Association between Interleukin-1 Polymorphisms and Susceptibility to Dental Peri-Implant Disease: A Meta-Analysis
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Search Strategy
2.3. Inclusion and Exclusion Criteria
2.4. Data Extraction
2.5. Quality of Assessment
2.6. Statistical Analyses
3. Results
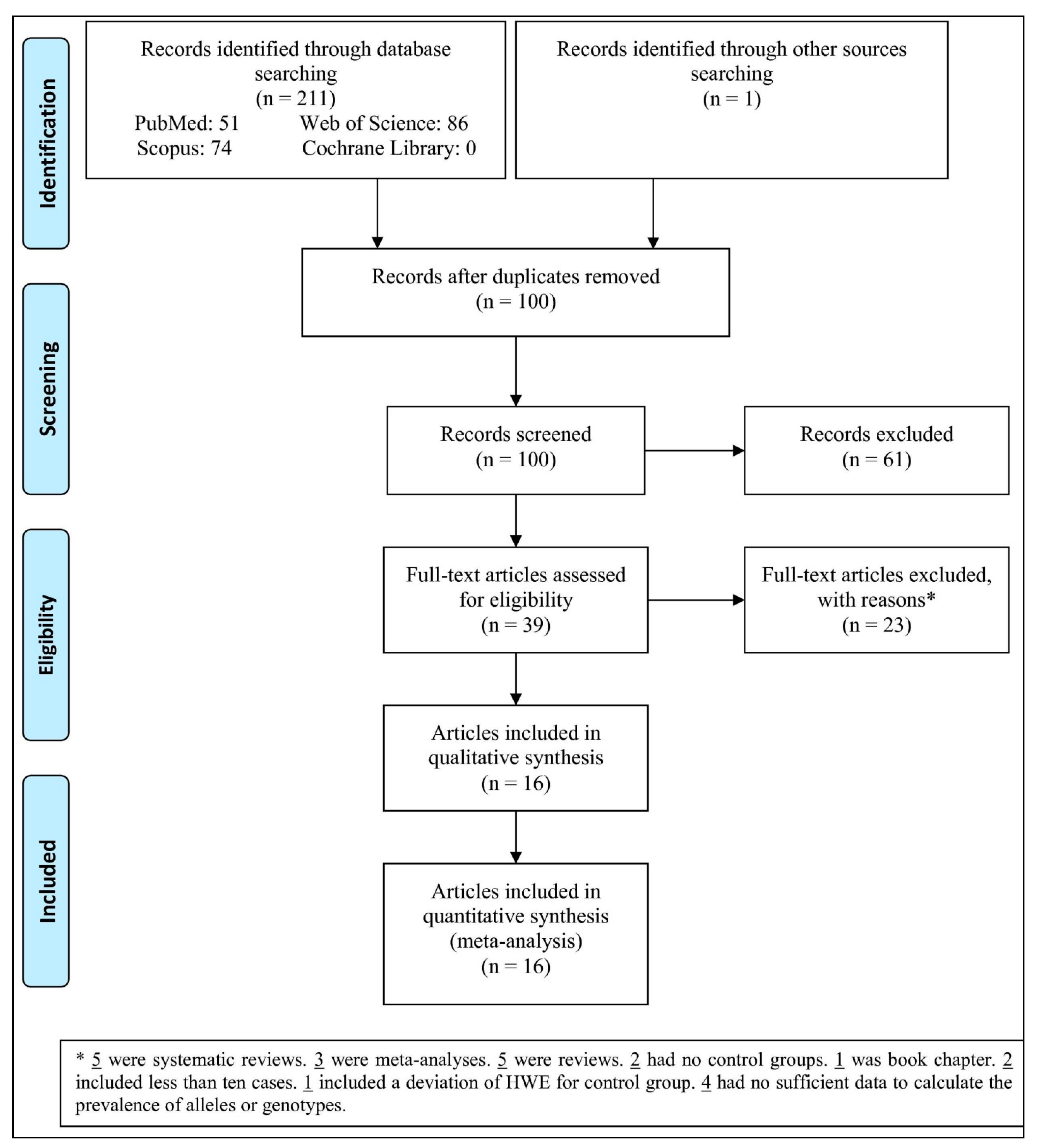
3.1. Study Selection
3.2. Characteristics of the Studies
3.3. Pooled Analyses
3.4. Subgroup Analysis
3.5. Meta-Regression
3.6. Sensitivity Analysis
3.7. Trial Sequential Analysis
3.8. Publication Bias
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Karoussis, I.K.; Kotsovilis, S.; Fourmousis, I. A comprehensive and critical review of dental implant prognosis in periodontally compromised partially edentulous patients. Clin. Oral Implant. Res. 2007, 18, 669–679. [Google Scholar] [CrossRef] [PubMed]
- Bornstein, M.M.; Halbritter, S.; Harnisch, H.; Weber, H.-P.; Buser, D. A retrospective analysis of patients referred for implant placement to a specialty clinic: Indications, surgical procedures, and early failures. Int. J. Oral Maxillofac. Implant. 2008, 23, 1109–1116. [Google Scholar]
- Dereka, X.; Mardas, N.; Chin, S.; Petrie, A.; Donos, N. A systematic review on the association between genetic predisposition and dental implant biological complications. Clin. Oral Implant. Res. 2012, 23, 775–788. [Google Scholar] [CrossRef] [PubMed]
- Kowalski, J.; Lapinska, B.; Nissan, J.; Lukomska-Szymanska, M. Factors Influencing Marginal Bone Loss around Dental Implants: A Narrative Review. Coatings 2021, 11, 865. [Google Scholar] [CrossRef]
- Heitz-Mayfield, L.J. Peri-implant diseases: Diagnosis and risk indicators. J. Clin. Periodontol. 2008, 35, 292–304. [Google Scholar] [CrossRef]
- Jamshidy, L.; Tadakamadla, S.K.; Choubsaz, P.; Sadeghi, M.; Tadakamadla, J. Association of IL-10 and TNF-α Polymorphisms with Dental Peri-Implant Disease Risk: A Meta-Analysis, Meta-Regression, and Trial Sequential Analysis. Int. J. Environ. Res. Public Health 2021, 18, 7697. [Google Scholar] [CrossRef]
- Mo, Y.-Y.; Zeng, X.-T.; Weng, H.; Cen, Y.; Zhao, Q.; Wen, X. Association between tumor necrosis factor-alpha G-308A polymorphism and dental peri-implant disease risk: A meta-analysis. Medicine 2016, 95, e4425. [Google Scholar] [CrossRef] [PubMed]
- Nicholls, J. The management of periodontal and peri implant disease. BDJ Team 2020, 7, 34–36. [Google Scholar] [CrossRef]
- Insua, A.; Monje, A.; Wang, H.L.; Inglehart, M. Patient-centered perspectives and understanding of peri-implantitis. J. Periodontol. 2017, 88, 1153–1162. [Google Scholar] [CrossRef] [PubMed]
- Liao, J.; Li, C.; Wang, Y.; Ten, M.; Sun, X.; Tian, A.; Zhang, Q.; Liang, X. Meta-analysis of the association between common interleukin-1 polymorphisms and dental implant failure. Mol. Biol. Rep. 2014, 41, 2789–2798. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Finkelstein, J. The relationship between single nucleotide polymorphisms and dental implant loss: A scoping review. Clin. Cosmet. Investig. Dent. 2019, 11, 131. [Google Scholar] [CrossRef] [PubMed]
- Casado, L.P.; Villas-Boas, R.; de Mello, W.; Duarte, L.M.E.; Granjeiro, M.J. Peri-implant disease and chronic periodontitis: Is interleukin-6 gene promoter polymorphism the common risk factor in a Brazilian population? Int. J. Oral Maxillofac. Implant. 2013, 28, 35–43. [Google Scholar] [CrossRef] [PubMed][Green Version]
- e Silva, R.C.; Reis, M.B.L.; Arid, J.; Flores, E.K.B.; Cruz, G.V.; Marañón-Vásquez, G.A.; de Souza, L.K.F.; Novaes, A.B., Jr.; de Queiroz, A.M.; Küchler, E.C. Association between Genetic Polymorphisms in RANK, RANKL and OPG and Peri-Implant Diseases in Patients from the Amazon Region. Braz. Dent. J. 2020, 31, 63–68. [Google Scholar] [CrossRef]
- Kornman, K.S. Interleukin 1 genetics, inflammatory mechanisms, and nutrigenetic opportunities to modulate diseases of aging. Am. J. Clin. Nutr. 2006, 83, 475S–483S. [Google Scholar] [CrossRef] [PubMed]
- Graves, D.T.; Cochran, D. The contribution of interleukin-1 and tumor necrosis factor to periodontal tissue destruction. J. Periodontol. 2003, 74, 391–401. [Google Scholar] [CrossRef] [PubMed]
