Proteasome Subunits Regulate Reproduction in Nilaparvata lugens and the Transovarial Transmission of Its Yeast-like Symbionts
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Insect Rearing
2.2. cDNA Synthesis
2.3. RNA Interference
2.4. Real-Time Quantitative PCR Analysis
2.5. Dissection Observations and Fertility Analysis
2.6. Immunofluorescence
3. Results
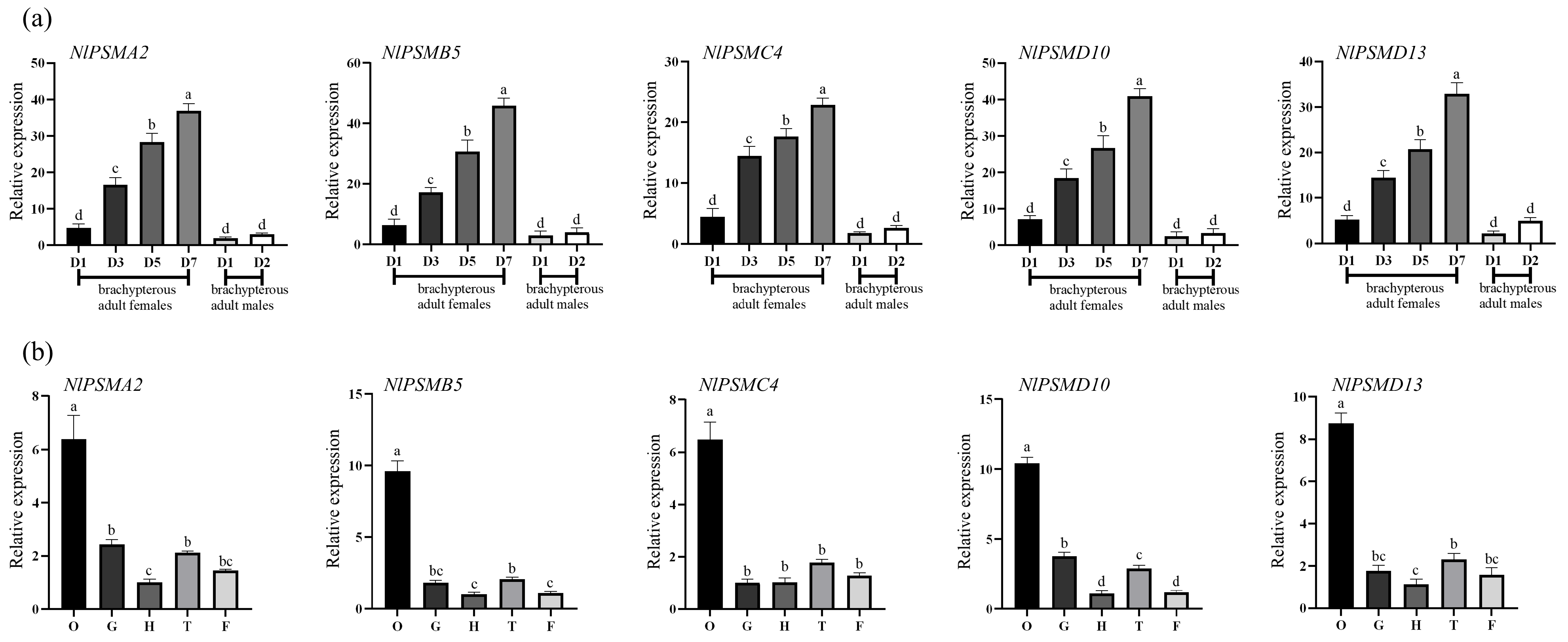
3.1. Developmental and Tissue-Specific Expression of NlPSMA2, NlPSMB5, NlPSMC4, NlPSMD10, and NlPSMD13
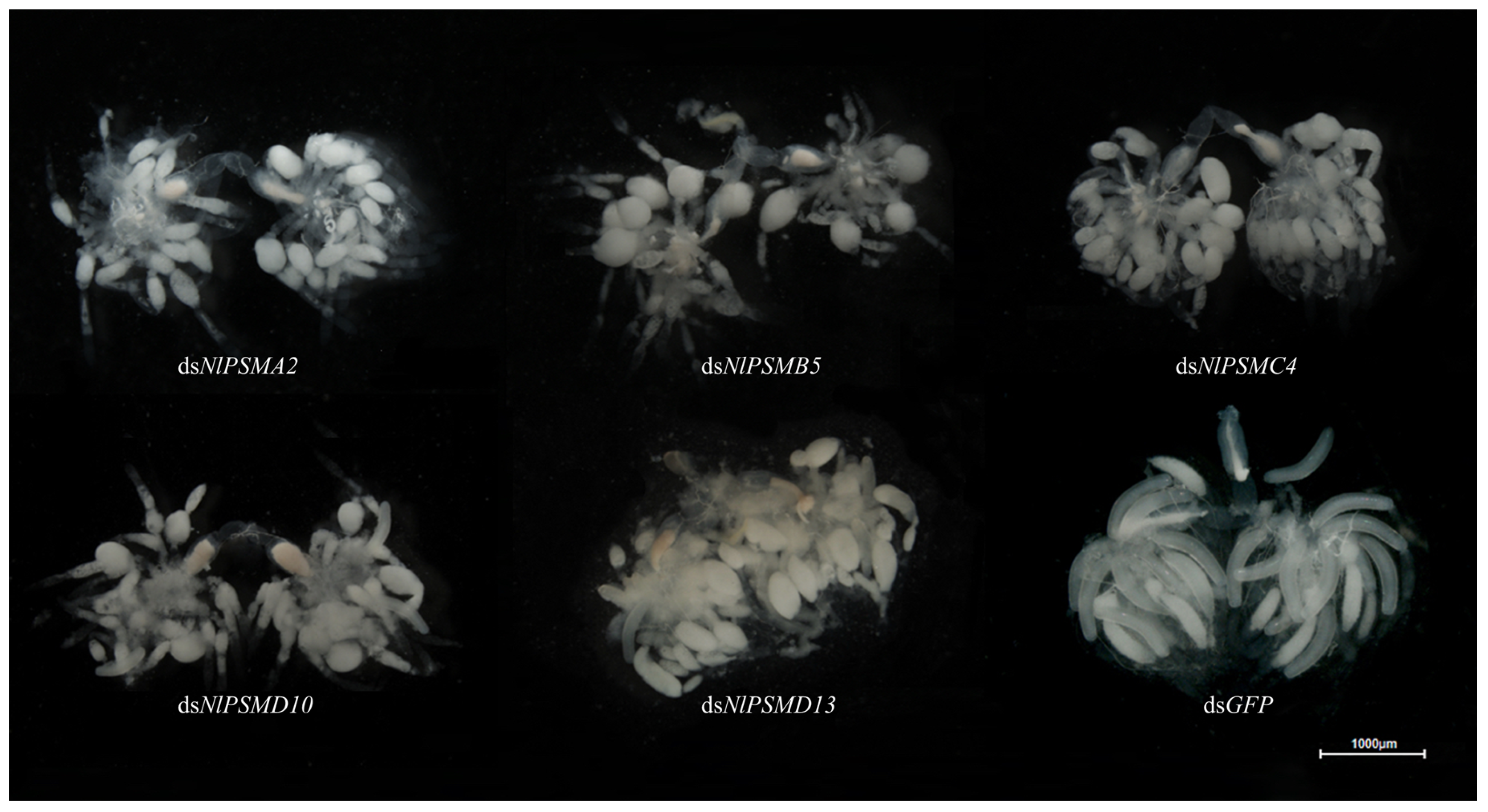
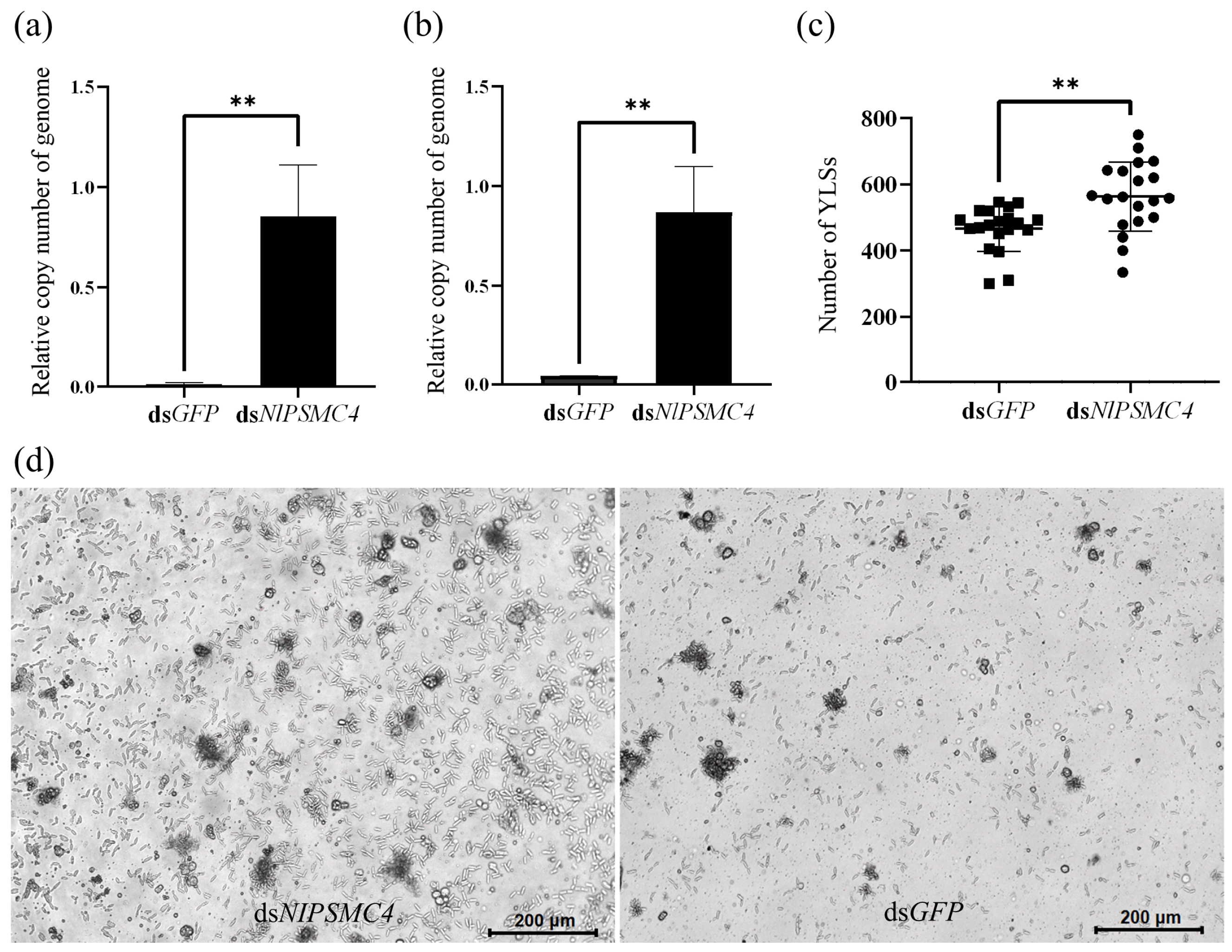
3.2. Function Validation of NlPSMA2, NlPSMB5, NlPSMC4, NlPSMD10, and NlPSMD13 by RNA Interference
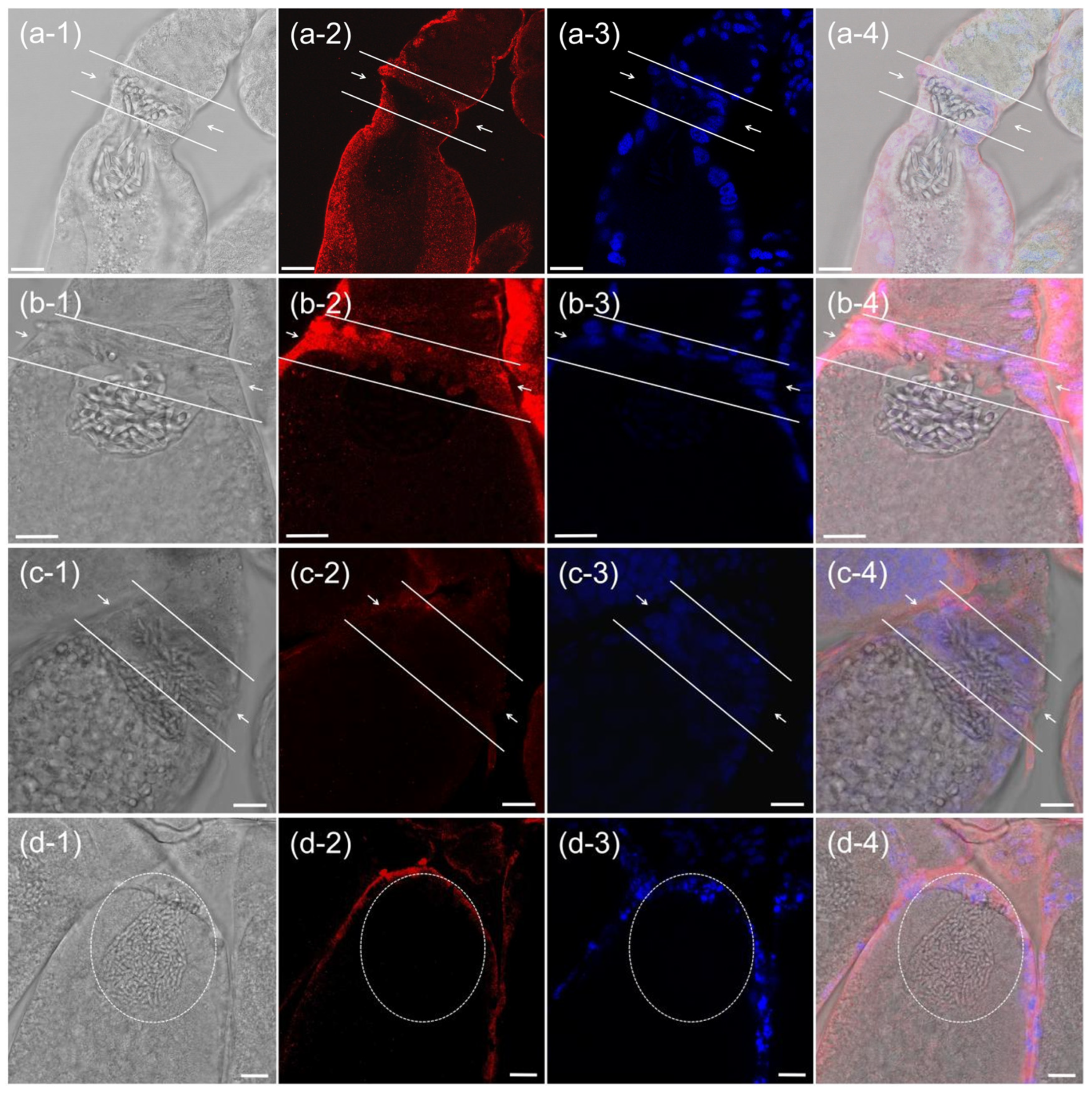
3.3. Immunofluorescence Analysis of PSMC4 Expression
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Iamba, K.; Dono, D. A Review on Brown Planthopper (Nilaparvata lugens Stål), a Major Pest of Rice in Asia and Pacific. Asian J. Res. Crop Sci. 2021, 6, 7–19. [Google Scholar] [CrossRef]
- Jeevanandham, N.; Raman, R.; Ramaiah, D.; Senthilvel, V.; Mookaiah, S.; Jegadeesan, R. Rice: Nilaparvata Lugens Stal Interaction—Current Status and Future Prospects of Brown Planthopper Management. J. Plant Dis. Prot. 2023, 130, 125–141. [Google Scholar] [CrossRef]
- Kumar, S.; Singh, H.; Patel, A.; Patel, J.N.; Kant, C. Brown Plant Hopper, Nilaparvata lugens (Stal) (Insecta: Delphacidae) a Major Insect of Rice in India: A Review. J. Entomol. Res. 2022, 46, 333–338. [Google Scholar] [CrossRef]
