Distribution of the Main Apis mellifera Mitochondrial DNA Lineages in Italy Assessed Using an Environmental DNA Approach
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Honey Samples
2.2. DNA Extraction
2.3. PCR and Sequencing Analyses
2.4. Data Analyses
3. Results
3.1. Description of the Main Honey-Derived Mitotype Patterns by Italian Region
3.2. Comparison of Mitotype Results between Honey Samples Produced by the Same Beekeepers
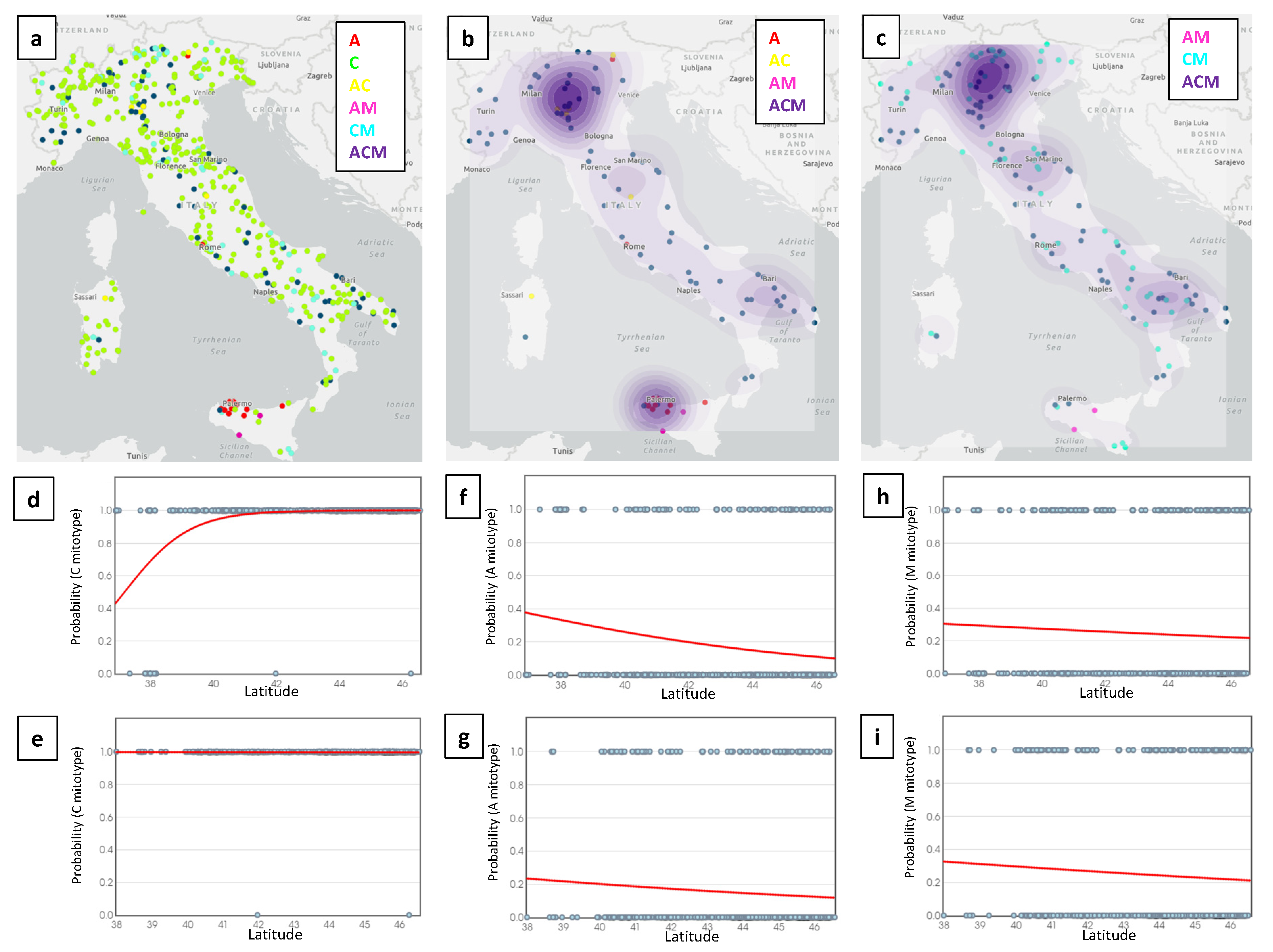
3.3. Distribution of Lineages over Geographic Coordinates
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Acknowledgments
Conflicts of Interest
References
- Jensen, A.B.; Palmer, K.A.; Boomsma, J.J.; Pedersen, B.V. Varying degrees of Apis mellifera ligustica introgression in protected populations of the black honeybee, Apis mellifera mellifera, in northwest Europe. Mol. Ecol. 2005, 14, 93–106. [Google Scholar] [CrossRef] [PubMed]
- Strange, J.P.; Garnery, L.; Sheppard, W.S. Morphological and molecular characterization of the Landes honey bee (Apis mellifera L.) ecotype for genetic conservation. J. Insect Conserv. 2008, 12, 527–537. [Google Scholar] [CrossRef]
- De la Rúa, P.; Jaffé, R.; Dall’Olio, R.; Muñoz, I.; Serrano, J. Biodiversity, conservation and current threats to European honeybees. Apidologie 2009, 40, 263–284. [Google Scholar] [CrossRef]
- Oleksa, A.; Chybicki, I.; Tofilski, A.; Burczyk, J. Nuclear and mitochondrial patterns of introgression into native dark bees (Apis mellifera mellifera) in Poland. J. Apic. Res. 2011, 50, 116–129. [Google Scholar] [CrossRef]
- Pinto, A.M.; Henriques, D.; Chávez-Galarza, J.; Kryger, P.; Garnery, L.; van der Zee, R.; Dahle, B.; Soland-Reckeweg, G.; de la Rúa, P.; Dall’Olio, R.; et al. Genetic integrity of the Dark European honey bee (Apis mellifera mellifera) from protected populations: A genome-wide assessment using SNPs and mtDNA sequence data. J. Apic. Res. 2014, 53, 269–278. [Google Scholar] [CrossRef]
- Fontana, P.; Costa, C.; Di Prisco, G.; Ruzzier, E.; Annoscia, D.; Battisti, A.; Caoduro, G.; Carpana, E.; Contessi, A.; Dal Lago, A.; et al. Appeal for biodiversity protection of native honey bee subspecies of Apis mellifera in Italy (San Michele all’Adige declaration. Bull. Insectol. 2018, 71, 257–271. [Google Scholar]
- Hassett, J.; Browne, K.A.; McCormack, G.P.; Moore, E.; Society, N.I.H.B.; Soland, G.; Geary, M. A significant pure population of the dark European honey bee (Apis mellifera mellifera) remains in Ireland. J. Apic. Res. 2018, 57, 337–350. [Google Scholar] [CrossRef]
- Meixner, M.D.; Costa, C.; Kryger, P.; Hatjina, F.; Bouga, M.; Ivanova, E.; Büchler, R. Conserving diversity and vitality for honey bee breeding. J. Apic. Res. 2010, 49, 85–92. [Google Scholar] [CrossRef]
- Requier, F.; Garnery, L.; Kohl, P.L.; Njovu, H.K.; Pirk, C.W.; Crewe, R.M.; Steffan-Dewenter, I. The conservation of native honey bees is crucial. Trends Ecol. Evol. 2019, 34, 789–798. [Google Scholar] [CrossRef]
