wkly. Rep. 2020, 69, 790–794. [Google Scholar] [CrossRef]
- United States Department of Health and Human Services (USDHHS) Social Determinants of Health. Available online: https://www.healthypeople.gov/2020/topics-objectives/topic/social-determinants-of-healthexternal (accessed on 1 December 2020).
- Emeruwa, U.N.; Ona, S.; Shaman, J.L.; Turitz, A.; Wright, J.D.; Gyamfi-Bannerman, C.; Melamed, A. Associations Between Built Environment, Neighborhood Socioeconomic Status, and SARS-CoV-2 Infection Among Pregnant Women in New York City. JAMA 2020, 324, 390–392. [Google Scholar] [CrossRef]
- Goyal, M.K.; Simpson, J.N.; Boyle, M.D.; Badolato, G.M.; Delaney, M.; McCarter, R.; Cora-Bramble, D. Racial and/or Ethnic and Socioeconomic Disparities of SARS-CoV-2 Infection Among Children. Pediatrics 2020, 146, 2020009951. [Google Scholar] [CrossRef]
- Hagan, L.M.; Williams, S.P.; Spaulding, A.C.; Toblin, R.L.; Figlenski, J.; Ocampo, J.; Ross, T.; Bauer, H.; Hutchinson, J.; Lucas, K.D.; et al. Mass Testing for SARS-CoV-2 in 16 Prisons and Jails—Six Jurisdictions, United States, April–May 2020. MMWR. Morb. Mortal. Wkly. Rep. 2020, 69, 1139–1143. [Google Scholar] [CrossRef]
- Mosites, E.; Parker, E.M.; Clarke, K.E.N.; Gaeta, J.M.; Baggett, T.P.; Imbert, E.; Sankaran, M.; Scarborough, A.; Huster, K.; Hanson, M.; et al. Assessment of SARS-CoV-2 Infection Prevalence in Homeless Shelters—Four U.S. Cities, March 27–April 15, 2020. MMWR. Morb. Mortal. Wkly. Rep. 2020, 69, 521–522. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention (CDC). People at Increased Risk. Available online: https://www.cdc.gov/coronavirus/2019-ncov/need-extra-precautions/index.html (accessed on 1 December 2020).
- Castagnoli, R.; Votto, M.; Licari, A.; Brambilla, I.; Bruno, R.; Perlini, S.; Rovida, F.; Baldanti, F.; Marseglia, G.L. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Infection in Children and Adolescents. JAMA Pediatr. 2020, 174, 882. [Google Scholar] [CrossRef]
- Jin, Y.-H.; Cai, L.; Cheng, Z.-S.; Cheng, H.; Deng, T.; Fan, Y.-P.; Fang, C.; Huang, D.; Huang, L.-Q.; Huang, Q.; et al. A rapid advice guideline for the diagnosis and treatment of 2019 novel coronavirus (2019-nCoV) infected pneumonia (standard version). Mil. Med. Res. 2020, 7, 4. [Google Scholar] [CrossRef] [PubMed]
- Burke, R.M.; Midgley, C.M.; Dratch, A.; Fenstersheib, M.; Haupt, T.; Holshue, M.; Ghinai, I.; Jarashow, M.C.; Lo, J.; McPherson, T.D.; et al. Active Monitoring of Persons Exposed to Patients with Confirmed COVID-19—United States, January–February 2020. MMWR. Morb. Mortal. Wkly. Rep. 2020, 69, 245–246. [Google Scholar] [CrossRef] [PubMed]
- Cheng, V.C.-C.; Wong, S.-C.; Chuang, V.W.-M.; So, S.Y.-C.; Chen, J.H.-K.; Sridhar, S.; To, K.K.-W.; Chan, J.F.-W.; Hung, I.F.-N.; Ho, P.-L.; et al. The role of community-wide wearing of face mask for control of coronavirus disease 2019 (COVID-19) epidemic due to SARS-CoV-2. J. Infect. 2020, 81, 107–114. [Google Scholar] [CrossRef]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Liu, J.; Liao, X.; Qian, S.; Yuan, J.; Wang, F.; Liu, Y.; Wang, Z.; Wang, F.-S.; Liu, L.; Zhang, Z. Community Transmission of Severe Acute Respiratory Syndrome Coronavirus 2, Shenzhen, China, 2020. Emerg. Infect. Dis. 2020, 26, 26. [Google Scholar] [CrossRef]
- Meselson, M. Droplets and Aerosols in the Transmission of SARS-CoV-2. New Engl. J. Med. 2020, 382, 2063. [Google Scholar] [CrossRef]
- World Health Organization (WHO). Scientific Brief: SARS-CoV-2 and Potential Airborne Transmission. Available online: https://www.cdc.gov/coronavirus/2019-ncov/more/scientific-brief-sars-cov-2.html (accessed on 1 December 2020).
- Centers for Disease Control (CDC). Animals & COVID-19. Available online: https://www.cdc.gov/coronavirus/2019-ncov/daily-life-coping/animals.html (accessed on 1 December 2020).
- Van Doremalen, N.; Miazgowicz, K.L.; Milne-Price, S.; Bushmaker, T.; Robertson, S.; Scott, D.; Kinne, J.; McLellan, J.S.; Zhu, J.; Munster, V.J. Host Species Restriction of Middle East Respiratory Syndrome Coronavirus through Its Receptor, Dipeptidyl Peptidase 4. J. Virol. 2014, 88, 9220–9232. [Google Scholar] [CrossRef]
- Fears, A.C.; Klimstra, W.B.; Duprex, P.; Hartman, A.; Weaver, S.C.; Plante, K.S.; Mirchandani, D.; Plante, J.A.; Aguilar, P.V.; Fernández, D.; et al. Persistence of Severe Acute Respiratory Syndrome Coronavirus 2 in Aerosol Suspensions. Emerg. Infect. Dis. 2020, 26, 2168–2171. [Google Scholar] [CrossRef]
- Aboubakr, H.A.; Sharafeldin, T.A.; Goyal, S.M. Stability of SARS-CoV-2 and other coronaviruses in the environment and on common touch surfaces and the influence of climatic conditions: A review. Transbound. Emerg. Dis. 2020, 13707. [Google Scholar] [CrossRef] [PubMed]
- Chin, A.W.H.; Chu, J.T.S.; Perera, M.R.A.; Hui, K.P.Y.; Yen, H.-L.; Chan, M.C.W.; Peiris, M.; Poon, L.L.M. Stability of SARS-CoV-2 in different environmental conditions. Lancet Microbe 2020, 1, e10. [Google Scholar] [CrossRef]
- Pastorino, B.; Touret, F.; Gilles, M.; de Lamballerie, X.; Charrel, R.N. Prolonged Infectivity of SARS-CoV-2 in Fomites. Emerg. Infect. Dis. 2020, 26, 2256–2257. [Google Scholar] [CrossRef]
- Biryukov, J.; Boydston, J.A.; Dunning, R.A.; Yeager, J.J.; Wood, S.; Reese, A.L.; Ferris, A.; Miller, D.; Weaver, W.; Zeitouni, N.E.; et al. Increasing Temperature and Relative Humidity Accelerates Inactivation of SARS-CoV-2 on Surfaces. mSphere 2020, 5, 00441-20. [Google Scholar] [CrossRef] [PubMed]
- Mondelli, M.U.; Colaneri, M.; Seminari, E.M.; Baldanti, F.; Bruno, R. Low risk of SARS-CoV-2 transmission by fomites in real-life conditions. Lancet Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Goldman, E. Exaggerated risk of transmission of COVID-19 by fomites. Lancet Infect. Dis. 2020, 20, 892–893. [Google Scholar] [CrossRef]
- Dibner, J. Fecal-oral transmission of COVID-19: Could hypochlorhydria play a role? J. Med. Virol. 2021, 93, 166–167. [Google Scholar] [CrossRef]
- Sun, J.; Zhu, A.; Li, H.; Zheng, K.; Zhuang, Z.; Chen, Z.; Shi, Y.; Zhang, Z.; Chen, S.-B.; Liu, X.; et al. Isolation of infectious SARS-CoV-2 from urine of a COVID-19 patient. Emerg. Microbes Infect. 2020, 9, 991–993. [Google Scholar] [CrossRef]
- Wang, J.; Feng, H.; Zhang, S.; Ni, Z.; Ni, L.; Chen, Y.; Zhuo, L.; Zhong, Z.; Qu, T. SARS-CoV-2 RNA detection of hospital isolation wards hygiene monitoring during the Coronavirus Disease 2019 outbreak in a Chinese hospital. Int. J. Infect. Dis. 2020, 94, 103–106. [Google Scholar] [CrossRef]
- Xiao, F.; Sun, J.; Xu, Y.; Li, F.; Huang, X.; Li, H.; Zhao, J.; Huang, J.; Zhao, J. Infectious SARS-CoV-2 in Feces of Patient with Severe COVID-19. Emerg. Infect. Dis. 2020, 26, 1920–1922. [Google Scholar] [CrossRef]
- Ahmed, W.; Angel, N.; Edson, J.; Bibby, K.; Bivins, A.; O’Brien, J.W.; Choi, P.M.; Kitajima, M.; Simpson, S.L.; Li, J.; et al. First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: A proof of concept for the wastewater surveillance of COVID-19 in the community. Sci. Total. Environ. 2020, 728, 138764. [Google Scholar] [CrossRef] [PubMed]
- Bhowmick, G.D.; Dhar, D.; Nath, D.; Ghangrekar, M.M.; Banerjee, R.; Das, S.; Chatterjee, J. Coronavirus disease 2019 (COVID-19) outbreak: Some serious consequences with urban and rural water cycle. NPJ Clean Water 2020, 3, 1–8. [Google Scholar] [CrossRef]
- Peng, L.; Liu, J.; Xu, W.; Luo, Q.; Chen, D.; Lei, Z.; Huang, Z.; Li, X.; Deng, K.; Lin, B.; et al. SARS-CoV-2 can be detected in urine, blood, anal swabs, and oropharyngeal swabs specimens. J. Med. Virol. 2020, 92, 1676–1680. [Google Scholar] [CrossRef] [PubMed]
- Chang, L.; Zhao, L.; Gong, H.; Wang, L.; Wang, L. Severe Acute Respiratory Syndrome Coronavirus 2 RNA Detected in Blood Donations. Emerg. Infect. Dis. 2020, 26, 1631–1633. [Google Scholar] [CrossRef]
- Chang, L.; Yan, Y.; Wang, L. Coronavirus Disease 2019: Coronaviruses and Blood Safety. Transfus. Med. Rev. 2020, 34, 75–80. [Google Scholar] [CrossRef]
- Hamer, S.A.; Pauvolid-Corrêa, A.; Zecca, I.B.; Davila, E.; Auckland, L.D.; Roundy, C.M.; Tang, W.; Torchetti, M.; Killian, M.L.; Jenkins-Moore, M.; et al. Natural SARS-CoV-2 infections, including virus isolation, among serially tested cats and dogs in households with confirmed human COVID-19 cases in Texas, USA. bioRxiv 2020. [Google Scholar] [CrossRef]
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Chen, X.; Deng, X.; Chen, Z.; Gong, H.; Yan, H.; Wu, Q.; Shi, H.; Lai, S.; Ajelli, M.; et al. Disease burden and clinical severity of the first pandemic wave of COVID-19 in Wuhan, China. Nat. Commun. 2020, 11, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Cleary, S.J.; Pitchford, S.C.; Amison, R.T.; Carrington, R.; Cabrera, C.L.R.; Magnen, M.; Looney, M.R.; Gray, E.; Page, C.P. Animal models of mechanisms of SARS-CoV-2 infection and COVID-19 pathology. Br. J. Pharmacol. 2020, 177, 4851–4865. [Google Scholar] [CrossRef]
- Sarkar, J.; Guha, R. Infectivity, virulence, pathogenicity, host-pathogen interactions of SARS and SARS-CoV-2 in experimental animals: A systematic review. Vet. Res. Commun. 2020, 44, 101–110. [Google Scholar] [CrossRef]
- Bartels, C.J.; Fakhri, A.Q.; Shams, M.H.; Briscoe, R.P.; Schreuder, B.E. Livestock mortality and offtake in sheep and goat flocks of livestock owners making use of services offered by paravets in West Afghanistan. Prev. Vet. Med. 2017, 146, 79–85. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization (WHO). Archived: WHO Timeline—COVID-19. Available online: https://www.who.int/news/item/27-04-2020-who-timeline---covid-19 (accessed on 1 December 2020).
