Effect of Host Genotype on Symbiont Titer in the Aphid—Buchnera Symbiosis
Abstract
: Obligate nutritional symbioses require balance between the energetic needs of the host and the symbiont. The resident symbiont population size within a host may have major impacts on host fitness, as both host and symbiont consume and supply metabolites in a shared metabolite pool. Given the massive genome degradation that is a hallmark of bacterial endosymbionts of insects, it is unclear at what level these populations are regulated, and how regulation varies among hosts within natural populations. We measured the titer of the endosymbiont Buchnera aphidicola from different clones of the pea aphid, Acyrthosiphon pisum, and found significant variation in titer, measured as Buchnera genomes per aphid genome, among aphid clones. Additionally, we found that titer can change with the age of the host, and that the number of bacteriocytes within an aphid is one factor likely controlling Buchnera titer. Buchnera titer measurements in clones from a sexual cross indicate that the symbiont genotype is not responsible for variation in titer and that this phenotype is likely non-heritable across sexual reproduction. Symbiont titer is more variable among lab-produced F1 aphid clones than among field-collected ones, suggesting that intermediate titer is favored in natural populations. Potentially, a low heritability of titer during the sexual phase may generate clones with extreme and maladaptive titers each season.1. Introduction
Bacterial endosymbionts are common associates of insects, often providing essential nutrients that are absent from the host diet [1]. Buchnera aphidicola, the primary symbiont of aphids, has become a model system for the study of these relationships. This member of the Gammaproteobacteria produces essential amino acids that are rare in the diet of aphids, phloem sap [2,3]. Early investigations of these symbionts suggested that the host controlled the titer of the symbionts [4], and later genomic studies revealed that reductive genome evolution in these symbionts resulted in the loss of many genes necessary for regulation of cell processes including division and growth [3,5]. These observations suggest that the symbiont is unable to control its own replication, and that the host plays a major role in regulating the titer of symbionts.
Despite the loss of regulatory mechanisms to control symbiont division and metabolism, variation in symbiont genotypes may contribute to differences in symbiont titer. Buchnera's high mutation rate [6], asexuality, and small population size of the symbiont [7,8] can lead to disruptions of processes that are likely essential to both symbiont and host [9]. Buchnera titer may also have been shaped by selection on the host, as the symbiont has little or no ability to alter gene expression in response to differences in amino acid content of phloem [10], which is known to vary among and between host plant species [11]. Symbiont titer may therefore be a mechanism for regulating the amino acid metabolism of the system.
A. pisum is a cyclical parthenogen, like many aphids, and undergoes several generations of clonal reproduction followed by a single generation of sexual reproduction [12]. In such life cycles, traits with epistatic or dominance genetic variance are heritable during clonal reproduction but not across the sexual reproductive phase [13,14]. Symbiont titer may be one such phenotype.
To assess variation in symbiont titer, we measured the titer of Buchnera in populations of the pea aphid, Acyrthosiphon pisum. To examine the underlying basis of the variation, we measured the number of bacteriocytes, the specialized cells in which Buchnera resides, between the clones, as well as the number of Buchnera within a bacteriocyte and the relationship of titer to amino acid requirements of the clones. Two clones with high and low titer were bred to produce F1 offspring, which were then screened for titer and bacteriocyte number.
2. Experimental Section
2.1. Aphid Clones
Parthenogenetic A. pisum females were collected from across the United States between 1998 and 2007 (Table S1). For each clone, a single female was used to establish clonal lineages maintained continuously under long day (16:8 L:D) conditions. Experimental aphid lines were kept in a growth chamber at 20 °C on Vicia faba seedlings in cup cages [15]. Short day conditions were used to induce sexual forms for clones 8–10–1 and 5A, which were reciprocally mated to yield F1 aphid clones, as described by Moran and Dunbar [16]. Individuals hatching from the sexually produced eggs were isolated and allowed to establish full sib clonal lineages under long day conditions. For all experiments, clones were divided into 3 sub-clones and allowed to reproduce for 3 generations prior to collection to control for maternal and environmental effects. Each experiment was replicated twice, beginning with the establishment of new subclones.
2.2. Bacteriocyte Counts
Adult viviparous females were placed on fresh V. faba seedlings and allowed to deposit nymphs for 12 hours, after which the adults were removed and the nymphs allowed to develop for 6 days, to their 4th instar. Fourth instar aphids were dissected in buffer A (250 mM sucrose, 35 mM Tris-HCl, 25 mM KCl, 10 mM MgCl2) in a watch glass. All bacteriocytes were identified, separated and counted under 6× magnification.
2.3. DNA Extractions
Adult viviparous A. pisum were placed on fresh V. faba seedlings and allowed to deposit nymphs for 12 hours. Nymphs were either allowed to develop for six days to their fourth instar or immediately collected at their first instar. Individual nymphs were collected in pestle tubes, frozen in liquid nitrogen, and crushed with a pestle. The resulting homogenized tissue was treated according to the Qiagen DNEasy kit.
To isolate DNA from individual bacteriocytes, single bacteriocytes from the aphids used for bacteriocyte counts were collected in pestle tubes, frozen in liquid nitrogen and crushed. Due to the small amount of starting material, the resulting homogenate was treated with lysis buffer [17] and then washed twice with phenol:chloroform:isoamyl alcohol 25:24:1, then once with chloroform. The DNA was then precipitated with sodium acetate and ethanol and resuspended in low TE (10 mM EDTA, 100 mM Tris-HCl). All DNA was treated with RNAse I at 37 °C for 30 minutes. Three aphids from each of the 3 subclones from each clone were used for the experiments.
2.4. Quantitative PCR
Buchnera titer was measured by comparing the number of Buchnera genomes to the number of aphid genomes using a single copy gene from both the aphid and the symbiont. This provided a rough correction for size differences between aphid clones, though some aphid cells are polyploid [18]. Aphid genomes were counted by assessing copy number of the gene encoding elongation factor 1-alpha (ef1α), while Buchnera genomes were counted by using the gene encoding adenosylmethionine-8-amino-7-oxononanoate aminotransferase (bioA). Primers used were ApEF1-alpha 107F 5′ - CTGATTGTGCCGTGCTTATTG - 3′, ApEF1-alpha 246R 5′ - TATGGTGGTTCAGTAGAGTCC - 3′, BuchAPS bioA 374F 5′ - AGTATTGGCAAGCATTAGGGC - 3′, BuchAPS bioA 526R 5′ - AAAAGAAGAAACTGGTCGTC - 3′. Standards of 107 copies were prepared according to the method of [10] for each gene. For each sample from 1st instar aphids, the number of copies of ef1α and bioA were compared on a Roche LightCycler using the FastStart DNA Masterplus SYBR Green I kit according to the kit instructions. Copy number was determined using LightCycler 3.0 software in comparison to the standards for each gene. For individual bacteriocytes, only the copy number of bioA was assessed, though the bacteriocytes are polyploid [18]. For each clone, three aphids from each subclone were tested, and each clone was measured twice, starting with the initiation of new subclones.
