Unraveling the Contact Network Patterns between Commercial Turkey Operation in North Carolina and the Distribution of Salmonella Species
Abstract
:1. Introduction
2. Results
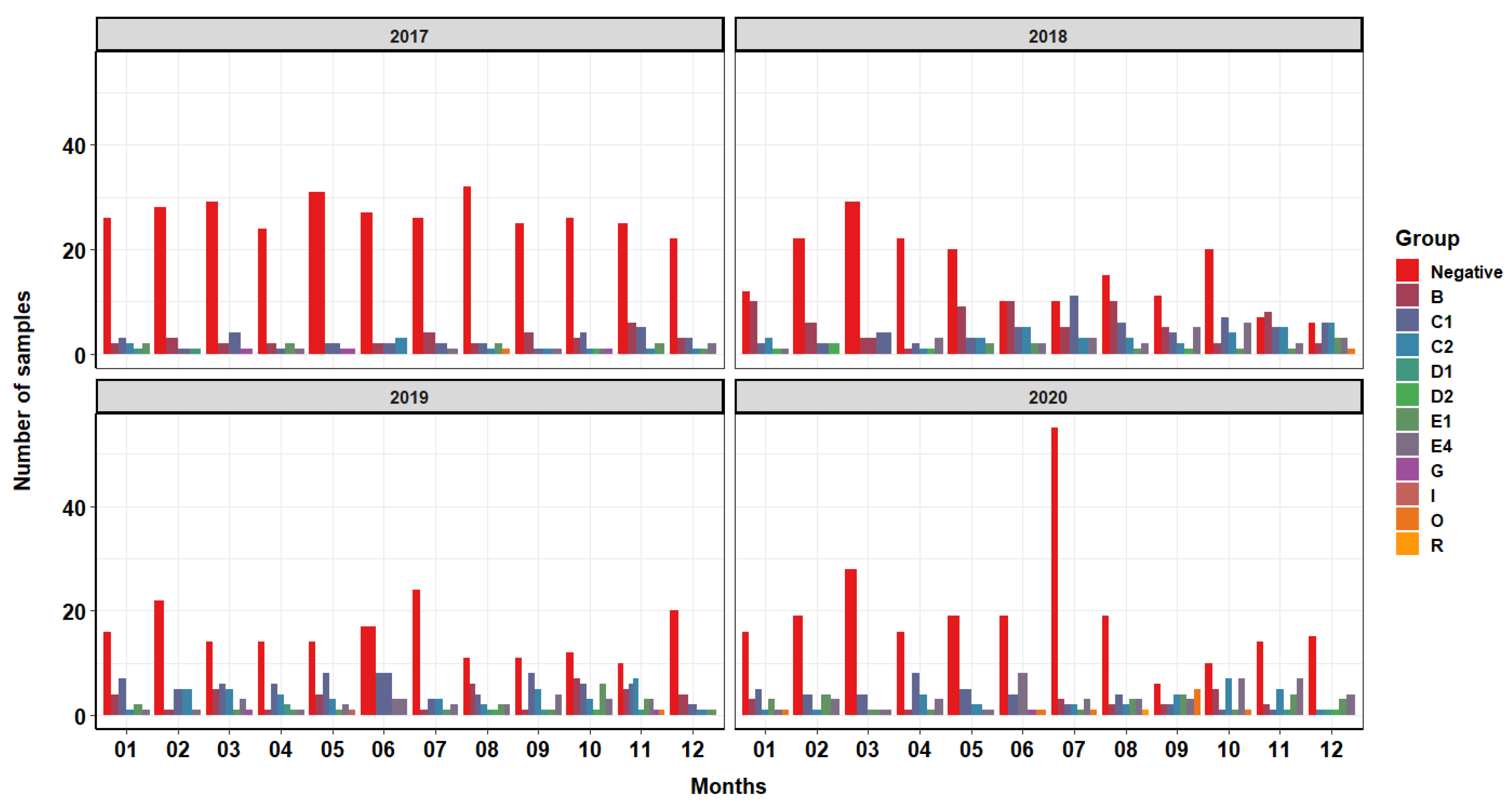
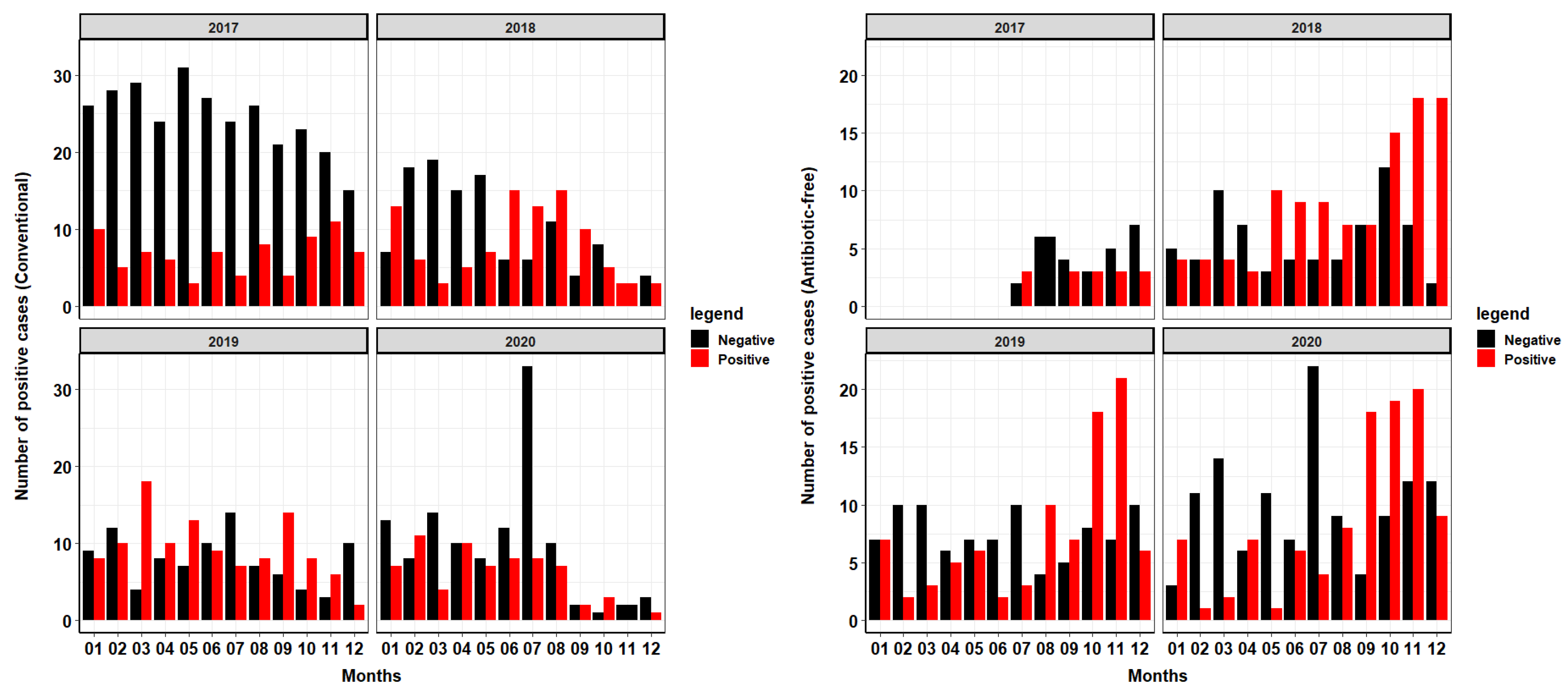
2.1. General Description of the Collected Database and the Distribution of Salmonella spp. Groups and Serotypes
Facility Age Influence on Salmonella
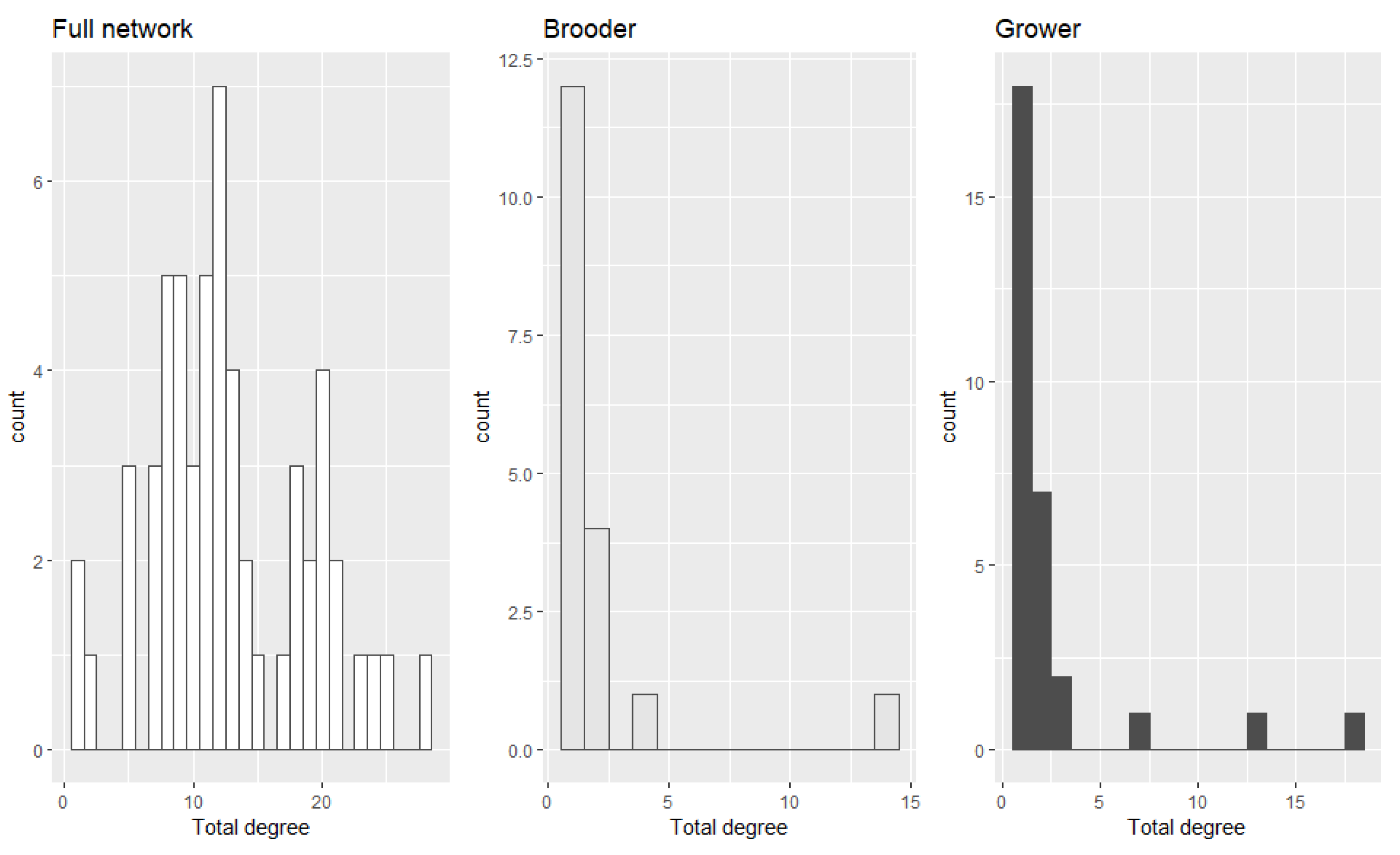
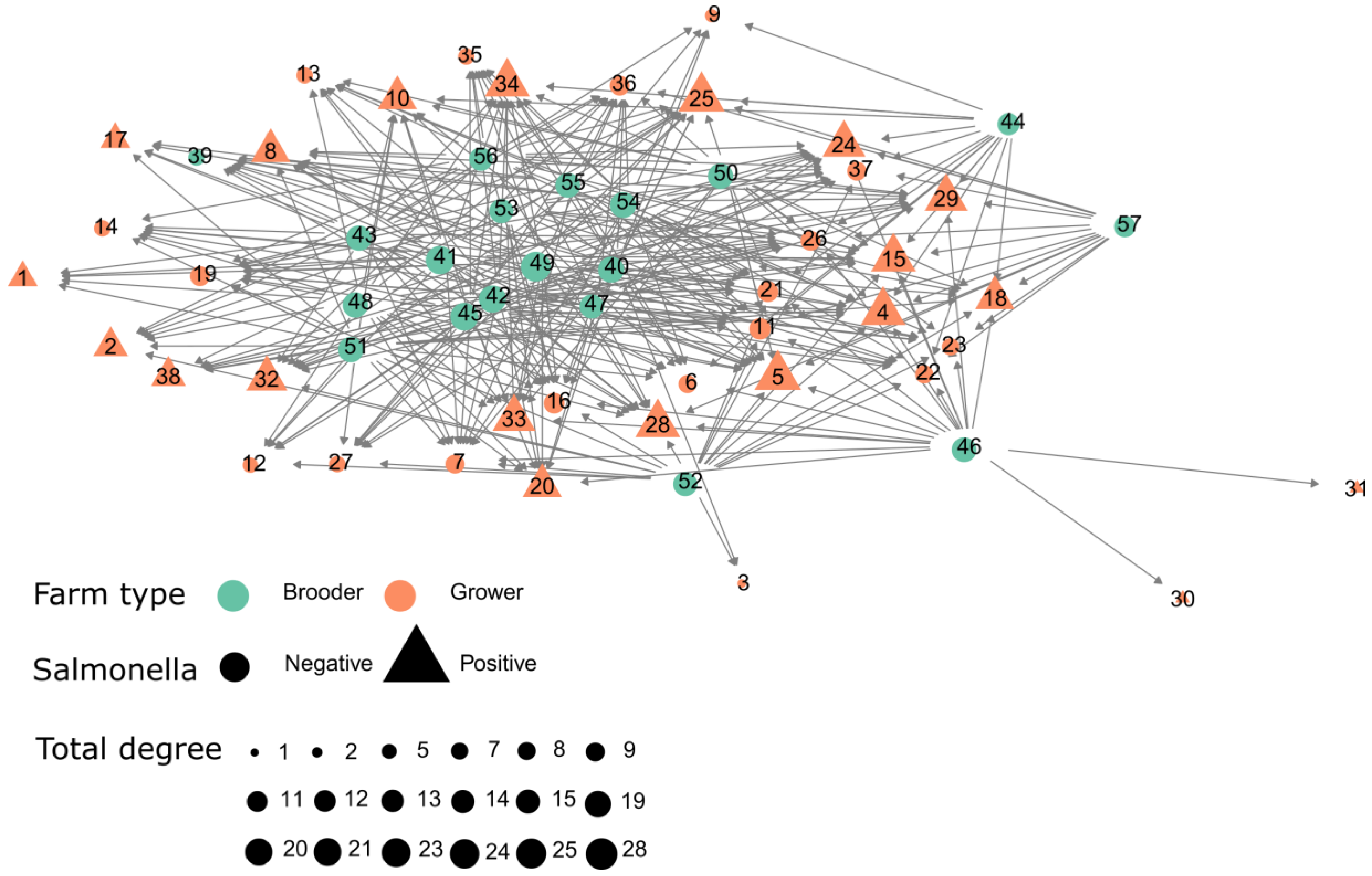
2.2. Between-Farm Contact Networks
The Contact Network for the Movements Involving Infected Movements between Brooder and Growout Farms
3. Discussion
4. Materials and Methods
4.1. General Description of Data and Mathematical Calculations Used to Aid in Data Interpretation
4.2. Contacts and Networks
4.2.1. Between Farm Contact Networks
4.2.2. The Contact Network for the Movements Involving Infected Movements
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Foley, S.L.; Lynne, A.M.; Nayak, R. Salmonella challenges: Prevalence in swine and poultry and potential pathogenicity of such isolates1,2. J. Anim. Sci. 2008, 86, E149–E162. [Google Scholar] [CrossRef]
