Mosquito Innate Immunity
Abstract
1. Introduction
2. The Immune System of Mosquitoes
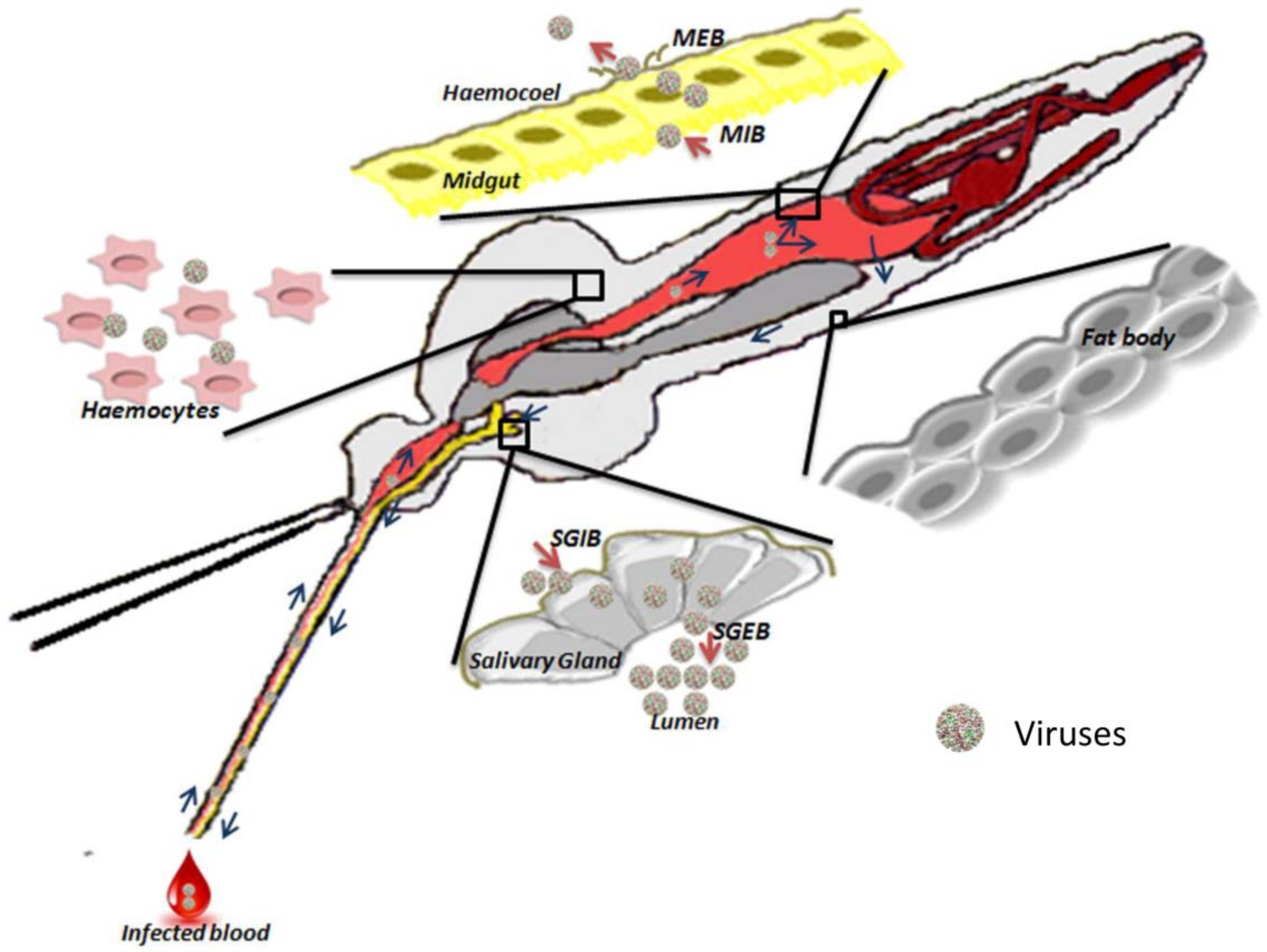
2.1. Physical Barriers
2.1.1. Midgut
2.1.2. Hemocele
2.1.3. Salivary Glands
2.2. Physiological Barriers to Protect Mosquitoes against Pathogens
2.2.1. Midgut-Infection Barrier
2.2.2. Midgut-Escape Barrier
2.2.3. Salivary Gland Infection Barrier and Salivary Gland Escape Barrier
2.3. Molecular Basis of Immunity
2.3.1. Recognition
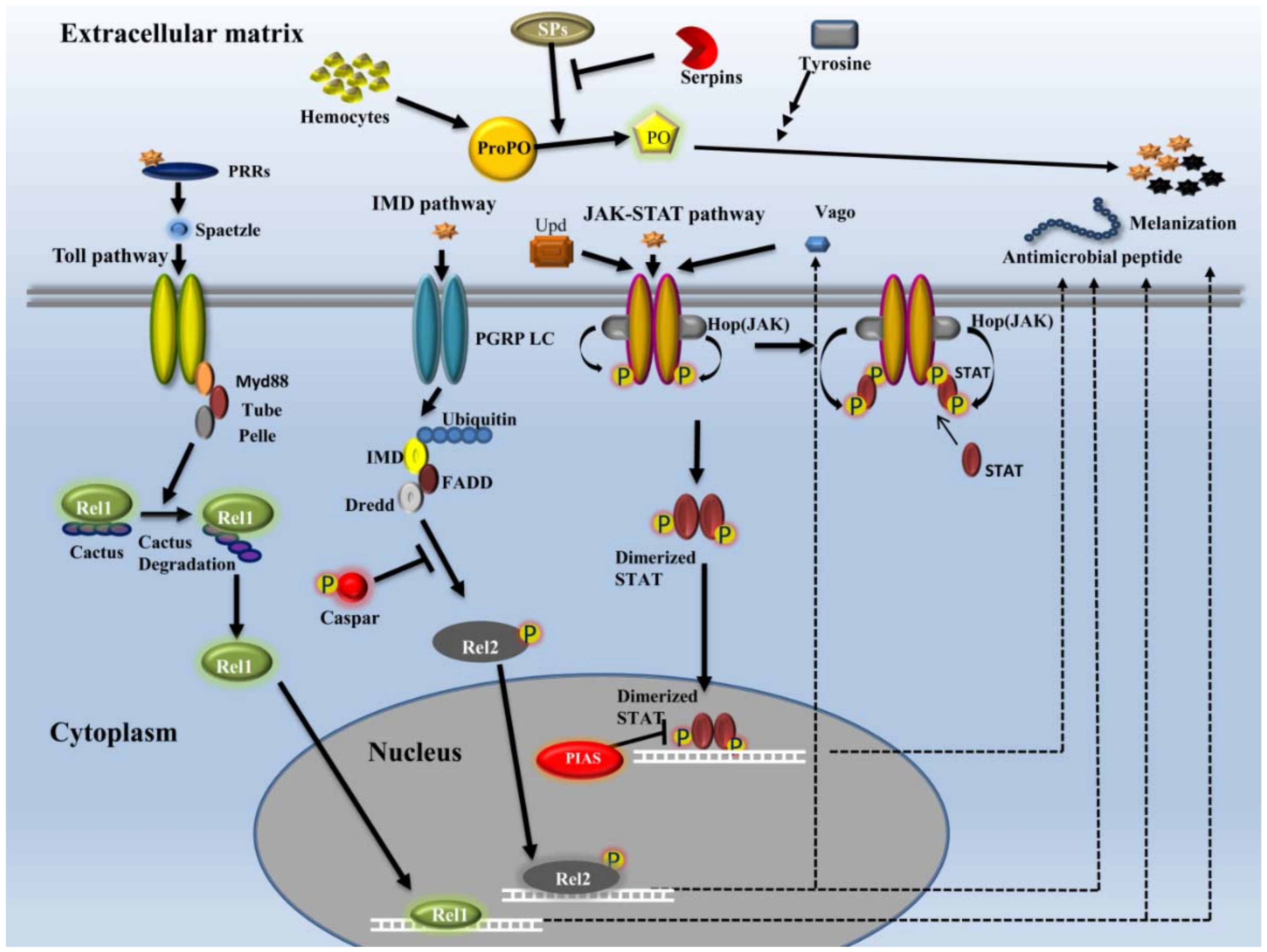
2.3.2. Signaling
- Toll pathway
- IMD pathway
- JAK-STAT pathway
- RNAi
2.3.3. Modulation
2.3.4. Effectors
- AMPs
- Reactive oxygen species/reactive nitrogen species
- Melanization
- Apoptosis
- Autophagy
- Phagocytosis
2.4. RNAi
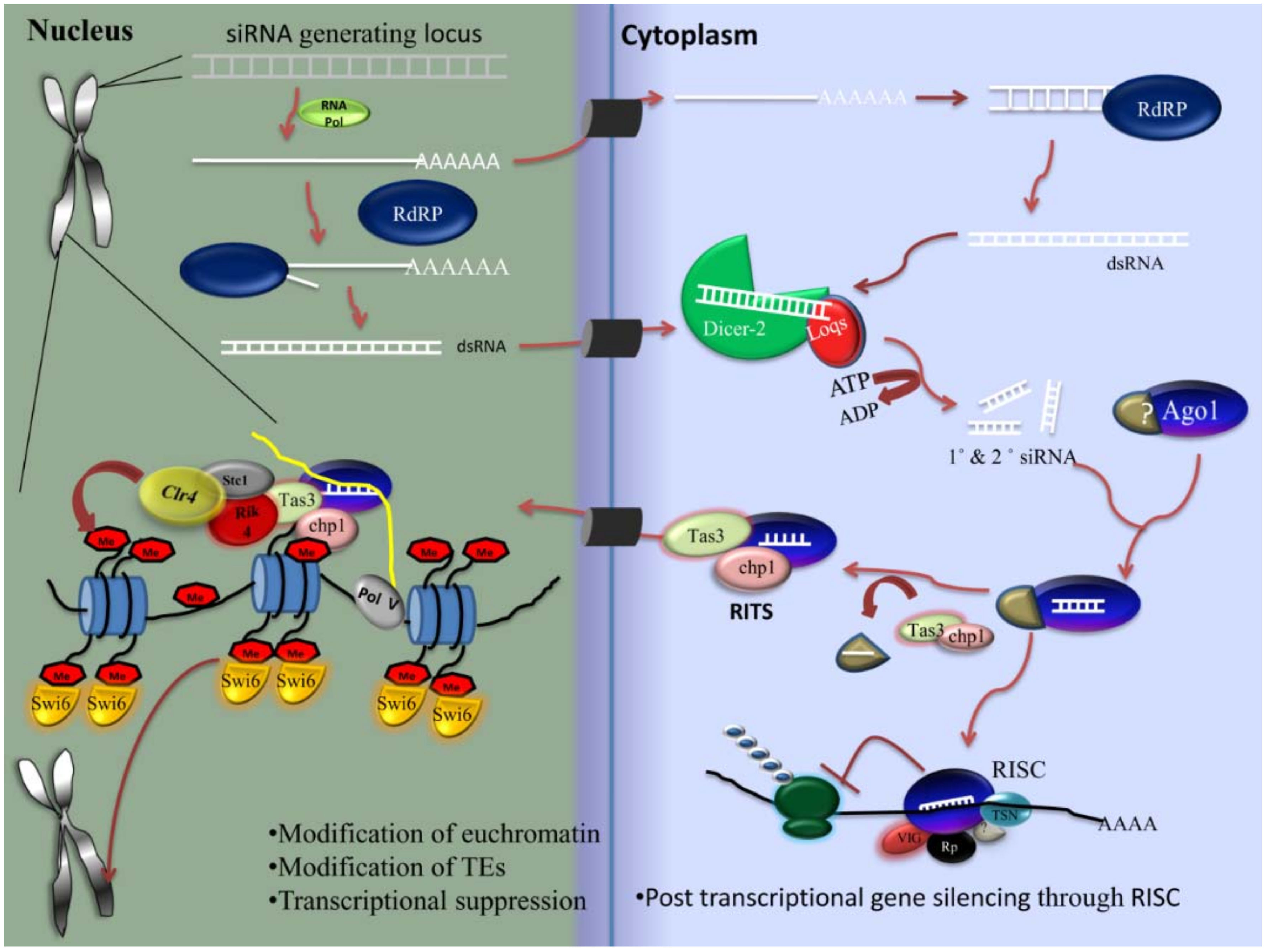
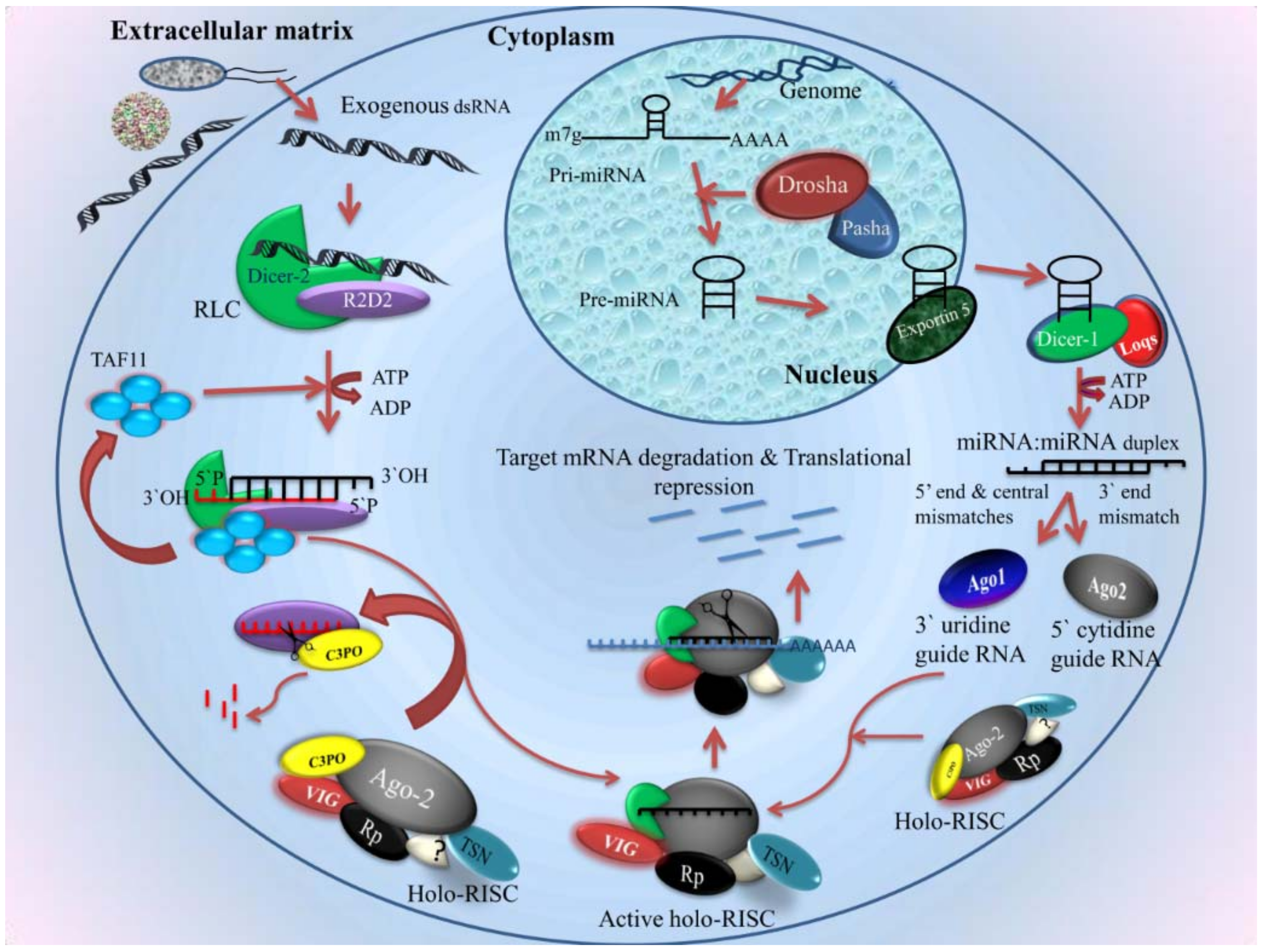
2.4.1. siRNA Pathway
- Biogenesis of siRNA
- Components of the siRNA pathway
- Mechanism
2.4.2. Micro RNA in Mosquito Immunity
2.4.3. PIWI Pathway
- Source of piRNA
- Components
- piRNA characteristics and generation mechanism
- Roles
2.4.4. Viral Counter Defenses to the RNAi Pathway
3. Future Perspectives
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Lai, S.-C.; Chen, C.-C.; Hou, R.F. Electron microscopic observations on wound-healing in larvae of the mosquito Armigeres subalbatus (Diptera: Culicidae). J. Med. Entomol. 2001, 38, 836–843. [Google Scholar] [CrossRef] [PubMed]
- Lai, S.-C.; Chen, C.-C.; Hou, R.F. Immunolocalization of prophenoloxidase in the process of wound healing in the mosquito Armigeres subalbatus (Diptera: Culicidae). J. Med. Entomol. 2002, 39, 266–274. [Google Scholar] [CrossRef] [PubMed]
- Strand, M.R. The insect cellular immune response. Insect Sci. 2008, 15, 1–14. [Google Scholar] [CrossRef]
- Blandin, S.; Shiao, S.-H.; Moita, L.F.; Janse, C.J.; Waters, A.P.; Kafatos, F.C.; Levashina, E.A. Complement-like protein TEP1 is a determinant of vectorial capacity in the malaria vector Anopheles gambiae. Cell 2004, 116, 661–670. [Google Scholar] [CrossRef]
- Das, S.; Dong, Y.; Garver, L.; Dimopoulos, G. Specificity of the Innate Immune System: A Closer Look at the Mosquito Pattern-Recognition Receptor Repertoire; Oxford University Press: New York, NY, USA, 2009. [Google Scholar]
- Levashina, E.A.; Moita, L.F.; Blandin, S.; Vriend, G.; Lagueux, M.; Kafatos, F.C. Conserved role of a complement-like protein in phagocytosis revealed by dsRNA knockout in cultured cells of the mosquito, Anopheles gambiae. Cell 2001, 104, 709–718. [Google Scholar] [CrossRef]
- Zhou, G.; Kohlhepp, P.; Geiser, D.; del Carmen Frasquillo, M.; Vazquez-Moreno, L.; Winzerling, J.J. Fate of blood meal iron in mosquitoes. J. Insect Physiol. 2007, 53, 1169–1178. [Google Scholar] [CrossRef] [PubMed]
- Abraham, E.G.; Jacobs-Lorena, M. Mosquito midgut barriers to malaria parasite development. Insect Biochem. Mol. Biol. 2004, 34, 667–671. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Ceron, L.; Rodriguez, M.H.; Chavez-Munguia, B.; Santillan, F.; Nettel, J.A.; Hernandez-Avila, J.E. Plasmodium vivax: Impaired escape of Vk210 phenotype ookinetes from the midgut blood bolus of Anopheles pseudopunctipennis. Exp. Parasitol. 2007, 115, 59–67. [Google Scholar] [CrossRef] [PubMed]
- Kato, N.; Mueller, C.R.; Fuchs, J.F.; McElroy, K.; Wessely, V.; Higgs, S.; Christensen, B.M. Evaluation of the function of a type I peritrophic matrix as a physical barrier for midgut epithelium invasion by mosquito-borne pathogens in Aedes aegypti. Vector-Borne Zoonotic Dis. 2008, 8, 701–712. [Google Scholar] [CrossRef] [PubMed]
- Dimopoulos, G. Insect immunity and its implication in mosquito–malaria interactions. Cell. Microbiol. 2003, 5, 3–14. [Google Scholar] [CrossRef] [PubMed]
- Salazar, M.I.; Richardson, J.H.; Sánchez-Vargas, I.; Olson, K.E.; Beaty, B.J. Dengue virus type 2: Replication and tropisms in orally infected Aedes aegypti mosquitoes. BMC Microbiol. 2007, 7, 9. [Google Scholar] [CrossRef] [PubMed]
- Devenport, M.; Alvarenga, P.H.; Shao, L.; Fujioka, H.; Bianconi, M.L.; Oliveira, P.L.; Jacobs-Lorena, M. Identification of the Aedes aegypti peritrophic matrix protein AeIMUCI as a heme-binding protein. Biochemistry 2006, 45, 9540–9549. [Google Scholar] [CrossRef] [PubMed]
- Dana, A.N.; Hong, Y.S.; Kern, M.K.; Hillenmeyer, M.E.; Harker, B.W.; Lobo, N.F.; Hogan, J.R.; Romans, P.; Collins, F.H. Gene expression patterns associated with blood-feeding in the malaria mosquito Anopheles gambiae. BMC Genom. 2005, 6, 5. [Google Scholar] [CrossRef] [PubMed]
- Jochim, R.C.; Teixeira, C.R.; Laughinghouse, A.; Mu, J.; Oliveira, F.; Gomes, R.B.; Elnaiem, D.-E.; Valenzuela, J.G. The midgut transcriptome of Lutzomyia longipalpis: Comparative analysis of cDNA libraries from sugar-fed, blood-fed, post-digested and Leishmania infantum chagasi-infected sand flies. BMC Genom. 2008, 9, 15. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, J.M.; Genta, F.A.; Sorgine, M.H.; Logullo, R.; Mesquita, R.D.; Paiva-Silva, G.O.; Majerowicz, D.; Medeiros, M.; Koerich, L.; Terra, W.R. An insight into the transcriptome of the digestive tract of the bloodsucking bug, Rhodnius prolixus. PLoS Negl. Trop. Dis. 2014, 8, e2594. [Google Scholar] [CrossRef] [PubMed]
- Anderson, J.M.; Sonenshine, D.E.; Valenzuela, J.G. Exploring the mialome of ticks: An annotated catalogue of midgut transcripts from the hard tick, Dermacentor variabilis (Acari: Ixodidae). BMC Genom. 2008, 9, 552. [Google Scholar] [CrossRef] [PubMed]
- Bonizzoni, M.; Dunn, W.A.; Campbell, C.L.; Olson, K.E.; Dimon, M.T.; Marinotti, O.; James, A.A. RNA-seq analyses of blood-induced changes in gene expression in the mosquito vector species, Aedes aegypti. BMC Genom. 2011, 12, 82. [Google Scholar] [CrossRef] [PubMed]
- Marinotti, O.; Nguyen, Q.K.; Calvo, E.; James, A.A.; Ribeiro, J. Microarray analysis of genes showing variable expression following a blood meal in Anopheles gambiae. Insect Mol. Biol. 2005, 14, 365–373. [Google Scholar] [CrossRef] [PubMed]
- Perner, J.; Provazník, J.; Schrenková, J.; Urbanová, V.; Ribeiro, J.M.; Kopáček, P. RNA-seq analyses of the midgut from blood-and serum-fed Ixodes ricinus ticks. Sci. Rep. 2016, 6, 36695. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Zhang, R.; Zhang, B.; Zhao, T.; Wang, P.; Liang, G.; Cheng, G. Blood meal acquisition enhances arbovirus replication in mosquitoes through activation of the GABAergic system. Nat. Commun. 2017, 8, 1262. [Google Scholar] [CrossRef] [PubMed]
- Smartt, C.T.; Shin, D.; Anderson, S.L. The Effect of West Nile Virus Infection on the Midgut Gene Expression of Culex pipiens quinquefasciatus Say (Diptera: Culicidae). Insects 2016, 7, 76. [Google Scholar] [CrossRef] [PubMed]
- Fukutani, K.F.; Kasprzykowski, J.I.; Paschoal, A.R.; Gomes, M.d.S.; Barral, A.; De Oliveira, C.I.; Ramos, P.I.P.; Queiroz, A.T.L.D. Meta-analysis of Aedes aegypti expression datasets: Comparing virus infection and blood fed transcriptomes to identify markers of virus presence. Front. Bioeng. Biotechnol. 2017, 5, 84. [Google Scholar] [CrossRef] [PubMed]
- Barletta, A.B.F.; Alves, L.R.; Silva, M.C.L.N.; Sim, S.; Dimopoulos, G.; Liechocki, S.; Maya-Monteiro, C.M.; Sorgine, M.H.F. Emerging role of lipid droplets in Aedes aegypti immune response against bacteria and Dengue virus. Sci. Rep. 2016, 6, 19928. [Google Scholar] [CrossRef] [PubMed]
- Peterson, T.M.; Gow, A.J.; Luckhart, S. Nitric oxide metabolites induced in Anopheles stephensi control malaria parasite infection. Free Radic. Biol. Med. 2007, 42, 132–142. [Google Scholar] [CrossRef] [PubMed]