- van Engelsdorp, D.; Meixner, M.D. A historical review of managed honey bee populations in Europe and the United States and the factors that may affect them. J. Invertebr. Pathol. 2010, 103, S80–S95. [Google Scholar] [CrossRef] [PubMed]
- De La Rúa, P.; Jiménez, Y.; Galián, J.; Serrano, J. Evaluation of the biodiversity of honey bee (Apis mellifera) populations from eastern Spain. J. Apic. Res. 2004, 43, 162–166. [Google Scholar] [CrossRef]
- De la Rúa, P.; Jaffé, R.; Muñoz, I.; Serrano, J.; Moritz, R.F.A.A.; Kraus, F.B. Conserving genetic diversity in the honeybee: Comments on Harpur et al. (2012). Mol. Ecol. 2013, 22, 3208–3210. [Google Scholar] [CrossRef]
- Muñoz, I.; De la Rúa, P. Wide genetic diversity in Old World honey bees threaten by introgression. Apidologie 2021, 52, 200–217. [Google Scholar] [CrossRef]
- Potts, S.G.; Roberts, S.P.M.; Dean, R.; Marris, G.; Brown, M.A.; Jones, R.; Neumann, P.; Settele, J. Declines of managed honey bees and beekeepers in Europe. J. Apic. Res. 2010, 49, 15–22. [Google Scholar] [CrossRef]
- Chauzat, M.P.; Cauquil, L.; Roy, L.; Franco, S.; Hendrikx, P.; Ribière-Chabert, M. Demographics of the European apicultural industry. PLoS ONE 2013, 8, e79018. [Google Scholar] [CrossRef] [PubMed]
- Themudo, G.E.; Rey-Iglesia, A.; Tascón, L.R.; Jensen, A.B.; da Fonseca, R.R.; Campos, P.F. Declining genetic diversity of European honeybees along the twentieth century. Sci. Rep. 2020, 10, 10520. [Google Scholar] [CrossRef] [PubMed]
- Le Conte, Y.; Navajas, M. Climate change: Impact on honey bee populations and diseases. Rev. Sci. Tech. Off. Int. Epiz. 2008, 27, 499–510. [Google Scholar]
- Reddy, P.R.; Verghese, A.; Rajan, V.V. Potential impact of climate change on honeybees (Apis spp.) and their pollination services. Pest. Man. Hort. Ecosyst. 2012, 18, 121–127. [Google Scholar]
- Flores, J.M.; Gil-Lebrero, S.; Gámiz, V.; Rodríguez, M.I.; Ortiz, M.A.; Quiles, F.J. Effect of the climate change on honey bee colonies in a temperate Mediterranean zone assessed through remote hive weight monitoring system in conjunction with exhaustive colonies assessment. Sci. Total Environ. 2019, 653, 1111–1119. [Google Scholar] [CrossRef] [PubMed]
- Vercelli, M.; Novelli, S.; Ferrazzi, P.; Lentini, G.; Ferracini, C. A Qualitative analysis of beekeepers’ perceptions and farm management adaptations to the impact of climate change on honey bees. Insects 2021, 12, 228. [Google Scholar] [CrossRef] [PubMed]
- Alpatov, W.W. Biometrical studies on variation and races of the honey bee (Apis mellifera L.). Q. Rev. Biol. 1929, 4, 1–58. [Google Scholar] [CrossRef]
- Goetze, G. Variabilitats- und Ziichtungsstudien an der Honigbiene mit besonderer Beriick- sichtigung der Lan-griilligkeit. Arch. Bienenkd. 1930, 11, 135–274. [Google Scholar]
- Tomassone, R.; Fresnaye, J. Étude d’une méthode biométrique et statistique permettant la discrimination et la clas-sification de populations d’abeilles (Apis mellifica L.). Apidologie 1971, 2, 49–65. [Google Scholar] [CrossRef][Green Version]