- De Morais, H.A.; dos Santos, A.P.; Nascimento, N.C.D.; Kmetiuk, L.B.; Barbosa, D.S.; Brandão, P.E.; Guimarães, A.M.S.; Pettan-Brewer, C.; Biondo, A.W. Natural Infection by SARS-CoV-2 in Companion Animals: A Review of Case Reports and Current Evidence of Their Role in the Epidemiology of COVID-19. Front. Vet. Sci. 2020, 7, 591216. [Google Scholar] [CrossRef] [PubMed]
- McAloose, D.; Laverack, M.; Wang, L.; Killian, M.L.; Caserta, L.C.; Yuan, F.; Mitchell, P.K.; Queen, K.; Mauldin, M.R.; Cronk, B.D.; et al. From People to Panthera: Natural SARS-CoV-2 Infection in Tigers and Lions at the Bronx Zoo. mBio 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Lam, H.M.; Ratmann, O.; Boni, M.F. Improved Algorithmic Complexity for the 3SEQ Recombination Detection Algorithm. Mol. Biol. Evol. 2018, 35, 247–251. [Google Scholar] [CrossRef]
- Ky, B.; Mann, D.L. COVID-19 Clinical Trials. JACC Cardio Oncol. 2020, 2, 254–269. [Google Scholar] [CrossRef]
- Wrapp, D.; Wang, N.; Corbett, K.S.; Goldsmith, J.A.; Hsieh, C.-L.; Abiona, O.; Graham, B.S.; McLellan, J.S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.-H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280. [Google Scholar] [CrossRef]
- Stopsack, K.H.; Mucci, L.A.; Antonarakis, E.S.; Nelson, P.S.; Kantoff, P.W. TMPRSS2 and COVID-19: Serendipity or Opportunity for Intervention? Cancer Discov. 2020, 10, 779–782. [Google Scholar] [CrossRef]
- Hammer, A.S.; Quaade, M.L.; Rasmussen, T.B.; Fonager, J.; Rasmussen, M.; Mundbjerg, K.; Lohse, L.; Strandbygaard, B.; Jørgensen, C.S.; Alfaro-Núñez, A.; et al. SARS-CoV-2 Transmission between Mink (Neovison vison) and Humans, Denmark. Emerg. Infect. Dis. 2021, 27, 547–551. [Google Scholar] [CrossRef]
- Koopmans, M. SARS-CoV-2 and the human-animal interface: Outbreaks on mink farms. Lancet Infect. Dis. 2021, 21, 18–19. [Google Scholar] [CrossRef]
- Jaimes, J.A.; André, N.M.; Chappie, J.S.; Millet, J.K.; Whittaker, G.R. Phylogenetic Analysis and Structural Modeling of SARS-CoV-2 Spike Protein Reveals an Evolutionary Distinct and Proteolytically Sensitive Activation Loop. J. Mol. Biol. 2020, 432, 3309–3325. [Google Scholar] [CrossRef] [PubMed]
- Choudhary, S.; Silakari, O. Scaffold morphing of arbidol (umifenovir) in search of multi-targeting therapy halting the interaction of SARS-CoV-2 with ACE2 and other proteases involved in COVID-19. Virus Res. 2020, 289, 198146. [Google Scholar] [CrossRef]
- Damas, J.; Hughes, G.M.; Keough, K.C.; Painter, C.A.; Persky, N.S.; Corbo, M.; Hiller, M.; Koepfli, K.-P.; Pfenning, A.R.; Zhao, H.; et al. Broad host range of SARS-CoV-2 predicted by comparative and structural analysis of ACE2 in vertebrates. Proc. Natl. Acad. Sci. USA 2020, 117, 22311–22322. [Google Scholar] [CrossRef] [PubMed]
- Lan, J.; Ge, J.; Yu, J.; Shan, S.; Zhou, H.; Fan, S.; Zhang, Q.; Shi, X.; Wang, Q.; Zhang, L.; et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 2020, 581, 215–220. [Google Scholar] [CrossRef]
- Garry, R.F. Mutations Arising in SARS-CoV-2 Spike on Sustained Human-to-Human Transmission and Human-to-Animal Passage. Available online: https://virological.org/t/mutations-arising-in-sars-cov-2-spike-on-sustained-human-to-human-transmission-and-human-to-animal-passage/578 (accessed on 22 January 2021).
- Tang, Y.-D.; Li, Y.-M.; Sun, J.; Zhang, H.-L.; Wang, T.-Y.; Sun, M.-X.; Yang, Y.-L.; Hu, X.-L.; Zhao, J.; Cai, X.-H. Cell entry of SARS-CoV-2 conferred by angiotensin-converting enzyme 2 (ACE2) of different species. bioRxiv 2020. [Google Scholar] [CrossRef]
- Kim, D.; Lee, J.-Y.; Yang, J.-S.; Kim, J.W.; Kim, V.N.; Chang, H. The Architecture of SARS-CoV-2 Transcriptome. Cell 2020, 181, 914–921.e10. [Google Scholar] [CrossRef] [PubMed]
- Rehman, S.U.; Shafique, L.; Ihsan, A.; Liu, Q. Evolutionary Trajectory for the Emergence of Novel Coronavirus SARS-CoV-2. Pathogens 2020, 9, 240. [Google Scholar] [CrossRef]
- Zhou, P.; Fan, H.; Lan, T.; Yáng, X.-L.; Shi, W.-F.; Zhang, W.; Zhu, Y.; Zhang, Y.-W.; Xie, Q.-M.; Mani, S.; et al. Fatal swine acute diarrhoea syndrome caused by an HKU2-related coronavirus of bat origin. Nature 2018, 556, 255–258. [Google Scholar] [CrossRef]
- Jung, K.; Saif, L.; Wang, Q. Porcine epidemic diarrhea virus (PEDV): An update on etiology, transmission, pathogenesis, and prevention and control. Virus Res. 2020, 286, 198045. [Google Scholar] [CrossRef]
- Lau, S.K.P.; Woo, P.C.Y.; Li, K.S.; Huang, Y.; Wang, M.; Lam, C.S.; Xu, H.; Guo, R.; Chan, K.-H.; Zheng, B.-J.; et al. Complete genome sequence of bat coronavirus HKU2 from Chinese horseshoe bats revealed a much smaller spike gene with a different evolutionary lineage from the rest of the genome. Virology 2007, 367, 428–439. [Google Scholar] [CrossRef]
- Wang, W.; Lin, X.-D.; Guo, W.-P.; Zhou, R.-H.; Wang, M.-R.; Wang, C.-Q.; Ge, S.; Mei, S.-H.; Li, M.-H.; Shi, M.; et al. Discovery, diversity and evolution of novel coronaviruses sampled from rodents in China. Virology 2015, 474, 19–27. [Google Scholar] [CrossRef]
- Corman, V.M.; Baldwin, H.J.; Tateno, A.F.; Zerbinati, R.M.; Annan, A.; Owusu, M.; Nkrumah, E.E.; Maganga, G.D.; Oppong, S.; Adu-Sarkodie, Y.; et al. Evidence for an Ancestral Association of Human Coronavirus 229E with Bats. J. Virol. 2015, 89, 11858–11870. [Google Scholar] [CrossRef] [PubMed]
- De Caro, N.; Martella, V.; Saif, L.; Buonavoglia, C. COVID-19 from veterinary medicine and one health perspectives: What animal coronaviruses have taught us. Res. Vet. Sci. 2020, 131, 21–23. [Google Scholar] [CrossRef] [PubMed]
- Lamers, M.M.; Smits, S.L.; Hundie, G.B.; Provacia, L.B.; Koopmans, M.P.G.; Osterhaus, A.D.M.E.; Haagmans, B.L.; Raj, V.S. Naturally occurring recombination in ferret coronaviruses revealed by complete genome characterization. J. Gen. Virol. 2016, 97, 2180–2186. [Google Scholar] [CrossRef] [PubMed]
- Lau, S.K.P.; Feng, Y.; Chen, H.; Luk, H.K.H.; Yang, W.-H.; Li, K.S.M.; Zhang, Y.-Z.; Huang, Y.; Song, Z.-Z.; Chow, F.W.-N.; et al. Severe Acute Respiratory Syndrome (SARS) Coronavirus ORF8 Protein Is Acquired from SARS-Related Coronavirus from Greater Horseshoe Bats through Recombination. J. Virol. 2015, 89, 10532–10547. [Google Scholar] [CrossRef] [PubMed]
- Corman, V.M.; Eckerle, I.; Memish, Z.A.; Liljander, A.M.; Dijkman, R.; Jonsdottir, H.; Ngeiywa, K.J.Z.J.; Kamau, E.; Younan, M.; Al Masri, M.; et al. Link of a ubiquitous human coronavirus to dromedary camels. Proc. Natl. Acad. Sci. USA 2016, 113, 9864–9869. [Google Scholar] [CrossRef] [PubMed]
- Kin, N.; Miszczak, F.; Diancourt, L.; Caro, V.; Moutou, F.; Vabret, A.; Gouilh, M.A. Comparative molecular epidemiology of two closely related coronaviruses, bovine coronavirus (BCoV) and human coronavirus OC43 (HCoV-OC43), reveals a different evolutionary pattern. Infect. Genet. Evol. 2016, 40, 186–191. [Google Scholar] [CrossRef]
- Woo, P.C.; Lau, S.K.; Lam, C.S.; Lau, C.C.; Tsang, A.K.; Lau, J.H.; Bai, R.; Teng, J.L.; Tsang, C.C.; Wang, M.; et al. Discovery of Seven Novel Mammalian and Avian Coronaviruses in the Genus Deltacoronavirus Supports Bat Coronaviruses as the Gene Source of Alphacoronavirus and Betacoronavirus and Avian Coronaviruses as the Gene Source of Gammacoronavirus and Deltacoronavirus. J. Virol. 2012, 86, 3995–4008. [Google Scholar] [CrossRef]