- Houk, E.; Hardy, J.; Chiles, R. Permeability of the midgut basal lamina in the mosquito, Culex tarsalis Coquillett (Insecta, Diptera). Acta Trop. 1981, 38, 163–171. [Google Scholar] [PubMed]
- Kantor, A.; Dong, S.; Held, N.; Ishimwe, E.; Passarelli, A.; Clem, R.; Franz, A. Identification and initial characterization of matrix metalloproteinases in the yellow fever mosquito, Aedes aegypti. Insect Mol. Biol. 2017, 26, 113–126. [Google Scholar] [CrossRef] [PubMed]
- Passarelli, A.L. Barriers to success: How baculoviruses establish efficient systemic infections. Virology 2011, 411, 383–392. [Google Scholar] [CrossRef] [PubMed]
- Hecker, H. Structure and function of midgut epithelial cells in Culicidae mosquitoes (Insecta, Diptera). Cell Tissue Res. 1977, 184, 321–341. [Google Scholar] [CrossRef] [PubMed]
- Houk, E.J. Midgut ultrastructure of Culex tarsalis (Diptera: Culcidae) before and after a bloodmeal. Tissue Cell 1977, 9, 103–118. [Google Scholar] [CrossRef]
- Okuda, K.; de Souza Caroci, A.; Ribolla, P.; De Bianchi, A.; Bijovsky, A. Functional morphology of adult female Culex quinquefasciatus midgut during blood digestion. Tissue Cell 2002, 34, 210–219. [Google Scholar] [CrossRef]
- Romoser, W.S.; Wasieloski, L.P., Jr.; Pushko, P.; Kondig, J.P.; Lerdthusnee, K.; Neira, M.; Ludwig, G.V. Evidence for arbovirus dissemination conduits from the mosquito (Diptera: Culicidae) midgut. J. Med. Entomol. 2004, 41, 467–475. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Manfredini, F.; Dimopoulos, G. Implication of the mosquito midgut microbiota in the defense against malaria parasites. PLoS Pathog. 2009, 5, e1000423. [Google Scholar] [CrossRef] [PubMed]
- Klowden, M.J. Making generalizations about vectors: Is there a physiology of “the mosquito”? Entomol. Res. 2007, 37, 1–13. [Google Scholar] [CrossRef]
- King, J.G.; Hillyer, J.F. Infection-induced interaction between the mosquito circulatory and immune systems. PLoS Pathog. 2012, 8, e1003058. [Google Scholar] [CrossRef] [PubMed]
- Bain, O.; Babayan, S. Behaviour of filariae: Morphological and anatomical signatures of their life style within the arthropod and vertebrate hosts. Filaria J. 2003, 2, 16. [Google Scholar] [CrossRef] [PubMed]
- Glenn, J.D.; King, J.G.; Hillyer, J.F. Structural mechanics of the mosquito heart and its function in bidirectional hemolymph transport. J. Exp. Biol. 2010, 213, 541–550. [Google Scholar] [CrossRef] [PubMed]
- Morahan, B.J.; Wang, L.; Coppel, R.L. No TRAP, no invasion. Trends Parasitol. 2009, 25, 77–84. [Google Scholar] [CrossRef] [PubMed]
- Bartholomay, L.; Mayhew, G.; Fuchs, J.; Rocheleau, T.; Erickson, S.; Aliota, M.; Christensen, B. Profiling infection responses in the haemocytes of the mosquito, Aedes aegypti. Insect Mol. Biol. 2007, 16, 761–776. [Google Scholar] [CrossRef] [PubMed]
- Castillo, J.; Robertson, A.; Strand, M. Characterization of hemocytes from the mosquitoes Anopheles gambiae and Aedes aegypti. Insect Biochem. Mol. Biol. 2006, 36, 891–903. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Martínez, S.; Lanz, H.; Rodrguez, M.H.; González-Ceron, L.; Tsutsumi, V. Cellular-mediated reactions to foreign organisms inoculated into the hemocoel of Anopheles albimanus (Diptera: Culicidae). J. Med. Entomol. 2002, 39, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Hillyer, J.F.; Schmidt, S.L.; Christensen, B.M. Rapid phagocytosis and melanization of bacteria and Plasmodium sporozoites by hemocytes of the mosquito Aedes aegypti. J. Parasitol. 2003, 89, 62–69. [Google Scholar] [CrossRef]
- Pinto, S.B.; Lombardo, F.; Koutsos, A.C.; Waterhouse, R.M.; McKay, K.; An, C.; Ramakrishnan, C.; Kafatos, F.C.; Michel, K. Discovery of Plasmodium modulators by genome-wide analysis of circulating hemocytes in Anopheles gambiae. Proc. Natl. Acad. Sci. USA 2009, 106, 21270–21275. [Google Scholar] [CrossRef] [PubMed]
- Moreira-Ferro, C.; Marinotti, O.; Bijovsky, A. Morphological and biochemical analyses of the salivary glands of the malaria vector, Anopheles darlingi. Tissue Cell 1999, 31, 264–273. [Google Scholar] [CrossRef] [PubMed]
- Orr, C.; Hudson, A.; West, A. The salivary glands of Aedes aegypti histological–histochemical studies. Can. J. Zool. 1961, 39, 265–272. [Google Scholar] [CrossRef]
- Marinotti, O.; James, A.A. An α-glucosidase in the salivary glands of the vector mosquito, Aedes aegypti. Insect Biochem. 1990, 20, 619–623. [Google Scholar] [CrossRef]
- Cao-Lormeau, V.-M. Dengue viruses binding proteins from Aedes aegypti and Aedes polynesiensis salivary glands. Virol. J. 2009, 6, 35. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, A.K.; Devenport, M.; Jethwaney, D.; Kalume, D.E.; Pandey, A.; Anderson, V.E.; Sultan, A.A.; Kumar, N.; Jacobs-Lorena, M. Malaria parasite invasion of the mosquito salivary gland requires interaction between the Plasmodium TRAP and the Anopheles saglin proteins. PLoS Pathog. 2009, 5, e1000265. [Google Scholar] [CrossRef] [PubMed]
- Korochkina, S.; Barreau, C.; Pradel, G.; Jeffery, E.; Li, J.; Natarajan, R.; Shabanowitz, J.; Hunt, D.; Frevert, U.; Vernick, K.D. A mosquito-specific protein family includes candidate receptors for malaria sporozoite invasion of salivary glands. Cell. Microbiol. 2006, 8, 163–175. [Google Scholar] [CrossRef] [PubMed]
- Phattanawiboon, B.; Jariyapan, N.; Roytrakul, S.; Paemanee, A.; Sor-Suwan, S.; Intakhan, N.; Chanmol, W.; Siriyasatien, P.; Saeung, A.; Choochote, W. Morphological and protein analyses of adult female salivary glands of Anopheles barbirostris species A1 (Diptera: Culicidae). Trop. Biomed. 2014, 31, 813–827. [Google Scholar] [PubMed]
- Clements, A. The adult salivary glands and their secretions. In The Biology of Mosquitoes; Chapman & Hall: London, UK, 1992; pp. 251–262. [Google Scholar]
- Bowers, D.F.; Abell, B.A.; Brown, D.T. Replication and tissue tropism of the alphavirus Sindbis in the mosquito Aedes albopictus. Virology 1995, 212, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Ciano, K.A.; Saredy, J.J.; Bowers, D.F. Heparan sulfate proteoglycan: An arbovirus attachment factor integral to mosquito salivary gland ducts. Viruses 2014, 6, 5182–5197. [Google Scholar] [CrossRef] [PubMed]
- Gaidamovich, S.Y.; Khutoretskaya, N.V.; Lvova, A.I.; Sveshnikova, N.A. Immunofluorescent staining study of the salivary glands of mosquitoes infected with group A arboviruses. Intervirology 1973, 1, 193–200. [Google Scholar] [CrossRef] [PubMed]
- Janzen, H.; Rhodes, A.; Doane, F. Chikungunya virus in salivary glands of Aedes aegypti (L.): An electron microscope study. Can. J. Microbiol. 1970, 16, 581–586. [Google Scholar] [CrossRef] [PubMed]
- Raquin, V.; Wannagat, M.; Zouache, K.; Legras-Lachuer, C.; Moro, C.V.; Mavingui, P. Detection of dengue group viruses by fluorescence in situ hybridization. Parasites Vectors 2012, 5, 243. [Google Scholar] [CrossRef] [PubMed]
- Tchankouo-Nguetcheu, S.; Bourguet, E.; Lenormand, P.; Rousselle, J.-C.; Namane, A.; Choumet, V. Infection by chikungunya virus modulates the expression of several proteins in Aedes aegypti salivary glands. Parasites Vectors 2012, 5, 264. [Google Scholar] [CrossRef] [PubMed]
- Girard, Y.A.; Schneider, B.S.; McGee, C.E.; Wen, J.; Han, V.C.; Popov, V.; Mason, P.W.; Higgs, S. Salivary gland morphology and virus transmission during long-term cytopathologic West Nile virus infection in Culex mosquitoes. Am. J. Trop. Med. Hyg. 2007, 76, 118–128. [Google Scholar] [PubMed]
- Mackenzie, J.M.; Jones, M.K.; Young, P.R. Immunolocalization of the dengue virus nonstructural glycoprotein NS1 suggests a role in viral RNA replication. Virology 1996, 220, 232–240. [Google Scholar] [CrossRef] [PubMed]