- Cosyn, J.; Christiaens, V.; Koningsveld, V.; Coucke, P.J.; De Coster, P.; De Paepe, A.; De Bruyn, H. An exploratory case-control study on the impact of IL-1 gene polymorphisms on early implant failure. Clin. Implant. Dent. Relat. Res. 2016, 18, 234–240. [Google Scholar] [CrossRef]
- Andreiotelli, M.; Koutayas, S.-O.; Madianos, P.N.; Strub, J.-R. Relationship between interleukin-1 genotype and peri-implantitis: A literature review. Quintessence Int. 2008, 39, 289–298. [Google Scholar] [PubMed]
- Huynh-Ba, G.; Lang, N.P.; Tonetti, M.; Zwahlen, M.; Salvi, G. Association of the composite IL-1 genotype with peri-implantitis: A systematic review. Clin. Oral Implant. Res. 2008, 19, 1154–1162. [Google Scholar] [CrossRef] [PubMed]
- Bormann, K.-H.; Stühmer, C.; Z’Graggen, M.; Kokemöller, H.; Rücker, M.; Gellrich, N.-C. IL-1 polymorphism and periimplantitis. A literature review. Schweiz. Mon. Fur Zahnmed. 2010, 120, 510–520. [Google Scholar]
- Kadkhodazadeh, M.; Tabari, Z.A.; Pourseyediyan, T.; Najafi, K.; Amid, R. Relationship between Genetic Polymorphisms with Periodontitis and Peri-Implantitis in the Iranian Population: A Literature Review. J. Long-Term Eff. Med Implant. 2016, 26, 183–190. [Google Scholar] [CrossRef] [PubMed]
- Junior, J.F.S.; Biguetti, C.C.; Matsumoto, M.A.; Kudo, G.A.H.; da Silva, R.B.P.; Saraiva, P.P.; Fakhouri, W.D. Can genetic factors compromise the success of dental implants? A systematic review and meta-analysis. Genes 2018, 9, 444. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, K.K.; Anwar, M.; Gupta, C.; Chand, P.; Singh, S.V. Association of interleukin-1 gene polymorphism and early crestal bone loss around submerged dental implants: A systematic review and meta-analysis. J. Indian Prosthodont. Soc. 2021, 21, 116. [Google Scholar]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement. Int. J. Surg. 2010, 8, 336–341. [Google Scholar] [CrossRef] [PubMed]
- Morgan, R.L.; Whaley, P.; Thayer, K.A.; Schünemann, H.J. Identifying the PECO: A framework for formulating good questions to explore the association of environmental and other exposures with health outcomes. Environ. Int. 2018, 121, 1027. [Google Scholar] [CrossRef] [PubMed]
- Wells, G.A.; Shea, B.; O’Connell, D.; Peterson, J.; Welch, V.; Losos, M.; Tugwell, P. The Newcastle-Ottawa Scale (NOS) for assessing the quality of nonrandomized studies in meta-analyses. Appl. Eng. Agric. 2014, 18, 727–734. [Google Scholar]
- Wetterslev, J.; Jakobsen, J.C.; Gluud, C. Trial sequential analysis in systematic reviews with meta-analysis. BMC Med Res. Methodol. 2017, 17, 39. [Google Scholar] [CrossRef] [PubMed]
- Imberger, G.; Thorlund, K.; Gluud, C.; Wetterslev, J. False-positive findings in Cochrane meta-analyses with and without application of trial sequential analysis: An empirical review. BMJ Open 2016, 6, e011890. [Google Scholar] [CrossRef] [PubMed]
- Rogers, M.A.; Figliomeni, L.; Baluchova, K.; Tan, A.E.; Davies, G.; Henry, P.J.; Price, P. Do interleukin-1 polymorphisms predict the development of jreiodontitis or the success of dental implants? J. Periodontal Res. 2002, 37, 37–41. [Google Scholar] [CrossRef] [PubMed]
- Shimpuku, H.; Nosaka, Y.; Kawamura, T.; Tachi, Y.; Shinohara, M.; Ohura, K. Genetic polymorphisms of the interleukin-1 gene and early marginal bone loss around endosseous dental implants. Clin. Oral Implant. Res. 2003, 14, 423–429. [Google Scholar] [CrossRef]
- Campos, M.I.; Santos, M.C.; Trevilatto, P.C.; Scarel-Caminaga, R.M.; Bezerra, F.J.; Line, S.R. Evaluation of the relationship between interleukin-1 gene cluster polymorphisms and early implant failure in non-smoking patients. Clin. Oral Implant. Res. 2005, 16, 94–201. [Google Scholar] [CrossRef]
- Laine, M.L.; Leonhardt, Å.; Roos-Jansåker, A.M.; Peña, A.S.; Van Winkelhoff, A.J.; Winkel, E.G.; Renvert, S. IL-1RN gene polymorphism is associated with peri-implantitis. Clin. Oral Implant. Res. 2006, 17, 380–385. [Google Scholar] [CrossRef]
- Lachmann, S.; Kimmerle-Müller, E.; Axmann, D.; Scheideler, L.; Weber, H.; Haas, R. Associations between peri-implant crevicular fluid volume, concentrations of crevicular inflammatory mediators, and composite IL-1A–889 and IL-1B+ 3954 genotype: A cross-sectional study on implant recall patients with and without clinical signs of peri-implantitis. Clin. Oral Implant. Res. 2007, 18, 212–223. [Google Scholar]