- Woo, P.C.Y.; Lau, S.K.P.; Lam, C.S.F.; Tsang, A.K.L.; Hui, S.-W.; Fan, R.Y.Y.; Martelli, P.; Yuen, K.-Y. Discovery of a Novel Bottlenose Dolphin Coronavirus Reveals a Distinct Species of Marine Mammal Coronavirus in Gammacoronavirus. J. Virol. 2014, 88, 1318–1331. [Google Scholar] [CrossRef]
- Wang, Y.; Cui, X.; Chen, X.; Yang, S.; Ling, Y.; Song, Q.; Zhu, S.; Sun, L.; Li, C.; Li, Y.; et al. A recombinant infectious bronchitis virus from a chicken with a spike gene closely related to that of a turkey coronavirus. Arch. Virol. 2020, 165, 703–707. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.-Z.; Holmes, E.C. A Genomic Perspective on the Origin and Emergence of SARS-CoV-2. Cell 2020, 181, 223–227. [Google Scholar] [CrossRef] [PubMed]
- Bonilla-Aldana, D.K.; Dhama, K.; Rodriguez-Morales, A.J. Revisiting the One Health Approach in the Context of COVID-19: A Look into the Ecology of this Emerging Disease. Adv. Anim. Vet. Sci. 2020, 8, 8. [Google Scholar] [CrossRef]
- Deng, J.; Jin, Y.; Liu, Y.; Sun, J.; Hao, L.; Bai, J.; Huang, T.; Lin, D.; Jin, Y.; Tian, K. Serological survey of SARS-CoV-2 for experimental, domestic, companion and wild animals excludes intermediate hosts of 35 different species of animals. Transbound. Emerg. Dis. 2020, 67, 1745–1749. [Google Scholar] [CrossRef] [PubMed]
- Luan, J.; Lu, Y.; Jin, X.; Zhang, L. Spike protein recognition of mammalian ACE2 predicts the host range and an optimized ACE2 for SARS-CoV-2 infection. Biochem. Biophys. Res. Commun. 2020, 526, 165–169. [Google Scholar] [CrossRef] [PubMed]
- Ehsani, S. COVID-19 and iron dysregulation: Distant sequence similarity between hepcidin and the novel coronavirus spike glycoprotein. Biol. Direct 2020, 15, 1–13. [Google Scholar] [CrossRef]
- Sang, E.R.; Tian, Y.; Gong, Y.; Miller, L.C.; Sang, Y. Integrate structural analysis, isoform diversity, and interferon-inductive propensity of ACE2 to predict SARS-CoV2 susceptibility in vertebrates. Heliyon 2020, 6, e04818. [Google Scholar] [CrossRef]
- Brook, C.E.; Dobson, A.P. Bats as ‘special’ reservoirs for emerging zoonotic pathogens. Trends Microbiol. 2015, 23, 172–180. [Google Scholar] [CrossRef] [PubMed]
- Skirmuntt, E.C.; Escalera-Zamudio, M.; Teeling, E.C.; Smith, A.; Katzourakis, A. The Potential Role of Endogenous Viral Elements in the Evolution of Bats as Reservoirs for Zoonotic Viruses. Annu. Rev. Virol. 2020, 7, 103–119. [Google Scholar] [CrossRef]
- Jo, W.K.; de Oliveira-Filho, E.F.; Rasche, A.; Greenwood, A.D.; Osterrieder, K.; Drexler, J.F. Potential zoonotic sources of SARS-CoV-2 infections. Transbound. Emerg. Dis. 2020, 13872. [Google Scholar] [CrossRef]
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N.; et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2020, 395, 565–574. [Google Scholar] [CrossRef]
- Touati, R.; Haddad-Boubaker, S.; Ferchichi, I.; Messaoudi, I.; Ouesleti, A.E.; Triki, H.; Lachiri, Z.; Kharrat, M. Comparative genomic signature representations of the emerging COVID-19 coronavirus and other coronaviruses: High identity and possible recombination between Bat and Pangolin coronaviruses. Genomics 2020, 112, 4189–4202. [Google Scholar] [CrossRef]
- Wong, G.; Bi, Y.-H.; Wang, Q.-H.; Chen, X.-W.; Zhang, Z.-G.; Yao, Y.-G. Zoonotic origins of human coronavirus 2019 (HCoV-19 /SARS-CoV-2): Why is this work important? Zool. Res. 2020, 41, 213–219. [Google Scholar] [CrossRef]
- Olival, K.J.; Cryan, P.M.; Amman, B.R.; Baric, R.; Blehert, D.S.; Brook, C.E.; Calisher, C.H.; Castle, K.T.; Coleman, J.T.H.; Daszak, P.; et al. Possibility for reverse zoonotic transmission of SARS-CoV-2 to free-ranging wildlife: A case study of bats. PLoS Pathog. 2020, 16, e1008758. [Google Scholar] [CrossRef]
- Li, X.; Zai, J.; Zhao, Q.; Nie, Q.; Li, Y.; Foley, B.T.; Chaillon, A. Evolutionary history, potential intermediate animal host, and cross-species analyses of SARS-CoV-2. J. Med. Virol. 2020, 92, 602–611. [Google Scholar] [CrossRef]
- Liu, P.; Jiang, J.-Z.; Wan, X.-F.; Hua, Y.; Li, L.; Zhou, J.; Wang, X.; Hou, F.; Chen, J.; Zou, J.; et al. Are pangolins the intermediate host of the 2019 novel coronavirus (SARS-CoV-2)? PLoS Pathog. 2020, 16, e1008421. [Google Scholar] [CrossRef]
- Zhang, Z.; Xiao, K.; Zhang, X.; Roy, A.; Shen, Y. Emergence of SARS-like coronavirus in China: An update. J. Infect. 2020, 80, e28–e29. [Google Scholar] [CrossRef]
- Wang, J.; Xu, X.; Zhou, X.; Chen, P.; Liang, H.; Li, X.; Zhong, W.; Hao, P. Molecular simulation of SARS-CoV-2 spike protein binding to pangolin ACE2 or human ACE2 natural variants reveals altered susceptibility to infection. J. Gen. Virol. 2020, 101, 921–924. [Google Scholar] [CrossRef]
- Goh, G.K.-M.; Dunker, A.K.; Foster, J.A.; Uversky, V.N. Shell Disorder Analysis Suggests That Pangolins Offered a Window for a Silent Spread of an Attenuated SARS-CoV-2 Precursor among Humans. J. Proteome Res. 2020, 19, 4543–4552. [Google Scholar] [CrossRef]
- Zhang, T.; Wu, Q.; Zhang, Z. Probable Pangolin Origin of SARS-CoV-2 Associated with the COVID-19 Outbreak. Curr. Biol. 2020, 30, 1346–1351.e2. [Google Scholar] [CrossRef]
- Liu, P.; Chen, W.; Chen, J.-P. Viral Metagenomics Revealed Sendai Virus and Coronavirus Infection of Malayan Pangolins (Manis javanica). Viruses 2019, 11, 979. [Google Scholar] [CrossRef]
- Wassenaar, T.M.; Zou, Y. 2019_nCoV/SARS-CoV-2: Rapid classification of betacoronaviruses and identification of Traditional Chinese Medicine as potential origin of zoonotic coronaviruses. Lett. Appl. Microbiol. 2020, 70, 342–348. [Google Scholar] [CrossRef]
- Cagliani, R.; Forni, D.; Clerici, M.; Sironi, M. Computational Inference of Selection Underlying the Evolution of the Novel Coronavirus, Severe Acute Respiratory Syndrome Coronavirus 2. J. Virol. 2020, 94, 94. [Google Scholar] [CrossRef]
- Stout, A.E.; André, N.M.; Jaimes, J.A.; Millet, J.K.; Whittaker, G.R. Coronaviruses in cats and other companion animals: Where does SARS-CoV-2/COVID-19 fit? Vet. Microbiol. 2020, 247, 108777. [Google Scholar] [CrossRef]
- Newman, A.; Smith, D.; Ghai, R.R.; Wallace, R.M.; Torchetti, M.K.; LoIacono, C.; Murrell, L.S.; Carpenter, A.; Moroff, S.; Rooney, J.A.; et al. First Reported Cases of SARS-CoV-2 Infection in Companion Animals—New York, March–April 2020. MMWR Morb. Mortal. Wkly. Rep. 2020, 69, 710–713. [Google Scholar] [CrossRef]
- Sailleau, C.; Dumarest, M.; Vanhomwegen, J.; Delaplace, M.; Caro, V.; Kwasiborski, A.; Hourdel, V.; Chevaillier, P.; Barbarino, A.; Comtet, L.; et al. First detection and genome sequencing of SARS-CoV-2 in an infected cat in France. Transbound. Emerg. Dis. 2020, 67, 2324–2328. [Google Scholar] [CrossRef]
- Zhang, Q.; Zhang, H.; Gao, J.; Huang, K.; Yang, Y.; Hui, X.; He, X.; Li, C.; Gong, W.; Zhang, Y.; et al. A serological survey of SARS-CoV-2 in cat in Wuhan. Emerg. Microbes Infect. 2020, 9, 2013–2019. [Google Scholar] [CrossRef]
- Thompson, H. A Cat Appears to Have Caught the Coronavirus, but It’s Complicated. Available online: https://www.sciencenews.org/article/cats-animals-pets-coronavirus-covid19 (accessed on 1 December 2020).