- Mackenzie, J.M.; Khromykh, A.A.; Jones, M.K.; Westaway, E.G. Subcellular localization and some biochemical properties of the flavivirus Kunjin nonstructural proteins NS2A and NS4A. Virology 1998, 245, 203–215. [Google Scholar] [CrossRef] [PubMed]
- Westaway, E.G.; Mackenzie, J.M.; Kenney, M.T.; Jones, M.K.; Khromykh, A.A. Ultrastructure of Kunjin virus-infected cells: Colocalization of NS1 and NS3 with double-stranded RNA, and of NS2B with NS3, in virus-induced membrane structures. J. Virol. 1997, 71, 6650–6661. [Google Scholar] [PubMed]
- Schneider, B.S.; Higgs, S. The enhancement of arbovirus transmission and disease by mosquito saliva is associated with modulation of the host immune response. Trans. R. Soc. Trop. Med. Hyg. 2008, 102, 400–408. [Google Scholar] [CrossRef] [PubMed]
- Thangamani, S.; Wikel, S.K. Differential expression of Aedes aegypti salivary transcriptome upon blood feeding. Parasites Vectors 2009, 2, 34. [Google Scholar] [CrossRef] [PubMed]
- Houk, E.; Obie, F.; Hardy, J. Peritrophic Membrane Formation and the Midgut Barrier to Arboviral Infection in the Mosquito, Culex tarsalis Coquillett (Insecta, Diptera); California University Berkeley School of Public Health: Berkeley, CA, USA, 1979. [Google Scholar]
- Hardy, J.L.; Houk, E.J.; Kramer, L.D.; Reeves, W.C. Intrinsic factors affecting vector competence of mosquitoes for arboviruses. Annu. Rev. Entomol. 1983, 28, 229–262. [Google Scholar] [CrossRef] [PubMed]
- Weaver, S.; Scott, T.; Lorenz, L.; Lerdthusnee, K.; Romoser, W. Togavirus-associated pathologic changes in the midgut of a natural mosquito vector. J. Virol. 1988, 62, 2083–2090. [Google Scholar] [PubMed]
- Kramer, L.D.; Hardy, J.L.; Presser, S.B.; Houk, E.J. Dissemination barriers for western equine encephalomyelitis virus in Culex tarsalis infected after ingestion of low viral doses. Am. J. Trop. Med. Hyg. 1981, 30, 190–197. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, J.; Oliveira, G.A.; Kotsyfakis, M.; Dixit, R.; Molina-Cruz, A.; Jochim, R.; Barillas-Mury, C. An epithelial serine protease, AgESP, is required for Plasmodium invasion in the mosquito Anopheles gambiae. PLoS ONE 2012, 7, e35210. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, J.; Brayner, F.A.; Alves, L.C.; Dixit, R.; Barillas-Mury, C. Hemocyte differentiation mediates innate immune memory in Anopheles gambiae mosquitoes. Science 2010, 329, 1353–1355. [Google Scholar] [CrossRef] [PubMed]
- Paulson, S.L.; Grimstad, P.R.; Craig, G.B. Midgut and salivary gland barriers to La Crosse virus dissemination in mosquitoes of the Aedes triseriatus group. Med. Vet. Entomol. 1989, 3, 113–123. [Google Scholar] [CrossRef] [PubMed]
- Scott, T.; Lorenz, L.; Weaver, S. Susceptibility of Aedes albopictus to infection with eastern equine encephalomyelitis virus. J. Am. Mosq. Control Assoc. 1990, 6, 274–278. [Google Scholar] [PubMed]
- Turell, M.; Mores, C.; Dohm, D.; Komilov, N.; Paragas, J.; Lee, J.; Shermuhemedova, D.; Endy, T.; Kodirov, A.; Khodjaev, S. Laboratory transmission of Japanese encephalitis and West Nile viruses by molestus form of Culex pipiens (Diptera: Culicidae) collected in Uzbekistan in 2004. J. Med. Entomol. 2006, 43, 296–300. [Google Scholar] [CrossRef] [PubMed]
- Romoser, W.; Turell, M.; Lerdthusnee, K.; Neira, M.; Dohm, D.; Ludwig, G.; Wasieloski, L. Pathogenesis of Rift Valley fever virus in mosquitoes—Tracheal conduits & the basal lamina as an extra-cellular barrier. In Infectious Diseases from Nature: Mechanisms of Viral Emergence and Persistence; Springer: Vienna, Austria, 2005; pp. 89–100. [Google Scholar]
- Dourado, L.; Ribeiro, L.; Brancalhao, R.; Tavares, J.; Borges, A.; Fernandez, M. Silkworm salivary glands are not susceptible to Bombyx mori nuclear polyhedrosis virus. Genet. Mol. Res. 2011, 10, 335–339. [Google Scholar] [CrossRef] [PubMed]
- Beaty, B.; Holterman, M.; Tabachnick, W.; Shope, R.; Rozhon, E.; Bishop, D. Molecular basis of bunyavirus transmission by mosquitoes: Role of the middle-sized RNA segment. Science 1981, 211, 1433–1435. [Google Scholar] [CrossRef] [PubMed]
- Grimstad, P.R.; Paulson, S.L.; Craig, G.B., Jr. Vector competence of Aedes hendersoni (Diptera: Culicidae) for La Crosse virus and evidence of a salivary-gland escape barrier. J. Med. Entomol. 1985, 22, 447–453. [Google Scholar] [CrossRef] [PubMed]
- Jupp, P. Culex theileri and Sindbis virus; salivary glands infection in relation to transmission. J. Am. Mosq. Control Assoc. 1985, 1, 374–376. [Google Scholar] [PubMed]
- Turell, M.J.; Britch, S.C.; Aldridge, R.L.; Kline, D.L.; Boohene, C.; Linthicum, K.J. Potential for mosquitoes (Diptera: Culicidae) from Florida to transmit Rift Valley fever virus. J. Med. Entomol. 2013, 50, 1111–1117. [Google Scholar] [CrossRef] [PubMed]
- Buchon, N.; Poidevin, M.; Kwon, H.-M.; Guillou, A.; Sottas, V.; Lee, B.-L.; Lemaitre, B. A single modular serine protease integrates signals from pattern-recognition receptors upstream of the Drosophila Toll pathway. Proc. Natl. Acad. Sci. USA 2009, 106, 12442–12447. [Google Scholar] [CrossRef] [PubMed]
- Cheng, G.; Liu, L.; Wang, P.; Zhang, Y.; Zhao, Y.O.; Colpitts, T.M.; Feitosa, F.; Anderson, J.F.; Fikrig, E. An in vivo transfection approach elucidates a role for Aedes aegypti thioester-containing proteins in flaviviral infection. PLoS ONE 2011, 6, e22786. [Google Scholar] [CrossRef] [PubMed]
- Lagueux, M.; Perrodou, E.; Levashina, E.A.; Capovilla, M.; Hoffmann, J.A. Constitutive expression of a complement-like protein in toll and JAK gain-of-function mutants of Drosophila. Proc. Natl. Acad. Sci. USA 2000, 97, 11427–11432. [Google Scholar] [CrossRef] [PubMed]
- Fraiture, M.; Baxter, R.H.; Steinert, S.; Chelliah, Y.; Frolet, C.; Quispe-Tintaya, W.; Hoffmann, J.A.; Blandin, S.A.; Levashina, E.A. Two mosquito LRR proteins function as complement control factors in the TEP1-mediated killing of Plasmodium. Cell Host Microbe 2009, 5, 273–284. [Google Scholar] [CrossRef] [PubMed]
- Povelones, M.; Waterhouse, R.M.; Kafatos, F.C.; Christophides, G.K. Leucine-rich repeat protein complex activates mosquito complement in defense against Plasmodium parasites. Science 2009, 324, 258–261. [Google Scholar] [CrossRef] [PubMed]
- Blandin, S.A.; Wang-Sattler, R.; Lamacchia, M.; Gagneur, J.; Lycett, G.; Ning, Y.; Levashina, E.A.; Steinmetz, L.M. Dissecting the genetic basis of resistance to malaria parasites in Anopheles gambiae. Science 2009, 326, 147–150. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, R.M.; Kriventseva, E.V.; Meister, S.; Xi, Z.; Alvarez, K.S.; Bartholomay, L.C.; Barillas-Mury, C.; Bian, G.; Blandin, S.; Christensen, B.M. Evolutionary dynamics of immune-related genes and pathways in disease-vector mosquitoes. Science 2007, 316, 1738–1743. [Google Scholar] [CrossRef] [PubMed]
- Ferguson, H.M.; Read, A.F. Why is the effect of malaria parasites on mosquito survival still unresolved? Trends Parasitol. 2002, 18, 256–261. [Google Scholar] [CrossRef]
- Baker, D.A.; Nolan, T.; Fischer, B.; Pinder, A.; Crisanti, A.; Russell, S. A comprehensive gene expression atlas of sex-and tissue-specificity in the malaria vector, Anopheles gambiae. BMC Genom. 2011, 12, 296. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Taylor, H.E.; Dimopoulos, G. AgDscam, a hypervariable immunoglobulin domain-containing receptor of the Anopheles gambiae innate immune system. PLoS Biol. 2006, 4, e229. [Google Scholar] [CrossRef] [PubMed]
- Osta, M.A.; Christophides, G.K.; Kafatos, F.C. Effects of mosquito genes on Plasmodium development. Science 2004, 303, 2030–2032. [Google Scholar] [CrossRef] [PubMed]