2.5. Statistical Analysis
Buchnera titer was determined by the ratio of Buchnera genomes to A. pisum genomes. The regression of Buchnera genome copies and aphid genome copies was linear but the slopes of the variables were unequal and the intercepts were non-zero, so ANCOVA analysis for unequal slopes was applied in JMP 8 (SAS). Buchnera genome copy number of individual bacteriocytes was log transformed and analyzed by ANOVA. Bacteriocyte counts were normally distributed and analyzed by MANOVA. Summary statistics are included in Table S2.
3. Results and Discussion
3.1. Buchnera Titer
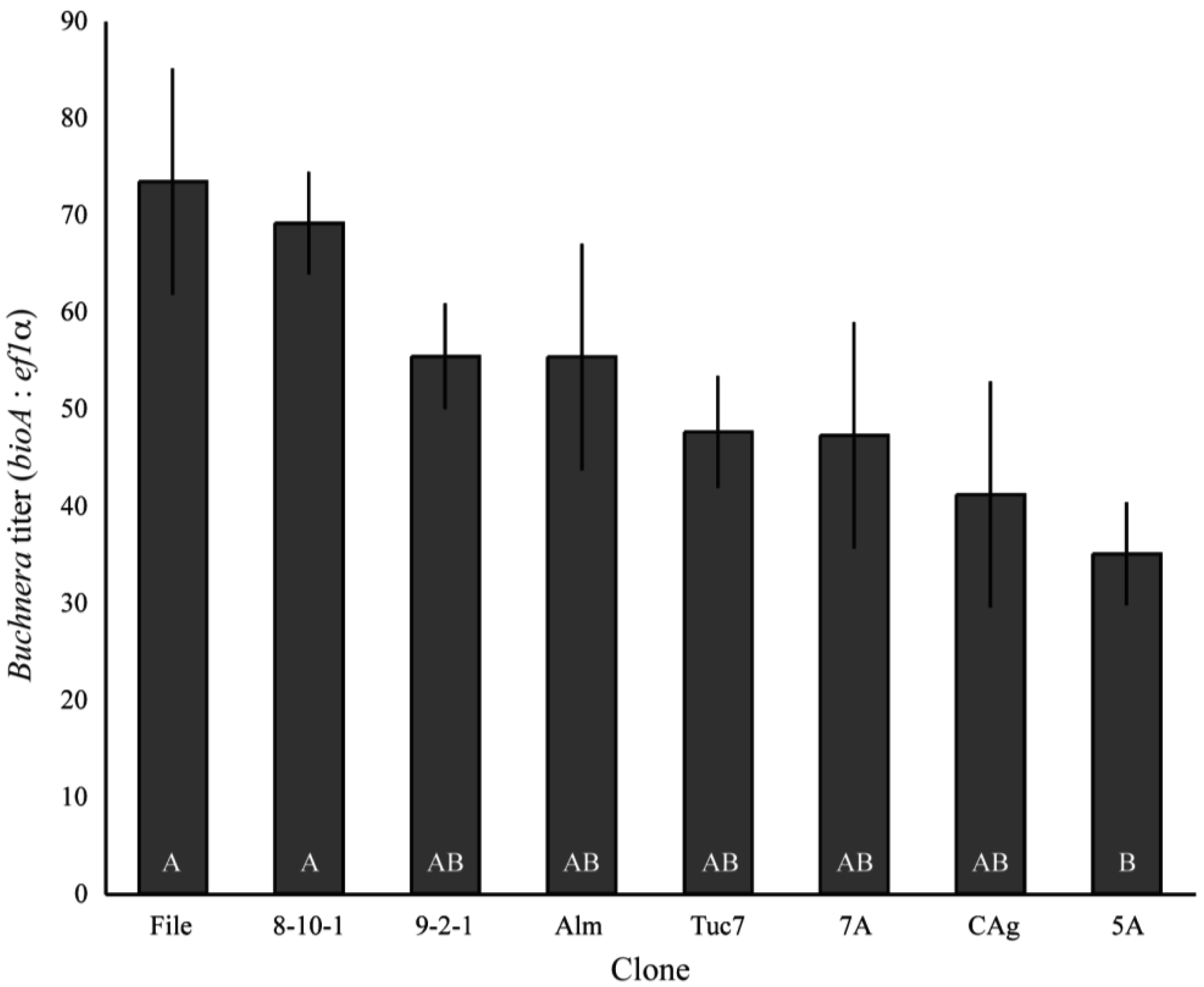
Buchnera titer (Buchnera genomes per aphid genome) varied significantly among field-collected clones of A. pisum in the first instar (F9,95 = 3.21, p = 0.0019, ANCOVA), with the lowest titer clone–5A–having an average of 35 Buchnera genomes per aphid genome, while the highest titer clone–File–had an average titer of 73.4 (Figure 1). The copy number of both ef1α and bioA varied significantly between clones (p < 0.0001 ANOVA), but the variation in ef1α copy number only explained 4.4% of the variance in Buchnera titer while bioA copy number explained 36.6% of the variance based on measuring effects of each variable independently.