- Lin, Y.-H.; Huang, P.; Lu, X.; Guan, D.-H.; Man, Y.; Wei, N.; Wang, Y.-Y.; Gong, P. The relationship between IL-1 gene polymorphism and marginal bone loss around dental implants. J. Oral Maxillofac. Surg. 2007, 65, 2340–2344. [Google Scholar] [CrossRef] [PubMed]
- Montes, C.C.; Alvim-Pereira, F.; De Castilhos, B.B.; Sakurai, M.L.L.; Olandoski, M.; Trevilatto, P.C. Analysis of the association of IL1B (C+ 3954T) and IL1RN (intron 2) polymorphisms with dental implant loss in a Brazilian population. Clin. Oral Implant. Res. 2009, 20, 208–217. [Google Scholar] [CrossRef]
- Dirschnabel, A.J.; Alvim-Pereira, F.; Alvim-Pereira, C.C.; Bernardino, J.F.; Rosa, E.A.R.; Trevilatto, P.C. Analysis of the association of IL1B (C-511T) polymorphism with dental implant loss and the clusterization phenomenon. Clin. Oral Implant. Res. 2011, 22, 1235–1241. [Google Scholar] [CrossRef] [PubMed]
- Hamdy, A.A.E.-M.M.; Ebrahem, M.A.E.-M. The Effect of Interleukin-1 Allele 2 Genotype (IL-1a− 889 and IL-1b+ 3954) on the Individual’s Susceptibility to Peri-Implantitis: Case-Control Study. J. Oral Implantol. 2011, 37, 325–334. [Google Scholar] [CrossRef] [PubMed]
- Melo, R.F.; Lopes, B.M.V.; Shibli, J.A.; Marcantonio, E., Jr.; Marcantonio, R.A.C.; Galli, G.M.T. Interleukin-1β and interleukin-6 expression and gene polymorphisms in subjects with peri-implant disease. Clin. Implant. Dent. Relat. Res. 2012, 14, 905–914. [Google Scholar] [CrossRef] [PubMed]
- Vaz, P.; Gallas, M.; Braga, A.; Sampaio-Fernandes, J.; Felino, A.; Tavares, P. IL1 gene polymorphisms and unsuccessful dental implants. Clin. Oral Implant. Res. 2012, 23, 1404–1413. [Google Scholar] [CrossRef] [PubMed]
- Jacobi-Gresser, E.; Huesker, K.; Schütt, S. Genetic and immunological markers predict titanium implant failure: A retrospective study. Int. J. Oral Maxillofac. Surg. 2013, 42, 537–543. [Google Scholar] [CrossRef] [PubMed]
- Petkovic-Curcin, A.; Zeljic, K.; Cikota-Aleksic, B.; Dakovic, D.; Tatic, Z.; Magic, Z. Association of Cytokine Gene Polymorphism with Peri-implantitis Risk. Int. J. Oral Maxillofac. Implant. 2017, 32, e241–e248. [Google Scholar] [CrossRef]
- Agrawal, K.K.; Chand, P.; Singh, S.V.; Singh, N.; Gupta, P.; Garg, R.K.; Chaurasia, A.; Anwar, M.; Kumar, A. Association of interleukin-1, interleukin-6, collagen type I alpha 1, and osteocalcin gene polymorphisms with early crestal bone loss around submerged dental implants: A nested case control study. J. Prosthet. Dent. 2021. [Google Scholar] [CrossRef]
- Saremi, L.; Shafizadeh, M.; Esmaeilzadeh, E.; Ghaffari, M.E.; Mahdavi, M.H.; Amid, R.; Kadkhodazadeh, M. Assessment of IL-10, IL-1ß and TNF-α gene polymorphisms in patients with peri-implantitis and healthy controls. Mol. Biol. Rep. 2021, 48, 2285–2290. [Google Scholar] [CrossRef] [PubMed]
- Moreira, P.; Costa, J.; Gomez, R.; Gollob, K.; Dutra, W. The IL1A (− 889) gene polymorphism is associated with chronic periodontal disease in a sample of Brazilian individuals. J. Periodontal Res. 2007, 42, 23–30. [Google Scholar] [CrossRef]
- Shirodaria, S.; Smith, J.; McKay, I.; Kennett, C.; Hughes, F. Polymorphisms in the IL-1A gene are correlated with levels of interleukin-1α protein in gingival crevicular fluid of teeth with severe periodontal disease. J. Dent. Res. 2000, 79, 1864–1869. [Google Scholar] [CrossRef]
- Dominici, R.; Cattaneo, M.; Malferrari, G.; Archi, D.; Mariani, C.; Grimaldi, L.; Biunno, I. Cloning and functional analysis of the allelic polymorphism in the transcription regulatory region of interleukin-1α. Immunogenetics 2002, 54, 82–86. [Google Scholar] [CrossRef]
- Amirisetty, R.; Patel, R.P.; Das, S.; Saraf, J.; Jyothy, A.; Munshi, A. Interleukin 1 β (+ 3954,− 511 and− 31) polymorphism in chronic periodontitis patients from North India. Acta Odontol. Scand. 2015, 73, 343–347. [Google Scholar] [CrossRef] [PubMed]
- Costa-Junior, F.; Alvim-Pereira, C.; Alvim-Pereira, F.; Trevilatto, P.; de Souza, A.; Santos, M.C.L. Influence of MMP-8 promoter polymorphism in early osseointegrated implant failure. Clin. Oral Investig. 2013, 17, 311–316. [Google Scholar] [CrossRef]