- Schnitger, A.K.; Yassine, H.; Kafatos, F.C.; Osta, M.A. Two C-type lectins cooperate to defend Anopheles gambiae against Gram-negative bacteria. J. Biol. Chem. 2009, 284, 17616–17624. [Google Scholar] [CrossRef] [PubMed]
- Dimopoulos, G.; Richman, A.; Müller, H.-M.; Kafatos, F.C. Molecular immune responses of the mosquito Anopheles gambiae to bacteria and malaria parasites. Proc. Natl. Acad. Sci. USA 1997, 94, 11508–11513. [Google Scholar] [CrossRef] [PubMed]
- Warr, E.; Das, S.; Dong, Y.; Dimopoulos, G. The Gram-Negative Bacteria-Binding Protein gene family: Its role in the innate immune system of Anopheles gambiae and in anti-Plasmodium defence. Insect Mol. Biol. 2008, 17, 39–51. [Google Scholar] [CrossRef] [PubMed]
- Ramirez, J.L.; Souza-Neto, J.; Cosme, R.T.; Rovira, J.; Ortiz, A.; Pascale, J.M.; Dimopoulos, G. Reciprocal tripartite interactions between the Aedes aegypti midgut microbiota, innate immune system and dengue virus influences vector competence. PLoS Negl. Trop. Dis. 2012, 6, e1561. [Google Scholar] [CrossRef] [PubMed]
- Xi, Z.; Ramirez, J.L.; Dimopoulos, G. The Aedes aegypti toll pathway controls dengue virus infection. PLoS Pathog. 2008, 4, e1000098. [Google Scholar] [CrossRef] [PubMed]
- Nüsslein-Volhard, C.; Wieschaus, E. Mutations affecting segment number and polarity in Drosophila. Nature 1980, 287, 795–801. [Google Scholar] [CrossRef] [PubMed]
- Michel, T.; Reichhart, J.-M.; Hoffmann, J.A.; Royet, J. Drosophila Toll is activated by Gram-positive bacteria through a circulating peptidoglycan recognition protein. Nature 2001, 414, 756–759. [Google Scholar] [CrossRef] [PubMed]
- Fehlbaum, P.; Bulet, P.; Michaut, L.; Lagueux, M.; Broekaert, W.F.; Hetru, C.; Hoffmann, J.A. Insect immunity. Septic injury of Drosophila induces the synthesis of a potent antifungal peptide with sequence homology to plant antifungal peptides. J. Biol. Chem. 1994, 269, 33159–33163. [Google Scholar] [PubMed]
- Lemaitre, B.; Nicolas, E.; Michaut, L.; Reichhart, J.-M.; Hoffmann, J.A. The dorsoventral regulatory gene cassette spätzle/Toll/cactus controls the potent antifungal response in Drosophila adults. Cell 1996, 86, 973–983. [Google Scholar] [CrossRef]
- Morisato, D.; Anderson, K.V. The spätzle gene encodes a component of the extracellular signaling pathway establishing the dorsal-ventral pattern of the Drosophila embryo. Cell 1994, 76, 677–688. [Google Scholar] [CrossRef]
- Schneider, D.S.; Jin, Y.; Morisato, D.; Anderson, K.V. A processed form of the Spatzle protein defines dorsal-ventral polarity in the Drosophila embryo. Development 1994, 120, 1243–1250. [Google Scholar] [PubMed]
- Shin, S.W.; Kokoza, V.; Bian, G.; Cheon, H.-M.; Kim, Y.J.; Raikhel, A.S. REL1, a homologue of Drosophila dorsal, regulates toll antifungal immune pathway in the female mosquito Aedes aegypti. J. Biol. Chem. 2005, 280, 16499–16507. [Google Scholar] [CrossRef] [PubMed]
- Bian, G.; Shin, S.W.; Cheon, H.-M.; Kokoza, V.; Raikhel, A.S. Transgenic alteration of Toll immune pathway in the female mosquito Aedes aegypti. Proc. Natl. Acad. Sci. USA 2005, 102, 13568–13573. [Google Scholar] [CrossRef] [PubMed]
- Weber, A.N.; Tauszig-Delamasure, S.; Hoffmann, J.A.; Lelièvre, E.; Gascan, H.; Ray, K.P.; Morse, M.A.; Imler, J.-L.; Gay, N.J. Binding of the Drosophila cytokine Spätzle to Toll is direct and establishes signaling. Nat. Immunol. 2003, 4, 794. [Google Scholar] [CrossRef] [PubMed]
- Parker, J.S.; Mizuguchi, K.; Gay, N.J. A family of proteins related to Spätzle, the toll receptor ligand, are encoded in the Drosophila genome. Proteins Struct. Funct. Bioinform. 2001, 45, 71–80. [Google Scholar] [CrossRef] [PubMed]
- De Gregorio, E.; Han, S.-J.; Lee, W.-J.; Baek, M.-J.; Osaki, T.; Kawabata, S.-I.; Lee, B.-L.; Iwanaga, S.; Lemaitre, B.; Brey, P.T. An immune-responsive Serpin regulates the melanization cascade in Drosophila. Dev. Cell 2002, 3, 581–592. [Google Scholar] [CrossRef]
- Frolet, C.; Thoma, M.; Blandin, S.; Hoffmann, J.A.; Levashina, E.A. Boosting NF-κB-dependent basal immunity of Anopheles gambiae aborts development of Plasmodium berghei. Immunity 2006, 25, 677–685. [Google Scholar] [CrossRef] [PubMed]
- Barletta, A.B.F.; Nascimento-Silva, M.C.L.; Talyuli, O.A.; Oliveira, J.H.M.; Pereira, L.O.R.; Oliveira, P.L.; Sorgine, M.H.F. Microbiota activates IMD pathway and limits Sindbis infection in Aedes aegypti. Parasites Vectors 2017, 10, 103. [Google Scholar] [CrossRef] [PubMed]
- Garver, L.S.; Bahia, A.C.; Das, S.; Souza-Neto, J.A.; Shiao, J.; Dong, Y.; Dimopoulos, G. Anopheles Imd pathway factors and effectors in infection intensity-dependent anti-Plasmodium action. PLoS Pathog. 2012, 8, e1002737. [Google Scholar] [CrossRef] [PubMed]
- Meister, S.; Kanzok, S.M.; Zheng, X.-l.; Luna, C.; Li, T.-R.; Hoa, N.T.; Clayton, J.R.; White, K.P.; Kafatos, F.C.; Christophides, G.K. Immune signaling pathways regulating bacterial and malaria parasite infection of the mosquito Anopheles gambiae. Proc. Natl. Acad. Sci. USA 2005, 102, 11420–11425. [Google Scholar] [CrossRef] [PubMed]
- Garver, L.S.; Dong, Y.; Dimopoulos, G. Caspar controls resistance to Plasmodium falciparum in diverse anopheline species. PLoS Pathog. 2009, 5, e1000335. [Google Scholar] [CrossRef] [PubMed]
- Meister, S.; Agianian, B.; Turlure, F.; Relógio, A.; Morlais, I.; Kafatos, F.C.; Christophides, G.K. Anopheles gambiae PGRPLC-mediated defense against bacteria modulates infections with malaria parasites. PLoS Pathog. 2009, 5, e1000542. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Das, S.; Cirimotich, C.; Souza-Neto, J.A.; McLean, K.J.; Dimopoulos, G. Engineered anopheles immunity to Plasmodium infection. PLoS Pathog. 2011, 7, e1002458. [Google Scholar] [CrossRef] [PubMed]
- Mitri, C.; Jacques, J.-C.; Thiery, I.; Riehle, M.M.; Xu, J.; Bischoff, E.; Morlais, I.; Nsango, S.E.; Vernick, K.D.; Bourgouin, C. Fine pathogen discrimination within the APL1 gene family protects Anopheles gambiae against human and rodent malaria species. PLoS Pathog. 2009, 5, e1000576. [Google Scholar] [CrossRef] [PubMed]
- Engel, P.; Moran, N.A. The gut microbiota of insects–diversity in structure and function. FEMS Microbiol. Rev. 2013, 37, 699–735. [Google Scholar] [CrossRef] [PubMed]
- Dostert, C.; Jouanguy, E.; Irving, P.; Troxler, L.; Galiana-Arnoux, D.; Hetru, C.; Hoffmann, J.A.; Imler, J.-L. The Jak-STAT signaling pathway is required but not sufficient for the antiviral response of drosophila. Nat. Immunol. 2005, 6, 946–953. [Google Scholar] [CrossRef] [PubMed]
- Agaisse, H.; Perrimon, N. The roles of JAK/STAT signaling in Drosophila immune responses. Immunol. Rev. 2004, 198, 72–82. [Google Scholar] [CrossRef] [PubMed]
- Gupta, L.; Molina-Cruz, A.; Kumar, S.; Rodrigues, J.; Dixit, R.; Zamora, R.E.; Barillas-Mury, C. The STAT pathway mediates late-phase immunity against Plasmodium in the mosquito Anopheles gambiae. Cell Host Microbe 2009, 5, 498–507. [Google Scholar] [CrossRef] [PubMed]
- Jupatanakul, N.; Sim, S.; Angleró-Rodríguez, Y.I.; Souza-Neto, J.; Das, S.; Poti, K.E.; Rossi, S.L.; Bergren, N.; Vasilakis, N.; Dimopoulos, G. Engineered Aedes aegypti JAK/STAT pathway-mediated immunity to dengue virus. PLoS Negl. Trop. Dis. 2017, 11, e0005187. [Google Scholar] [CrossRef] [PubMed]
- Souza-Neto, J.A.; Sim, S.; Dimopoulos, G. An evolutionary conserved function of the JAK-STAT pathway in anti-dengue defense. Proc. Natl. Acad. Sci. USA 2009, 106, 17841–17846. [Google Scholar] [CrossRef] [PubMed]