- Scharff, R.L. Food Attribution and Economic Cost Estimates for Meat- and Poultry-Related Illnesses. J. Food Prot. 2020, 83, 959–967. [Google Scholar] [CrossRef] [PubMed]
- Vandeplas, S.; Dauphin, R.D.; Beckers, Y.; Thonart, P.; Théwis, A. Salmonella in Chicken: Current and Developing Strategies to Reduce Contamination at Farm Level. J. Food Prot. 2010, 73, 774–785. [Google Scholar] [CrossRef] [PubMed]
- Brenner, F.W.; Villar, R.G.; Angulo, F.J.; Tauxe, R.; Swaminathan, B. Salmonella Nomenclature. J. Clin. Microbiol. 2000, 38, 2465–2467. [Google Scholar] [CrossRef] [Green Version]
- Swayne, D.E. (Ed.) Diseases of Poultry, 14th ed.; Wiley-Blackwell: Hoboken, NJ, USA, 2020. [Google Scholar]
- Peng, Y.; Deng, X.Y.; Harrison, M.A.; Alali, W.Q. Salmonella Levels Associated with Skin of Turkey Parts. J. Food Prot. 2016, 79, 801–805. [Google Scholar] [CrossRef]
- Yang, Q.; Zhao, X.; Lemey, P.; Suchard, M.A.; Bi, Y.; Shi, W.; Liu, D.; Qi, W.; Zhang, G.; Stenseth, N.C.; et al. Assessing the role of live poultry trade in community-structured transmission of avian influenza in China. Proc. Natl. Acad. Sci. USA 2020, 117, 5949–5954. [Google Scholar] [CrossRef] [Green Version]