- Ruttner, F.; Tassencourt, L.; Louveaux, J. Biometrical-statistical analysis of the geographic variability of Apis mellifera L. Apidologie 1978, 9, 363–381. [Google Scholar] [CrossRef]
- Ruttner, F. Biogeography and Taxonomy of Honeybees; Springer: Berlin, Germany, 1988. [Google Scholar] [CrossRef]
- Cornuet, J.M.; Garnery, L. Mitochondrial DNA variability in honeybees and its phylogeographic implications. Apidologie 1991, 22, 627–642. [Google Scholar] [CrossRef]
- Garnery, L.; Cornuet, J.M.; Solignac, M. Evolutionary history of the honey bee Apis mellifera inferred from mitochondrial DNA analysis. Mol. Ecol. 1992, 1, 145–154. [Google Scholar] [CrossRef] [PubMed]
- Arias, M.C.; Sheppard, W.S. Molecular phylogenetics of honey bee subspecies (Apis mellifera L.) inferred from mitochondrial DNA sequence. Mol. Phylogenetics Evol. 1996, 5, 557–566. [Google Scholar] [CrossRef] [PubMed]
- Wallberg, A.; Han, F.; Wellhagen, G.; Dahle, B.; Kawata, M.; Haddad, N.; Simões, Z.L.P.; Allsopp, M.H.; Kandemir, I.; De la Rúa, P.; et al. A worldwide survey of genome sequence variation provides insight into the evolutionary history of the honeybee Apis mellifera. Nat. Genet. 2014, 46, 1081–1088. [Google Scholar] [CrossRef] [PubMed]
- Tihelka, E.; Cai, C.; Pisani, D.; Donoghue, P.C.J. Mitochondrial genomes illuminate the evolutionary history of the Western honey bee (Apis mellifera). Sci. Rep. 2020, 10, 14515. [Google Scholar] [CrossRef] [PubMed]
- Cridland, J.M.; Tsutsui, N.D.; Ramírez, S.R. The complex demographic history and evolutionary origin of the western honey bee, Apis mellifera. Genome Biol. Evol. 2017, 9, 457–472. [Google Scholar] [CrossRef]
- Smith, D.R.; Palopoli, M.F.; Taylor, B.R.; Garnery, L.; Cornuet, J.-M.; Solignac, M.; Brown, W.M. Geographical overlap of two mitochondrial genomes in Spanish honeybees (Apis mellifera iberica). J. Hered. 1991, 82, 96–100. [Google Scholar] [CrossRef] [PubMed]
- Miguel, I.; Iriondo, M.; Garnery, L.; Sheppard, W.S.; Estonba, A. Gene flow within the M evolutionary lineage of Apis mellifera: Role of the Pyrenees, isolation by distance and post-glacial re-colonization routes in the western Europe. Apidologie 2007, 38, 141–155. [Google Scholar] [CrossRef]
- Garnery, L.; Mosshine, E.H.; Oldroyd, B.P.; Cornuet, J.M. Mitochondrial DNA variation in Moroccan and Spanish honey bee populations. Mol. Ecol. 1995, 4, 465–472. [Google Scholar] [CrossRef]
- Garnery, L.; Franck, P.; Baudry, E.; Vautrin, D.; Cornuet, J.M.; Solignac, M. Genetic diversity of the west European honey bee (Apis mellifera mellifera and A. m. iberica) I. Mitochondrial DNA. Genet. Sel. Evol. 1998, 30, S31–S47. [Google Scholar] [CrossRef]
- Cánovas, F.; De la Rúa, P.; Serrano, J.; Galián, J. Geographical patterns of mitochondrial DNA variation in Apis mellifera iberiensis (Hymenoptera: Apidae). J. Zool. Syst. Evol. Res. 2007, 46, 24–30. [Google Scholar] [CrossRef]