- Pociot, F.; Mølvig, J.; Wogensen, L.; Worsaae, H.; Nerup, J. A Taql polymorphism in the human interleukin-1β (IL-1β) gene correlates with IL-1β secretion in vitro. Eur. J. Clin. Investig. 1992, 22, 396–402. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.; Song, Q.-B.; Yao, P.-F.; Hu, Q.-L.; Hu, P.-J.; Zeng, Z.-R.; Pang, R.-P. No relationship between IL-1B gene polymorphism and gastric acid secretion in younger healthy volunteers. World J. Gastroenterol. WJG 2005, 11, 6549. [Google Scholar] [CrossRef] [PubMed]
- Buchs, N.; Di Giovine, F.S.; Silvestri, T.; Vannier, E.; Duff, G.W.; Miossec, P. IL-1B and IL-1Ra gene polymorphisms and disease severity in rheumatoid arthritis: Interaction with their plasma levels. Genes Immun. 2001, 2, 222–228. [Google Scholar] [CrossRef] [PubMed]
- Goiato, M.C.; Dos Santos, D.; Santiago, J.J.; Moreno, A.; Pellizzer, E.P. Longevity of dental implants in type IV bone: A systematic review. Int. J. Oral Maxillofac. Surg. 2014, 43, 1108–1116. [Google Scholar] [CrossRef] [PubMed]
- Batista, V.E.d.S.; Junior, J.F.S.; Almeida, D.A.d.F.; Lopes, L.F.d.T.P.; Verri, F.R.; Pellizzer, E.P. The effect of offset implant configuration on bone stress distribution: A systematic review. J. Prosthodont. 2015, 24, 93–99. [Google Scholar] [CrossRef] [PubMed]
- Wilson, T.G.W., Jr.; Nunn, M. The Relationship Between the Interleukin–1 Periodontal Genotype and Implant Loss. Initial Data. J. Periodontol. 1999, 70, 724–729. [Google Scholar] [CrossRef] [PubMed]
- Strietzel, F.P.; Reichart, P.A.; Kale, A.; Kulkarni, M.; Wegner, B.; Küchler, I. Smoking interferes with the prognosis of dental implant treatment: A systematic review and meta-analysis. J. Clin. Periodontol. 2007, 34, 523–544. [Google Scholar] [CrossRef] [PubMed]
| Genetic Model | First Author, Publication Year | Case | Control | Weight | Odds Ratio | ||
|---|---|---|---|---|---|---|---|
| Events | Total | Events | Total | M-H, Fixed, 95% CI | |||
| T vs. C | Rogers, 2002 [28] | 10 | 38 | 16 | 62 | 8.8% | 1.03 [0.41, 2.57] |
| Shimpuku, 2003 [29] | 4 | 34 | 4 | 44 | 3.0% | 1.33 [0.31, 5.77] | |
| Campos, 2005 [30] | 17 | 56 | 21 | 68 | 12.9% | 0.98 [0.45, 2.10] | |
| Laine, 2006 [31] | 49 | 142 | 33 | 98 | 25.0% | 1.04 [0.60, 1.79] | |
| Melo, 2012 [37] | 25 | 32 | 51 | 62 | 7.4% | 0.77 [0.27, 2.23] | |
| Jacobi-Gresser, 2013 [39] | 28 | 82 | 32 | 136 | 15.5% | 1.69 [0.92, 3.08] | |
| Cosyn, 2016 [16] | 10 | 24 | 4 | 26 | 2.2% | 3.93 [1.03, 14.99] | |
| Agrawal, 2021 [41] | 98 | 136 | 89 | 126 | 25.2% | 1.07 [0.63, 1.83] | |
| Subtotal (95% CI) | 544 | 622 | 100.0% | 1.19 [0.92, 1.55] | |||
| Total events | 241 | 250 | |||||
| Heterogeneity: Chi2 = 5.74, df = 7 (p = 0.57); I2 = 0% Test for overall effect: Z = 1.30 (p = 0.19) | |||||||
| TT vs. CC | Rogers, 2002 [28] | 0 | 9 | 3 | 21 | 12.2% | 0.28 [0.01, 5.96] |
| Shimpuku, 2003 [29] | 1 | 15 | 0 | 18 | 2.4% | 3.83 [0.14, 101.07] | |
| Campos, 2005 [30] | 4 | 19 | 5 | 23 | 21.0% | 0.96 [0.22, 4.23] | |
| Laine, 2006 [31] | 2 | 26 | 3 | 22 | 17.6% | 0.53 [0.08, 3.49] | |
| Melo, 2012 [37] | 9 | 9 | 21 | 22 | 3.8% | 1.33 [0.05, 35.60] | |
| Jacobi-Gresser, 2013 [39] | 3 | 19 | 3 | 42 | 9.2% | 2.44 [0.44, 13.38] | |
| Cosyn, 2016 [16] | 2 | 6 | 0 | 9 | 1.6% | 10.56 [0.41, 268.69] | |
| Agrawal, 2021 [41] | 37 | 44 | 32 | 38 | 32.1% | 0.99 [0.30, 3.25] | |
| Subtotal (95% CI) | 147 | 195 | 100.0% | 1.18 [0.62, 2.25] | |||
| Total events | 58 | 67 | |||||
| Heterogeneity: Chi2 = 4.67, df = 7 (p = 0.70); I2 = 0% Test for overall effect: Z = 0.51 (p = 0.61) | |||||||
| CT vs. CC | Rogers, 2002 [28] | 10 | 19 | 10 | 28 | 9.6% | 2.00 [0.61, 6.55] |
| Shimpuku, 2003 [29] | 2 | 16 | 4 | 22 | 7.4% | 0.64 [0.10, 4.03] | |
| Campos, 2005 [30] | 9 | 24 | 11 | 29 | 15.6% | 0.98 [0.32, 3.00] | |
| Laine, 2006 [31] | 45 | 69 | 27 | 46 | 28.3% | 1.32 [0.61, 2.84] | |
| Melo, 2012 [37] | 7 | 7 | 9 | 10 | 1.3% | 2.37 [0.08, 66.88] | |
| Jacobi-Gresser, 2013 [39] | 22 | 38 | 26 | 65 | 20.3% | 2.06 [0.91, 4.65] | |
| Cosyn, 2016 [16] | 6 | 10 | 4 | 13 | 3.5% | 3.38 [0.60, 19.01] | |
| Agrawal, 2021 [41] | 24 | 31 | 25 | 31 | 14.2% | 0.82 [0.24, 2.80] | |
| Subtotal (95% CI) | 214 | 244 | 100.0% | 1.45 [0.97, 2.16] | |||
| Total events | 125 | 116 | |||||