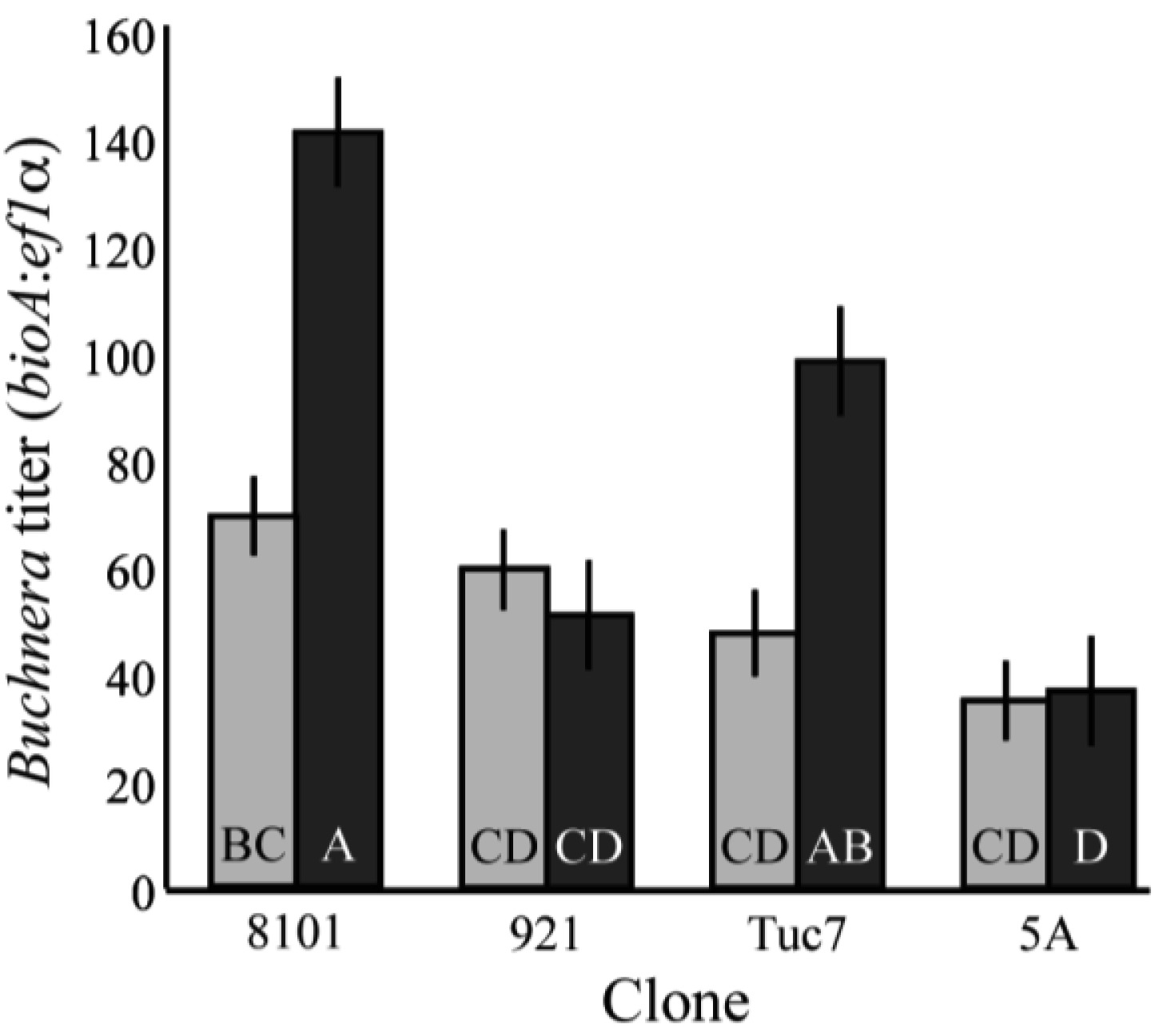
Buchnera titer also varies across the development of the host. Four clones were tested, and there were significant changes in the titer of Buchnera from 1st to 4th instar (F7,97 = 10.37, p < 0.0001; ANCOVA, Figure 2), with clones 8-10-1 and Tuc7 increasing significantly in titer (p = 0.0005 and p = 0.027, respectively student's t-test), and the titer in clones 5A and 9-2-1 not changing significantly (p = 0.82, p = 0.33, respectively student's t-test). Titer varied significantly between the clones at the 4th instar (Figure 2), with clones 8-10-1 and Tuc7 exhibiting similar titer and clones 5A and 9-2-1 having significantly lower titer (p < 0.05, Tukey's HSD). The copy number of both Buchnera genomes and aphid genomes varied significantly between the 4th instars of the clones tested. In 4th instar aphids, Buchnera genome copy number explained 62.9% of the variation in titer (p < 0.0001, ANOVA) while the aphid genome accounted for 18.5% (p < 0.0001, ANOVA) based on measuring effects of each variable independently.
3.2. Titer in F1 Clones
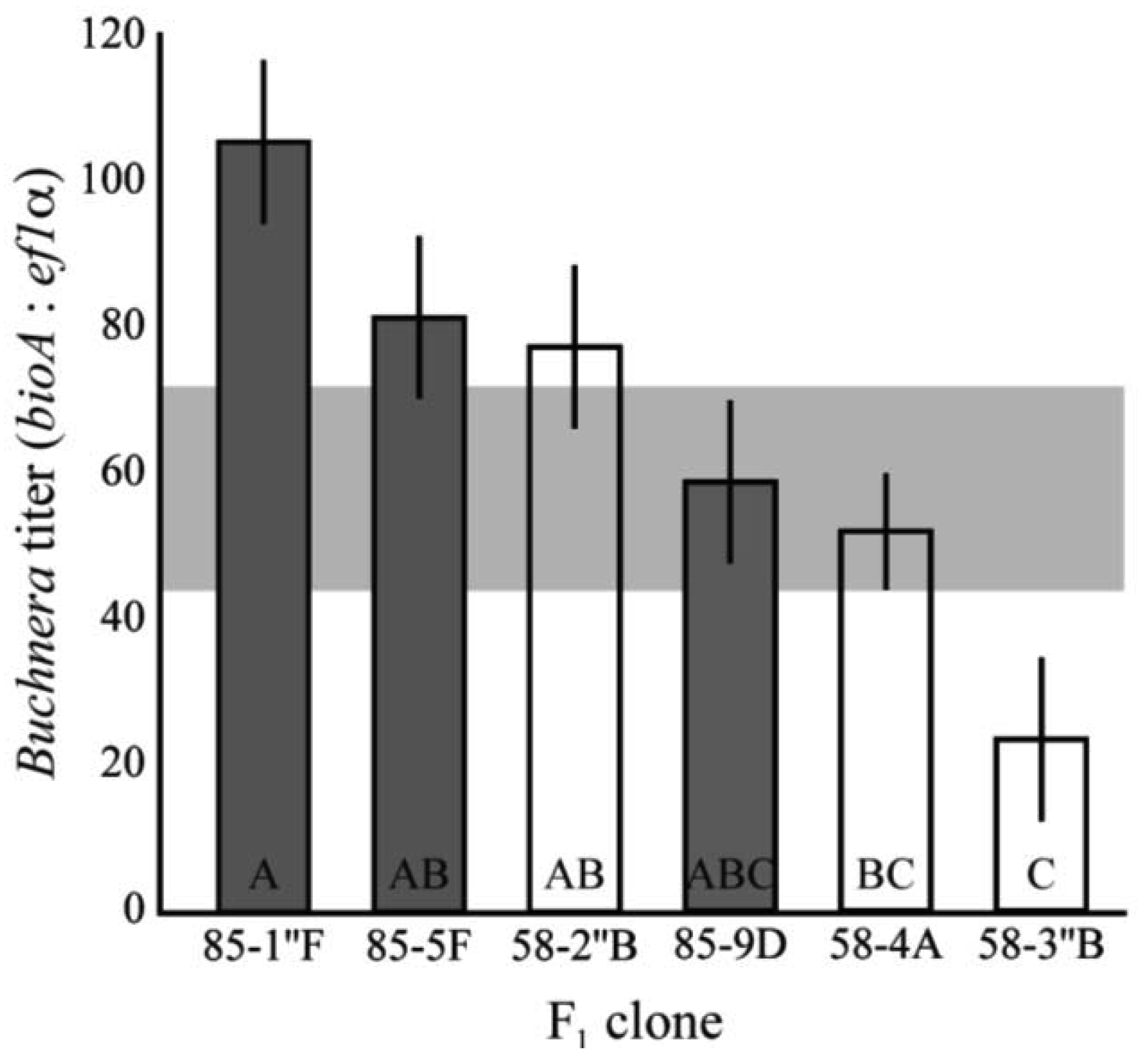
To determine the pattern of heritability in titer, clones 5A and 8-10-1 were reciprocally mated to produce a panel of full-sib clones with distinct genotypes. The Buchnera titer of 1st instar aphids were significantly different among F1 clones (F6,93 = 2.9, p = 0.0004, ANCOVA, Figure 3). The clones also had significantly higher and lower average titers than those observed in the lab clones–those established from field collected asexual females–or their parental clones (F1,15 = 8.48, p = 0.0042, ANCOVA). Titers ranged from 23.4 Buchnera genomes per aphid genome in clone 58-3″B and 105.6 in clone 85-1″F. These values were significantly lower and higher than those observed in the parental clones (p < 0.05, Tukey's HSD).