- Pinto, M.A.; Henriques, D.; Neto, M.; Guedes, H.; Muñoz, I.; Azevedo, J.C.; De la Rúa, P. Maternal diversity patterns of Ibero-Atlantic populations reveal further complexity of Iberian honeybees. Apidologie 2013, 44, 430–439. [Google Scholar] [CrossRef]
- Chávez-Galarza, J.; Garnery, L.; Henriques, D.; Neves, C.J.; Loucif-Ayad, W.; Jonhston, J.S.; Pinto, M.A. Mitochondrial DNA variation of Apis mellifera iberiensis: Further insights from a large-scale study using sequence data of the tRNA leu-cox2 intergenic region. Apidologie 2017, 48, 533–544. [Google Scholar] [CrossRef]
- Sinacori, A.; Rinderer, T.E.; Lancaster, V.; Sheppard, W.S. A morphological and mitochondrial assessment of Apis mellifera from Palermo, Italy. Apidologie 1998, 29, 481–490. [Google Scholar] [CrossRef]
- Franck, P.; Garnery, L.; Celebrano, G.; Solignac, M.; Cornuet, J.M. Hybrid origins of honeybees from Italy (Apis mellifera ligustica) and sicily (A. m. sicula). Mol. Ecol. 2000, 9, 907–921. [Google Scholar] [CrossRef]
- Muñoz, I.; Dall’Olio, R.; Lodesani, M.; De la Rúa, P. Estimating introgression in Apis mellifera siciliana populations: Are the conservation islands really effective? Insect Conserv. Divers. 2014, 7, 563–571. [Google Scholar] [CrossRef]
- Franck, P.; Garnery, L.; Solignac, M.; Cornuet, J.M. The origin of west European subspecies of honeybees (Apis mellifera): New insights from microsatellite and mitochondrial data. Evolution 1998, 52, 1119–1134. [Google Scholar] [CrossRef]
- Rortais, A.; Arnold, G.; Alburaki, M.; Legout, H.; Garnery, L. Review of the Dra I COI-COII test for the conservation of the black honeybee (Apis mellifera mellifera). Conserv. Genet. Resour. 2011, 3, 383–391. [Google Scholar] [CrossRef]
- Sušnik, S.; Kozmus, P.; Poklukar, J.; Meglic, V. Molecular characterisation of indigenous Apis mellifera carnica in Slovenia. Apidologie 2004, 35, 623–636. [Google Scholar] [CrossRef]
- Muñoz, I.; Dall’Olio, R.; Lodesani, M.; De la Rúa, P. Population genetic structure of coastal Croatian honeybees (Apis mellifera carnica). Apidologie 2009, 40, 617–626. [Google Scholar] [CrossRef]
- Nedić, N.; Stanisavljević, L.; Mladenović, M.; Stanisavljević, J. Molecular characterization of the honeybee Apis mellifera carnica in Serbia. Arch. Biol. Sci. 2009, 61, 587–598. [Google Scholar] [CrossRef]
- Vecchi, A. Sulla distribuzione geografica dell’Apis mellifica ligustica Spin. in Italia. Bollettino del Laboratorio di Zoologia Generale e Agraria Della R. Scuola Superiore d’agricoltura in Portici 1927, 20, 150–168. [Google Scholar]
- Marletto, F.; Manino, A.; Pedrini, P. Indagini biometriche su popolazioni di Apis mellifera L. delle Alpi occidentali. L’Apicoltore Moderno 1984, 75, 213–223. [Google Scholar]
- Marletto, F.; Manino, A.; Pedrini, P. Integrazione fra sottospecie di Apis mellifera L. in Liguria. L’Apicoltore Moderno 1984, 75, 159–163. [Google Scholar]
- Nazzi, F. Morphometric analysis of honey bees from an area of racial hybridization in northeastern Italy. Apidologie 1992, 23, 89–96. [Google Scholar] [CrossRef]