- Rawlings, N.D.; Barrett, A.J. [2] Families of serine peptidases. Methods Enzymol. 1994, 244, 19–61. [Google Scholar] [PubMed]
- Cerenius, L.; Söderhäll, K. The prophenoloxidase-activating system in invertebrates. Immunol. Rev. 2004, 198, 116–126. [Google Scholar] [CrossRef] [PubMed]
- Ligoxygakis, P.; Pelte, N.; Hoffmann, J.A.; Reichhart, J.-M. Activation of Drosophila Toll during fungal infection by a blood serine protease. Science 2002, 297, 114–116. [Google Scholar] [CrossRef] [PubMed]
- Dimopoulos, G.; Richman, A.; Della Torre, A.; Kafatos, F.C.; Louis, C. Identification and characterization of differentially expressed cDNAs of the vector mosquito, Anopheles gambiae. Proc. Natl. Acad. Sci. USA 1996, 93, 13066–13071. [Google Scholar] [CrossRef] [PubMed]
- Volz, J.; Osta, M.A.; Kafatos, F.C.; Müller, H.-M. The roles of two clip domain serine proteases in innate immune responses of the malaria vector Anopheles gambiae. J. Biol. Chem. 2005, 280, 40161–40168. [Google Scholar] [CrossRef] [PubMed]
- Nakhleh, J.; Christophides, G.K.; Osta, M.A. The serine protease homolog CLIPA14 modulates the intensity of the immune response in the mosquito Anopheles gambiae. J. Biol. Chem. 2017, 292, 18217–18226. [Google Scholar] [CrossRef] [PubMed]
- Patston, P.A. Serpins and other serine protease inhibitors. Immunol. Today 2000, 21, 354. [Google Scholar] [CrossRef]
- Abraham, E.G.; Pinto, S.B.; Ghosh, A.; Vanlandingham, D.L.; Budd, A.; Higgs, S.; Kafatos, F.C.; Jacobs-Lorena, M.; Michel, K. An immune-responsive serpin, SRPN6, mediates mosquito defense against malaria parasites. Proc. Natl. Acad. Sci. USA 2005, 102, 16327–16332. [Google Scholar] [CrossRef] [PubMed]
- Chalk, R.; Albuquerque, C.M.; Ham, P.J.; Townson, H. Full sequence and characterization of two insect defensins: Immune peptides from the mosquito Aedes aegypti. Proc. R. Soc. Lond. B Biol. Sci. 1995, 261, 217–221. [Google Scholar] [CrossRef] [PubMed]
- Cho, W.-L.; Fu, Y.-C.; Chen, C.-C.; Ho, C.-M. Cloning and characterization of cDNAs encoding the antibacterial peptide, defensin A, from the mosquito, Aedes aegypti. Insect Biochem. Mol. Biol. 1996, 26, 395–402. [Google Scholar] [CrossRef]
- Lowenberger, C.; Bulet, P.; Charlet, M.; Hetru, C.; Hodgeman, B.; Christensen, B.M.; Hoffmann, J.A. Insect immunity: Isolation of three novel inducible antibacterial defensins from the vector mosquito, Aedes aegypti. Insect Biochem. Mol. Biol. 1995, 25, 867–873. [Google Scholar] [CrossRef]
- Xiao, X.; Liu, Y.; Zhang, X.; Wang, J.; Li, Z.; Pang, X.; Wang, P.; Cheng, G. Complement-related proteins control the flavivirus infection of Aedes aegypti by inducing antimicrobial peptides. PLoS Pathog. 2014, 10, e1004027. [Google Scholar] [CrossRef] [PubMed]
- Christophides, G.K.; Zdobnov, E.; Barillas-Mury, C.; Birney, E.; Blandin, S.; Blass, C.; Brey, P.T.; Collins, F.H.; Danielli, A.; Dimopoulos, G. Immunity-related genes and gene families in Anopheles gambiae. Science 2002, 298, 159–165. [Google Scholar] [CrossRef] [PubMed]
- Lowenberger, C.; Smartt, C.; Bulet, P.; Ferdig, M.; Severson, D.; Hoffmann, J.; Christensen, B. Insect immunity: Molecular cloning, expression, and characterization of cDNAs and genomic DNA encoding three isoforms of insect defensin in Aedes aegypti. Insect Mol. Biol. 1999, 8, 107–118. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.-J.; Fuchs, J.F.; Mayhew, G.F.; Helen, E.Y.; Christensen, B.M. Tissue-enriched expression profiles in Aedes aegypti identify hemocyte-specific transcriptome responses to infection. Insect Biochem. Mol. Biol. 2012, 42, 729–738. [Google Scholar] [CrossRef] [PubMed]
- Blandin, S.; Moita, L.F.; Köcher, T.; Wilm, M.; Kafatos, F.C.; Levashina, E.A. Reverse genetics in the mosquito Anopheles gambiae: Targeted disruption of the Defensin gene. EMBO Rep. 2002, 3, 852–856. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Zhu, Y.; Pang, X.; Xiao, X.; Zhang, R.; Cheng, G. Regulation of Antimicrobial Peptides in Aedes aegypti Aag2 Cells. Front. Cell. Infect. Microbiol. 2017, 7, 22. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Aguilar, R.; Xi, Z.; Warr, E.; Mongin, E.; Dimopoulos, G. Anopheles gambiae immune responses to human and rodent Plasmodium parasite species. PLoS Pathog. 2006, 2, e52. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Christophides, G.K.; Cantera, R.; Charles, B.; Han, Y.S.; Meister, S.; Dimopoulos, G.; Kafatos, F.C.; Barillas-Mury, C. The role of reactive oxygen species on Plasmodium melanotic encapsulation in Anopheles gambiae. Proc. Natl. Acad. Sci. USA 2003, 100, 14139–14144. [Google Scholar] [CrossRef] [PubMed]
- Molina-Cruz, A.; DeJong, R.J.; Charles, B.; Gupta, L.; Kumar, S.; Jaramillo-Gutierrez, G.; Barillas-Mury, C. Reactive oxygen species modulate Anopheles gambiae immunity against bacteria and Plasmodium. J. Biol. Chem. 2008, 283, 3217–3223. [Google Scholar] [CrossRef] [PubMed]
- Hillyer, J.F.; Estévez-Lao, T.Y. Nitric oxide is an essential component of the hemocyte-mediated mosquito immune response against bacteria. Dev. Comp. Immunol. 2010, 34, 141–149. [Google Scholar] [CrossRef] [PubMed]
- Luckhart, S.; Li, K. Transcriptional complexity of the Anopheles stephensi nitric oxide synthase gene. Insect Biochem. Mol. Biol. 2001, 31, 249–256. [Google Scholar] [CrossRef]
- Luckhart, S.; Rosenberg, R. Gene structure and polymorphism of an invertebrate nitric oxide synthase gene. Gene 1999, 232, 25–34. [Google Scholar] [CrossRef]
- Lim, J.; Gowda, D.C.; Krishnegowda, G.; Luckhart, S. Induction of nitric oxide synthase in Anopheles stephensi by Plasmodium falciparum: Mechanism of signaling and the role of parasite glycosylphosphatidylinositols. Infect. Immun. 2005, 73, 2778–2789. [Google Scholar] [CrossRef] [PubMed]
- Luckhart, S.; Vodovotz, Y.; Cui, L.; Rosenberg, R. The mosquito Anopheles stephensi limits malaria parasite development with inducible synthesis of nitric oxide. Proc. Natl. Acad. Sci. USA 1998, 95, 5700–5705. [Google Scholar] [CrossRef] [PubMed]
- De Almeida Oliveira, G.; Lieberman, J.; Barillas-Mury, C. Epithelial nitration by a peroxidase/NOX5 system mediates mosquito antiplasmodial immunity. Science 2012, 335, 856–859. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.S.; Barillas-Mury, C. Implications of Time Bomb model of ookinete invasion of midgut cells. Insect Biochem. Mol. Biol. 2002, 32, 1311–1316. [Google Scholar] [CrossRef]
- Akira, S.; Takeda, K. Toll-like receptor signalling. Nat. Rev. Immunol. 2004, 4, 499–511. [Google Scholar] [CrossRef] [PubMed]
- Beerntsen, B.T.; James, A.A.; Christensen, B.M. Genetics of mosquito vector competence. Microbiol. Mol. Biol. Rev. 2000, 64, 115–137. [Google Scholar] [CrossRef] [PubMed]
- Cho, W.; Liu, H.; Lee, C.; Kuo, C.; Chang, T.; Liu, C.; Chen, C. Molecular cloning, characterization and tissue expression of prophenoloxidase cDNA from the mosquito Armigeres subalbatus inoculated with Dirofilaria immitis microfilariae. Insect Mol. Biol. 1998, 7, 31–40. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.R.; Yao, R.; Han, Q.; Christensen, B.M.; Li, J. Identification and molecular characterization of a prophenoloxidase involved in Aedes aegypti chorion melanization. Insect Mol. Biol. 2005, 14, 185–194. [Google Scholar] [CrossRef] [PubMed]
- Taft, A.; Chen, C.; Li, J.; Christensen, B. Molecular cloning of two prophenoloxidase genes from the mosquito Aedes aegypti. Insect Mol. Biol. 2001, 10, 97–103. [Google Scholar] [CrossRef] [PubMed]