- Guinat, C.; Durand, B.; Vergne, T.; Corre, T.; Rautureau, S.; Scoizec, A.; Lebouquin-Leneveu, S.; Guérin, J.-L.; Paul, M.C. Role of Live-Duck Movement Networks in Transmission of Avian Influenza, France, 2016–2017. Emerg. Infect. Dis. 2020, 26, 472–480. [Google Scholar] [CrossRef] [Green Version]
- Heyndrickx, M.; Vandekerchove, D.; Herman, L.; Rollier, I.; Grijspeerdt, K.; De Zutter, L. Routes for salmonella contamination of poultry meat: Epidemiological study from hatchery to slaughterhouse. Epidemiol. Infect. 2002, 129, 253–265. [Google Scholar] [CrossRef] [PubMed]
- Moyen, N.; Ahmed, G.; Gupta, S.; Tenzin, T.; Khan, R.; Khan, T.; Debnath, N.; Yamage, M.; Pfeiffer, D.; Fournie, G. A large-scale study of a poultry trading network in Bangladesh: Implications for control and surveillance of avian influenza viruses. BMC Vet. Res. 2018, 14, 12. [Google Scholar] [CrossRef] [Green Version]
- Brown, V.R.; Miller, R.S.; McKee, S.C.; Ernst, K.H.; Didero, N.M.; Maison, R.M.; Grady, M.J.; Shwiff, S.A. Risks of introduction and economic consequences associated with African swine fever, classical swine fever and foot-and-mouth disease: A review of the literature. Transbound. Emerg. Dis. 2020, 68, 1910–1965. [Google Scholar] [CrossRef]