- Utzeri, V.J.; Ribani, A.; Fontanesi, L. Authentication of honey based on a DNA method to differentiate Apis mellifera subspecies: Application to Sicilian honey bee (A. m. siciliana) and Iberian honey bee (A. m. iberiensis) honeys. Food Control. 2018, 91, 294–301. [Google Scholar] [CrossRef]
- Utzeri, V.J.; Ribani, A.; Schiavo, G.; Bertolini, F.; Bovo, S.; Fontanesi, L. Application of next generation semiconductor based sequencing to detect the botanical composition of monofloral, polyfloral and honeydew honey. Food Control. 2018, 86, 342–349. [Google Scholar] [CrossRef]
- Utzeri, V.J.; Schiavo, G.; Ribani, A.; Bertolini, F.; Bovo, S.; Fontanesi, L. A next generation sequencing approach for targeted Varroa destructor (Acari: Varroidae) mitochondrial DNA analysis based on honey derived environmental DNA. J. Invertebr. Pathol. 2019, 161, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Ribani, A.; Utzeri, V.J.; Taurisano, V.; Fontanesi, L. Honey as a source of environmental DNA for the detection and monitoring of honey bee pathogens and parasites. Vet. Sci. 2020, 7, 113. [Google Scholar] [CrossRef]
- Ribani, A.; Utzeri, V.J.; Taurisano, V.; Galuppi, R.; Fontanesi, L. Analysis of honey environmental DNA indicates that the honey bee (Apis mellifera L.) trypanosome parasite Lotmaria passim is widespread in the apiaries of the North of Italy. J. Invertebr. Pathol. 2021, 107628. [Google Scholar] [CrossRef] [PubMed]
- Cornuet, J.M.; Garnery, L.; Solignac, M. Putative origin and function of the intergenic region between COI and COII of Apis mellifera L. mitochondrial DNA. Genetics 1991, 128, 393–403. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Meixner, M.; Pinto, M.; Bouga, M.; Kryger, P.; Ivanova, E.; Fuchs, S. Standard methods for characterizing subspecies and ecotypes of Apis mellifera. J. Apic. Res. 2013, 52, 1–28. [Google Scholar] [CrossRef]
- Péntek-Zakar, E.; Oleksa, A.; Borowik, T.; Kusza, S. Population structure of honey bees in the Carpathian Basin (Hungary) confirms introgression from surrounding subspecies. Ecol. Evol. 2015, 5, 5456–5467. [Google Scholar] [CrossRef]
- Oleksa, A.; Kusza, S.; Tofilski, A. Mitochondrial DNA suggests the introduction of honeybees of African ancestry to East-Central Europe. Insects 2021, 12, 410. [Google Scholar] [CrossRef]
- Meixner, M.D.; Sheppard, W.S.; Poklukar, J. Asymmetrical distribution of a mitochondrial DNA polymorphism between 2 introgressing honey bee subspecies. Apidologie 1993, 24, 147–153. [Google Scholar] [CrossRef]
- Hall, H.G.; Smith, D.R. Distinguishing African and European honeybee matrilines using amplified mitochondrial DNA. Proc. Natl. Acad. Sci. USA 1991, 88, 4548–4552. [Google Scholar] [CrossRef]
- Abrahamovich, A.H.; Atela, O.; De la Rúa, P.; Galián, J. Assessment of the mitochondrial origin of honey bees from Argentina. J. Apic. Res. 2007, 46, 191–194. [Google Scholar] [CrossRef]