- Garigliany, M.; van Laere, A.-S.; Clercx, C.; Giet, D.; Escriou, N.; Huon, C.; van der Werf, S.; Eloit, M.; Desmecht, D. SARS-CoV-2 Natural Transmission from Human to Cat, Belgium, March 2020. Emerg. Infect. Dis. 2020, 26, 3069–3071. [Google Scholar] [CrossRef]
- Agriculture Fisheries and Conservation (AFCD). Pet Cat Tests Positive for COVID-19 Virus; Agriculture Fisheries and Conservation: Hong Kong, China, 2020. [Google Scholar]
- Bosco-Lauth, A.M.; Hartwig, A.E.; Porter, S.M.; Gordy, P.W.; Nehring, M.; Byas, A.D.; Vandewoude, S.; Ragan, I.K.; Maison, R.M.; Bowen, R.A. Experimental infection of domestic dogs and cats with SARS-CoV-2: Pathogenesis, transmission, and response to reexposure in cats. Proc. Natl. Acad. Sci. USA 2020, 117, 26382–26388. [Google Scholar] [CrossRef]
- Segalés, J.; Puig, M.; Rodon, J.; Avila-Nieto, C.; Carrillo, J.; Cantero, G.; Terrón, M.T.; Cruz, S.; Parera, M.; Noguera-Julián, M.; et al. Detection of SARS-CoV-2 in a cat owned by a COVID-19−affected patient in Spain. Proc. Natl. Acad. Sci. USA 2020, 117, 24790–24793. [Google Scholar] [CrossRef]
- Halfmann, P.J.; Hatta, M.; Chiba, S.; Maemura, T.; Fan, S.; Takeda, M.; Kinoshita, N.; Hattori, S.-I.; Sakai-Tagawa, Y.; Iwatsuki-Horimoto, K.; et al. Transmission of SARS-CoV-2 in Domestic Cats. New Engl. J. Med. 2020, 383, 592–594. [Google Scholar] [CrossRef]
- Gaudreault, N.N.; Trujillo, J.D.; Carossino, M.; Meekins, D.A.; Morozov, I.; Madden, D.W.; Indran, S.V.; Bold, D.; Balaraman, V.; Kwon, T.; et al. SARS-CoV-2 infection, disease and transmission in domestic cats. Emerg. Microbes Infect. 2020, 9, 2322–2332. [Google Scholar] [CrossRef]
- Muñoz-Fontela, C.; Dowling, W.E.; Funnell, S.G.P.; Gsell, P.-S.; Riveros-Balta, A.X.; Albrecht, R.A.; Andersen, H.; Baric, R.S.; Carroll, M.W.; Cavaleri, M.; et al. Animal models for COVID-19. Nature 2020, 586, 509–515. [Google Scholar] [CrossRef]
- Patterson, E.I.; Elia, G.; Grassi, A.; Giordano, A.; Desario, C.; Medardo, M.; Smith, S.L.; Anderson, E.R.; Prince, T.; Patterson, G.T.; et al. Evidence of exposure to SARS-CoV-2 in cats and dogs from households in Italy. Nat. Commun. 2020, 11, 1–5. [Google Scholar] [CrossRef]
- Bartlett, S.L.; Diel, D.G.; Wang, L.; Zec, S.; Laverack, M.; Martins, M.; Caserta, L.C.; Killian, M.L.; Terio, K.; Olmstead, C.; et al. SARS-CoV-2 infection and longitudinal fecal screening in Malayan tigers (Panthera tigris jacksoni), Amur tigers (Panthera tigris altaica), and African lions (Panthera leo krugeri) at the Bronx zoo, New York, USA. J. Zoo Wildl. Med. 2021, 51, 733–744. [Google Scholar] [CrossRef]
- Wise, A.G.; Kiupel, M.; Maes, R. Molecular characterization of a novel coronavirus associated with epizootic catarrhal enteritis (ECE) in ferrets. Virology 2006, 349, 164–174. [Google Scholar] [CrossRef]
- Chu, Y.-K.; Ali, G.D.; Jia, F.; Li, Q.; Kelvin, D.; Couch, R.C.; Harrod, K.S.; Hutt, J.A.; Cameron, C.; Weiss, S.R.; et al. The SARS-CoV ferret model in an infection–challenge study. Virology 2008, 374, 151–163. [Google Scholar] [CrossRef]
- Martina, B.E.E.; Haagmans, B.L.; Kuiken, T.; Fouchier, R.A.M.; Rimmelzwaan, G.F.; van Amerongen, G.; Peiris, J.S.M.; Lim, W.; Osterhaus, A.D.M.E. SARS virus infection of cats and ferrets. Nat. Cell Biol. 2003, 425, 915. [Google Scholar] [CrossRef]
- Richard, M.; Kok, A.; de Meulder, D.; Bestebroer, T.M.; Lamers, M.M.; Okba, N.M.A.; van Vlissingen, M.F.; Rockx, B.; Haagmans, B.L.; Koopmans, M.; et al. SARS-CoV-2 is transmitted via contact and via the air between ferrets. Nat. Commun. 2020, 11, 3496. [Google Scholar] [CrossRef]
- Ter-Meulen, J.; Bakker, A.B.H.; Brink, E.N.V.D.; Weverling, G.J.; Martina, B.E.E.; Haagmans, B.L.; Kuiken, T.; de Kruif, J.; Preiser, W.; Spaan, W.; et al. Human monoclonal antibody as prophylaxis for SARS coronavirus infection in ferrets. Lancet 2004, 363, 2139–2141. [Google Scholar] [CrossRef]
- Weingartl, H.; Czub, M.; Czub, S.; Neufeld, J.; Marszal, P.; Gren, J.; Smith, G.; Jones, S.; Proulx, R.; Deschambault, Y.; et al. Immunization with Modified Vaccinia Virus Ankara-Based Recombinant Vaccine against Severe Acute Respiratory Syndrome Is Associated with Enhanced Hepatitis in Ferrets. J. Virol. 2004, 78, 12672–12676. [Google Scholar] [CrossRef]
- Kim, Y.-I.; Kim, S.-G.; Kim, S.-M.; Kim, E.-H.; Park, S.-J.; Yu, K.-M.; Chang, J.-H.; Lee, S.; Casel, M.A.B.; Um, J.; et al. Infection and Rapid Transmission of SARS-CoV-2 in Ferrets. Cell Host Microbe 2020, 27, 704–709. [Google Scholar] [CrossRef] [PubMed]
- Skarlupka, A.L.; Ross, T.M. Immune Imprinting in the Influenza Ferret Model. Vaccines 2020, 8, 173. [Google Scholar] [CrossRef] [PubMed]
- Belser, J.A.; Eckert, A.M.; Tumpey, T.M.; Maines, T.R. Complexities in Ferret Influenza Virus Pathogenesis and Transmission Models. Microbiol. Mol. Biol. Rev. 2016, 80, 733–744. [Google Scholar] [CrossRef]
- Sawatzki, K.; Hil, N.J.; Puryear, W.B.; Foss, A.D.; Stone, J.J.; Runstadler, J.A. Ferrets not infected by SARS-CoV-2 in a high-exposure domestic setting. bioRxiv 2020. [Google Scholar] [CrossRef]
- Oreshkova, N.; Molenaar, R.J.; Vreman, S.; Harders, F.; Munnink, B.B.O.; Hakze, R.; Gerhards, N.; Tolsma, P.; Bouwstra, R.; Sikkema, R.; et al. SARS-CoV2 infection in farmed mink, Netherlands. bioRxiv 2020. [Google Scholar] [CrossRef]
- United States Department of Agriculture (USDA). Dutch Mink Industry to Close in 2021 Due to Coronavirus. Available online: https://apps.fas.usda.gov/newgainapi/api/Report/DownloadReportByFileName?fileName=Dutch Mink Industry to Close in 2021 Due to Coronavirus _The Hague_Netherlands_08-28-2020 (accessed on 1 December 2020).
- European Centre for Disease Prevention and Control Detection Detection of new SARS-CoV-2 Variants Related to Mink. Available online: https://www.ecdc.europa.eu/sites/default/files/documents/RRA-SARS-CoV-2-in-mink-12-nov-2020.pdf (accessed on 20 December 2020).
- USDA (United States Department of Agriculture). USDA Confirms SARS-CoV-2 in Mink in Utah. Available online: https://www.aphis.usda.gov/aphis/newsroom/stakeholder-info/sa_by_date/sa-2020/sa-08/sare-cov-2-mink (accessed on 22 January 2021).
- International Society for Infectious Diseases. PRO/AH/EDR> COVID-19 Update (209): Netherlands (NB) Farmed Mink, Animal-to-Human, Cat, Epid. Available online: https://promedmail.org/promed-post/?id=20200525.7375359 (accessed on 22 January 2021).
