Experimental Induction of Tenacibaculosis in Atlantic Salmon (Salmo salar L.) Using Tenacibaculum maritimum, T. dicentrarchi, and T. finnmarkense
Abstract
:1. Introduction
2. Results
2.1. Bacterial Isolate Collection
2.2. Clinical Signs of Infection
2.3. Kaplan–Meier Analysis
2.4. qPCR Regression Modeling
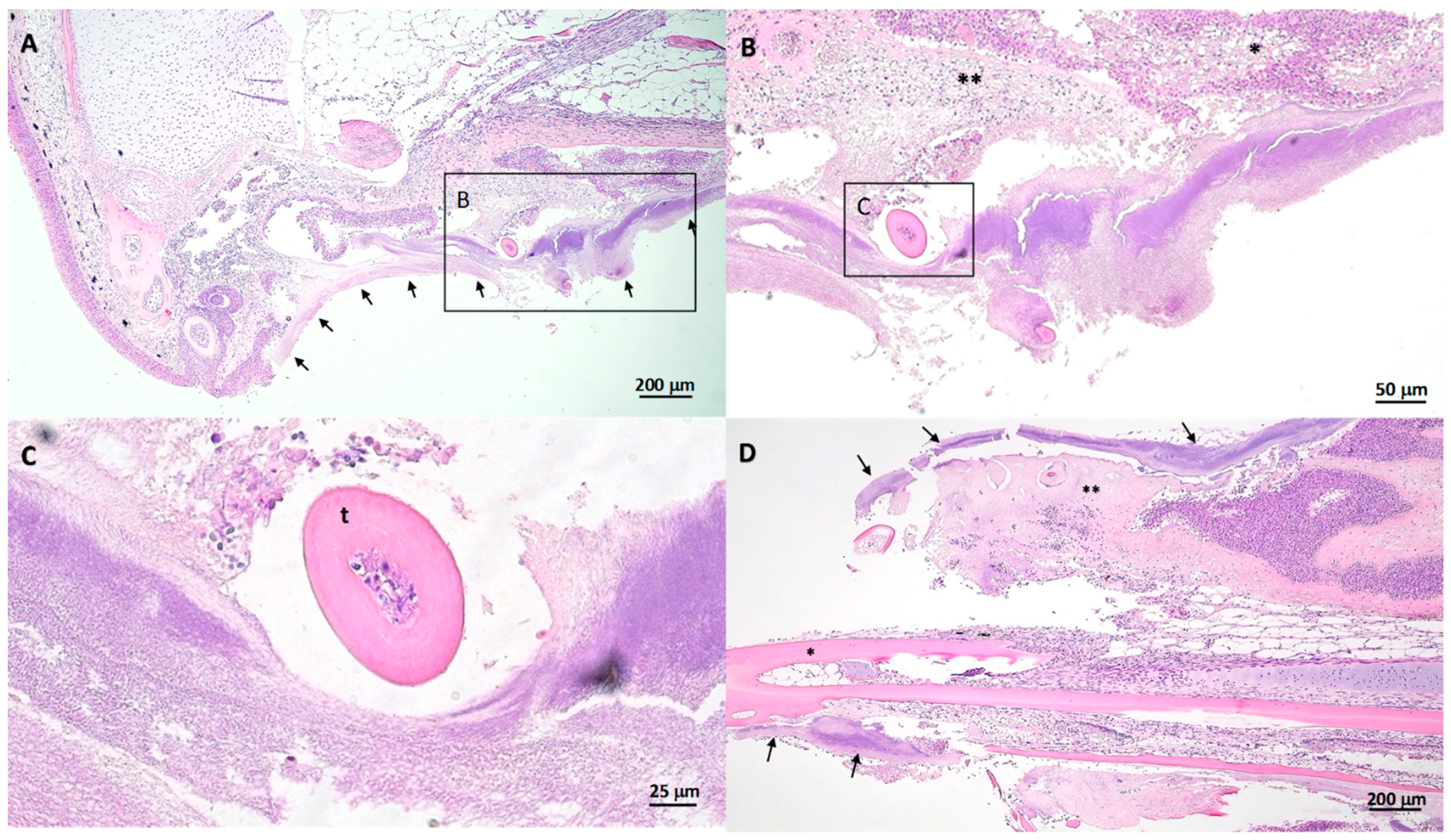
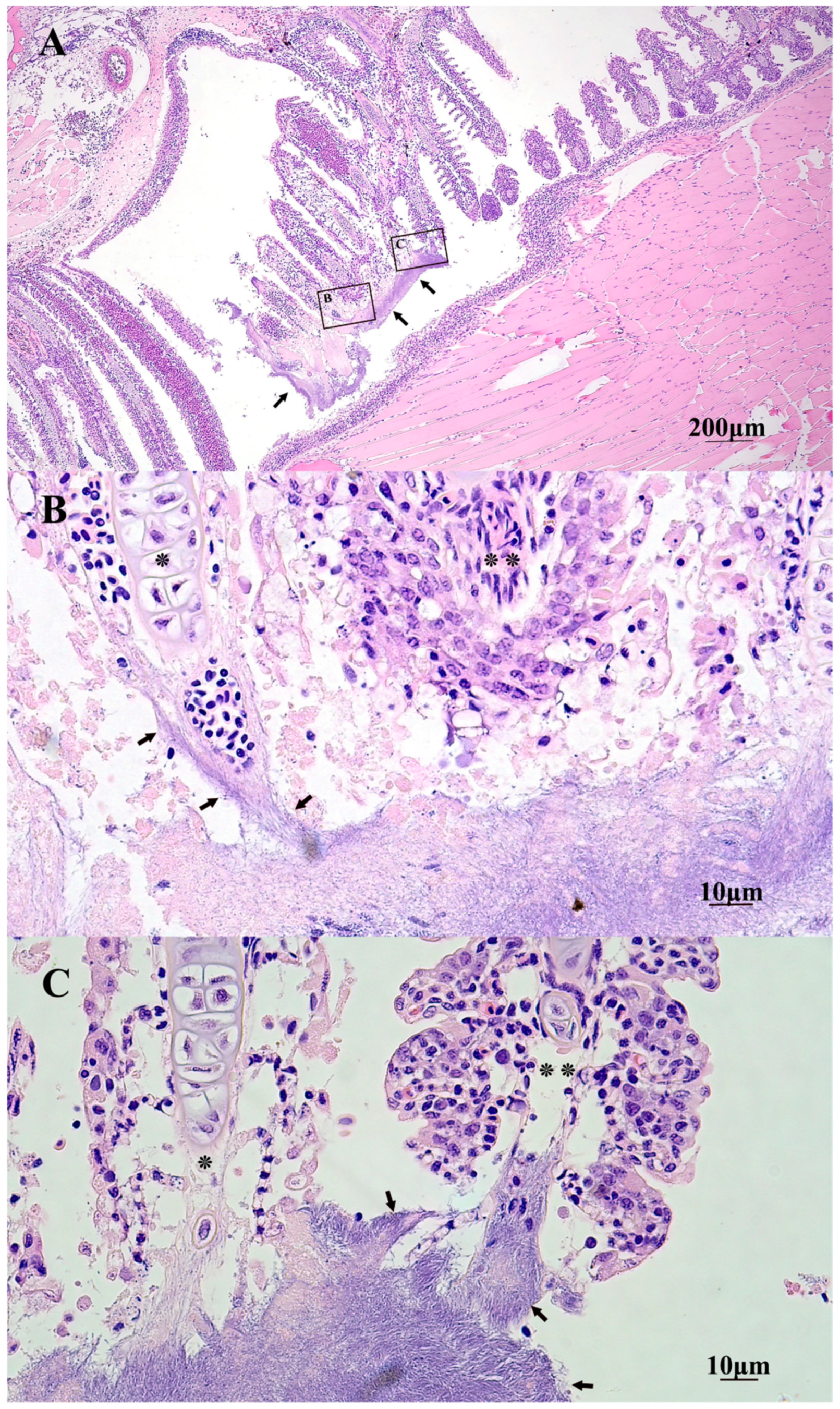
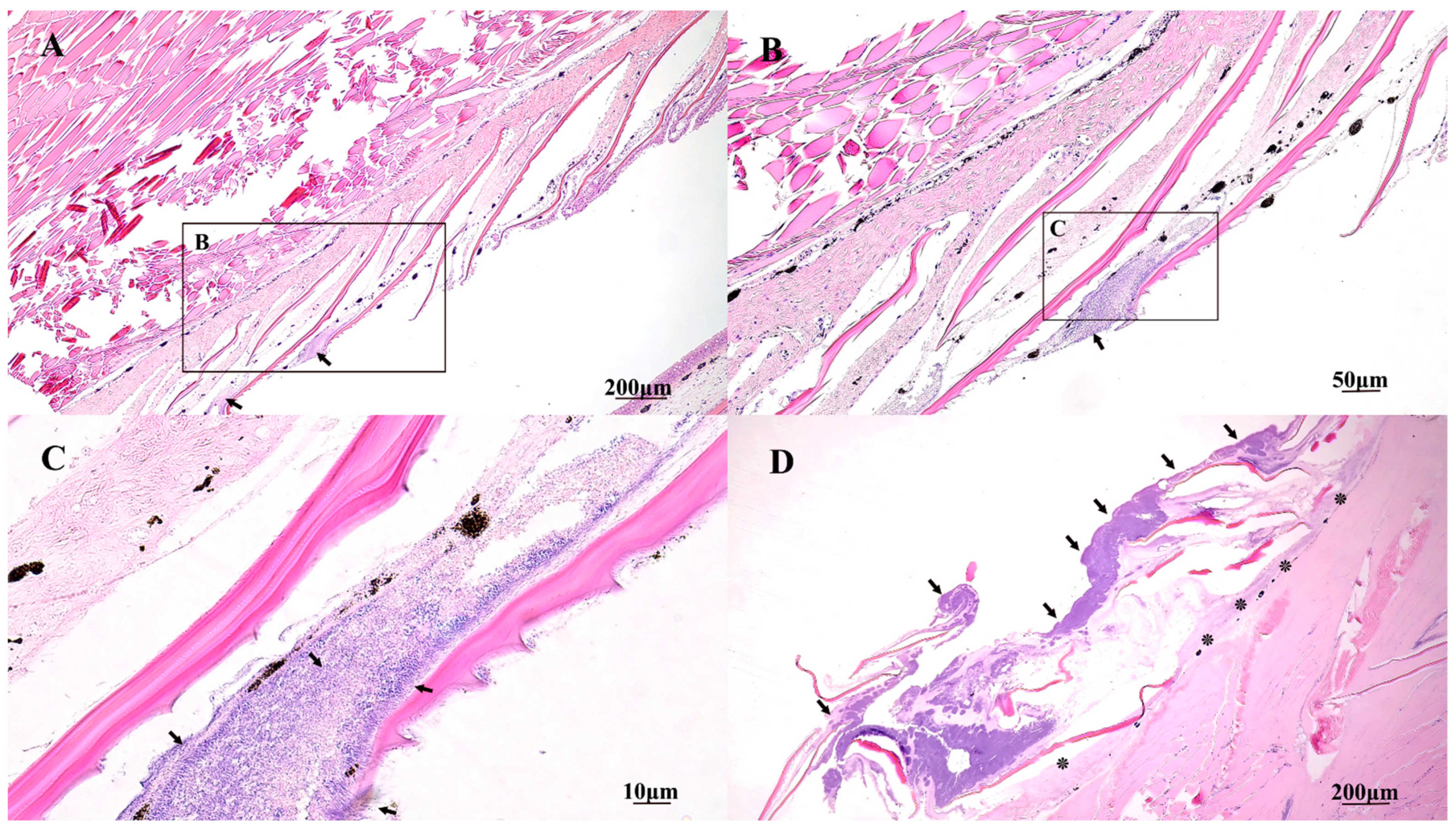
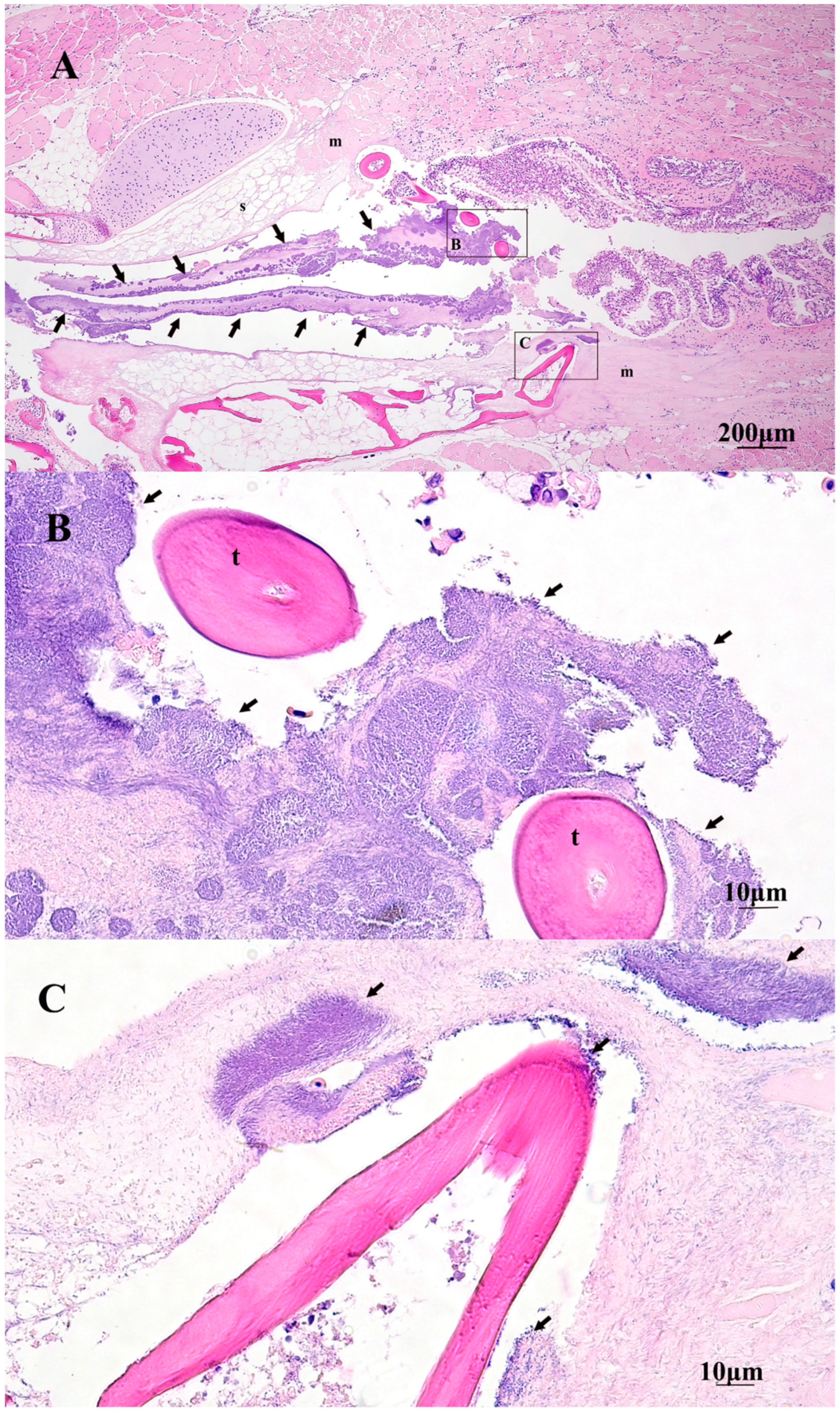
2.5. Gross and Histopathology
3. Discussion
4. Materials and Methods
4.1. Fish Husbandry
4.2. Bacterial Propagation
4.3. Direct and Cohabitation Bath Exposures
4.4. Pre- and Post-Exposure Sampling
4.5. Quantitative PCR
4.6. Histopathology
4.7. Statistics