- Spötter, A.; Gupta, P.; Nürnberg, G.; Reinsch, N.; Bienefeld, K. Development of a 44K SNP assay focussing on the analysis of a varroa-specific defence behaviour in honey bees (Apis mellifera carnica). Mol. Ecol. Res. 2012, 12, 323–332. [Google Scholar] [CrossRef] [PubMed]
- Muñoz, I.; Henriques, D.; Johnston, J.S.; Chávez-Galarza, J.; Kryger, P.; Pinto, M.A. Reduced SNP panels for genetic identification and introgression analysis in the dark honey bee (Apis mellifera mellifera). PLoS ONE 2015, 10, e0124365. [Google Scholar] [CrossRef]
- Henriques, D.; Parejo, M.; Vignal, A.; Wragg, D.; Wallberg, A.; Webster, M.T.; Pinto, M.A. Developing reduced SNP assays from whole-genome sequence data to estimate introgression in an organism with complex genetic patterns, the Iberian honeybee (Apis mellifera iberiensis). Evol. Appl. 2018, 11, 1270–1282. [Google Scholar] [CrossRef] [PubMed]
- Henriques, D.; Browne, K.A.; Barnett, M.W.; Parejo, M.; Kryger, P.; Freeman, T.C.; Muñoz, I.; Garnery, L.; Highet, F.; Jonhston, J.S.; et al. High sample throughput genotyping for estimating C-lineage introgression in the dark honeybee: An accurate and cost-effective SNP-based tool. Sci. Rep. 2018, 8, 8552. [Google Scholar] [CrossRef] [PubMed]
- Jones, J.C.; Du, Z.G.; Bernstein, R.; Meyer, M.; Hoppe, A.; Schilling, E.; Ableitner, M.; Juling, K.; Dick, R.; Strauss, A.S.; et al. Tool for genomic selection and breeding to evolutionary adaptation: Development of a 100K single nucleotide polymorphism array for the honey bee. Ecol. Evol. 2020, 10, 6246–6256. [Google Scholar] [CrossRef] [PubMed]
- Bovo, S.; Utzeri, V.J.; Ribani, A.; Cabbri, R.; Fontanesi, L. Shotgun sequencing of honey DNA can describe honey bee derived environmental signatures and the honey bee hologenome complexity. Sci. Rep. 2020, 10, 9279. [Google Scholar] [CrossRef] [PubMed]
- Winston, M.L.; Otis, G.W.; Taylor, O.R. Absconding behaviour of the Africanized honeybee in South America. J. Apic. Res. 1979, 18, 85–94. [Google Scholar] [CrossRef]
- Winston, M.L. Killer Bees. The Africanized Honey Bee in the Americas; Harvard University Press: Cambridge, MA, USA, 1992; ISBN 067450352X. [Google Scholar]
- Villa, J.D. Defensive behaviour of Africanized and European honeybees at two elevations in Colombia. J. Apic. Res. 1988, 27, 141–145. [Google Scholar] [CrossRef]
- Guzmán-Novoa, E.; Page, R.E., Jr. Selective breeding of honey bees (Hymenoptera: Apidae) in Africanized areas. J. Econ. Entomol. 1999, 92, 521–525. [Google Scholar] [CrossRef]
- Livanis, G.; Moss, C.B. The effect of Africanized honey bees on honey production in the United States: An informational approach. Ecol. Econ. 2010, 69, 895–904. [Google Scholar] [CrossRef]
- Commission Delegated Regulation (EU) 2020/688 of 17 December 2019 Supplementing Regulation (EU) 2016/429 of the European Parliament and of the Council, as Regards Animal Health Requirements for Movements within the Union of Terrestrial Animals and Hatching Eggs. OJ L 174. 3 June 2020, pp. 140–210. Available online: https://eur-lex.europa.eu/legal-content/EN/TXT/?uri=CELEX:32020R0688 (accessed on 10 June 2021).