- Fujita, D.; Kohli, A.; Horgan, F.G. Rice Resistance to Planthoppers and Leafhoppers. Crit. Rev. Plant Sci. 2013, 32, 162–191. [Google Scholar] [CrossRef]
- Ling, Y.; Weilin, Z. Genetic and Biochemical Mechanisms of Rice Resistance to Planthopper. Plant Cell Rep. 2016, 35, 1559–1572. [Google Scholar] [CrossRef] [PubMed]
- Xue, J.; Zhou, X.; Zhang, C.X.; Yu, L.L.; Fan, H.W.; Wang, Z.; Xu, H.J.; Xi, Y.; Zhu, Z.R.; Zhou, W.W.; et al. Genomes of the Rice Pest Brown Planthopper and Its Endosymbionts Reveal Complex Complementary Contributions for Host Adaptation. Genome Biol. 2014, 15, 521. [Google Scholar] [CrossRef]
- Fahad, S.; Nie, L.; Hussain, S.; Khan, F.; Khan, F.A.; Saud, S.; Muhammad, H.; Li, L.; Liu, X.; Tabassum, A. Rice Pest Management and Biological Control. In Sustainable Agriculture Reviews: Cereals; Springer: Cham, Switzerland, 2015; pp. 85–106. [Google Scholar]
- Norton, G.W.; Heong, K.L.; Johnson, D.; Savary, S. Rice Pest Management: Issues and Opportunities. In Rice in the Global Economy: Strategic Research and Policy Issues for Food Security; IRRI: Los Banos, Philippines, 2010. [Google Scholar]
- Padmakumari, A.P.; Suneetha, K.; Sundaram, R.M. Current Status of Host PlantResistance to Insects in Rice and Future Perspectives. In PlantResistance to Insects in Major Field Crops; Springer Nature: Singapore, 2024; pp. 69–122. [Google Scholar]