- Müller, H.-M.; Dimopoulos, G.; Blass, C.; Kafatos, F.C. A Hemocyte-like Cell Line Established from the Malaria Vector Anopheles gambiae Expresses Six Prophenoloxidase Genes. J. Biol. Chem. 1999, 274, 11727–11735. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Christensen, B.; Chen, C. Molecular cloning of a second prophenoloxidase cDNA from the mosquito Armigeres subalbatus: Prophenoloxidase expression in blood-fed and microfilariae-inoculated mosquitoes. Insect Mol. Biol. 2001, 10, 87–96. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Kambris, Z.; Lemaitre, B.; Hashimoto, C. A serpin that regulates immune melanization in the respiratory system of Drosophila. Dev. Cell 2008, 15, 617–626. [Google Scholar] [CrossRef] [PubMed]
- Bartholomay, L.; Fuchs, J.; Cheng, L.L.; Beck, E.; Vizioli, J.; Lowenberger, C.; Christensen, B. Reassessing the role of defensin in the innate immune response of the mosquito, Aedes aegypti. Insect Mol. Biol. 2004, 13, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Bauer, H.; Gromer, S.; Urbani, A.; Schnölzer, M.; Schirmer, R.H.; Müller, H.M. Thioredoxin reductase from the malaria mosquito Anopheles gambiae. FEBS J. 2003, 270, 4272–4281. [Google Scholar]
- Barreaux, A.; Barreaux, P.; Koella, J. Overloading the immunity of the mosquito Anopheles gambiae with multiple immune challenges. Parasites Vectors 2016, 9, 210. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Jiang, G.; Chan, G.; Santos, C.P.; Severson, D.W.; Xiao, L. Michelob_x is the missing inhibitor of apoptosis protein antagonist in mosquito genomes. EMBO Rep. 2005, 6, 769–774. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Clem, R.J. Defining the core apoptosis pathway in the mosquito disease vector Aedes aegypti: The roles of iap1, ark, dronc, and effector caspases. Apoptosis 2011, 16, 105–113. [Google Scholar] [CrossRef] [PubMed]
- Bryant, B.; Blair, C.D.; Olson, K.E.; Clem, R.J. Annotation and expression profiling of apoptosis-related genes in the yellow fever mosquito, Aedes aegypti. Insect Biochem. Mol. Biol. 2008, 38, 331–345. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Clem, R.J. The role of IAP antagonist proteins in the core apoptosis pathway of the mosquito disease vector Aedes aegypti. Apoptosis 2011, 16, 235–248. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Gort, T.; Boyle, D.L.; Clem, R.J. Effects of manipulating apoptosis on Sindbis virus infection of Aedes aegypti mosquitoes. J. Virol. 2012, 86, 6546–6554. [Google Scholar] [CrossRef] [PubMed]
- Eng, M.W.; van Zuylen, M.N.; Severson, D.W. Apoptosis-related genes control autophagy and influence DENV-2 infection in the mosquito vector, Aedes aegypti. Insect Biochem. Mol. Biol. 2016, 76, 70–83. [Google Scholar] [CrossRef] [PubMed]
- Da Silva, J.B.; De Albuquerque, C.M.; De Araujo, E.C.; Peixoto, C.A.; Hurd, H. Immune defense mechanisms of Culex quinquefasciatus (Diptera: Culicidae) against Candida albicans infection. J. Invertebr. Pathol. 2000, 76, 257–262. [Google Scholar] [CrossRef] [PubMed]
- Hillyer, J.F.; Barreau, C.; Vernick, K.D. Efficiency of salivary gland invasion by malaria sporozoites is controlled by rapid sporozoite destruction in the mosquito haemocoel. Int. J. Parasitol. 2007, 37, 673–681. [Google Scholar] [CrossRef] [PubMed]
- Moita, L.F.; Vriend, G.; Mahairaki, V.; Louis, C.; Kafatos, F.C. Integrins of Anopheles gambiae and a putative role of a new β integrin, BINT2, in phagocytosis of E. coli. Insect Biochem. Mol. Biol. 2006, 36, 282–290. [Google Scholar] [CrossRef] [PubMed]
- Moita, L.F.; Wang-Sattler, R.; Michel, K.; Zimmermann, T.; Blandin, S.; Levashina, E.A.; Kafatos, F.C. In vivo identification of novel regulators and conserved pathways of phagocytosis in A. gambiae. Immunity 2005, 23, 65–73. [Google Scholar] [CrossRef] [PubMed]
- Sandiford, S.L.; Dong, Y.; Pike, A.; Blumberg, B.J.; Bahia, A.C.; Dimopoulos, G. Cytoplasmic actin is an extracellular insect immune factor which is secreted upon immune challenge and mediates phagocytosis and direct killing of bacteria, and is a Plasmodium Antagonist. PLoS Pathog. 2015, 11, e1004631. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, N.; Dasaradhi, P.; Mohmmed, A.; Malhotra, P.; Bhatnagar, R.K.; Mukherjee, S.K. RNA interference: Biology, mechanism, and applications. Microbiol. Mol. Biol. Rev. 2003, 67, 657–685. [Google Scholar] [CrossRef] [PubMed]
- Ding, S.-W.; Li, H.; Lu, R.; Li, F.; Li, W.-X. RNA silencing: A conserved antiviral immunity of plants and animals. Virus Res. 2004, 102, 109–115. [Google Scholar] [CrossRef] [PubMed]
- Bronkhorst, A.W.; van Rij, R.P. The long and short of antiviral defense: Small RNA-based immunity in insects. Curre. Opin. Virol. 2014, 7, 19–28. [Google Scholar] [CrossRef] [PubMed]
- Goic, B.; Stapleford, K.A.; Frangeul, L.; Doucet, A.J.; Gausson, V.; Blanc, H.; Schemmel-Jofre, N.; Cristofari, G.; Lambrechts, L.; Vignuzzi, M. Virus-derived DNA drives mosquito vector tolerance to arboviral infection. Nat. Commun. 2016, 7, 12410. [Google Scholar] [CrossRef] [PubMed]
- Goic, B.; Vodovar, N.; Mondotte, J.A.; Monot, C.; Frangeul, L.; Blanc, H.; Gausson, V.; Vera-Otarola, J.; Cristofari, G.; Saleh, M.-C. RNA-mediated interference and reverse transcription control the persistence of RNA viruses in the insect model Drosophila. Nat. Immunol. 2013, 14, 396. [Google Scholar] [CrossRef] [PubMed]
- Deshpande, G.; Calhoun, G.; Schedl, P. Drosophila argonaute-2 is required early in embryogenesis for the assembly of centric/centromeric heterochromatin, nuclear division, nuclear migration, and germ-cell formation. Genes Dev. 2005, 19, 1680–1685. [Google Scholar] [CrossRef] [PubMed]
- Shao, H.; Huang, Y.; Zhang, L.; Yuan, K.; Chu, Y.; Dou, Z.; Jin, C.; Garcia-Barrio, M.; Liu, X.; Yao, X. Spatiotemporal dynamics of Aurora B-PLK1-MCAK signaling axis orchestrates kinetochore bi-orientation and faithful chromosome segregation. Sci. Rep. 2015, 5, 12204. [Google Scholar] [CrossRef] [PubMed]
- Bellés, X. Beyond Drosophila: RNAi in vivo and functional genomics in insects. Annu. Rev. Entomol. 2010, 55, 111–128. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.; Tej, S.S.; Luo, S.; Haudenschild, C.D.; Meyers, B.C.; Green, P.J. Elucidation of the small RNA component of the transcriptome. Science 2005, 309, 1567–1569. [Google Scholar] [CrossRef] [PubMed]
- Matzke, M.A.; Mosher, R.A. RNA-directed DNA methylation: An epigenetic pathway of increasing complexity. Nat. Rev. Genet. 2014, 15, 394–408. [Google Scholar] [CrossRef] [PubMed]
- Wierzbicki, A.T.; Ream, T.S.; Haag, J.R.; Pikaard, C.S. RNA polymerase V transcription guides ARGONAUTE4 to chromatin. Nat. Genet. 2009, 41, 630–634. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.; Simanshu, D.K.; Ma, J.-B.; Park, J.-E.; Heo, I.; Kim, V.N.; Patel, D.J. A phosphate-binding pocket within the platform-PAZ-connector helix cassette of human Dicer. Mol. Cell 2014, 53, 606–616. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Kolb, F.A.; Jaskiewicz, L.; Westhof, E.; Filipowicz, W. Single processing center models for human Dicer and bacterial RNase III. Cell 2004, 118, 57–68. [Google Scholar] [CrossRef] [PubMed]
- Ding, S.-W. RNA-based antiviral immunity. Nat. Rev. Immunol. 2010, 10, 632–644. [Google Scholar] [CrossRef] [PubMed]
- Okamura, K.; Ishizuka, A.; Siomi, H.; Siomi, M.C. Distinct roles for Argonaute proteins in small RNA-directed RNA cleavage pathways. Genes Dev. 2004, 18, 1655–1666. [Google Scholar] [CrossRef] [PubMed]
- Cenik, E.S.; Fukunaga, R.; Lu, G.; Dutcher, R.; Wang, Y.; Hall, T.M.T.; Zamore, P.D. Phosphate and R2D2 restrict the substrate specificity of Dicer-2, an ATP-driven ribonuclease. Mol. Cell 2011, 42, 172–184. [Google Scholar] [CrossRef] [PubMed]