- Halasa, T.; Bøtner, A.; Mortensen, S.; Christensen, H.; Toft, N.; Boklund, A. Control of African swine fever epidemics in industrialized swine populations. Vet. Microbiol. 2016, 197, 142–150. [Google Scholar] [CrossRef]
- Halasa, T.; Bøtner, A.; Mortensen, S.; Christensen, H.; Wulff, S.B.; Boklund, A. Modeling the Effects of Duration and Size of the Control Zones on the Consequences of a Hypothetical African Swine Fever Epidemic in Denmark. Front. Vet. Sci. 2018, 5, 49. [Google Scholar] [CrossRef]
- Beltrán-Alcrudo, D.; Gallardo, M.A.A.C.; Kramer, S.A.; Penrith, M.L.; Kamata, A.; Wiersma, L. African Swine Fever: Detection and Diagnosis: A Manual for Veterinarians; Springer Publishing: New York City, NY, USA, 2017. [Google Scholar]
- Guinat, C.; Porphyre, T.; Gogin, A.; Dixon, L.; Pfeiffer, D.U.; Gubbins, S. Inferring Within-Herd Transmission Parameters for African Swine Fever Virus Using Mortality Data from Outbreaks in the Russian Federation. Transbound. Emerg. Dis. 2018, 65, e264–e271. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Anderson, P.; Hume, M.; Byrd, J.; Hernandez, C.; Stevens, S.; Stringfellow, K.; Caldwell, D. Molecular analysis of Salmonella serotypes at different stages of commercial turkey processing. Poult. Sci. 2010, 89, 2030–2037. [Google Scholar] [CrossRef] [PubMed]