- Yu, J.B.; Lv, X.; Liu, Q.; Tu, J.Y.; Yu, X.P.; Xu, Y.P. Death-Associated Protein-1 Plays a Role in the Reproductive Development of Nilaparvata lugens and the Transovarial Transmission of Its Yeast-Like Symbiont. Insects 2024, 15, 425. [Google Scholar] [CrossRef]
- Pan, H.B.; Li, M.Y.; Wu, W.; Wang, Z.L.; Yu, X.P. Host-Plant Induced Shifts in Microbial Community Structure in Small Brown Planthopper, Laodelphax striatellus (Homoptera: Delphacidae). J. Econ. Entomol. 2021, 114, 937–946. [Google Scholar] [CrossRef]
- Horgan, F.G.; Peñalver Cruz, A.; Arida, A.; Ferrater, J.B.; Bernal, C.C. Adaptation by the Brown Planthopper to Resistant Rice: A Test of Female-Derived Virulence and the Role of Yeast-like Symbionts. Insects 2021, 12, 908. [Google Scholar] [CrossRef]
- Nan, G.H.; Xu, Y.P.; Yu, Y.W.; Zhao, C.X.; Zhang, C.X.; Yu, X.P. Oocyte Vitellogenesis Triggers the Entry of Yeast-like Symbionts into the Oocyte of Brown Planthopper (Hemiptera: Delphacidae). Ann. Entomol. Soc. Am. 2016, 109, 753–758. [Google Scholar] [CrossRef]
- Tanahashi, N.; Suzuki, M.; Fujiwara, T.; Takahashi, E.; Shimbara, N.; Chung, C.H.; Tanaka, K. Chromosomal Localization and Immunological Analysis of a Family of Human 26S Proteasomal ATPases. Biochem. Biophys. Res. Commun. 1998, 243, 229–232. [Google Scholar] [CrossRef]
- Huo, L.J.; Fan, H.Y.; Zhong, Z.S.; Chen, D.Y.; Schatten, H.; Sun, Q.Y. Ubiquitin–Proteasome Pathway Modulates Mouse Oocyte Meiotic Maturation and Fertilization via Regulation of MAPK Cascade and Cyclin B1 Degradation. Mech. Dev. 2004, 121, 1275–1287. [Google Scholar] [CrossRef] [PubMed]