| Heterogeneity: Chi2 = 4.11, df = 7 (p = 0.77); I2 = 0% Test for overall effect: Z = 1.81 (p = 0.07) | |||||||
| TT + CT vs. CC | Rogers, 2002 [28] | 10 | 19 | 13 | 31 | 10.7% | 1.54 [0.49, 4.85] |
| Shimpuku, 2003 [29] | 3 | 17 | 4 | 22 | 6.6% | 0.96 [0.18, 5.03] | |
| Campos, 2005 [30] | 13 | 28 | 16 | 34 | 17.7% | 0.97 [0.36, 2.66] | |
| Laine, 2006 [31] | 47 | 71 | 30 | 49 | 27.4% | 1.24 [0.58, 2.64] | |
| Melo, 2012 [37] | 16 | 16 | 30 | 31 | 1.4% | 1.62 [0.06, 42.12] | |
| Jacobi-Gresser, 2013 [39] | 25 | 41 | 29 | 68 | 19.4% | 2.10 [0.95, 4.63] | |
| Cosyn, 2016 [16] | 8 | 12 | 4 | 13 | 2.9% | 4.50 [0.84, 24.18] | |
| Agrawal, 2021 [41] | 61 | 68 | 57 | 63 | 13.9% | 0.92 [0.29, 2.89] | |
| Subtotal (95% CI) | 272 | 311 | 100.0% | 1.43 [0.98, 2.10] | |||
| Total events | 183 | 183 | |||||
| Heterogeneity: Chi2 = 4.21, df = 7 (p = 0.76); I2 = 0% Test for overall effect: Z = 1.83 (p = 0.07) | |||||||
| TT vs. CC + CT | Rogers, 2002 [28] | 0 | 19 | 3 | 31 | 7.7% | 0.21 [0.01, 4.27] |
| Shimpuku, 2003 [29] | 1 | 17 | 0 | 22 | 1.2% | 4.09 [0.16, 106.89] | |
| Campos, 2005 [30] | 4 | 28 | 5 | 34 | 11.3% | 0.97 [0.23, 4.01] | |
| Laine, 2006 [31] | 2 | 71 | 3 | 49 | 10.1% | 0.44 [0.07, 2.76] | |
| Melo, 2012 [37] | 9 | 16 | 21 | 31 | 18.3% | 0.61 [0.18, 2.12] | |
| Jacobi-Gresser, 2013 [39] | 3 | 41 | 3 | 68 | 6.1% | 1.71 [0.33, 8.90] | |
| Cosyn, 2016 [16] | 2 | 12 | 0 | 13 | 1.1% | 6.43 [0.28, 148.77] | |
| Agrawal, 2021 [41] | 37 | 68 | 32 | 63 | 44.2% | 1.16 [0.58, 2.30] | |
| Subtotal (95% CI) | 272 | 311 | 100.0% | 1.02 [0.64, 1.63] | |||
| Total events | 58 | 67 | |||||
| Heterogeneity: Chi2 = 5.03, df = 7 (p = 0.66); I2 = 0% Test for overall effect: Z = 0.08 (p = 0.94) | |||||||
| Genetic Model | First Author, Publication Year | Case | Control | Weight | Odds Ratio | ||
|---|---|---|---|---|---|---|---|
| Events | Total | Events | Total | M-H, Random, 95% CI | |||
| T vs. C | Shimpuku, 2003 [29] | 22 | 34 | 19 | 44 | 10.6% | 2.41 [0.96, 6.07] |
| Campos, 2005 [30] | 21 | 56 | 30 | 68 | 12.6% | 0.76 [0.37, 1.57] | |
| Laine, 2006 [31] | 51 | 142 | 35 | 98 | 14.6% | 1.01 [0.59, 1.73] | |
| Lin, 2007 [33] | 37 | 58 | 19 | 60 | 12.2% | 3.80 [1.77, 8.16] | |
| Dirschnabel, 2011 [35] | 83 | 184 | 144 | 370 | 16.4% | 1.29 [0.90, 1.84] | |
| Melo, 2012 [37] | 25 | 32 | 51 | 62 | 9.4% | 0.77 [0.27, 2.23] | |
| Cosyn, 2016 [16] | 16 | 28 | 18 | 28 | 9.2% | 0.74 [0.25, 2.17] | |
| Agrawal, 2021 [41] | 40 | 136 | 61 | 126 | 14.9% | 0.44 [0.27, 0.74] | |
| Subtotal (95% CI) | 670 | 856 | 100.0% | 1.10 [0.69, 1.75] | |||
| Total events | 295 | 377 | |||||
| Heterogeneity: Tau2 = 0.32; Chi2 = 27.86, df = 7 (p = 0.0002); I2 = 75% Test for overall effect: Z = 0.38 (p = 0.70) | |||||||
| TT vs. CC | Shimpuku, 2003 [29] | 8 | 11 | 3 | 9 | 8.8% | 5.33 [0.78, 36.33] |
| Campos, 2005 [30] | 7 | 21 | 7 | 18 | 13.4% | 0.79 [0.21, 2.92] | |
| Laine, 2006 [31] | 10 | 40 | 5 | 24 | 14.4% | 1.27 [0.37, 4.28] | |
| Lin, 2007 [33] | 14 | 20 | 6 | 15 | 12.6% | 3.50 [0.86, 14.30] | |
| Dirschnabel, 2011 [35] | 21 | 51 | 28 | 97 | 20.1% | 1.73 [0.85, 3.51] | |
| Melo, 2012 [37] | 2 | 6 | 3 | 13 | 7.6% | 1.67 [0.20, 14.05] | |
| Cosyn, 2016 [16] | 5 | 8 | 6 | 8 | 7.5% | 0.56 [0.06, 4.76] | |
| Agrawal, 2021 [41] | 7 | 42 | 13 | 28 | 15.6% | 0.23 [0.08, 0.69] | |
| Subtotal (95% CI) | 199 | 212 | 100.0% | 1.20 [0.59, 2.42] | |||
| Total events | 74 | 71 | |||||
| Heterogeneity: Tau2 = 0.51; Chi2 = 15.17, df = 7 (p = 0.03); I2 = 54% Test for overall effect: Z = 0.50 (p = 0.61) | |||||||
| CT vs. CC | Shimpuku, 2003 [29] | 6 | 9 | 13 | 19 | 3.4% | 0.92 [0.17, 5.00] |
| Campos, 2005 [30] | 7 | 21 | 16 | 27 | 11.4% | 0.34 [0.10, 1.13] | |
| Laine, 2006 [31] | 31 | 61 | 25 | 44 | 17.5% | 0.79 [0.36, 1.71] | |
| Lin, 2007 [33] | 9 | 15 | 15 | 24 | 5.6% | 0.90 [0.24, 3.38] | |
| Dirschnabel, 2011 [35] | 41 | 71 | 88 | 157 | 28.3% | 1.07 [0.61, 1.89] | |
| Melo, 2012 [37] | 10 | 14 | 18 | 28 | 4.2% | 1.39 [0.34, 5.60] | |
| Cosyn, 2016 [16] | 6 | 9 | 6 | 8 | 2.6% | 0.67 [0.08, 5.54] | |
| Agrawal, 2021 [41] | 26 | 61 | 35 | 50 | 27.0% | 0.32 [0.14, 0.70] | |
| Subtotal (95% CI) | 261 | 357 | 100.0% | 0.72 [0.52, 1.01] | |||
| Total events | 136 | 216 | |||||