- Meixner, M.D.; Büchler, R.; Costa, C.; Francis, R.M.; Hatjina, F.; Kryger, P.; Uzunov, A.; Carreck, N.L. Honey bee genotypes and the environment. J. Apic. Res. 2014, 53, 183–187. [Google Scholar] [CrossRef]
| Regions 1 | No. of Samples 2 | No. of Samples with Different Profiles 2,3 | Frequency of Honey Samples with the Indicated Lineage 2,4 | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Only A | Only C | AC | AM | CM | ACM | A | C | M | ||
| Piedmont (Piemonte) | 56 | 0 | 46 | 0 | 0 | 3 | 7 | 0.125 | 1.000 | 0.179 |
| Valle d’Aosta | 8 | 0 | 7 | 0 | 0 | 1 | 0 | 0.000 | 1.000 | 0.125 |
| Liguria | 7 | 0 | 6 | 0 | 0 | 1 | 0 | 0.000 | 1.000 | 0.143 |
| Lombardy (Lombardia) | 69 | 0 | 48 | 3 | 0 | 6 | 12 | 0.217 | 1.000 | 0.261 |
| Trentino-Alto Adige | 20 | 1 | 8 | 0 | 0 | 8 | 3 | 0.200 | 0.950 | 0.550 |
| Veneto | 41 | 0 | 28 | 1 | 0 | 7 | 5 | 0.146 | 1.000 | 0.293 |
| Friuli-Venezia Giulia | 22 | 0 | 20 | 0 | 0 | 2 | 0 | 0.000 | 1.000 | 0.091 |
| Emilia-Romagna | 63 (100) | 0 (0) | 52 (83) | 0 (0) | 0 (0) | 4 (7) | 7 (10) | 0.111 (0.110) | 1.000 (1.000) | 0.175 (0.170) |
| Tuscany (Toscana) | 37 | 0 | 29 | 0 | 0 | 4 | 4 | 0.108 | 1.000 | 0.216 |
| Umbria | 15 | 0 | 12 | 1 | 0 | 1 | 1 | 0.133 | 1.000 | 0.133 |
| Marche | 20 | 0 | 16 | 0 | 0 | 0 | 4 | 0.200 | 1.000 | 0.200 |
| Lazio | 26 | 1 | 18 | 0 | 0 | 4 | 3 | 0.154 | 0.962 | 0.269 |
| Abruzzo | 19 | 0 | 16 | 0 | 0 | 1 | 2 | 0.105 | 1.000 | 0.158 |
| Molise | 9 | 0 | 6 | 0 | 0 | 2 | 1 | 0.111 | 1.000 | 0.333 |
| Campania | 26 | 0 | 19 | 0 | 0 | 2 | 5 | 0.192 | 1.000 | 0.269 |
| Puglia | 34 | 0 | 23 | 0 | 0 | 2 | 9 | 0.265 | 1.000 | 0.324 |
| Basilicata | 20 | 0 | 12 | 0 | 0 | 5 | 3 | 0.150 | 1.000 | 0.400 |
| Calabria | 9 | 0 | 5 | 0 | 0 | 2 | 2 | 0.222 | 1.000 | 0.444 |
| Sicily (Sicilia) | 26 | 12 | 6 | 0 | 2 | 4 | 2 | 0.615 | 0.462 | 0.231 |
| Sardinia | 23 (43) | 0 (0) | 19 (36) | 1 (1) | 0 (0) | 2 (3) | 1 (3) | 0.087 (0.093) | 1.000 (1.000) | 0.130 (0.140) |
| Italy | 550 (607) | 14 (14) | 396 (444) | 6 (6) | 2 (2) | 61 (65) | 71 (76) | 0.169 (0.161) | 0.971 (0.974) | 0.240 (0.232) |
| Mitotypes 1 | Latitude 2 | Constant 3 | Chi-Square 4 | Odd Ratio (95% CI) 5 |
|---|---|---|---|---|
| Only C (P + Sa + Si) | 0.130 (0.038); 0.0007 | −4.665 (1.657); 0.005 | 11.398; 0.0007 | 1.139 (1.056, 1.227) |