- Monchâtre-Leroy, E.; Boué, F.; Boucher, J.-M.; Renault, C.; Moutou, F.; Gouilh, M.A.; Umhang, G. Identification of Alpha and Beta Coronavirus in Wildlife Species in France: Bats, Rodents, Rabbits, and Hedgehogs. Viruses 2017, 9, 364. [Google Scholar] [CrossRef]
- Gryseels, S.; de Bruyn, L.; Gyselings, R.; Calvignac-Spencer, S.; Leendertz, F.; Leirs, H. Risk of human-to-wildlife transmission of SARS-CoV-2. Preprints 2020. [Google Scholar] [CrossRef]
- Fagre, A.; Lewis, J.; Eckley, M.; Zhan, S.; Rocha, S.M.; Sexton, N.R.; Burke, B.; Geiss, B.; Peersen, O.; Kading, R.; et al. SARS-CoV-2 infection, neuropathogenesis and transmission among deer mice: Implications for reverse zoonosis to New World rodents. bioRxiv 2020. [Google Scholar] [CrossRef]
- Bedford, N.L.; Hoekstra, H.E. Peromyscus mice as a model for studying natural variation. eLife 2015, 4, e06813. [Google Scholar] [CrossRef]
- Bron, G.M.; Malavé, C.M.; Boulerice, J.T.; Osorio, J.E.; Rocke, T.E. Plague-Positive Mouse Fleas on Mice Before Plague Induced Die-Offs in Black-Tailed and White-Tailed Prairie Dogs. Vect. Borne Zoonotic Dis. 2019, 19, 486–493. [Google Scholar] [CrossRef]
- Davis, S.; Calvet, E. Fluctuating Rodent Populations and Risk to Humans from Rodent-Borne Zoonoses. Vect. Borne Zoonotic Dis. 2005, 5, 305–314. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Wang, H.; Tang, X.; Fang, S.; Ma, D.; Du, C.; Wang, Y.; Pan, H.; Yao, W.; Zhang, R.; et al. SARS-CoV-2 and Three Related Coronaviruses Utilize Multiple ACE2 Orthologs and Are Potently Blocked by an Improved ACE2-Ig. J. Virol. 2020, 94. [Google Scholar] [CrossRef] [PubMed]
- Dagotto, G.; Yu, J.; Barouch, D.H. Approaches and Challenges in SARS-CoV-2 Vaccine Development. Cell Host Microbe 2020, 28, 364–370. [Google Scholar] [CrossRef]
- Jiang, S.; Hillyer, C.; Du, L. Neutralizing Antibodies against SARS-CoV-2 and Other Human Coronaviruses. Trends Immunol. 2020, 41, 355–359. [Google Scholar] [CrossRef] [PubMed]
- Tian, J.H.; Patel, N.; Haupt, R.; Zhou, H.; Weston, S.; Hammond, H.; Lague, J.; Portnoff, A.D.; Norton, J.; Guebre-Xabier, M.; et al. SARS-CoV-2 spike glycoprotein vaccine candidate NVX-CoV2373 elicits immunogenicity in baboons and protection in mice. bioRxiv 2020. [Google Scholar] [CrossRef]
- Dinnon, K.H.; Leist, S.R.; Schäfer, A.; Edwards, C.E.; Martinez, D.R.; Montgomery, S.A.; West, A.; Yount, B.L., Jr.; Hou, Y.J.; Adams, L.E.; et al. A mouse-adapted model of SARS-CoV-2 to test COVID-19 countermeasures. Nature 2020, 586, 560–566. [Google Scholar] [CrossRef]
- Younes, S.; Younes, N.; Shurrab, F.; Nasrallah, G.K. Severe acute respiratory syndrome coronavirus-2 natural animal reservoirs and experimental models: Systematic review. Rev. Med. Virol. 2020, e2196. [Google Scholar] [CrossRef]
- Roberts, A.; Vogel, L.; Guarner, J.; Hayes, N.; Murphy, B.; Zaki, S.; Subbarao, K. Severe Acute Respiratory Syndrome Coronavirus Infection of Golden Syrian Hamsters. J. Virol. 2005, 79, 503–511. [Google Scholar] [CrossRef]
- Imai, M.; Iwatsuki-Horimoto, K.; Hatta, M.; Loeber, S.; Halfmann, P.J.; Nakajima, N.; Watanabe, T.; Ujie, M.; Takahashi, K.; Ito, M.; et al. Syrian hamsters as a small animal model for SARS-CoV-2 infection and countermeasure development. Proc. Natl. Acad. Sci. USA 2020, 117, 16587–16595. [Google Scholar] [CrossRef]
- Sia, S.F.; Yan, L.-M.; Chin, A.W.H.; Fung, K.; Choy, K.-T.; Wong, A.Y.L.; Kaewpreedee, P.; Perera, R.A.P.M.; Poon, L.L.M.; Nicholls, J.M.; et al. Pathogenesis and transmission of SARS-CoV-2 in golden hamsters. Nat. Cell Biol. 2020, 583, 834–838. [Google Scholar] [CrossRef]
- Corman, V.M.; Kallies, R.; Philipps, H.; Göpner, G.; Müller, M.A.; Eckerle, I.; Brünink, S.; Drosten, C.; Drexler, J.F. Characterization of a Novel Betacoronavirus Related to Middle East Respiratory Syndrome Coronavirus in European Hedgehogs. J. Virol. 2014, 88, 717–724. [Google Scholar] [CrossRef]
- Tsoleridis, T.; Onianwa, O.; Horncastle, E.; Dayman, E.; Zhu, M.; Danjittrong, T.; Wachtl, M.; Behnke, J.M.; Chapman, S.; Strong, V.; et al. Discovery of Novel Alphacoronaviruses in European Rodents and Shrews. Viruses 2016, 8, 84. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Lin, X.-D.; Liao, Y.; Guan, X.-Q.; Guo, W.-P.; Xing, J.-G.; Holmes, E.C.; Zhang, Y.-Z. Discovery of a Highly Divergent Coronavirus in the Asian House Shrew from China Illuminates the Origin of the Alphacoronaviruses. J. Virol. 2017, 91, e00764-17. [Google Scholar] [CrossRef]
- Niller, H.H.; Angstwurm, K.; Rubbenstroth, D.; Schlottau, K.; Ebinger, A.; Giese, S.; Wunderlich, S.; Banas, B.; Forth, L.F.; Hoffmann, D.; et al. Zoonotic spillover infections with Borna disease virus 1 leading to fatal human encephalitis, 1999–2019: An epidemiological investigation. Lancet Infect. Dis. 2020, 20, 467–477. [Google Scholar] [CrossRef]
- Johne, R.; Tausch, S.H.; Grützke, J.; Falkenhagen, A.; Patzina-Mehling, C.; Beer, M.; Höper, D.; Ulrich, R.G. Distantly Related Rotaviruses in Common Shrews, Germany, 2004–2014. Emerg. Infect. Dis. 2019, 25, 2310–2314. [Google Scholar] [CrossRef]
- Wang, B.; Cai, C.-L.; Chun-Lin, C.; Zhang, W.; Zhu, Y.; Chen, W.-H.; Zhuo, F.; Shi, Z.-L.; Yang, X.-L. Detection and characterization of three zoonotic viruses in wild rodents and shrews from Shenzhen city, China. Virol. Sin. 2017, 32, 290–297. [Google Scholar] [CrossRef]
- Pangjai, D.; Maruyama, S.; Boonmar, S.; Kabeya, H.; Sato, S.; Nimsuphan, B.; Petkanchanapong, W.; Wootta, W.; Wangroongsarb, P.; Boonyareth, M.; et al. Prevalence of zoonotic Bartonella species among rodents and shrews in Thailand. Comp. Immunol. Microbiol. Infect. Dis. 2014, 37, 109–114. [Google Scholar] [CrossRef]
- Chomel, B.B.; Riley, P.Y. Hedgehog Zoonoses. Emerg. Infect. Dis. 2005, 11, 1146. [Google Scholar] [CrossRef]
- Lau, S.K.P.; Luk, H.K.H.; Wong, A.C.P.; Fan, R.Y.Y.; Lam, C.S.F.; Li, K.S.M.; Ahmed, S.S.; Chow, F.W.-N.; Cai, J.-P.; Zhu, X.; et al. Identification of a Novel Betacoronavirus (Merbecovirus) in Amur Hedgehogs from China. Viruses 2019, 11, 980. [Google Scholar] [CrossRef]
- Johansen, M.D.; Irving, A.; Montagutelli, X.; Tate, M.D.; Rudloff, I.; Nold, M.F.; Hansbro, N.G.; Kim, R.Y.; Donovan, C.; Liu, G.; et al. Animal and translational models of SARS-CoV-2 infection and COVID-19. Mucosal Immunol. 2020, 13, 877–891. [Google Scholar] [CrossRef]
- Bloch, J.I.; Boyer, D. Grasping Primate Origins. Science 2002, 298, 1606–1610. [Google Scholar] [CrossRef]
- Xue-Long, J.; Chen, S.-Y.; Nie, W.-H.; Jiang, X.-L.; Yao, Y.-G. Evaluating the Phylogenetic Position of Chinese Tree Shrew (Tupaia belangeri chinensis) Based on Complete Mitochondrial Genome: Implication for Using Tree Shrew as an Alternative Experimental Animal to Primates in Biomedical Research. J. Genet. Genom. 2012, 39, 131–137. [Google Scholar] [CrossRef]
- Yuan, B.; Yang, C.; Xia, X.; Zanin, M.; Wong, S.; Yang, F.; Chang, J.; Mai, Z.; Zhao, J.; Zhang, Y.-H.; et al. The tree shrew is a promising model for the study of influenza B virus infection. Virol. J. 2019, 16, 77. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.-F.; Zhao, J.; Zhu, Y.-T.; Wang, Y.; Liu, R.; Zhao, S.; Li, R.-F.; Yang, C.-G.; Li, J.-Q.; Zhong, N. The tree shrew provides a useful alternative model for the study of influenza H1N1 virus. Virol. J. 2013, 10, 111. [Google Scholar] [CrossRef] [PubMed]
- Li, F. Structure, Function, and Evolution of Coronavirus Spike Proteins. Annu. Rev. Virol. 2016, 3, 237–261. [Google Scholar] [CrossRef] [PubMed]
- Xie, Z.-C.; Riezu-Boj, J.-I.; Lasarte, J.-J.; Guillen, J.; Su, J.-H.; Civeira, M.-P.; Prieto, J. Transmission of Hepatitis C Virus Infection to Tree Shrews. Virology 1998, 244, 513–520. [Google Scholar] [CrossRef] [PubMed]
- Yan, R.Q.; Su, J.J.; Huang, D.R.; Gan, Y.C.; Yang, C.; Huang, G.H. Human hepatitis B virus and hepatocellular carcinoma I. Experimental infection of tree shrews with hepatitis B virus. J. Cancer Res. Clin. Oncol. 1996, 122, 283–288. [Google Scholar] [CrossRef]
- Li, J.-P.; Liao, Y.; Zhang, Y.; Wang, J.-J.; Wang, L.-C.; Feng, K.; Li, Q.-H.; Liu, L.-D. Experimental infection of tree shrews (Tupaia belangeri) with Coxsackie virus A16. Zool. Res. 2014, 35, 485–491. [Google Scholar]
- Pu, T.; Ding, C.; Li, Y.; Liu, X.; Li, H.; Duan, J.; Zhang, H.; Bi, Y.; Cun, W. Evaluate severe acute respiratory syndrome coronavirus 2 infectivity by pseudoviral particles. J. Med. Virol. 2020, 92, 1609–1614. [Google Scholar] [CrossRef]
- Xu, L.; Yu, D.-D.; Ma, Y.-H.; Yao, Y.-L.; Luo, R.-H.; Feng, X.-L.; Cai, H.-R.; Han, J.-B.; Wang, X.-H.; Li, M.-H.; et al. COVID-19-like symptoms observed in Chinese tree shrews infected with SARS-CoV-2. Zool. Res. 2020, 41, 517–526. [Google Scholar] [CrossRef]
- Zhao, Y.; Wang, J.; Kuang, D.; Xu, J.; Yang, M.; Ma, C.; Zhao, S.; Li, J.; Long, H.; Ding, K.; et al. Susceptibility of tree shrew to SARS-CoV-2 infection. Sci. Rep. 2020, 10, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.J.; Chen, W.X. Viral haemorrhagic disease in rabbits: A review. Vet. Res. Commun. 1989, 13, 205–212. [Google Scholar] [CrossRef] [PubMed]
- Yugo, D.M.; Cossaboom, C.M.; Meng, X.-J. Naturally Occurring Animal Models of Human Hepatitis E Virus Infection. ILAR J. 2014, 55, 187–199. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Luan, J.; Cui, H.; Zhang, L. ACE2 isoform diversity predicts the host susceptibility of SARS-CoV-2. Transbound. Emerg. Dis. 2020, 13773. [Google Scholar] [CrossRef]
- Kumar, A.; Pandey, S.N.; Pareek, V.; Narayan, R.K.; Faiq, M.A.; Kumari, C. Predicting susceptibility for SARS-CoV-2 infection in domestic and wildlife animals using ACE2 protein sequence homology. Zoo Biol. 2020, 21576. [Google Scholar] [CrossRef]
- Preziuso, S. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Exhibits High Predicted Binding Affinity to ACE2 from Lagomorphs (Rabbits and Pikas). Animals 2020, 10, 1460. [Google Scholar] [CrossRef]
- Mykytyn, A.Z.; Lamers, M.M.; Okba, N.M.; Breugem, T.I.; Schipper, D.; van den Doel, P.B.; van Run, P.; van Amerongen, G.; de Waal, L.; Koopmans, M.; et al. Susceptibility of rabbits to SARS-CoV-2. bioRxiv 2020. [Google Scholar] [CrossRef]
- Yekta, R.; Vahid-Dastjerdi, L.; Norouzbeigi, S.; Mortazavian, A.M. Food products as potential carriers of SARS-CoV-2. Food Control. 2021, 123, 107754. [Google Scholar] [CrossRef]
- Sit, T.H.C.; Brackman, C.J.; Ip, S.M.; Tam, K.W.S.; Law, P.Y.T.; To, E.M.W.; Yu, V.Y.T.; Sims, L.D.; Tsang, D.N.C.; Chu, D.K.W.; et al. Infection of dogs with SARS-CoV-2. Nat. Cell Biol. 2020, 586, 776–778. [Google Scholar] [CrossRef]
- Agriculture Fisheries and Conservation Department (AFCD). Detection of Low Level COVID-19 Virus in Pet Dog. Available online: https://www.info.gov.hk/gia/general/202002/202028/P2020022800013.htm.3 (accessed on 1 December 2020).