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Nowlan, J.P.; Lumsden, J.S.; Russell, S. Advancements in characterizing Tenacibaculum infections in Canada. Pathogens 2020, 9, 1029. [Google Scholar] [CrossRef] [PubMed]
- Avendaño-Herrera, R.; Toranzo, A.E.; Magariños, B. Tenacibaculosis infection in marine fish caused by Tenacibaculum maritmum: A review. Dis. Aquat. Org. 2006, 71, 255–266. [Google Scholar] [CrossRef]
- Avendaño-Herrera, R.; Collarte, C.; Saldarriaga-Córdoba, M.; Irgang, R. New salmonid hosts for Tenacibaculum species: Expansion of tenacibaculosis in Chilean aquaculture. J. Fish Dis. 2020, 43, 1077–1108. [Google Scholar] [CrossRef]
- Avendaño-Herrera, R.; Irgang, R.; Sandoval, C.; Moreno-Lira, P.; Houel, A.; Duchaud, E.; Ilardi, P. Isolation, characterization and virulence potential of Tenacibaculum dicentrarchi in salmonid cultures in Chile. Transbound. Emerg. Dis. 2016, 63, 121–126. [Google Scholar] [CrossRef] [PubMed]
- Frisch, K.; Småge, S.B.; Johansen, R.; Duesund, H.; Brevik, Ø.J.; Nylund, A. Pathology of experimentally induced mouthrot caused by Tenacibaculum maritimum in Atlantic salmon smolts. PLoS ONE 2018, 13, e0206951. [Google Scholar] [CrossRef] [PubMed]
- Frisch, K.; Vallestad, C.; Duesund, H.; Brevik, Ø.J.; Klevan, A.; Nylund, A. Experimental induction of mouthrot in Atlantic salmon smolts using Tenacibaculum maritimum from Western Canada. J. Fish Dis. 2018, 41, 1247–1258. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nowlan, J.P.; Lumsden, J.S.; Russell, S. Quantitative PCR for Tenacibaculum dicentrarchi and T. finnmarkense. J. Fish Dis. 2021, 44, 655–659. [Google Scholar] [CrossRef]