- Okamura, K.; Robine, N.; Liu, Y.; Liu, Q.; Lai, E.C. R2D2 organizes small regulatory RNA pathways in Drosophila. Mol. Cell. Biol. 2011, 31, 884–896. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Rand, T.A.; Kalidas, S.; Du, F.; Kim, H.-E.; Smith, D.P.; Wang, X. R2D2, a bridge between the initiation and effector steps of the Drosophila RNAi pathway. Science 2003, 301, 1921–1925. [Google Scholar] [CrossRef] [PubMed]
- Balakrishna Pillai, A.; Nagarajan, U.; Mitra, A.; Krishnan, U.; Rajendran, S.; Hoti, S.; Mishra, R. RNA interference in mosquito: Understanding immune responses, double-stranded RNA delivery systems and potential applications in vector control. Insect Mol. Biol. 2017, 26, 127–139. [Google Scholar] [CrossRef] [PubMed]
- Dana, H.; Chalbatani, G.M.; Mahmoodzadeh, H.; Karimloo, R.; Rezaiean, O.; Moradzadeh, A.; Mehmandoost, N.; Moazzen, F.; Mazraeh, A.; Marmari, V. Molecular Mechanisms and Biological Functions of siRNA. IJBS 2017, 13, 48. [Google Scholar] [PubMed]
- Betancur, J.G.; Tomari, Y. Dicer is dispensable for asymmetric RISC loading in mammals. RNA 2012, 18, 24–30. [Google Scholar] [CrossRef] [PubMed]
- Iwasaki, S.; Kobayashi, M.; Yoda, M.; Sakaguchi, Y.; Katsuma, S.; Suzuki, T.; Tomari, Y. Hsc70/Hsp90 chaperone machinery mediates ATP-dependent RISC loading of small RNA duplexes. Mol. Cell 2010, 39, 292–299. [Google Scholar] [CrossRef] [PubMed]
- Liang, C.; Wang, Y.; Murota, Y.; Liu, X.; Smith, D.; Siomi, M.C.; Liu, Q. TAF11 assembles the RISC loading complex to enhance RNAi efficiency. Mol. Cell 2015, 59, 807–818. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, D.S.; Hutvágner, G.; Du, T.; Xu, Z.; Aronin, N.; Zamore, P.D. Asymmetry in the assembly of the RNAi enzyme complex. Cell 2003, 115, 199–208. [Google Scholar] [CrossRef]
- Liu, Y.; Ye, X.; Jiang, F.; Liang, C.; Chen, D.; Peng, J.; Kinch, L.N.; Grishin, N.V.; Liu, Q. C3PO, an endoribonuclease that promotes RNAi by facilitating RISC activation. Science 2009, 325, 750–753. [Google Scholar] [CrossRef] [PubMed]
- Tolia, N.H.; Joshua-Tor, L. Slicer and the argonautes. Nat. Chem. Biol. 2006, 3, 36. [Google Scholar] [CrossRef] [PubMed]
- Song, J.-J.; Smith, S.K.; Hannon, G.J.; Joshua-Tor, L. Crystal structure of Argonaute and its implications for RISC slicer activity. Science 2004, 305, 1434–1437. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Carmell, M.A.; Rivas, F.V.; Marsden, C.G.; Thomson, J.M.; Song, J.-J.; Hammond, S.M.; Joshua-Tor, L.; Hannon, G.J. Argonaute2 is the catalytic engine of mammalian RNAi. Science 2004, 305, 1437–1441. [Google Scholar] [CrossRef] [PubMed]
- Orban, T.I.; Izaurralde, E. Decay of mRNAs targeted by RISC requires XRN1, the Ski complex, and the exosome. RNA 2005, 11, 459–469. [Google Scholar] [CrossRef] [PubMed]
- Saj, A.; Lai, E.C. Control of microRNA biogenesis and transcription by cell signaling pathways. Curr. Opin. Genet. Dev. 2011, 21, 504–510. [Google Scholar] [CrossRef] [PubMed]
- Yeom, K.-H.; Lee, Y.; Han, J.; Suh, M.R.; Kim, V.N. Characterization of DGCR8/Pasha, the essential cofactor for Drosha in primary miRNA processing. Nucleic Acids Res. 2006, 34, 4622–4629. [Google Scholar] [CrossRef] [PubMed]
- Ruby, J.G.; Jan, C.H.; Bartel, D.P. Intronic microRNA precursors that bypass Drosha processing. Nature 2007, 448, 83–86. [Google Scholar] [CrossRef] [PubMed]
- Winter, J.; Jung, S.; Keller, S.; Gregory, R.I.; Diederichs, S. Many roads to maturity: MicroRNA biogenesis pathways and their regulation. Nat. Cell Biol. 2009, 11, 228–234. [Google Scholar] [CrossRef] [PubMed]
- Friedman, R.C.; Farh, K.K.-H.; Burge, C.B.; Bartel, D.P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 2009, 19, 92–105. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [PubMed]
- Ørom, U.A.; Nielsen, F.C.; Lund, A.H. MicroRNA-10a binds the 5′ UTR of ribosomal protein mRNAs and enhances their translation. Mol. Cell 2008, 30, 460–471. [Google Scholar] [CrossRef] [PubMed]
- Kozomara, A.; Griffiths-Jones, S. miRBase: Annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 2013, 42, D68–D73. [Google Scholar] [CrossRef] [PubMed]
- Blair, C.D.; Olson, K.E. The role of RNA interference (RNAi) in arbovirus-vector interactions. Viruses 2015, 7, 820–843. [Google Scholar] [CrossRef] [PubMed]
- Keene, K.M.; Foy, B.D.; Sanchez-Vargas, I.; Beaty, B.J.; Blair, C.D.; Olson, K.E. RNA interference acts as a natural antiviral response to O’nyong-nyong virus (Alphavirus; Togaviridae) infection of Anopheles gambiae. Proc. Natl. Acad. Sci. USA 2004, 101, 17240–17245. [Google Scholar] [CrossRef] [PubMed]
- Berry, B.; Deddouche, S.; Kirschner, D.; Imler, J.-L.; Antoniewski, C. Viral suppressors of RNA silencing hinder exogenous and endogenous small RNA pathways in Drosophila. PLoS ONE 2009, 4, e5866. [Google Scholar] [CrossRef] [PubMed]
- Hussain, M.; Abraham, A.M.; Asgari, S. An ascovirus-encoded RNase III autoregulates its expression and suppresses RNA interference-mediated gene silencing. J. Virol. 2010, 84, 3624–3630. [Google Scholar] [CrossRef] [PubMed]
- Obbard, D.J.; Jiggins, F.M.; Halligan, D.L.; Little, T.J. Natural selection drives extremely rapid evolution in antiviral RNAi genes. Curr. Biol. 2006, 16, 580–585. [Google Scholar] [CrossRef] [PubMed]
- Sabin, L.R.; Zhou, R.; Gruber, J.J.; Lukinova, N.; Bambina, S.; Berman, A.; Lau, C.-K.; Thompson, C.B.; Cherry, S. Ars2 regulates both miRNA-and siRNA-dependent silencing and suppresses RNA virus infection in Drosophila. Cell 2009, 138, 340–351. [Google Scholar] [CrossRef] [PubMed]
- Hussain, M.; Walker, T.; O’Neill, S.L.; Asgari, S. Blood meal induced microRNA regulates development and immune associated genes in the Dengue mosquito vector, Aedes aegypti. Insect Biochem. Mol. Biol. 2013, 43, 146–152. [Google Scholar] [CrossRef] [PubMed]
- Lucas, K.J.; Zhao, B.; Liu, S.; Raikhel, A.S. Regulation of physiological processes by microRNAs in insects. Curr. Opin. Insect Sci. 2015, 11, 1–7. [Google Scholar] [CrossRef] [PubMed]
- McFarlane, M.; Arias-Goeta, C.; Martin, E.; O’Hara, Z.; Lulla, A.; Mousson, L.; Rainey, S.M.; Misbah, S.; Schnettler, E.; Donald, C.L. Characterization of Aedes aegypti innate-immune pathways that limit Chikungunya virus replication. PLoS Negl. Trop. Dis. 2014, 8, e2994. [Google Scholar] [CrossRef] [PubMed]
- Pan, X.; Zhou, G.; Wu, J.; Bian, G.; Lu, P.; Raikhel, A.S.; Xi, Z. Wolbachia induces reactive oxygen species (ROS)-dependent activation of the Toll pathway to control dengue virus in the mosquito Aedes aegypti. Proc. Natl. Acad. Sci. USA 2012, 109, E23–E31. [Google Scholar] [CrossRef] [PubMed]
- Saldaña, M.A.; Etebari, K.; Hart, C.E.; Widen, S.G.; Wood, T.G.; Thangamani, S.; Asgari, S.; Hughes, G.L. Zika virus alters the microRNA expression profile and elicits an RNAi response in Aedes aegypti mosquitoes. PLoS Negl. Trop. Dis. 2017, 11, e0005760. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Vargas, I.; Scott, J.C.; Poole-Smith, B.K.; Franz, A.W.; Barbosa-Solomieu, V.; Wilusz, J.; Olson, K.E.; Blair, C.D. Dengue virus type 2 infections of Aedes aegypti are modulated by the mosquito’s RNA interference pathway. PLoS Pathog. 2009, 5, e1000299. [Google Scholar] [CrossRef] [PubMed]
- Yan, H.; Zhou, Y.; Liu, Y.; Deng, Y.; Puthiyakunnon, S.; Chen, X. MiR-252 of the Asian tiger mosquito Aedes albopictus regulates dengue virus replication by suppressing the expression of the dengue virus envelope protein. J. Med. Virol. 2014, 86, 1428–1436. [Google Scholar] [CrossRef] [PubMed]
- Weick, E.M.; Miska, E.A. piRNAs: From biogenesis to function. Development 2014, 141, 3458–3471. [Google Scholar] [CrossRef] [PubMed]