- Goumenou, M.; Spandidos, D.A.; Tsatsakis, A. [Editorial] Possibility of transmission through dogs being a contributing factor to the extreme Covid-19 outbreak in North Italy. Mol. Med. Rep. 2020, 21, 2293–2295. [Google Scholar] [CrossRef]
- Delong, J. Dutch Minister Confirms Dog, Three Cats Have Caught Novel Coronavirus. Available online: https://www.reporter.am/dutch-minister-confirms-dog-three-cats-have-caught-novel-coronavirus/ (accessed on 1 December 2020).
- Miller, L.; Pacheco, G.J.; Janak, J.C.; Grimm, R.C.; Dierschke, N.A.; Baker, J.; Orman, J.A. Causes of Death in Military Working Dogs During Operation Iraqi Freedom and Operation Enduring Freedom, 2001–2013. Mil. Med. 2018, 183, e467–e474. [Google Scholar] [CrossRef] [PubMed]
- Rooney, N.J.; Clark, C.C.; Casey, R.A. Minimizing fear and anxiety in working dogs: A review. J. Vet. Behav. 2016, 16, 53–64. [Google Scholar] [CrossRef]
- Guan, Y.J.; Zheng, B.J.; He, Y.Q.; Liu, X.L.; Zhuang, Z.X.; Cheung, C.L.; Luo, S.W.; Li, P.H.; Zhang, L.J.; Butt, K.M.; et al. Isolation and Characterization of Viruses Related to the SARS Coronavirus from Animals in Southern China. Science 2003, 302, 276–278. [Google Scholar] [CrossRef] [PubMed]
- Freuling, C.; Müller, T.; Mettenleiter, T. Vaccines against pseudorabies virus (PrV). Vet. Microbiol. 2017, 206, 3–9. [Google Scholar] [CrossRef]
- Zhai, X.; Sun, J.; Yan, Z.; Zhang, J.; Zhao, J.; Zhao, Z.; Gao, Q.; He, W.-T.; Veit, M.; Su, S. Comparison of Severe Acute Respiratory Syndrome Coronavirus 2 Spike Protein Binding to ACE2 Receptors from Human, Pets, Farm Animals, and Putative Intermediate Hosts. J. Virol. 2020, 94. [Google Scholar] [CrossRef]
- Wang, W.; Yang, L.; Wronski, T.; Chen, S.; Hu, Y.; Huang, S. Captive breeding of wildlife resources—China’s revised supply-side approach to conservation. Wildl. Soc. Bull. 2019, 43, 425–435. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Sun, J.; Zhu, J.; Ding, X.; Lan, T.; Zhu, L.; Xiang, R.; Ding, P.; Wang, H.; Wang, X.; et al. Single-cell screening of SARS-CoV-2 target cells in pets, livestock, poultry and wildlife. bioRxiv 2020. [Google Scholar] [CrossRef]
- Mathavarajah, S.; Dellaire, G. Lions, tigers and kittens too: ACE2 and susceptibility to COVID-19. Evol. Med. Public Health 2020, 2020, 109–113. [Google Scholar] [CrossRef] [PubMed]
- Davoust, B.; Levasseur, A.; Mediannikov, O. Studies of nonhuman primates: Key sources of data on zoonoses and microbiota. New Microbes New Infect. 2018, 26, S104–S108. [Google Scholar] [CrossRef] [PubMed]
- Estes, J.D.; Wong, S.W.; Brenchley, J.M. Nonhuman primate models of human viral infections. Nat. Rev. Immunol. 2018, 18, 390–404. [Google Scholar] [CrossRef]
- Devaux, C.A.; Mediannikov, O.; Medkour, H.; Raoult, D. Infectious Disease Risk Across the Growing Human-Non Human Primate Interface: A Review of the Evidence. Front. Public Health 2019, 7, 305. [Google Scholar] [CrossRef] [PubMed]
- Bao, L.; Deng, W.; Gao, H.; Xiao, C.; Liu, J.; Xue, J.; Lv, Q.; Liu, J.; Yu, P.; Xu, Y.; et al. Reinfection could not occur in SARS-CoV-2 infected rhesus macaques. bioRxiv 2020. [Google Scholar] [CrossRef]
- Munster, V.J.; Feldmann, F.; Williamson, B.N.; van Doremalen, N.; Pérez-Pérez, L.; Schulz, J.; Meade-White, K.; Okumura, A.; Callison, J.; Brumbaugh, B.; et al. Respiratory disease in rhesus macaques inoculated with SARS-CoV-2. Nat. Cell Biol. 2020, 585, 268–272. [Google Scholar] [CrossRef]
- Yu, Z.; Cheng, K.; Sun, W.; Xin, Y.; Cai, J.; Ma, R.; Zhao, Q.; Li, L.; Huang, J.; Sang, X.; et al. Lowly pathogenic avian influenza (H9N2) infection in Plateau pika (Ochotona curzoniae), Qinghai Lake, China. Vet. Microbiol. 2014, 173, 132–135. [Google Scholar] [CrossRef]
- Rockx, B.; Kuiken, T.; Herfst, S.; Bestebroer, T.; Lamers, M.M.; Munnink, B.B.O.; de Meulder, D.; van Amerongen, G.; Brand, J.V.D.; Okba, N.M.A.; et al. Comparative pathogenesis of COVID-19, MERS, and SARS in a nonhuman primate model. Science 2020, 368, 1012–1015. [Google Scholar] [CrossRef]
- Woolsey, C.; Borisevich, V.; Prasad, A.N.; Agans, K.N.; Deer, D.J.; Dobias, N.S.; Heymann, J.C.; Foster, S.L.; Levine, C.B.; Medina, L.; et al. Establishment of an African green monkey model for COVID-19 and protection against re-infection. Nat. Immunol. 2021, 22, 86–98. [Google Scholar] [CrossRef]
- Gong, S.-R.; Bao, L.-L. The battle against SARS and MERS coronaviruses: Reservoirs and Animal Models. Anim. Model. Exp. Med. 2018, 1, 125–133. [Google Scholar] [CrossRef]
- Lu, S.; Zhao, Y.; Yu, W.; Yang, Y.; Gao, J.; Wang, J.; Kuang, D.; Yang, M.; Yang, J.; Ma, C.; et al. Comparison of nonhuman primates identified the suitable model for COVID-19. Signal. Transduct. Target. Ther. 2020, 5, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Jones, B.E.; Brown-Augsburger, P.L.; Corbett, K.S.; Westerndorf, K.; Davies, J.; Cujec, T.P.; Wiethoff, C.M.; Blackbourne, J.L.; Heinz, B.A.; Foster, D.; et al. LY-CoV555, a rapidly isolated potent neutralizing antibody, provides protection in a non-human primate model of SARS-CoV-2 infection. bioRxiv 2020. [Google Scholar] [CrossRef]
- Mercado, N.B.; Zahn, R.; Wegmann, F.; Loos, C.; Chandrashekar, A.; Yu, J.; Liu, J.; Peter, L.; Mcmahan, K.; Tostanoski, L.H.; et al. Single-shot Ad26 vaccine protects against SARS-CoV-2 in rhesus macaques. Nature 2020, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Gibbons, A. Captive Gorillas Test Positive for Coronavirus. Available online: https://www.sciencemag.org/news/2021/01/captive-gorillas-test-positive-coronavirus (accessed on 28 January 2021).
- San Diego Zoo Gorilla Troop at the San Diego Zoo Safari Park Test Positive for COVID-19: The Great Apes Continue to be Observed Closely by the San Diego Zoo Global Veterinary Team. Available online: https://zoo.sandiegozoo.org/pressroom/news-releases/gorilla-troop-san-diego-zoo-safari-park-test-positive-covid-19 (accessed on 28 January 2021).