As Buchnera are maternally transmitted, aphids from the two matrilines have different Buchnera genotypes. There was no significant difference observed in the Buchnera titer of F1 clones from the two matrilines, suggesting that differences between the genomes of Buchnera from 5A and 8-10-1 are not responsible for the differences in symbiont titer between these two clones (F1,93 = 0.046, p = 0.83, ANCOVA).
3.3. Bacteriocyte Quantification and Buchnera per Bacteriocyte
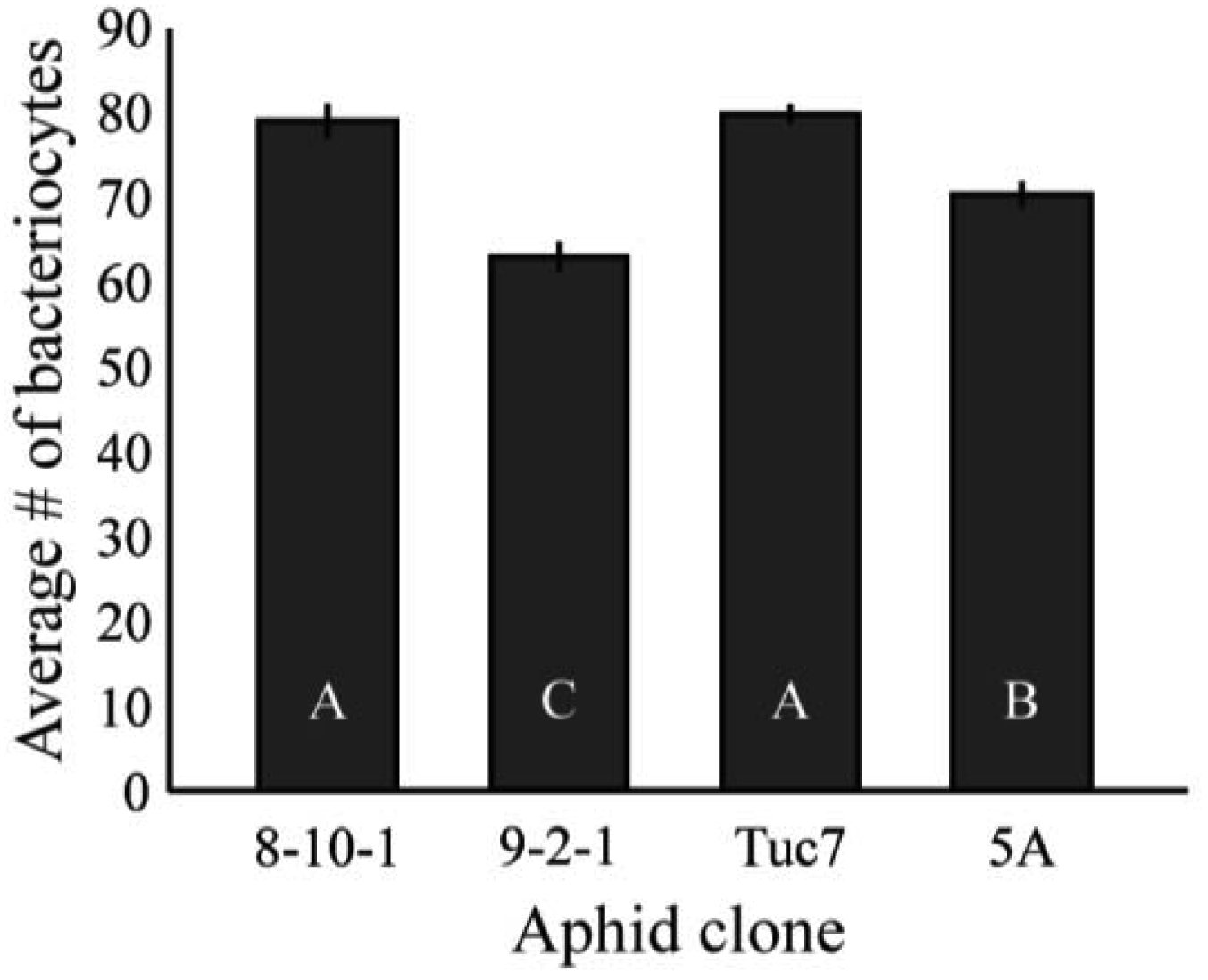
Figure 4 shows the average number of bacteriocytes per aphid at the 4th instar. Bacteriocyte counts varied significantly between clones tested (F3,172 = 21.3, p < 0.0001, ANOVA). These data are mostly consistent with the Buchnera titer of 4th instar aphids (Figure 2), with Tuc7 and 8-10-1 having similar numbers of bacteriocytes per aphid while 9-2-1 and 5A have significantly fewer (p < 0.05, Tukey's HSD). Clone 9-2-1 has fewer bacteriocytes than clone 5A (p < 0.05, Tukey's HSD), though these clones do not differ in the titer of Buchnera at the 4th instar (Figure 2). While variation in Buchnera titer in 1st instars is not consistently reflected in measures of number of bacteriocytes or the number of Buchnera per bacteriocyte, the symbiont titer of 4th instar aphids recapitulates the patterns observed in the number of bacteriocytes in the clones tested, suggesting that control of bacteriocyte number is one mechanism by which the symbiont populations within an individual is regulated by the host.