| Heterogeneity: Chi2 = 8.58, df = 7 (p = 0.28); I2 = 18% Test for overall effect: Z = 1.91 (p = 0.06) | |||||||
| TT + CT vs. CC | Shimpuku, 2003 [29] | 14 | 17 | 16 | 22 | 2.8% | 1.75 [0.37, 8.33] |
| Campos, 2005 [30] | 14 | 28 | 23 | 34 | 11.7% | 0.48 [0.17, 1.34] | |
| Laine, 2006 [31] | 41 | 71 | 30 | 49 | 16.8% | 0.87 [0.41, 1.82] | |
| Lin, 2007 [33] | 23 | 29 | 21 | 30 | 4.8% | 1.64 [0.50, 5.40] | |
| Dirschnabel, 2011 [35] | 62 | 92 | 116 | 185 | 28.2% | 1.23 [0.73, 2.08] | |
| Melo, 2012 [37] | 12 | 16 | 21 | 31 | 4.0% | 1.43 [0.37, 5.56] | |
| Cosyn, 2016 [16] | 11 | 14 | 12 | 14 | 2.9% | 0.61 [0.09, 4.37] | |
| Agrawal, 2021 [41] | 33 | 68 | 48 | 63 | 28.8% | 0.29 [0.14, 0.62] | |
| Subtotal (95% CI) | 335 | 428 | 100.0% | 0.84 [0.61, 1.14] | |||
| Total events | 210 | 287 | |||||
| Heterogeneity: Chi2 = 13.40, df = 7 (p = 0.06); I2 = 48% Test for overall effect: Z = 1.15 (p = 0.25) | |||||||
| TT vs. CC + CT | Shimpuku, 2003 [29] | 8 | 17 | 3 | 22 | 3.0% | 5.63 [1.20, 26.41] |
| Campos, 2005 [30] | 7 | 28 | 7 | 34 | 10.2% | 1.29 [0.39, 4.24] | |
| Laine, 2006 [31] | 10 | 71 | 5 | 49 | 11.0% | 1.44 [0.46, 4.52] | |
| Lin, 2007 [33] | 14 | 29 | 6 | 30 | 6.6% | 3.73 [1.18, 11.83] | |
| Dirschnabel, 2011 [35] | 21 | 92 | 28 | 185 | 31.0% | 1.66 [0.88, 3.12] | |
| Melo, 2012 [37] | 2 | 16 | 3 | 31 | 3.9% | 1.33 [0.20, 8.92] | |
| Cosyn, 2016 [16] | 5 | 14 | 6 | 14 | 8.3% | 0.74 [0.16, 3.39] | |
| Agrawal, 2021 [41] | 7 | 68 | 13 | 63 | 26.1% | 0.44 [0.16, 1.19] | |
| Subtotal (95% CI) | 335 | 428 | 100.0% | 1.45 [1.00, 2.09] | |||
| Total events | 74 | 71 | |||||
| Heterogeneity: Chi2 = 12.03, df = 7 (p = 0.10); I2 = 42% Test for overall effect: Z = 1.95 (p = 0.05) | |||||||
| Genetic Model | First Author, Publication Year | Case | Control | Weight | Odds Ratio | ||
|---|---|---|---|---|---|---|---|
| Events | Total | Events | Total | M-H, Fixed, 95% CI | |||
| T vs. C | Rogers, 2002 [28] | 11 | 38 | 13 | 62 | 22.2% | 1.54 [0.61, 3.89] |
| Campos, 2005 [30] | 13 | 56 | 14 | 68 | 30.7% | 1.17 [0.50, 2.74] | |
| Jacobi-Gresser, 2013 [39] | 24 | 82 | 28 | 136 | 47.1% | 1.60 [0.85, 3.00] | |
| Subtotal (95% CI) | 176 | 266 | 100.0% | 1.45 [0.93, 2.27] | |||
| Total events | 48 | 55 | |||||
| Heterogeneity: Chi2 = 0.35, df = 2 (p = 0.84); I2 = 0% Test for overall effect: Z = 1.64 (p = 0.10) | |||||||
| TT vs. CC | Rogers, 2002 [28] | 0 | 8 | 1 | 20 | 27.8% | 0.76 [0.03, 20.74] |
| Campos, 2005 [30] | 3 | 21 | 2 | 24 | 52.4% | 1.83 [0.28, 12.19] | |
| Jacobi-Gresser, 2013 [39] | 2 | 21 | 1 | 42 | 19.8% | 4.32 [0.37, 50.58] | |
| Subtotal (95% CI) | 50 | 86 | 100.0% | 2.03 [0.54, 7.55] | |||
| Total events | 5 | 4 | |||||
| Heterogeneity: Chi2 = 0.71, df = 2 (p = 0.70); I2 = 0% Test for overall effect: Z = 1.05 (p = 0.29) | |||||||
| CT vs. CC | Rogers, 2002 [28] | 11 | 19 | 11 | 30 | 18.7% | 2.38 [0.73, 7.69] |
| Campos, 2005 [30] | 7 | 25 | 10 | 32 | 32.8% | 0.86 [0.27, 2.70] | |
| Jacobi-Gresser, 2013 [39] | 20 | 39 | 26 | 67 | 48.5% | 1.66 [0.75, 3.68] | |
| Subtotal (95% CI) | 83 | 129 | 100.0% | 1.53 [0.87, 2.70] | |||
| Total events | 38 | 47 | |||||
| Heterogeneity: Chi2 = 1.56, df = 2 (p = 0.46); I2 = 0% Test for overall effect: Z = 1.47 (p = 0.14) | |||||||
| TT + CT vs. CC | Rogers, 2002 [28] | 11 | 19 | 12 | 31 | 19.0% | 2.18 [0.68, 6.96] |
| Campos, 2005 [30] | 10 | 28 | 12 | 34 | 34.5% | 1.02 [0.36, 2.90] | |
| Jacobi-Gresser, 2013 [39] | 22 | 41 | 27 | 68 | 46.6% | 1.76 [0.80, 3.85] | |
| Subtotal (95% CI) | 88 | 133 | 100.0% | 1.58 [0.91, 2.74] | |||
| Total events | 43 | 51 | |||||
| Heterogeneity: Chi2 = 1.04, df = 2 (p = 0.59); I2 = 0% Test for overall effect: Z = 1.64 (p = 0.10) | |||||||
| TT vs. CC + CT | Rogers, 2002 [28] | 0 | 19 | 1 | 31 | 32.6% | 0.52 [0.02, 13.46] |
| Campos, 2005 [30] | 3 | 28 | 2 | 34 | 46.7% | 1.92 [0.30, 12.38] | |
| Jacobi-Gresser, 2013 [39] | 2 | 41 | 1 | 68 | 20.7% | 3.44 [0.30, 39.13] | |
| Subtotal (95% CI) | 88 | 133 | 100.0% | 1.78 [0.49, 6.42] | |||
| Total events | 5 | 4 | |||||
| Heterogeneity: Chi2 = 0.84, df = 2 (p = 0.66); I2 = 0% Test for overall effect: Z = 0.88 (p = 0.38) | |||||||
| Genetic Model | First Author, Publication Year | Case | Control | Weight | Odds Ratio | ||