- Negrey, J.D.; Reddy, R.B.; Scully, E.J.; Phillips-Garcia, S.; Owens, L.A.; Langergraber, K.E.; Mitani, J.C.; Thompson, M.E.; Wrangham, R.W.; Muller, M.N.; et al. Simultaneous outbreaks of respiratory disease in wild chimpanzees caused by distinct viruses of human origin. Emerg. Microbes Infect. 2019, 8, 139–149. [Google Scholar] [CrossRef]
- Callaway, E. Labs rush to study coronavirus in transgenic animals—Some are in short supply. Nat. Cell Biol. 2020, 579, 183. [Google Scholar] [CrossRef]
- Cohen, J. From mice to monkeys, animals studied for coronavirus answers. Science 2020, 368, 221–222. [Google Scholar] [CrossRef] [PubMed]
- Mohd, H.A.; Al-Tawfiq, J.A.; Memish, Z.A. Middle East Respiratory Syndrome Coronavirus (MERS-CoV) origin and animal reservoir. Virol. J. 2016, 13, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Alshukairi, A.N.; Zheng, J.; Zhao, J.; Nehdi, A.; Baharoon, S.A.; Layqah, L.; Bokhari, A.; Al Johani, S.M.; Samman, N.; Boudjelal, M.; et al. High Prevalence of MERS-CoV Infection in Camel Workers in Saudi Arabia. mBio 2018, 9, e01985-18. [Google Scholar] [CrossRef] [PubMed]
- Azhar, E.I.; El-Kafrawy, S.A.; Farraj, S.A.; Hassan, A.M.; Al-Saeed, M.S.; Hashem, A.M.; Madani, T.A. Evidence for Camel-to-Human Transmission of MERS Coronavirus. N. Engl. J. Med. 2014, 370, 2499–2505. [Google Scholar] [CrossRef]
- Kandeil, A.; Gomaa, M.; Shehata, M.; El-Taweel, A.; Kayed, A.E.; Abiadh, A.; Jrijer, J.; Moatasim, Y.; Kutkat, O.; Bagato, O.; et al. Middle East respiratory syndrome coronavirus infection in non-camelid domestic mammals. Emerg. Microbes Infect. 2019, 8, 103–108. [Google Scholar] [CrossRef]
- Fung, S.-Y.; Yuen, K.-S.; Ye, Z.-W.; Chan, C.-P.; Jin, D.-Y. A tug-of-war between severe acute respiratory syndrome coronavirus 2 and host antiviral defence: Lessons from other pathogenic viruses. Emerg. Microbes Infect. 2020, 9, 558–570. [Google Scholar] [CrossRef]
- So, R.T.Y.; Chu, D.K.W.; Miguel, E.; Perera, R.A.P.M.; Oladipo, J.O.; Fassi-Fihri, O.; Aylet, G.; Ko, R.L.W.; Zhou, Z.; Cheng, M.-S.; et al. Diversity of Dromedary Camel Coronavirus HKU23 in African Camels Revealed Multiple Recombination Events among Closely Related Betacoronaviruses of the Subgenus Embecovirus. J. Virol. 2019, 93. [Google Scholar] [CrossRef]
- Gai, J.; Ma, L.; Li, G.; Zhu, M.; Qiao, P.; Li, X.; Zhang, H.; Zhang, Y.; Chen, Y.; Gong, R.; et al. A potent neutralizing nanobody against SARS-CoV-2 with inhaled delivery potential. bioRxiv 2020. [Google Scholar] [CrossRef]
- Crameri, G.; Durr, P.A.; Klein, R.; Foord, A.; Yu, M.; Riddell, S.; Haining, J.; Johnson, D.; Hemida, M.G.; Barr, J.; et al. Experimental Infection and Response to Rechallenge of Alpacas with Middle East Respiratory Syndrome Coronavirus. Emerg. Infect. Dis. 2016, 22, 1071–1074. [Google Scholar] [CrossRef]
- Reusken, C.; Schilp, C.; Raj, V.S.; de Bruin, E.; Kohl, R.H.; Farag, E.A.; Haagmans, B.L.; Al-Romaihi, H.; Le Grange, F.; Bosch, B.-J.; et al. MERS-CoV Infection of Alpaca in a Region Where MERS-CoV is Endemic. Emerg. Infect. Dis. 2016, 22, 1129–1131. [Google Scholar] [CrossRef] [PubMed]
- Hanke, L.; Perez, L.V.; Sheward, D.J.; Das, H.; Schulte, T.; Moliner-Morro, A.; Corcoran, M.; Achour, A.; Hedestam, G.B.K.; Hällberg, B.M.; et al. An alpaca nanobody neutralizes SARS-CoV-2 by blocking receptor interaction. Nat. Commun. 2020, 11, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Nieto, G.E.V.; Jara, R.; Himelreichs, J.; Salinas, C.; Pinto, T.; Cheuquemilla, Y.; Margolles, Y.; del Rey, N.L.G.; Chacon, Z.M.; Cuevas, A.; et al. Fast isolation of sub-nanomolar affinity alpaca nanobody against the Spike RBD of SARS-CoV-2 by combining bacterial display and a simple single-step density gradient selection. bioRxiv 2020. [Google Scholar] [CrossRef]
- Saif, L.J. Bovine Respiratory Coronavirus. Vet. Clin. North. Am. Food Anim. Pr. 2010, 26, 349–364. [Google Scholar] [CrossRef]
- Clark, M. Bovine coronavirus. Br. Vet. J. 1993, 149, 51–70. [Google Scholar] [CrossRef]
- Qiu, Y.; Zhao, Y.-B.; Wang, Q.; Li, J.-Y.; Zhou, Z.-J.; Liao, C.-H.; Ge, X.-Y. Predicting the angiotensin converting enzyme 2 (ACE2) utilizing capability as the receptor of SARS-CoV-2. Microbes Infect. 2020, 22, 221–225. [Google Scholar] [CrossRef] [PubMed]
- Ulrich, L.; Wernike, K.; Hoffmann, D.; Mettenleiter, T.C.; Beer, M. Experimental Infection of Cattle with SARS-CoV-2. Emerg. Infect. Dis. 2020, 26, 2979–2981. [Google Scholar] [CrossRef] [PubMed]
- Meekins, D.A.; Morozov, I.; Trujillo, J.D.; Gaudreault, N.N.; Bold, D.; Carossino, M.; Artiaga, B.L.; Indran, S.V.; Kwon, T.; Balaraman, V.; et al. Susceptibility of swine cells and domestic pigs to SARS-CoV-2. Emerg. Microbes Infect. 2020, 9, 2278–2288. [Google Scholar] [CrossRef] [PubMed]
- Vergara-Alert, J.; Rodon, J.; Carrillo, J.; Te, N.; Izquierdo-Useros, N.; de la Concepción, M.L.R.; Ávila-Nieto, C.; Guallar, V.; Valencia, A.; Cantero, G.; et al. Pigs are not susceptible to SARS-CoV-2 infection but are a model for viral immunogenicity studies. Transbound. Emerg. Dis. 2020, 13861. [Google Scholar] [CrossRef]
- De Wit, E.; Feldmann, F.; Horne, E.; Martellaro, C.; Haddock, E.; Bushmaker, T.; Rosenke, K.; Okumura, A.; Rosenke, R.; Saturday, G.; et al. Domestic Pig Unlikely Reservoir for MERS-CoV. Emerg. Infect. Dis. 2017, 23, 985–988. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Vlasova, A.N.; Kenney, S.P.; Saif, L.J. Emerging and re-emerging coronaviruses in pigs. Curr. Opin. Virol. 2019, 34, 39–49. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.-S.; Sun, Y.-L.; Liu, C.-Y.; Yang, P.-C.; Chang, L.C.; Cheng, I.-C.; Mao, S.J.; Huang, M.-C. Complete nucleotide sequence of pig (Sus scrofa) mitochondrial genome and dating evolutionary divergence within Artiodactyla. Gene 1999, 236, 107–114. [Google Scholar] [CrossRef]
- Opriessnig, T.; Huang, Y. Coronavirus disease 2019 (COVID-19) outbreak: Could pigs be vectors for human infections? Xenotransplantation 2020, 27, e12591. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, Q.; Liu, S.; Zhang, X.; Jiang, W.; Wang, K.; Wang, S.; Peng, C.; Hou, G.; Li, J.; Yu, X.; et al. Surveillance and Taxonomic Analysis of the Coronavirus Dominant in Pigeons in China. Transbound. Emerg. Dis. 2020, 13541. [Google Scholar] [CrossRef] [PubMed]
- Suarez, D.L.; Pantin-Jackwood, M.J.; Swayne, D.E.; Lee, S.A.; Deblois, S.M.; Spackman, E. Lack of Susceptibility to SARS-CoV-2 and MERS-CoV in Poultry. Emerg. Infect. Dis. 2020, 26, 3074–3076. [Google Scholar] [CrossRef]
- Schlottau, K.; Rissmann, M.; Graaf, A.; Schön, J.; Sehl, J.; Wylezich, C.; Höper, D.; Mettenleiter, T.C.; Balkema-Buschmann, A.; Harder, T.; et al. SARS-CoV-2 in fruit bats, ferrets, pigs, and chickens: An experimental transmission study. Lancet Microbe 2020, 1, e218–e225. [Google Scholar] [CrossRef]
- Liu, Z.; Xiao, X.; Wei, X.; Li, J.; Yang, J.; Tan, H.; Zhu, J.; Zhang, Q.; Wu, J.; Liu, L. Composition and divergence of coronavirus spike proteins and host ACE2 receptors predict potential intermediate hosts of SARS-CoV-2. J. Med. Virol. 2020, 92, 595–601. [Google Scholar] [CrossRef]
- Luan, J.; Jin, X.; Lu, Y.; Zhang, L. SARS-CoV-2 spike protein favors ACE2 from Bovidae and Cricetidae. J. Med. Virol. 2020, 92, 1649–1656. [Google Scholar] [CrossRef]
- Ji, W.; Wang, W.; Zhao, X.; Zai, J.; Li, X. Cross-species transmission of the newly identified coronavirus 2019-nCoV. J. Med. Virol. 2020, 92, 433–440. [Google Scholar] [CrossRef]
- Zhang, C.; Zheng, W.; Huang, X.; Bell, E.W.; Zhou, X.; Zhang, Y. Protein Structure and Sequence Reanalysis of 2019-nCoV Genome Refutes Snakes as Its Intermediate Host and the Unique Similarity between Its Spike Protein Insertions and HIV-1. J. Proteome Res. 2020, 19, 1351–1360. [Google Scholar] [CrossRef]
- Hogan, C.A.; Sahoo, M.K.; Pinsky, B.A. Sample Pooling as a Strategy to Detect Community Transmission of SARS-CoV-2. JAMA 2020, 323, 1967. [Google Scholar] [CrossRef] [PubMed]