- Higuchi, C.; Shimizu, N.; Shin, S.W.; Morita, K.; Nagai, K.; Anzai, M.; Matsumoto, K. Ubiquitin-Proteasome System Modulates Zygotic Genome Activation in Early Mouse Embryos and Influences Full-term Development. J. Reprod. Dev. 2018, 64, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Sakao, Y.; Kawai, T.; Takeuchi, O.; Copeland, N.G.; Gilbert, D.J.; Jenkins, N.A.; Takeda, K.; Akira, S. Mouse Proteasomal ATPases Psmc3 and Psmc4: Genomic Organization and Gene Targeting. Genomics 2000, 67, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Faria-Reis, A.; Santos-Araújo, S.; Pereira, J.; Rios, T.; Majerowicz, D.; Gondim, K.C.; Ramos, I. Silencing of the 20S Proteasomal Subunit-A6 Triggers Full Oogenesis Arrest and Increased mRNA Levels of the Selective Autophagy Adaptor Protein p62/SQSTM1 in the Ovary of the Vector Rhodnius prolixus. PLoS Neglected Trop. Dis. 2023, 17, e0011380. [Google Scholar] [CrossRef]
- Wang, W.; Yang, R.R.; Peng, L.Y.; Zhang, L.; Yao, Y.L.; Bao, Y.Y. Proteolytic Activity of the Proteasome is Required for Female Insect Reproduction. Open Biol. 2021, 11, 200251. [Google Scholar] [CrossRef]
- Cheng, X.; Wang, W.; Zhang, L.; Yang, R.R.; Ma, Y.; Bao, Y.Y. ATPase Subunits of the 26S Proteasome Are Important for Oocyte Maturation in the Brown Planthopper. Insect Mol. Biol. 2022, 31, 317–333. [Google Scholar] [CrossRef]