- Ku, H.Y.; Lin, H. PIWI proteins and their interactors in piRNA biogenesis, germline development and gene expression. Natl. Sci. Rev. 2014, 1, 205–218. [Google Scholar] [CrossRef] [PubMed]
- Shirayama, M.; Seth, M.; Lee, H.C.; Gu, W.; Ishidate, T.; Conte, D., Jr.; Mello, C.C. piRNAs initiate an epigenetic memory of nonself RNA in the C. elegans germline. Cell 2012, 150, 65–77. [Google Scholar] [CrossRef] [PubMed]
- Miesen, P.; Girardi, E.; van Rij, R.P. Distinct sets of PIWI proteins produce arbovirus and transposon-derived piRNAs in Aedes aegypti mosquito cells. Nucleic Acids Res. 2015, 43, 6545–6556. [Google Scholar] [CrossRef] [PubMed]
- Vodovar, N.; Bronkhorst, A.W.; van Cleef, K.W.; Miesen, P.; Blanc, H.; van Rij, R.P.; Saleh, M.C. Arbovirus-derived piRNAs exhibit a ping-pong signature in mosquito cells. PLoS ONE 2012, 7, e30861. [Google Scholar] [CrossRef] [PubMed]
- Petit, M.; Mongelli, V.; Frangeul, L.; Blanc, H.; Jiggins, F.; Saleh, M.-C. piRNA pathway is not required for antiviral defense in Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 2016, 113, E4218–E4227. [Google Scholar] [CrossRef] [PubMed]
- Ross, R.J.; Weiner, M.M.; Lin, H. PIWI proteins and PIWI-interacting RNAs in the soma. Nature 2014, 505, 353–359. [Google Scholar] [CrossRef] [PubMed]
- Varjak, M.; Maringer, K.; Watson, M.; Sreenu, V.B.; Fredericks, A.C.; Pondeville, E.; Donald, C.L.; Sterk, J.; Kean, J.; Vazeille, M.; et al. Aedes aegypti Piwi4 Is a Noncanonical PIWI Protein Involved in Antiviral Responses. mSphere 2017, 2. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Dong, Y.; Gu, J.; Puthiyakunnon, S.; Wu, Y.; Chen, X.G. Developmental piRNA profiles of the invasive vector mosquito Aedes albopictus. Parasit Vectors 2016, 9, 524. [Google Scholar] [CrossRef] [PubMed]
- Girardi, E.; Miesen, P.; Pennings, B.; Frangeul, L.; Saleh, M.C.; van Rij, R.P. Histone-derived piRNA biogenesis depends on the ping-pong partners Piwi5 and Ago3 in Aedes aegypti. Nucleic Acids Res. 2017, 45, 4881–4892. [Google Scholar] [CrossRef] [PubMed]
- Miesen, P.; Ivens, A.; Buck, A.H.; van Rij, R.P. Small RNA Profiling in Dengue Virus 2-Infected Aedes Mosquito Cells Reveals Viral piRNAs and Novel Host miRNAs. PLoS Negl. Trop. Dis. 2016, 10, e0004452. [Google Scholar] [CrossRef] [PubMed]
- Campbell, C.L.; Black, W.C.t.; Hess, A.M.; Foy, B.D. Comparative genomics of small RNA regulatory pathway components in vector mosquitoes. BMC Genom. 2008, 9, 425. [Google Scholar] [CrossRef] [PubMed]
- Morazzani, E.M.; Wiley, M.R.; Murreddu, M.G.; Adelman, Z.N.; Myles, K.M. Production of virus-derived ping-pong-dependent piRNA-like small RNAs in the mosquito soma. PLoS Pathog. 2012, 8, e1002470. [Google Scholar] [CrossRef] [PubMed]
- Ernst, C.; Odom, D.T.; Kutter, C. The emergence of piRNAs against transposon invasion to preserve mammalian genome integrity. Nat. Commun. 2017, 8, 1411. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, T.; Lin, H. Posttranscriptional regulation of gene expression by Piwi proteins and piRNAs. Mol. Cell 2014, 56, 18–27. [Google Scholar] [CrossRef] [PubMed]
- Schnettler, E.; Donald, C.L.; Human, S.; Watson, M.; Siu, R.W.; McFarlane, M.; Fazakerley, J.K.; Kohl, A.; Fragkoudis, R. Knockdown of piRNA pathway proteins results in enhanced Semliki Forest virus production in mosquito cells. J. Gen. Virol. 2013, 94, 1680–1689. [Google Scholar] [CrossRef] [PubMed]
- Palatini, U.; Miesen, P.; Carballar-Lejarazu, R.; Ometto, L.; Rizzo, E.; Tu, Z.; van Rij, R.P.; Bonizzoni, M. Comparative genomics shows that viral integrations are abundant and express piRNAs in the arboviral vectors Aedes aegypti and Aedes albopictus. BMC Genom. 2017, 18, 512. [Google Scholar] [CrossRef] [PubMed]
- Olson, K.E.; Bonizzoni, M. Nonretroviral integrated RNA viruses in arthropod vectors: An occasional event or something more? Curr. Opin. Insect Sci. 2017, 22, 45–53. [Google Scholar] [CrossRef] [PubMed]
- Myles, K.M.; Wiley, M.R.; Morazzani, E.M.; Adelman, Z.N. Alphavirus-derived small RNAs modulate pathogenesis in disease vector mosquitoes. Proc. Natl. Acad. Sci. USA 2008, 105, 19938–19943. [Google Scholar] [CrossRef] [PubMed]
- Gammon, D.B.; Mello, C.C. RNA interference-mediated antiviral defense in insects. Curr. Opin. Insect Sci. 2015, 8, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Kakumani, P.K.; Rajgokul, K.; Ponia, S.S.; Kaur, I.; Mahanty, S.; Medigeshi, G.R.; Banerjea, A.C.; Chopra, A.P.; Malhotra, P.; Mukherjee, S.K. Dengue NS3, an RNAi suppressor, modulates the human miRNA pathways through its interacting partner. Biochem. J. 2015, 471, 89–99. [Google Scholar] [CrossRef] [PubMed]
- Mathur, K.; Anand, A.; Dubey, S.K.; Sanan-Mishra, N.; Bhatnagar, R.K.; Sunil, S. Analysis of chikungunya virus proteins reveals that non-structural proteins nsP2 and nsP3 exhibit RNA interference (RNAi) suppressor activity. Sci. Rep. 2016, 6, 38065. [Google Scholar] [CrossRef] [PubMed]
- Weinheimer, I.; Jiu, Y.; Rajamäki, M.-L.; Matilainen, O.; Kallijärvi, J.; Cuellar, W.J.; Lu, R.; Saarma, M.; Holmberg, C.I.; Jäntti, J. Suppression of RNAi by dsRNA-degrading RNaseIII enzymes of viruses in animals and plants. PLoS Pathog. 2015, 11, e1004711. [Google Scholar] [CrossRef] [PubMed]
- Samuel, G.H.; Wiley, M.R.; Badawi, A.; Adelman, Z.N.; Myles, K.M. Yellow fever virus capsid protein is a potent suppressor of RNA silencing that binds double-stranded RNA. Proc. Natl. Acad. Sci. USA 2016, 113, 13863–13868. [Google Scholar] [CrossRef] [PubMed]
- Van Mierlo, J.T.; Overheul, G.J.; Obadia, B.; van Cleef, K.W.; Webster, C.L.; Saleh, M.-C.; Obbard, D.J.; van Rij, R.P. Novel Drosophila viruses encode host-specific suppressors of RNAi. PLoS Pathog. 2014, 10, e1004256. [Google Scholar] [CrossRef] [PubMed]
- Chao, J.A.; Lee, J.H.; Chapados, B.R.; Debler, E.W.; Schneemann, A.; Williamson, J.R. Dual modes of RNA-silencing suppression by Flock House virus protein B2. Nat. Struct. Mol. Biol. 2005, 12, 952. [Google Scholar] [CrossRef] [PubMed]
- Lakatos, L.; Csorba, T.; Pantaleo, V.; Chapman, E.J.; Carrington, J.C.; Liu, Y.P.; Dolja, V.V.; Calvino, L.F.; López-Moya, J.J.; Burgyán, J. Small RNA binding is a common strategy to suppress RNA silencing by several viral suppressors. EMBO J. 2006, 25, 2768–2780. [Google Scholar] [CrossRef] [PubMed]
- Azevedo, J.; Garcia, D.; Pontier, D.; Ohnesorge, S.; Yu, A.; Garcia, S.; Braun, L.; Bergdoll, M.; Hakimi, M.A.; Lagrange, T. Argonaute quenching and global changes in Dicer homeostasis caused by a pathogen-encoded GW repeat protein. Genes Dev. 2010, 24, 904–915. [Google Scholar] [CrossRef] [PubMed]
- Samuel, G.H.; Adelman, Z.N.; Myles, K.M. Antiviral Immunity and Virus-Mediated Antagonism in Disease Vector Mosquitoes. Trends Microbiol. 2018, 26, 447–461. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kumar, A.; Srivastava, P.; Sirisena, P.; Dubey, S.K.; Kumar, R.; Shrinet, J.; Sunil, S. Mosquito Innate Immunity. Insects 2018, 9, 95. https://doi.org/10.3390/insects9030095
Kumar A, Srivastava P, Sirisena P, Dubey SK, Kumar R, Shrinet J, Sunil S. Mosquito Innate Immunity. Insects. 2018; 9(3):95. https://doi.org/10.3390/insects9030095
Chicago/Turabian StyleKumar, Ankit, Priyanshu Srivastava, PDNN Sirisena, Sunil Kumar Dubey, Ramesh Kumar, Jatin Shrinet, and Sujatha Sunil. 2018. "Mosquito Innate Immunity" Insects 9, no. 3: 95. https://doi.org/10.3390/insects9030095
APA StyleKumar, A., Srivastava, P., Sirisena, P., Dubey, S. K., Kumar, R., Shrinet, J., & Sunil, S. (2018). Mosquito Innate Immunity. Insects, 9(3), 95. https://doi.org/10.3390/insects9030095