- Sanville, B.; Corbett, R.; Pidcock, W.; Hardin, K.; Sebat, C.; Nguyen, M.-V.; Thompson, G.R.; Haczku, A.; Schivo, M.; Cohen, S. A Community-transmitted Case of Severe Acute Respiratory Distress Syndrome (SARS) Due to SARS-CoV-2 in the United States. Clin. Infect. Dis. 2020, 71, 2222–2226. [Google Scholar] [CrossRef] [PubMed]
- Munir, K.; Ashraf, S.; Munir, I.; Khalid, H.; Muneer, M.A.; Mukhtar, N.; Amin, S.; Imran, M.A.; Chaudhry, U.; Zaheer, M.U.; et al. Zoonotic and Reverse Zoonotic Events of SARS-CoV-2 and their Impact on Global Health. Emerg. Microbes Infect. 2020, 9, 1–35. [Google Scholar] [CrossRef]
- Franklin, A.B.; Bevins, S.N. Spillover of SARS-CoV-2 into novel wild hosts in North America: A conceptual model for perpetuation of the pathogen. Sci. Total. Environ. 2020, 733, 139358. [Google Scholar] [CrossRef]
- Xia, C.; Lam, S.S.; Sonne, C. Ban unsustainable mink production. Science 2020, 370, 539. [Google Scholar] [CrossRef] [PubMed]
- Meyer, B.; Müller, M.A.; Corman, V.M.; Reusken, C.B.; Ritz, D.; Godeke, G.-J.; Lattwein, E.; Kallies, S.; Siemens, A.; van Beek, J.; et al. Antibodies against MERS Coronavirus in Dromedary Camels, United Arab Emirates, 2003 and 2013. Emerg. Infect. Dis. 2014, 20, 552–559. [Google Scholar] [CrossRef] [PubMed]
- Peck, K.M.; Burch, C.L.; Heise, M.T.; Baric, R. Coronavirus Host Range Expansion and Middle East Respiratory Syndrome Coronavirus Emergence: Biochemical Mechanisms and Evolutionary Perspectives. Annu. Rev. Virol. 2015, 2, 95–117. [Google Scholar] [CrossRef]
- Killerby, M.E.; Biggs, H.M.; Midgley, C.M.; Gerber, S.I.; Watson, J.T. Middle East Respiratory Syndrome Coronavirus Transmission. Emerg. Infect. Dis. 2020, 26, 191–198. [Google Scholar] [CrossRef] [PubMed]
- Dickman, A.; Hinks, A.; Macdonald, E.A.; Burnham, D.; Macdonald, D.W. Priorities for global felid conservation. Conserv. Biol. 2015, 29, 854–864. [Google Scholar] [CrossRef]
- Rabinowitz, P.; Scotch, M.; Conti, L. Human and Animal Sentinels for Shared Health Risks. Vet. Ital. 2010, 45, 23–24. [Google Scholar]
- Lee, C.T.; Slavinski, S.; Schiff, C.; Merlino, M.; Daskalakis, D.; Liu, D.; Rakeman, J.L.; Misener, M.; Thompson, C.; Leung, Y.L.; et al. Outbreak of Influenza A(H7N2) Among Cats in an Animal Shelter With Cat-to-Human Transmission—New York City, 2016. Clin. Infect. Dis. 2017, 65, 1927–1929. [Google Scholar] [CrossRef] [PubMed]
- Kwan, J.L.; Kluh, S.; Madon, M.B.; Nguyen, D.V.; Barker, C.M.; Reisen, W.K. Sentinel Chicken Seroconversions Track Tangential Transmission of West Nile Virus to Humans in the Greater Los Angeles Area of California. Am. J. Trop. Med. Hyg. 2010, 83, 1137–1145. [Google Scholar] [CrossRef] [PubMed]
- Ziegler, U.; Jöst, H.; Müller, K.; Fischer, D.; Rinder, M.; Tietze, D.T.; Danner, K.-J.; Becker, N.; Skuballa, J.; Hamann, H.-P.; et al. Epidemic Spread of Usutu Virus in Southwest Germany in 2011 to 2013 and Monitoring of Wild Birds for Usutu and West Nile Viruses. Vect. Borne Zoonotic Dis. 2015, 15, 481–488. [Google Scholar] [CrossRef] [PubMed]
| Genus | Species | Recombinant Events | Hosts |
|---|---|---|---|
| Alphacoronavirus | SADS-CoV | HKU2-related bat COVs from horseshoe bats (Rhinolophus sinicus, Rhinolopus pusillus, Rhinolopus rex, Rhinolopus affinus) with swine acute diarrhea syndrome coronavirus (SADS-CoV) [128]. | Domestic pigs |
| S INDEL PEDV | Transmissible gastroenteritis virus mutant with recombination of G1a CV777-lineage classical and the G2 strain of porcine epidemic diarrhea virus (PEDV) [129]. | Domestic pigs | |
| Rhinolophus bat coronavirus HKU2 | Evidence of previous recombinant events with HKU2 from horseshoe bats shares 15-amino acid peptide corresponding within the RBM of the spike protein of SARS-CoV [130]. | Horseshoe bats (Rhinolophus sinicus) | |
| Lucheng Rn rat coronavirus (LRNV) | Recombinant origin due to its N gene sequence more closely related to the genus Betacoronavirus than Alphacoronavirus. Also formed a divergent lineage in S gene tree with horseshoe bat coronavirus HKU2 [131]. | Brown rats (Rattus norvegicus) | |
| HCoV-229E | Recombination event between alpaca HCoV-229E and bat 229E-related CoVs [132]. | Noack’s round-leaf bats (Hipposideros cf. ruber) and Aba round leaf bats (Hipposideros abae) | |
| Canine Coronavirus type 1 (CCoV-II) | Recombination event involving CCoV-II and porcine transmissible gastroenteritis virus (TGEV) [133] | Domestic dogs | |
| Ferret Coronavirus (FRCoVs) | Comparison of FRCoV with ferret systemic coronavirus and ferret enteric coronavirus revealed that recombination occurred in the spike, 3c, and envelope genes occurred between different FRCoVs [134]. | Domesticated ferrets (Mustela putorius) | |
| Betacoronavirus | SARS-CoV-2 | Horseshoe bats (Rhinolophus spp.) (RaTG13) and potentially Malayan pangolins (Manis javanica) (PCoV) [27]. | Humans and cats (Felidae) |
| SARS-CoV | SARS-CoV-like viruses from horseshoe bats (Rhinolophus macrotis, Rousettus leschenaulti, Rhinolophus pearsoni, Rhinolophus pussilus) and humans [135]. | Horseshoe bats (Rhinolophus spp.), Himalayan palm civets (Paguma larvata), raccoon dogs (Nyctereutes procyonoides) | |
| MERS-CoV | Humans and dromedary camels (Camelus dromedarius) [136]. | Bats (Neoromicia capensis, Vespertilio superans), dromedary camels (Camelus dromedarius), and European hedgehogs (Erinaceus europaeus) | |
| Bovine CoV (BCoV) | There is a putative recombinant detection only in the BCoV strain HEC 4408 with human CoV OC43 [137]. | Waterbuck (Kobus ellypsiprimus), sambar deer (Cervus unicolor), white-tailed deer (Odocoileus virginianus), elk (Cervus elephus), giraffe (Giraffa camelopardalis), and sable antelopes (Hipotragus niger) | |
| Deltacoronavirus | Wigeon coronavirus HKU20 | NS7b of WiCoV HKU20 and CMCoV HKU21, and NS7d of WiCoV HKU20, were also found to be homologous to the NS3b of IBV and hypothetical protein of goose coronavirus, respectively [138]. | Wigeons (Mareca spp.) |
| Bulbul Coronavirus HKU11 | NS7a of NHCoV HKU19 was found to be homologous to the NS7a of BuCoV HKU11, ThCoV HKU12, and MunCoV HKU13 [138]. | Bulbuls (Pycnonotidae) | |
| Porcine coronavirus HKU15 | PorCoV HKU15 contains a stem-loop II motif, a conserved RNA element downstream of N and upstream of the poly(A) tail, similar to those in Infectious Bronchitis Virus, TCoV, SARSr-Rh-BatCoV, and SARS-CoV [138]. | Domestic pigs | |
| Gammacoronavirus | Beluga whale CoV (BWCoV) | Beluga whale BWCoV SW1 (Delphinapterus leucas) and bottlenose dolphin (Tursiops truncatus) BDCoV HKU22 exhibit high similarity [139]. | Beluga whales (Delphinapterus leucas) |
| Infectious bronchitis virus (IBV) | Evidence of recombinant events can be seen from the high similarity in IBV strains of chicken and turkey CoVs [140]. | Turkey (Meleagris spp.), Goose (Anser spp.), Duck (Anas spp.), and Pigeons (Columbidae) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hedman, H.D.; Krawczyk, E.; Helmy, Y.A.; Zhang, L.; Varga, C. Host Diversity and Potential Transmission Pathways of SARS-CoV-2 at the Human-Animal Interface. Pathogens 2021, 10, 180. https://doi.org/10.3390/pathogens10020180
Hedman HD, Krawczyk E, Helmy YA, Zhang L, Varga C. Host Diversity and Potential Transmission Pathways of SARS-CoV-2 at the Human-Animal Interface. Pathogens. 2021; 10(2):180. https://doi.org/10.3390/pathogens10020180
Chicago/Turabian StyleHedman, Hayden D., Eric Krawczyk, Yosra A. Helmy, Lixin Zhang, and Csaba Varga. 2021. "Host Diversity and Potential Transmission Pathways of SARS-CoV-2 at the Human-Animal Interface" Pathogens 10, no. 2: 180. https://doi.org/10.3390/pathogens10020180
APA StyleHedman, H. D., Krawczyk, E., Helmy, Y. A., Zhang, L., & Varga, C. (2021). Host Diversity and Potential Transmission Pathways of SARS-CoV-2 at the Human-Animal Interface. Pathogens, 10(2), 180. https://doi.org/10.3390/pathogens10020180