- Ji, J.L.; Han, S.J.; Zhang, R.J.; Yu, J.B.; Li, Y.B.; Yu, X.P.; Liu, G.F.; Xu, Y.P. Inter-Alpha-Trypsin Inhibitor Heavy Chain 4 Plays an Important Role in the Development and Reproduction of Nilaparvata Lugens. Insects 2022, 13, 303. [Google Scholar] [CrossRef]
- Cuanalo-Contreras, K.; Mukherjee, A.; Soto, C. Role of Protein Misfolding and Proteostasis Deficiency in Protein Misfolding Diseases and Aging. Int. J. Cell Biol. 2013, 2013, 638083. [Google Scholar] [CrossRef]
- Nandi, D.; Tahiliani, P.; Kumar, A.; Chandu, D. The Ubiquitin-Proteasome System. J. Biosci. 2006, 31, 137–155. [Google Scholar] [CrossRef]
- Li, X.J.; Li, S. Proteasomal Dysfunction in Aging and Huntington Disease. Neurobiol. Dis. 2011, 43, 4–8. [Google Scholar] [CrossRef]
- Adams, J. The Proteasome: Structure, Function, and Role in the Cell. Cancer Treat. Rev. 2003, 29, 3–9. [Google Scholar] [CrossRef]
- Liu, K.; Zhang, S.; Gong, Y.; Zhu, P.; Shen, W.; Zhang, Q. PSMC4 Promotes Prostate Carcinoma Progression by Regulating the CBX3–EGFR-PI3K-AKT-mTOR Pathway. J. Cell. Mol. Med. 2023, 27, 2437–2447. [Google Scholar] [CrossRef] [PubMed]
- Kao, T.J.; Wu, C.C.; Phan, N.N.; Liu, Y.H.; Ta, H.D.K.; Anuraga, G.; Wu, Y.F.; Lee, K.H.; Chuang, J.Y.; Wang, C.Y. Prognoses and Genomic Analyses of Proteasome 26S Subunit, ATPase (PSMC) Family Genes in Clinical Breast Cancer. Aging 2021, 13, 17970. [Google Scholar] [CrossRef]
- Hossain, S.; Lash, E.; Veri, A.O.; Cowen, L.E. Functional Connections between Cell Cycle and Proteostasis in the Regulation of Candida albicans Morphogenesis. Cell Rep. 2021, 34, 108781. [Google Scholar] [CrossRef]