- Bourassa, D.V.; Holmes, J.M.; Cason, J.A.; Cox, N.A.; Rigsby, L.L.; Buhr, R.J. Prevalence and Serogroup Diversity of Salmonella for Broiler Neck Skin, Whole Carcass Rinse, and Whole Carcass Enrichment Sampling Methodologies following Air or Immersion Chilling†. J. Food Prot. 2015, 78, 1938–1944. [Google Scholar] [CrossRef]
- Santos, F.B.O.; Li, X.; Payne, J.B.; Sheldon, B.W. Estimation of Most Probable Number Salmonella Populations on Commercial North Carolina Turkey Farms. J. Appl. Poult. Res. 2005, 14, 700–708. [Google Scholar] [CrossRef]
- Gupta, R.; Kaur, D.; Chopra, S.; Nagra, S.; Rai, D.; Patil, R. Performance analysis of the broiler chicks under different cooling devices during hot-dry summer. Indian J. Anim. Res. 2014, 48, 480. [Google Scholar] [CrossRef]
- Dannemiller, K.C.; Weschler, C.J.; Peccia, J. Fungal and bacterial growth in floor dust at elevated relative humidity levels. Indoor Air 2016, 27, 354–363. [Google Scholar] [CrossRef]
- Van Hoorebeke, S.; Van Immerseel, F.; De Vylder, J.; Ducatelle, R.; Haesebrouck, F.; Pasmans, F.; de Kruif, A.; Dewulf, J. The age of production system and previous Salmonella infections on-farm are risk factors for low-level Salmonella infections in laying hen flocks. Poult. Sci. 2010, 89, 1315–1319. [Google Scholar] [CrossRef] [PubMed]