| Only C (P) | 0.052 (0.050); 0.302 | −1.2179 (2.197); 0.579 | 1.056; 0.304 | 1.053 (0.955, 1.162) |
| C (P + Sa + Si) | 0.974 (0.184); <0.0001 | −36.189 (7.197); <0.0001 | 58.965; <0.0001 | 2.647 (1.845, 3.798) |
| C (P) | −0.064 (0.370); 0.864 | 8.318 (16.343); 0.611 | 0.031; 0.861 | 0.938 (0.454, 1.938) |
| A (P + Sa + Si) | −0.176 (0.045); 0.0001 | 5.979 (1.919); 0.002 | 15.235; 0.0001 | 0.839 (0.768, 0.916) |
| A (P) | −0.096 (0.060); 0.112 | 2.458 (2.635); 0.351 | 2.478; 0.116 | 0.909 (0.807, 1.023) |
| M (P + Sa + Si) | −0047 (0.040); 0.240 | 0.913 (1.743); 0.600 | 1.365; 0.243 | 0.954 (0.882, 1.032) |
| M (P) | −0.069 (0.051); 0.176 | 1.890 (2.226); 0.396 | 1.812; 0.178 | 0.934 (0.845,1.031) |
| ACM (P + Sa + Si) | −0.040 (0.051); 0.432 | −0.166 (2.216); 0.940 | 0.609; 0.435 | 0.961 (0.869, 1.062) |
| ACM (P) | −0.127 (0.062); 0.043 | 3.677 (2.716); 0.176 | 4.031; 0.447 | 0.881 (0.780, 0.996) |
| Multiple mitotypes (P + Sa + Si) | −0.038 (0.040); 0.338 | 0.578 (1.725); 0.738 | 0.911; 0.340 | 0.963 (0.890, 1.041) |
| Multiple mitotypes (P) | −0.0535 (0.050); 0.289 | 1.273 (2.207); 0.564 | 1.118; 0.290 | 0.948 (0.859,1.046) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Utzeri, V.J.; Ribani, A.; Taurisano, V.; Banqué, C.H.i.; Fontanesi, L. Distribution of the Main Apis mellifera Mitochondrial DNA Lineages in Italy Assessed Using an Environmental DNA Approach. Insects 2021, 12, 620. https://doi.org/10.3390/insects12070620
Utzeri VJ, Ribani A, Taurisano V, Banqué CHi, Fontanesi L. Distribution of the Main Apis mellifera Mitochondrial DNA Lineages in Italy Assessed Using an Environmental DNA Approach. Insects. 2021; 12(7):620. https://doi.org/10.3390/insects12070620
Chicago/Turabian StyleUtzeri, Valerio Joe, Anisa Ribani, Valeria Taurisano, Carles Hernández i Banqué, and Luca Fontanesi. 2021. "Distribution of the Main Apis mellifera Mitochondrial DNA Lineages in Italy Assessed Using an Environmental DNA Approach" Insects 12, no. 7: 620. https://doi.org/10.3390/insects12070620
APA StyleUtzeri, V. J., Ribani, A., Taurisano, V., Banqué, C. H. i., & Fontanesi, L. (2021). Distribution of the Main Apis mellifera Mitochondrial DNA Lineages in Italy Assessed Using an Environmental DNA Approach. Insects, 12(7), 620. https://doi.org/10.3390/insects12070620