- Chandrasekaran, A.; Adkins, L.J.; Seltzer, H.M.; Pant, K.; Tryban, S.T.; Molloy, C.T.; Weinberg, J.B. Age-Dependent Effects of Immunoproteasome Deficiency on Mouse Adenovirus Type 1 Pathogenesis. J. Virol. 2019, 93, e00569-19. [Google Scholar] [CrossRef]

| Primers | Primer Sequence (5′-3′) |
|---|---|
| dsNlPSMA2-F | GGATCCTAATACGACTCACTATAGGGATCGGCCCTATCTGTTC |
| dsNlPSMA2-R | GGATCCTAATACGACTCACTATAGGGCATCACACACACCCACC |
| dsNlPSMB5-F | GGATCCTAATACGACTCACTATAGGGCATGGGCCTCTCCATGG |
| dsNlPSMB5-R | GGATCCTAATACGACTCACTATAGGCTCGGAAATTTTGATCCA |
| dsNlPSMC4-F | GGATCCTAATACGACTCACTATAGGCAACTAGAGTTTTTGGCT |
| dsNlPSMC4-R | GGATCCTAATACGACTCACTATAGGGCATTGCTGTGTTTGTGC |
| dsNlPSMD10-F | GGATCCTAATACGACTCACTATAGGGCATTTCGAAATCGTGAA |
| dsNlPSMD10-R | GGATCCTAATACGACTCACTATAGGGCATGAGACCTAGTGAGG |
| dsNlPSMD13-F | GGATCCTAATACGACTCACTATAGGGCTCAAATCGAAGAACTC |
| dsNlPSMD13-R | GGATCCTAATACGACTCACTATAGGGAGAACTTTGCATAGTGC |
| dsGFP-F | GGATCCTAATACGACTCACTATAGGGATACGTGCAGGAGAGGAC |
| dsGFP-R | GGATCCTAATACGACTCACTATAGGGCAGATTGTGTGGACAGG |
| Primers | Primer Sequence (5′-3′) |
|---|---|
| NlPSMA2-qF | CACCGTCCGTAGGAATAAAAGC |
| NlPSMA2-qR | GGACCCATACCACTGTAGACCATT |
| NlPSMB5-qF | GCTTTAGCAGATGTATGTGGAATG |
| NlPSMB5-qR | ACCTGATTTTGACGGGTTTTC |
| NlPSMC4-qF | TGGAACTGCCGCTCACTC |
| NlPSMC4-qR | CCCTCGCCCAGGTATTTT |
| NlPSMD10-qF | AAGCCGTTCCGAAGTAGC |
| NlPSMD10-qR | CAGCCAAATCAAGAGGTGTT |
| NlPSMD13-qF | CTCTTCGCTATCTCGGCTGTA |