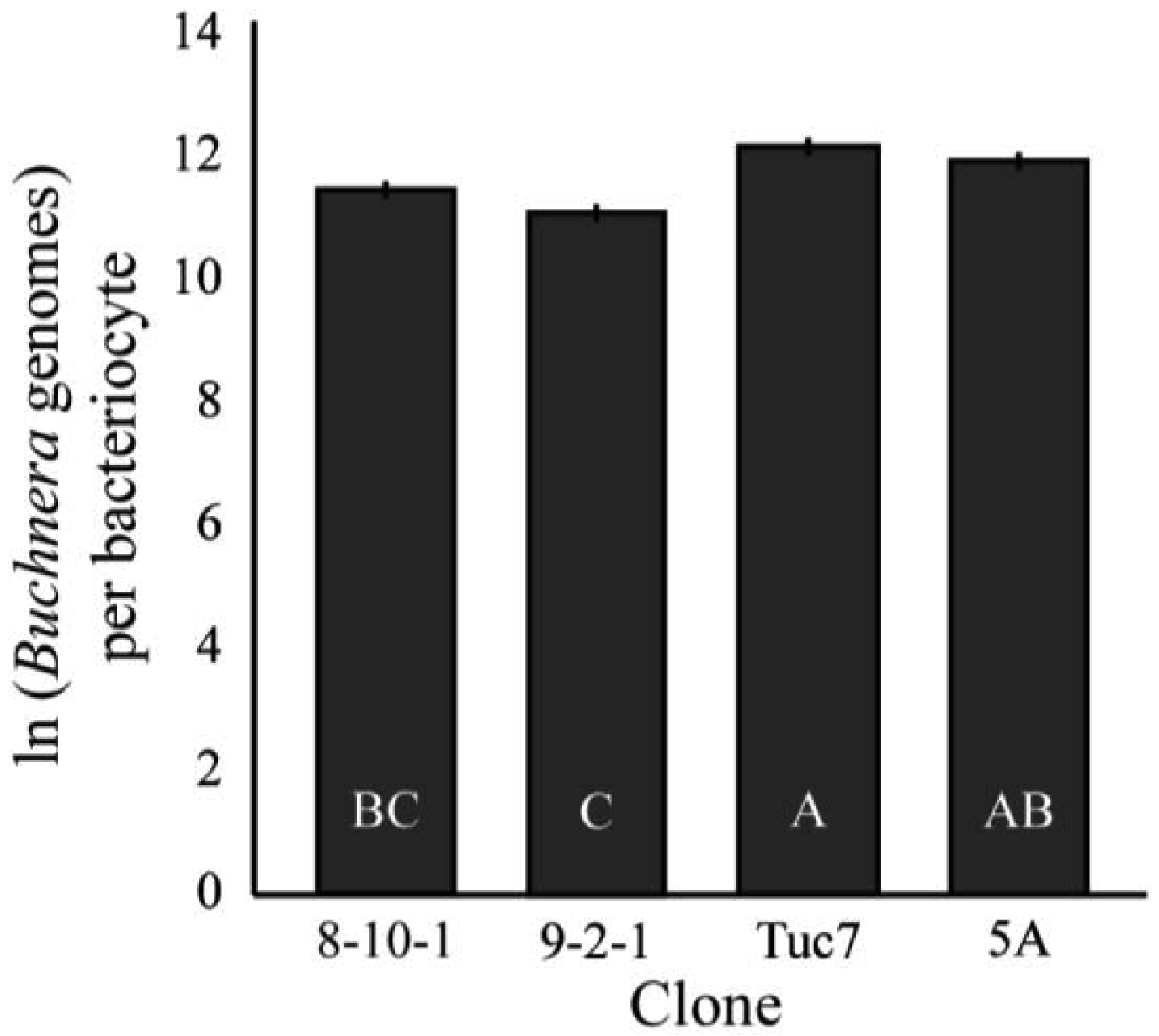
The number of Buchnera genomes per bacteriocyte was significantly different among the clones examined (F3,94 = 10.24, p < 0.0001, MANOVA, Figure 5). 9-2-1 had the fewest Buchnera per bacteriocyte, significantly fewer than all the clones except 8-10-1 (p < 0.05, Tukey's HSD). Tuc7 and 5A had an equivalent number of Buchnera per bacteriocyte, while 8-10-1 had fewer than Tuc7, but the difference between 5A and 8-10-1 was not significant.
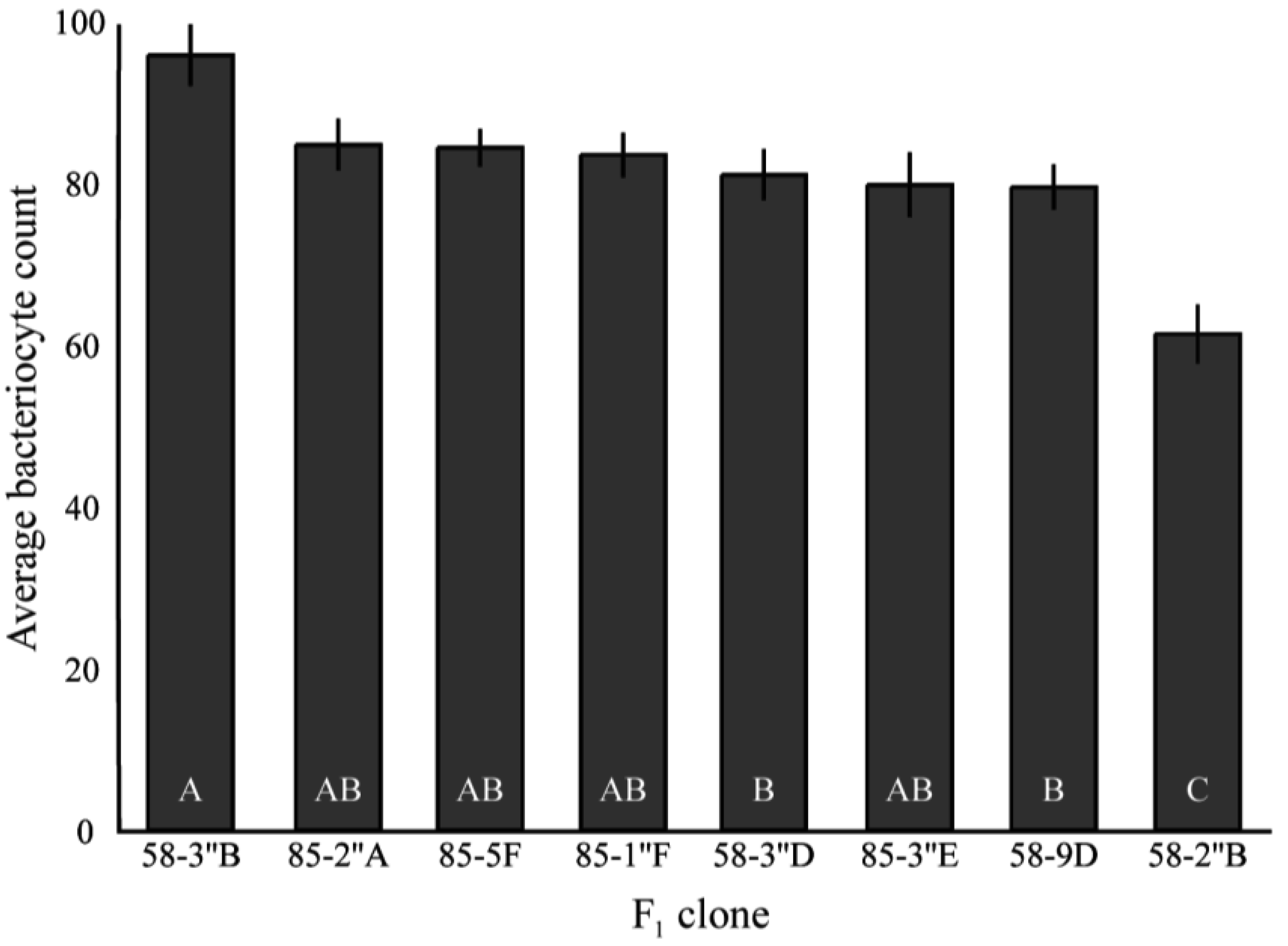
Together, results for bacteriocyte numbers are consistent with the differences in total Buchnera titers in three of the four clones examined. 5A and 8-10-1 exhibited the largest difference in 1st instar Buchnera titer, and 8-10-1 had significantly more bacteriocytes than 5A. In these clones, there was no significant difference between the numbers of Buchnera genomes per bacteriocyte, suggesting that the number of bacteriocytes may be responsible for the difference in Buchnera titer between the clones. Tuc7 had more bacteriocytes than 5A and more Buchnera per bacteriocyte than 8-10-1, though it had intermediate 1st instar titer compared to these clones. While there was a strong correlation between bacteriocyte numbers and Buchnera titer for the four clones tested (R2 = 0.62), the relationship was not