- Nowlan, J.P.; Britney, S.R.; Lumsden, J.S.; Russell, S. Application of quantitative-PCR to monitor netpen sites in British Columbia (Canada) for Tenacibaculum species. Pathogens 2021, 10, 414. [Google Scholar] [CrossRef]
- Småge, S.B.; Frisch, K.; Vold, V.; Duesund, H.; Brevik, Ø.J.; Olsen, R.H.; Nylund, A. Induction of tenacibaculosis in Atlantic salmon smolts using Tenacibaculum finnmarkense and the evaluation of a whole cell inactivated vaccine. Aquaculture 2018, 495, 858–864. [Google Scholar] [CrossRef]
- Wynne, J.W.; Thakur, K.K.; Slinger, J.; Samsing, F.; Milligan, B.; Powell, J.F.; Siah, A. Microbiome profiling reveals a microbial dysbiosis during a natural outbreak of tenacibaculosis (Yellow mouth) in Atlantic salmon. Front. Microb. 2020, 11, 586387. [Google Scholar] [CrossRef] [PubMed]
- Slinger, J.; Adams, M.B.; Wynne, J.W. Bacteriomic profiling of branchial lesions induced by Neoparamoeba perurans challenge reveals commensal dysbiosis and an association with Tenacibaculum dicentrarchi in AGD-affected Atlantic salmon (Salmo salar L.). Microorganisms 2020, 8, 1189. [Google Scholar] [CrossRef] [PubMed]
- Hipra Laboratories. (N.D.). ICTHIOVAC® TM. Available online: https://www.hipra.com/portal/en/hipra/animalhealth/products/detail-global/icthiovac-tm (accessed on 19 March 2021).
- Hewison, T.; Ness, M. Tenacibaculum maritimum—A Fish Farm Perspective. Tenacibaculum maritimum: Current Knowledge & Future Directions. Workshop Report and Synopsis. Maritime Heritage Centre, Campbell River, Canada, 2015. Available online: http://docplayer.net/37754558-Tenacibaculum-maritimum-current-knowledge-future-directions.html (accessed on 19 March 2021).