- Davies, R.H.; Breslin, M. Persistence of Salmonella Enteritidis Phage Type 4 in the environment and arthropod vectors on an empty free-range chicken farm. Environ. Microbiol. 2003, 5, 79–84. [Google Scholar] [CrossRef]
- Cárdenas, N.C.; Galvis, J.O.A.; Farinati, A.A.; Grisi-Filho, J.H.H.; Diehl, G.N.; Machado, G.; Grisi-Filho, J.H.H.; Cardenas, N. Burkholderia mallei: The dynamics of networks and disease transmission. Transbound. Emerg. Dis. 2018, 66, 715–728. [Google Scholar] [CrossRef] [PubMed]
- Wasserman, S.; Faust, K. Social Network Analysis Methods and Applications; Cambridge University Press: Cambridge, UK, 2012. [Google Scholar]
- Craft, M.E. Infectious disease transmission and contact networks in wildlife and livestock. Philos. Trans. R. Soc. B Biol. Sci. 2015, 370, 20140107. [Google Scholar] [CrossRef] [PubMed]
- VanderWaal, K.; Enns, E.; Picasso-Risso, C.; Packer, C.; Craft, M.E. Evaluating empirical contact networks as potential transmission pathways for infectious diseases. J. R. Soc. Interface 2016, 13, 20160166. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2017; Available online: https://www.r-project.org (accessed on 20 November 2021).