significant (F1,4 = 3.39, p = 0.2, ANCOVA) though a lack of power likely contributed to this. Significant variation in bacteriocyte numbers was also seen in the F1 clones (F5,106 = 8.00, p < 0.0001, ANOVA, Figure S1), though no comparison could be made between the Buchnera titer and bacteriocyte number in these clones, as they were measured at 1st and 4th instars, respectively.
3.4. Buchnera Titer and Amino Acid Requirements of Clones
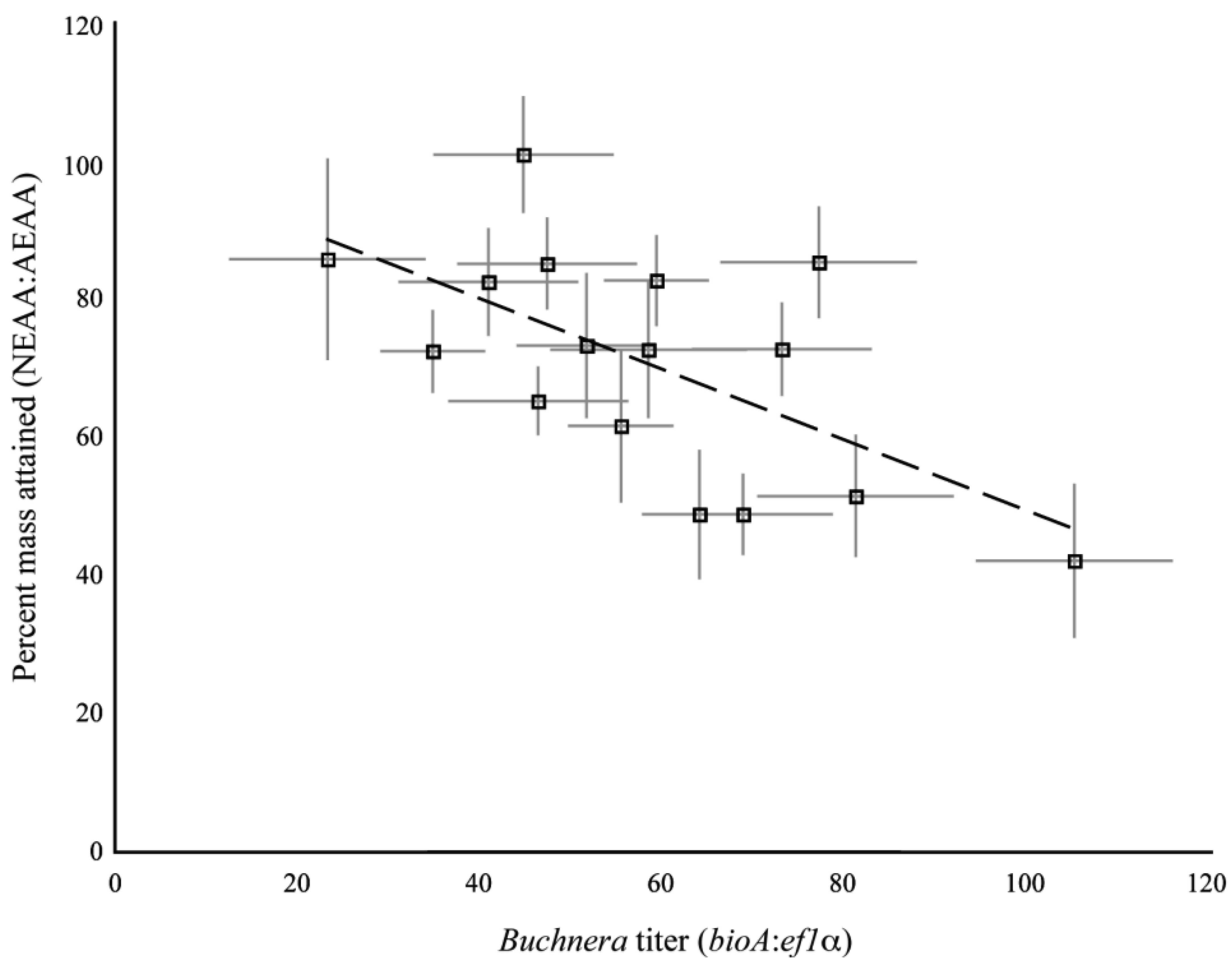
The clones used in the present study were previously assayed for their dietary requirements of essential amino acids [9]. In that study, we measured the mass of aphids reared on artificial diets with and without essential amino acids. We used the ratio of aphid mass on the diet without essential amino acids to the mass of aphids reared on diet with all the essential amino acids as a measure of the dietary requirement for essential amino acids for each clone. As different individuals were measured for the previous study and the current study, we compared the mean amino acid requirement of a clone against the mean 1st instar Buchnera titer as measured in the current study. We found a significant, positive association between amino acid requirements and Buchnera titer for the set of clones tested (R2 = 0.38, F1,14 = 8.71, p = 0.01, ANCOVA, Figure S2).