| NlPSMD13-qR | GCCACTCATTGGGTGTATTTT |
| Entomomyces delphacidicola-qF | TCCCTCTGTGGAACCCCA |
| Entomomyces delphacidicola-qR | GGCGGTCCTAGAAACCAACA |
| Moesziomyces sp.-qF | TGATGCCCCTTAGATGTTCCG |
| Moesziomyces sp.-qR | CACAAGTTTACCCAGTCATTTCG |
| Nl18S-qF | GTAACCCGCTGAACCTCC |
| Nl18S-qR | GTCCGAAGACCTCACTAAATCA |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lv, X.; Tu, J.-Y.; Liu, Q.; Wu, Z.-Q.; Lin, C.; Zhou, T.; Yu, X.-P.; Xu, Y.-P. Proteasome Subunits Regulate Reproduction in Nilaparvata lugens and the Transovarial Transmission of Its Yeast-like Symbionts. Insects 2025, 16, 895. https://doi.org/10.3390/insects16090895
Lv X, Tu J-Y, Liu Q, Wu Z-Q, Lin C, Zhou T, Yu X-P, Xu Y-P. Proteasome Subunits Regulate Reproduction in Nilaparvata lugens and the Transovarial Transmission of Its Yeast-like Symbionts. Insects. 2025; 16(9):895. https://doi.org/10.3390/insects16090895
Chicago/Turabian StyleLv, Xin, Jia-Yu Tu, Qian Liu, Zhi-Qiang Wu, Chen Lin, Tao Zhou, Xiao-Ping Yu, and Yi-Peng Xu. 2025. "Proteasome Subunits Regulate Reproduction in Nilaparvata lugens and the Transovarial Transmission of Its Yeast-like Symbionts" Insects 16, no. 9: 895. https://doi.org/10.3390/insects16090895
APA StyleLv, X., Tu, J.-Y., Liu, Q., Wu, Z.-Q., Lin, C., Zhou, T., Yu, X.-P., & Xu, Y.-P. (2025). Proteasome Subunits Regulate Reproduction in Nilaparvata lugens and the Transovarial Transmission of Its Yeast-like Symbionts. Insects, 16(9), 895. https://doi.org/10.3390/insects16090895