|---|---|---|---|---|---|---|---|
| Events | Total | Events | Total | M-H, Random, 95% CI | |||
| T vs. C | Shimpuku, 2003 [29] | 1 | 34 | 2 | 44 | 6.0% | 0.64 [0.06, 7.33] |
| Laine, 2006 [31] | 40 | 142 | 30 | 98 | 20.6% | 0.89 [0.51, 1.56] | |
| Lin, 2007 [33] | 7 | 58 | 2 | 60 | 10.4% | 3.98 [0.79, 20.03] | |
| Montes, 2009 [34] | 40 | 180 | 78 | 352 | 21.8% | 1.00 [0.65, 1.55] | |
| Melo, 2012 [37] | 13 | 32 | 16 | 62 | 16.9% | 1.97 [0.79, 4.87] | |
| Cosyn, 2016 [16] | 10 | 28 | 1 | 28 | 7.3% | 15.00 [1.76, 127.54] | |
| Saremi, 2021 [42] | 20 | 100 | 7 | 178 | 17.0% | 6.11 [2.48, 15.03] | |
| Subtotal (95% CI) | 574 | 822 | 100.0% | 2.04 [1.02, 4.08] | |||
| Total events | 131 | 136 | |||||
| Heterogeneity: Tau2 = 0.53; Chi2 = 22.46, df = 6 (p = 0.0010); I2 = 73% Test for overall effect: Z = 2.01 (p = 0.04) | |||||||
| TT vs. CC | Shimpuku, 2003 [29] | 0 | 16 | 0 | 20 | Not estimable | |
| Laine, 2006 [31] | 4 | 39 | 5 | 29 | 27.7% | 0.55 [0.13, 2.26] | |
| Lin, 2007 [33] | 0 | 22 | 0 | 28 | Not estimable | ||
| Montes, 2009 [34] | 2 | 54 | 8 | 114 | 25.7% | 0.51 [0.10, 2.49] | |
| Melo, 2012 [37] | 2 | 7 | 3 | 21 | 20.7% | 2.40 [0.31, 18.55] | |
| Cosyn, 2016 [16] | 2 | 8 | 0 | 13 | 12.3% | 10.38 [0.43, 249.04] | |
| Saremi, 2021 [42] | 4 | 38 | 0 | 82 | 13.6% | 21.52 [1.13, 410.64] | |
| Subtotal (95% CI) | 184 | 307 | 100.0% | 1.73 [0.45, 6.58] | |||
| Total events | 14 | 16 | |||||
| Heterogeneity: Tau2 = 1.16; Chi2 = 8.40, df = 4 (p = 0.08); I2 = 52% Test for overall effect: Z = 0.80 (p = 0.42) | |||||||
| CT vs. CC | Shimpuku, 2003 [29] | 1 | 17 | 2 | 22 | 3.5% | 0.63 [0.05, 7.53] |
| Laine, 2006 [31] | 32 | 67 | 20 | 44 | 26.7% | 1.10 [0.51, 2.35] | |
| Lin, 2007 [33] | 7 | 29 | 2 | 30 | 3.2% | 4.45 [0.84, 23.61] | |
| Montes, 2009 [34] | 36 | 88 | 62 | 168 | 53.2% | 1.18 [0.70, 2.01] | |
| Melo, 2012 [37] | 9 | 14 | 10 | 28 | 5.0% | 3.24 [0.85, 12.36] | |
| Cosyn, 2016 [16] | 6 | 12 | 1 | 14 | 1.0% | 13.00 [1.27, 133.28] | |
| Saremi, 2021 [42] | 12 | 46 | 7 | 89 | 7.5% | 4.13 [1.50, 11.40] | |
| Subtotal (95% CI) | 273 | 395 | 100.0% | 1.68 [1.18, 2.39] | |||
| Total events | 103 | 104 | |||||
| Heterogeneity: Chi2 = 11.73, df = 6 (p = 0.07); I2 = 49% Test for overall effect: Z = 2.89 (p = 0.004) | |||||||
| TT + CT vs. CC | Shimpuku, 2003 [29] | 1 | 17 | 2 | 22 | 6.3% | 0.63 [0.05, 7.53] |
| Laine, 2006 [31] | 36 | 71 | 25 | 49 | 20.6% | 0.99 [0.48, 2.05] | |
| Lin, 2007 [33] | 7 | 29 | 2 | 30 | 10.8% | 4.45 [0.84, 23.61] | |
| Montes, 2009 [34] | 38 | 90 | 70 | 176 | 23.1% | 1.11 [0.66, 1.85] | |
| Melo, 2012 [37] | 11 | 16 | 13 | 31 | 14.3% | 3.05 [0.85, 10.90] | |
| Cosyn, 2016 [16] | 8 | 14 | 1 | 14 | 7.1% | 17.33 [1.75, 171.66] | |
| Saremi, 2021 [42] | 16 | 50 | 7 | 89 | 17.7% | 5.51 [2.08, 14.60] | |
| Subtotal (95% CI) | 287 | 411 | 100.0% | 2.27 [1.11, 4.64] | |||
| Total events | 117 | 120 | |||||
| Heterogeneity: Tau2 = 0.51; Chi2 = 17.02, df = 6 (p = 0.009); I2 = 65% Test for overall effect: Z = 2.23 (p = 0.03) | |||||||
| TT vs. CC + CT | Shimpuku, 2003 [29] | 0 | 17 | 0 | 22 | Not estimable | |
| Laine, 2006 [31] | 4 | 71 | 5 | 49 | 41.6% | 0.53 [0.13, 2.06] | |
| Lin, 2007 [33] | 0 | 29 | 0 | 30 | Not estimable | ||
| Montes, 2009 [34] | 2 | 90 | 8 | 176 | 39.5% | 0.48 [0.10, 2.30] | |
| Melo, 2012 [37] | 2 | 16 | 3 | 31 | 13.3% | 1.33 [0.20, 8.92] | |
| Cosyn, 2016 [16] | 2 | 14 | 0 | 14 | 3.1% | 5.80 [0.25, 132.56] | |
| Saremi, 2021 [42] | 4 | 50 | 0 | 89 | 2.5% | 17.32 [0.91, 328.70] | |
| Subtotal (95% CI) | 287 | 411 | 100.0% | 1.19 [0.58, 2.45] | |||
| Total events | 14 | 16 | |||||
| Heterogeneity: Chi2 = 6.85, df = 4 (p = 0.14); I2 = 42% Test for overall effect: Z = 0.47 (p = 0.63) | |||||||
| Genetic Model | First Author, Publication Year | Case | Control | Weight | Odds Ratio | ||
|---|---|---|---|---|---|---|---|
| Events | Total | Events | Total | M-H, Random, 95% CI | |||
| A2 vs. A1 | Campos, 2005 [30] | 19 | 54 | 17 | 66 | 47.5% | 1.56 [0.71, 3.43] |
| Petkovic-Curcin, 2017 [40] | 18 | 68 | 50 | 128 | 52.5% | 0.56 [0.29, 1.07] | |
| Subtotal (95% CI) | 122 | 194 | 100.0% | 0.91 [0.34, 2.49] | |||
| Total events | 37 | 67 | |||||
| Heterogeneity: Tau2 = 0.39; Chi2 = 3.91, df = 1 (p = 0.05); I2 = 74% Test for overall effect: Z = 0.18 (p = 0.86) | |||||||