- Hesami, S.; Allen, K.J.; Metcalf, D.; Ostland, V.E.; MacInnes, J.I.; Lumsden, J.S. Phenotypic and genotypic analysis of Flavobacterium psychrophilum isolates from Ontario salmonids with bacterial coldwater disease. Can. J. Microb. 2008, 54, 619–629. [Google Scholar] [CrossRef] [PubMed]
- Samen, F.A.E.; Secor, G.A.; Gudmestad, N.C. Variability in virulence among asexual progenies of Phytophthora infestans. Phytopathology 2003, 93, 293–304. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Levipan, H.A.; Avendaño-Herrera, R. Different phenotypes of mature biofilm in Flavobacterium psychrophilum share a potential for virulence that differs from planktonic state. Front. Cell. Infect. Microbiol. 2017, 7, 76. [Google Scholar] [CrossRef] [PubMed]
- Wakabayashi, H.; Hikida, M.; Masumura, K. Flexibacter maritimus sp. nov., a pathogen of marine fishes. Int. J. Syst. Evol. Microbiol. 1986, 36, 396–398. [Google Scholar] [CrossRef] [Green Version]
- Bauer, M.A.; Kainz, K.; Carmona-Gutierrez, D.; Madeo, F. Microbial wars: Competition in ecological niches and within the microbiome. Microb. Cell 2018, 5, 215. [Google Scholar] [CrossRef]
- Gjessing, M.C.; Thoen, E.; Tengs, T.; Skotheim, S.A.; Dale, O.B. Salmon gill poxvirus, a recently characterized infectious agent of multifactorial gill disease in freshwater- and seawater-reared Atlantic salmon. J. Fish Dis. 2017, 40, 1253–1265. [Google Scholar] [CrossRef] [PubMed]
- Orieux, N.; Bourdineaud, J.P.; Douet, D.G.; Daniel, P.; Le Henaff, M. Quantification of Flavobacterium psychrophilum in rainbow trout, Oncorhynchus mykiss (Walbaum), tissues by qPCR. J. Fish Dis. 2011, 34, 811–821. [Google Scholar] [CrossRef]
- Sou, M.; Soh, M.; Nursyafiq, A.M.; Yan, N.S.; László, O.; Henning, S. Insights into the microbiome of farmed Asian sea bass (Lates calcarifer) with symptoms of tenacibaculosis and description of Tenacibaculum singaporense sp. nov. Antonie van Leeuwenhoek 2020, 113, 737–752. [Google Scholar] [CrossRef]
- Faílde, L.D.; Losada, A.P.; Bermúdez, R.; Santos, Y.; Quiroga, M.I. Tenacibaculum maritimum infection: Pathology and immunohistochemistry in experimentally challenged turbot (Psetta maxima L.). Microb. Pathog. 2013, 65, 82–88. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Pascual, D.; Lunazzi, A.; Magdelenat, G.; Rouy, Z.; Roulet, A.; Lopez-Roques, C.; Duchaud, E. The complete genome sequence of the fish pathogen Tenacibaculum maritimum provides insights into virulence mechanisms. Front. Microbiol. 2017, 8, 1542. [Google Scholar] [CrossRef] [PubMed]
- Fringuelli, E.; Savage, P.D.; Gordon, A.; Baxter, E.J.; Rodger, H.D.; Graham, D.A. Development of a quantitative real-time PCR for the detection of Tenacibaculum maritimum and its application to field samples. J. Fish Dis. 2012, 35, 579–590. [Google Scholar] [CrossRef] [PubMed]
- Nowlan, J. Quantitative PCR detection of Tenacibaculum maritimum and Tenacibaculum dicentrarchi at Netpen Sites in British Columbia (Canada). Masters’ Dissertation, Unviersity of Guelph, Guelph, ON, Canada, 2020. [Google Scholar]
- RStudio Team. RStudio: Integrated Development for R. Version 3.6.1 (2019-07-05) “Action of the Toes”. RStudio, PBC, Boston, MA, USA. Available online: http://www.rstudio.com/ (accessed on 19 March 2021).