| Group | Number of Positive | Proportion (%) |
|---|---|---|
| C1 | 194 | 28.67 |
| B | 165 | 24.37 |
| C2 | 116 | 17.13 |
| E4 | 101 | 14.92 |
| E1 | 63 | 9.31 |
| O | 12 | 1.77 |
| D1, D2 | 9 | 1.33 |
| G | 6 | 0.89 |
| I, R | 1 | 0.15 |
| Serovar | Group | Number of Positive | Proportion (%) |
|---|---|---|---|
| Infantis | C1 | 179 | 26.44 |
| Senftenberg | E4 | 86 | 12.76 |
| Albany | C2 | 74 | 10.93 |
| Schwarzengrund | B | 59 | 8.71 |
| Uganda | E1 | 56 | 8.27 |
| Agona | B | 40 | 5.91 |
| Muenchen | C2 | 28 | 4.13 |
| 1,4,5,12 | B | 26 | 3.84 |
| Typhimurium | B | 17 | 2.51 |
| Liverpool | E4 | 16 | 2.36 |
| Reading | B | 15 | 2.21 |
| Hadar | C2 | 10 | 1.47 |
| Berta, Ouakam, Rissen | * | 9 | 1.33 |
| Alachua | O | 8 | 1.18 |
| Anatum | E1 | 6 | 0.88 |
| Lillie, Worthington | * | 5 | 0.74 |
| Heidelberg, Rough O | * | 4 | 0.59 |
| 6,7:r:-, Derby, Kiambu | * | 2 | 0.29 |
| 16:d:-, 6,8:z10:-, Arizoniae, Johannesburg, Kentucky, Newport | * | 1 | 0.15 |
| Parameter | Network Metric at Farm Level |
|---|---|
| Nodes | 56 |
| Edges | 358 |
| Sum of moved batches | 888 |
| Mean degree | 12.78 |
| Centralization | 0.14 |
| Max value of in degree | 14 |
| Max value of out-degree | 28 |
| Max size of GWCC | 56 (100%) |
| Max size of GSCC | 1 (1.78%) |
| Mean betweenness | 0 |
| Parameter | Definition | Reference |
|---|---|---|
| Nodes | The unit of interest in network analysis. For example, premises or slaughterhouses. | [24] |
| Edge | The link between two nodes in a network. | [24] |
| Degree (k) | A number of unique contacts to and from a specific premise. When the directionality is considered, the ingoing and outgoing contacts are defined: out-degree is the number of contacts originating from a specific premise, and in-degree is the number of contacts coming into a specific premise. | [24] |
| Movements | The number of movements as batches are recorded over a certain period of time. | [24] |
| Betweenness | The extent to which a node lies on a path connecting other pairs of nodes, defined by the number of geodesics (shortest paths) going through a node. | [24] |
| Giant weakly connected component (GWCC) | The proportion of nodes that are connected in the largest component when directionality of movement is ignored. | [24] |
| Giant strongly connected component (GSCC) | The proportion of the nodes that are connected in the largest component when directionality of movement is considered. | [24] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ellington, C.; Hebron, C.; Crespo, R.; Machado, G. Unraveling the Contact Network Patterns between Commercial Turkey Operation in North Carolina and the Distribution of Salmonella Species. Pathogens 2021, 10, 1539. https://doi.org/10.3390/pathogens10121539
Ellington C, Hebron C, Crespo R, Machado G. Unraveling the Contact Network Patterns between Commercial Turkey Operation in North Carolina and the Distribution of Salmonella Species. Pathogens. 2021; 10(12):1539. https://doi.org/10.3390/pathogens10121539
Chicago/Turabian StyleEllington, Cameron, Claude Hebron, Rocio Crespo, and Gustavo Machado. 2021. "Unraveling the Contact Network Patterns between Commercial Turkey Operation in North Carolina and the Distribution of Salmonella Species" Pathogens 10, no. 12: 1539. https://doi.org/10.3390/pathogens10121539
APA StyleEllington, C., Hebron, C., Crespo, R., & Machado, G. (2021). Unraveling the Contact Network Patterns between Commercial Turkey Operation in North Carolina and the Distribution of Salmonella Species. Pathogens, 10(12), 1539. https://doi.org/10.3390/pathogens10121539