| A2A2 vs. A1A1 | Campos, 2005 [30] | 3 | 14 | 2 | 20 | 53.4% | 2.45 [0.35, 17.08] |
| Petkovic-Curcin, 2017 [40] | 0 | 22 | 11 | 36 | 46.6% | 0.05 [0.00, 0.88] | |
| Subtotal (95% CI) | 36 | 56 | 100.0% | 0.40 [0.01, 23.82] | |||
| Total events | 3 | 13 | |||||
| Heterogeneity: Tau2 = 7.19; Chi2 = 5.57, df = 1 (p = 0.02); I2 = 82% Test for overall effect: Z = 0.44 (p = 0.66) | |||||||
| A1A2 vs. A1A1 | Campos, 2005 [30] | 13 | 24 | 13 | 31 | 46.7% | 1.64 [0.56, 4.79] |
| Petkovic-Curcin, 2017 [40] | 12 | 34 | 28 | 53 | 53.3% | 0.49 [0.20, 1.18] | |
| Subtotal (95% CI) | 58 | 84 | 100.0% | 0.86 [0.26, 2.81] | |||
| Total events | 25 | 41 | |||||
| Heterogeneity: Tau2 = 0.48; Chi2 = 2.91, df = 1 (p = 0.09); I2 = 66% Test for overall effect: Z = 0.25 (p = 0.80) | |||||||
| A2A2 + A1A2 vs. A1A1 | Campos, 2005 [30] | 16 | 27 | 15 | 33 | 48.1% | 1.75 [0.62, 4.88] |
| Petkovic-Curcin, 2017 [40] | 13 | 34 | 39 | 64 | 51.9% | 0.40 [0.17, 0.93] | |
| Subtotal (95% CI) | 61 | 97 | 100.0% | 0.81 [0.19, 3.45] | |||
| Total events | 29 | 54 | |||||
| Heterogeneity: Tau2 = 0.86; Chi2 = 4.71, df = 1 (p = 0.03); I2 = 79% Test for overall effect: Z = 0.29 (p = 0.77) | |||||||
| A2A2 vs. A1A1 + A1A2 | Campos, 2005 [30] | 3 | 27 | 2 | 33 | 18.7% | 1.94 [0.30, 12.53] |
| Petkovic-Curcin, 2017 [40] | 3 | 34 | 11 | 64 | 81.3% | 0.47 [0.12, 1.80] | |
| Subtotal (95% CI) | 61 | 97 | 100.0% | 0.74 [0.26, 2.09] | |||
| Total events | 6 | 13 | |||||
| Heterogeneity: Chi2 = 1.47, df = 1 (p = 0.23); I2 = 32% Test for overall effect: Z = 0.57 (p = 0.57) | |||||||
| The Composite Genotype | First Author, Publication Year | Case | Control | Weight | Odds Ratio | ||
|---|---|---|---|---|---|---|---|
| Events | Total | Events | Total | M-H, Fixed, 95% CI | |||
| IL−1A (−889) and IL−1B (+3953) | Rogers, 2002 [28] | 9 | 19 | 11 | 31 | 20.7% | 1.64 [0.51, 5.23] |
| Campos, 2005 [30] | 8 | 28 | 10 | 34 | 30.3% | 0.96 [0.32, 2.89] | |
| Vaz, 2012 [38] | 25 | 55 | 27 | 100 | 49.1% | 2.25 [1.13, 4.49] | |
| Total (95% CI) | 102 | 165 | 100.0% | 1.73 [1.03, 2.92] | |||
| Total events | 42 | 48 | |||||
| Heterogeneity: Chi2 = 1.67, df = 2 (p = 0.43); I2 = 0% Test for overall effect: Z = 2.08 (p = 0.04) | |||||||
| IL−1A (−889) and IL−1B (+3954) | Laine, 2006 [31] | 34 | 71 | 22 | 49 | 40.5% | 1.13 [0.54, 2.34] |
| Lachmann, 2007 [32] | 6 | 11 | 8 | 18 | 28.1% | 1.50 [0.33, 6.77] | |
| Hamdy, 2011 [36] | 17 | 25 | 5 | 25 | 31.4% | 8.50 [2.34, 30.91] | |
| Total (95% CI) | 107 | 92 | 100.0% | 2.31 [0.65, 8.17] | |||
| Total events | 57 | 35 | |||||
| Heterogeneity: Tau2 = 0.89; Chi2 = 7.19, df = 2 (p = 0.03); I2 = 72% Test for overall effect: Z = 1.29 (p = 0.20) | |||||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mohammadi, H.; Roochi, M.M.; Sadeghi, M.; Garajei, A.; Heidar, H.; Meybodi, A.A.; Dallband, M.; Mostafavi, S.; Mostafavi, M.; Salehi, M.; et al. Association between Interleukin-1 Polymorphisms and Susceptibility to Dental Peri-Implant Disease: A Meta-Analysis. Pathogens 2021, 10, 1600. https://doi.org/10.3390/pathogens10121600
Mohammadi H, Roochi MM, Sadeghi M, Garajei A, Heidar H, Meybodi AA, Dallband M, Mostafavi S, Mostafavi M, Salehi M, et al. Association between Interleukin-1 Polymorphisms and Susceptibility to Dental Peri-Implant Disease: A Meta-Analysis. Pathogens. 2021; 10(12):1600. https://doi.org/10.3390/pathogens10121600
Chicago/Turabian StyleMohammadi, Hady, Mehrnoush Momeni Roochi, Masoud Sadeghi, Ata Garajei, Hosein Heidar, Ali Aghaie Meybodi, Mohsen Dallband, Sarton Mostafavi, Melina Mostafavi, Mojtaba Salehi, and et al. 2021. "Association between Interleukin-1 Polymorphisms and Susceptibility to Dental Peri-Implant Disease: A Meta-Analysis" Pathogens 10, no. 12: 1600. https://doi.org/10.3390/pathogens10121600
APA StyleMohammadi, H., Roochi, M. M., Sadeghi, M., Garajei, A., Heidar, H., Meybodi, A. A., Dallband, M., Mostafavi, S., Mostafavi, M., Salehi, M., Tadakamadla, J., Sadeghi-Bahmani, D., & Brand, S. (2021). Association between Interleukin-1 Polymorphisms and Susceptibility to Dental Peri-Implant Disease: A Meta-Analysis. Pathogens, 10(12), 1600. https://doi.org/10.3390/pathogens10121600