| Room | RAS Tank | Specific to | Comparison * | Mean LNOB ± SD | KW α | df | p |
|---|---|---|---|---|---|---|---|
| 1 | 1 & 2 | Tdic | Int. and Ext. tissues | Int: 4.66 ± 0.909 Ext: 5.96 ± 1.69 | 4.5 | 1 | 0.034 |
| 2 | 1–4 | Tmar | Int. and Ext. tissues | Int: 4.51 ± 0.758 Ext: 6.33 ± 1.39 | 72 | 1 | <2.2 × 10−16 |
| 1 | 1 & 2 | Tdic | E and C fish | E: 5.49 ± 1.58 C: 5.80 ± 1.89 | 0.66 | 1 | 0.41 |
| 2 | 1–4 | Tmar | E and C fish | E: 5.71 ± 1.48 C: 5.75 ± 1.52 | 0.021 | 1 | 0.88 |
| Room | ET#:Treatment | Final Concentration | Number Exposed | Duration of Exposure (hr) | Post-exposure RAS Tank (T) Location | Number in Each RAS Tank * |
|---|---|---|---|---|---|---|
| 1 | ET1: T. dicentrarchi TdChD04 | 2.5 × 107 | 40 | 5 | T1 & T2 | 20E & 20C |
| 1 | ET2: T. finnmarkense Tsp.2 | 2.5 × 107 | 40 | 5 | T3 & T4 | 20E & 20C |
| 1 | ET3: Sham-Control (medium only) | 0 | 40 | 5 | T5 | 40E |
| 2 | ET1: T. maritimum NLF-15 | 2.5 × 106 | 41 | 5 | T1 & T2 | 20/21E α & 20C |
| 2 | ET2: T. maritimum NLF-15 | 2.5 × 108 | 40 | 5 | T3 & T4 | 20E & 20C |
| 2 | ET3: Sham-Control (medium only) | 0 | 40 | 5 | T5 | 40E |
| Specific to | Assay Equation | Reference for Assay |
|---|---|---|
| T. maritimum | x = LOG((((10(Mean Cq−34.4)/−3.2))/10)/100)*c*v)*33.3) * | [24] |
| T. dicentrarchi | x = LOG((((10(Mean Cq−36.9)/−3.4))/10)/100)*c*v)*33.3) | [7,25] |
| T. finnmarkense | x = LOG((((10(Mean Cq−38.4)/−3.6))/10)/100)*c*v)*33.3) | [7,25] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nowlan, J.P.; Britney, S.R.; Lumsden, J.S.; Russell, S. Experimental Induction of Tenacibaculosis in Atlantic Salmon (Salmo salar L.) Using Tenacibaculum maritimum, T. dicentrarchi, and T. finnmarkense. Pathogens 2021, 10, 1439. https://doi.org/10.3390/pathogens10111439
Nowlan JP, Britney SR, Lumsden JS, Russell S. Experimental Induction of Tenacibaculosis in Atlantic Salmon (Salmo salar L.) Using Tenacibaculum maritimum, T. dicentrarchi, and T. finnmarkense. Pathogens. 2021; 10(11):1439. https://doi.org/10.3390/pathogens10111439
Chicago/Turabian StyleNowlan, Joseph P., Scott R. Britney, John S. Lumsden, and Spencer Russell. 2021. "Experimental Induction of Tenacibaculosis in Atlantic Salmon (Salmo salar L.) Using Tenacibaculum maritimum, T. dicentrarchi, and T. finnmarkense" Pathogens 10, no. 11: 1439. https://doi.org/10.3390/pathogens10111439
APA StyleNowlan, J. P., Britney, S. R., Lumsden, J. S., & Russell, S. (2021). Experimental Induction of Tenacibaculosis in Atlantic Salmon (Salmo salar L.) Using Tenacibaculum maritimum, T. dicentrarchi, and T. finnmarkense. Pathogens, 10(11), 1439. https://doi.org/10.3390/pathogens10111439