4. Conclusions
Control of symbiont titer varies among clonal lines of pea aphid. The titer of Buchnera, defined as number of Buchnera genomes relative to aphid genomes, can also vary with age of the aphid, increasing significantly between the 1st and 4th instar in two of the four clones tested. Part of the variation in Buchnera titer may be due to differences in the number of bacteriocytes between clones. There is significant variation in the number of Buchnera genomes per bacteriocyte, but this was not paralleled by variation in overall Buchnera titer. In both the current study and previous research, the number of bacteriocytes has been shown to vary between A. pisum clones [19].
A moderate positive association was found between the essential amino acid requirements of the clones and the titer of Buchnera. Previous work has shown that deleterious mutations accumulate in the genome of Buchnera [20], including the amino acid biosynthesis genes of Buchnera [9], though recent studies have revealed that some classes of deleterious mutations can be overcome by translational slippage [21,22]. It is possible that the increased number of Buchnera genomes in clones with dietary amino acid requirements is a compensatory change that increases the number of functional transcripts produced from the inactivated gene.
The mechanistic basis for variation in symbiont titer is unclear, though several mechanisms may contribute. Bacteriocyte development involves the interactions of multiple genes during development, and differences in the expression of these genes or timing of expression between clones could impact titer [18]. Additionally, host lysozyme-like genes expressed in the bacteriocytes have been shown to degrade Buchnera and bacteriocytes in post-reproductive aphids [23]. While the current study focused on pre-reproductive aphids, it is possible that variation in expression of lysozyme genes between clones contributes to differences in symbiont titer. Recent theoretical work has suggested that the aphid can manipulate the metabolism of Buchnera by regulating the supply of precursor metabolites [24], and variation in the supply of metabolites may have a profound impact on Buchnera's replication and division. Variation in the DNA sequence of the genes involved in these processes, their expression, or their interaction could all impact symbiont titer.
The lack of a maternal effect in the titer of F1 clones indicates that Buchnera is not primarily responsible for the observed wide variation in titer among aphid clones. This finding is consistent with previous work indicating that Buchnera lacks basic regulatory mechanisms, as well as the absence of many genes involved in cell cycle control within the symbiont genomes. However, the differences between the Buchnera genomes of the parental clones are minimal, with a total of 9 point mutations [6], and it is possible that additional genomic differences between Buchnera strains could further affect titer. Nonetheless, we observed wide variation in titer attributable to host genotype.
Selection appears to favor clones with an intermediate titer within a 2-fold range as evidenced by the titers observed in field-collected clones. These observations, along with the 4-fold variation in F1 symbiont titer, indicates that extremely high or low symbiont levels are likely maladaptive. The extensive variation seen among F1 clones, which displayed titers much higher and much lower than either parent, suggests epistatic or dominance effects on titer, or the occurrence of high levels of heterozygosity in the parental genotypes at loci affecting titer. Because only six F1 clones were tested, from a single cross, we cannot estimate heritability of this trait. During clonal reproduction, selection may remove clones with extreme titers from the population, resulting in the intermediate titer observed in field-collected clones. However, the phenotype appears non-heritable across phases of sexual reproduction, resulting in reappearance of clones with extreme titers each spring, following the annual sexual generation of most populations. It is remarkable that symbiont titer, which is likely an important factor in aphid fitness, exhibits such variation after sexual reproduction.
| Clone | Collection Date | Collection Location | Collected by | Host Plant |
|---|---|---|---|---|
| 8.10.1 | 2001 | Cayuga Co., NY | J. Russell | Medicago sativa |
| 9.2.1 | 2001 | Cayuga Co., NY | J. Russell | Medicago sativa |
| Tuc7 | 2007 | Tucson, AZ | N. Moran | Medicago sativa |
| 5A | 1999 | Madison, WI | N. Moran | Medicago lupulina |
| Alm | 2008 | Whitman Co., WA | S. Eigenbrode | Pisum sativum |
| File | 2008 | Whitman Co., WA | S. Eigenbrode | Pisum sativum |
| Cag | 2007 | Walnut Creek, CA | N. Moran | Medicago lupulina |
| 7A | 2001 | Cayuga Co., NY | J. Russell | Medicago sativa |
| Factor | F | p |
|---|---|---|
| Titer of field-collected clones | ||
| bioA copy #1 | F9, 95 = 5.26 | < 0.0001 |
| ef1α copy #1 | F9, 95 = 5.80 | < 0.0001 |
| Clone2 | F9, 95 = 3.21 | 0.0019 |
| Titer of F1 aphid clones | ||
| Clone2 | F6, 93 = 2.90 | 0.0004 |
| [Matriline] Clone2 | F1, 93 = 0.046 | 0.83 |
| bioA copy #1 | F18, 93 = 3.13 | 0.0002 |
| ef1α copy #1 | F18, 93 = 2.13 | 0.0099 |
| Titer of 4th instar aphids | ||
| Clone2 | F3, 97 = 23.12 | < 0.0001 |
| Instar2 | F1, 97 = 24.19 | < 0.0001 |
| Clone × Instar2 | F7, 97 = 10.37 | < 0.0001 |
| Bacteriocyte counts | ||
| Field collected clones3 | F3,172 = 21.31 | <0.0001 |
| F1 clones3 | F5,106 = 8.00 | <0.0001 |
| Buchnera per bacteriocyte | ||
| Clone1 | F3, 94 = 10.24 | < 0.0001 |
| [Bacteriocyte] Clone1 | F16, 94 = 0.498 | 0.942 |
1ANOVA,2ANCOVA,3MANOVA
Acknowledgments
The authors would like to thank N. Gerardo and S. Eigenbrode for aphid collections, and K. Hammond for assistance with aphid culture maintenance. M. van Passel, P. Degnan and A. Hansen provided feedback on the manuscript. This work was supported by NSF 0723472 grant to N.A.M and a research award from the Center for Insect Science at the University of Arizona to K.J.V.
References
- Baumann, P. Biology of bacteriocyte-associated endosymbionts of plant sap-sucking insects. Annu. Rev. Microbiol. 2005, 59, 155–189. [Google Scholar]
- Sandstrom, J.; Moran, N. How nutritionally imbalanced is phloem sap for aphids? Entomol. Exp. Appl. 1999, 91, 203–210. [Google Scholar]
- Shigenobu, S.; Watanabe, H.; Hattori, M.; Sakaki, Y.; Ishikawa, H. Genome sequence of the endocellular bacterial symbiont of aphids Buchnera sp. APS. Nature 2000, 407, 81–86. [Google Scholar]
- Buchner, P. Animal Symbiosis with Plant Microorganisms; Interscience Inc.: New York, NY, USA, 1965. [Google Scholar]
- Ochman, H.; Moran, N.A. Genes lost and genes found: Evolution of bacterial pathogenesis and symbiosis. Science 2001, 292, 1096–1098. [Google Scholar]
- Moran, N.A.; McLaughlin, H.J.; Sorek, R. The dynamics and time scale of ongoing genomic erosion in symbiotic bacteria. Science 2009, 323, 379–382. [Google Scholar]
- Mira, A.; Moran, N.A. Estimating population size and transmission bottlenecks in maternally transmitted endosymbiotic bacteria. Microb. Ecol. 2002, 44, 137–143. [Google Scholar]
- Moran, N.A. Accelerated evolution and Muller's rachet in endosymbiotic bacteria. Proc. Natl. Acad. Sci. USA 1996, 93, 2873–2878. [Google Scholar]
- Vogel, K.J.; Moran, N.A. Sources of variation in dietary requirements in an obligate nutritional symbiosis. Proc. Biol. Sci. 2011, 278, 115–121. [Google Scholar]
- Moran, N.A.; Dunbar, H.E.; Wilcox, J.L. Regulation of transcription in a reduced bacterial genome: Nutrient-provisioning genes of the obligate symbiont Buchnera aphidicola. J. Bacteriol. 2005, 187, 4229–4237. [Google Scholar]
- Sandstrom, J.; Pettersson, J. Amino acid composition of phloem sap and the relation to intraspecific variation in pea aphid (Acyrthosiphon pisum) performance. J. Insect Physiol. 1994, 40, 947–955. [Google Scholar]
- Blackman, R.L.; Eastop, V.F. Aphids on the World's Crops: An Identification and Information Guide, 2nd ed.; Wiley: Chichester, UK, 2000. [Google Scholar]
- Deng, H.W.; Lynch, M. Change of genetic architecture in response to sex. Genetics 1996, 143, 203–212. [Google Scholar]
- Pfrender, M.E.; Lynch, M. Quantitative genetic variation in Daphnia: Temporal changes in genetic architecture. Evolution 2000, 54, 1502–1509. [Google Scholar]
- Oliver, K.M.; Russell, J.A.; Moran, N.A.; Hunter, M.S. Facultative bacterial symbionts in aphids confer resistance to parasitic wasps. Proc. Natl. Acad. Sci. USA 2003, 100, 1803–1807. [Google Scholar]
- Moran, N.A.; Dunbar, H.E. Sexual acquisition of beneficial symbionts in aphids. Proc. Natl. Acad. Sci. USA 2006, 103, 12803–12806. [Google Scholar]
- Fukatsu, T.; Ishikawa, H. A novel eukaryotic extracellular symbiont in an aphid, Astegopteryx styraci (Homoptera, Aphididae, Hormaphidinae). J. Insect Physiol. 1992, 38, 765–773. [Google Scholar]
- Braendle, C.; Miura, T.; Bickel, R.; Shingleton, A.W.; Kambhampati, S.; Stern, D.L. Developmental origin and evolution of bacteriocytes in the aphid-Buchnera symbiosis. PLoS Biol. 2003, 1, E21. [Google Scholar]
- Wilkinson, T.L.; Douglas, A.E. Host cell allometry and regulation of the symbiosis between pea aphids, Acyrthosiphon pisum, and bacteria Buchnera. J. Insect Physiol. 1998, 44, 629–635. [Google Scholar]
- Shigenobu, S.; Watanabe, H.; Sakaki, Y.; Ishikawa, H. Accumulation of species-specific amino acid replacements that cause loss of particular protein functions in Buchnera, an endocellular bacterial symbiont. J. Mol. Evol. 2001, 53, 377–386. [Google Scholar]
- Tamas, I.; Wernegreen, J.J.; Nystedt, B.; Kauppinen, S.N.; Darby, A.C.; Gomez-Valero, L.; Lundin, D.; Poole, A.M.; Andersson, S.G.E. Endosymbiont gene functions impaired and rescued by polymerase infidelity at poly(a) tracts. Proc. Natl. Acad. Sci. USA 2008, 105, 14934–14939. [Google Scholar]
- Wernegreen, J.J.; Kauppinen, S.N.; Degnan, P.H. Slip into something more functional: Selection maintains ancient frameshifts in homopolymeric sequences. Mol. Biol. Evol. 2010, 27, 833–839. [Google Scholar]
- Nishikori, K.; Morioka, K.; Kubo, T.; Morioka, M. Age- and morph-dependent activation of the lysosomal system and Buchnera degradation in aphid endosymbiosis. J. Insect Physiol. 2009, 55, 351–357. [Google Scholar]
- MacDonald, S.J.; Thomas, G.H.; Douglas, A.E. Genetic and metabolic determinants of nutritional phenotype in an insect-bacterial symbiosis. Mol. Ecol. 2011, 20, 2073–2084. [Google Scholar]
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Vogel, K.J.; Moran, N.A. Effect of Host Genotype on Symbiont Titer in the Aphid—Buchnera Symbiosis. Insects 2011, 2, 423-434. https://doi.org/10.3390/insects2030423
Vogel KJ, Moran NA. Effect of Host Genotype on Symbiont Titer in the Aphid—Buchnera Symbiosis. Insects. 2011; 2(3):423-434. https://doi.org/10.3390/insects2030423
Chicago/Turabian StyleVogel, Kevin J., and Nancy A. Moran. 2011. "Effect of Host Genotype on Symbiont Titer in the Aphid—Buchnera Symbiosis" Insects 2, no. 3: 423-434. https://doi.org/10.3390/insects2030423
APA StyleVogel, K. J., & Moran, N. A. (2011). Effect of Host Genotype on Symbiont Titer in the Aphid—Buchnera Symbiosis. Insects, 2(3), 423-434. https://doi.org/10.3390/insects2030423




