Distribution and Genetic Variability of Bemisia tabaci Cryptic Species (Hemiptera: Aleyrodidae) in Italy
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
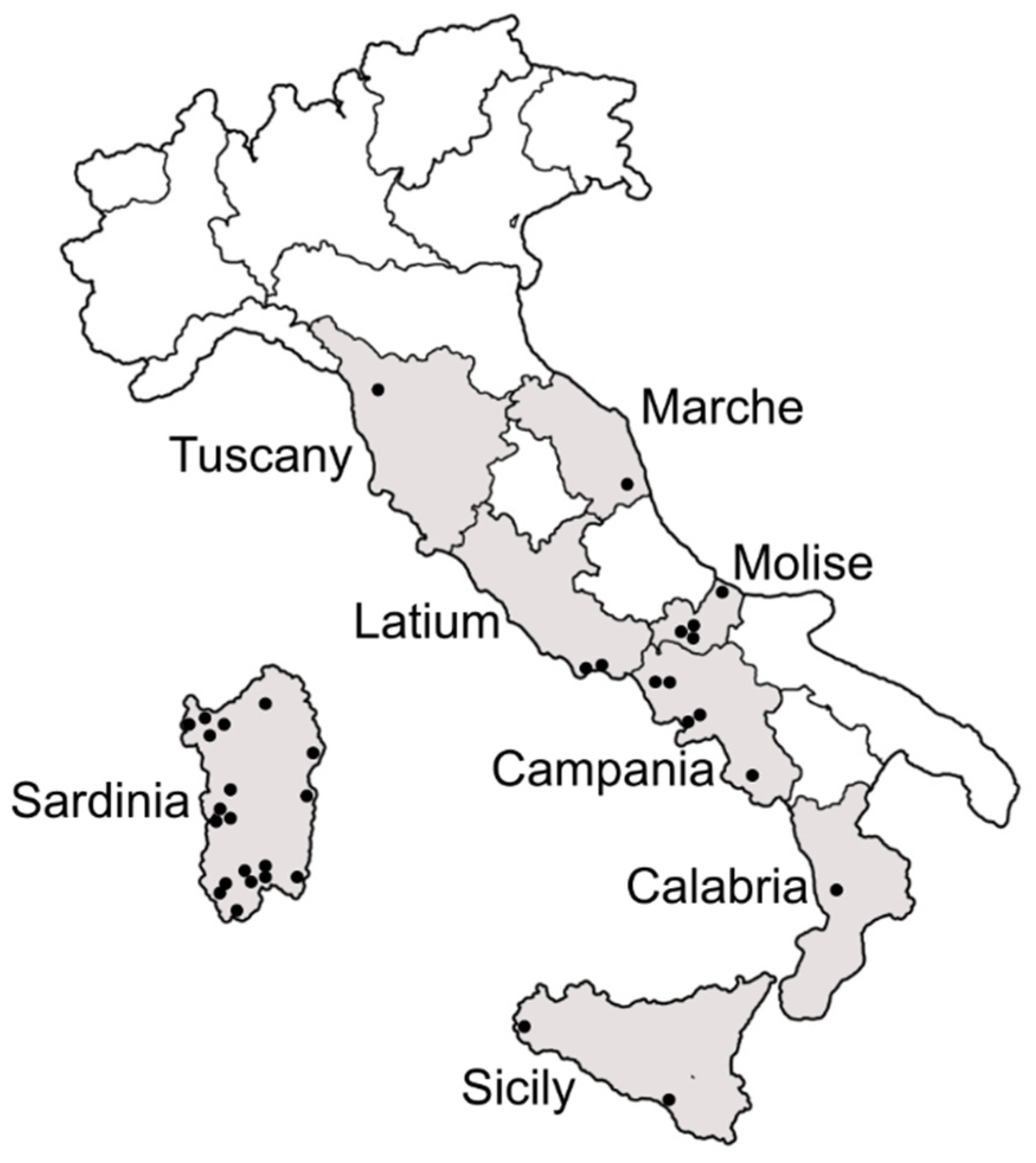
2.1. Sample Collection
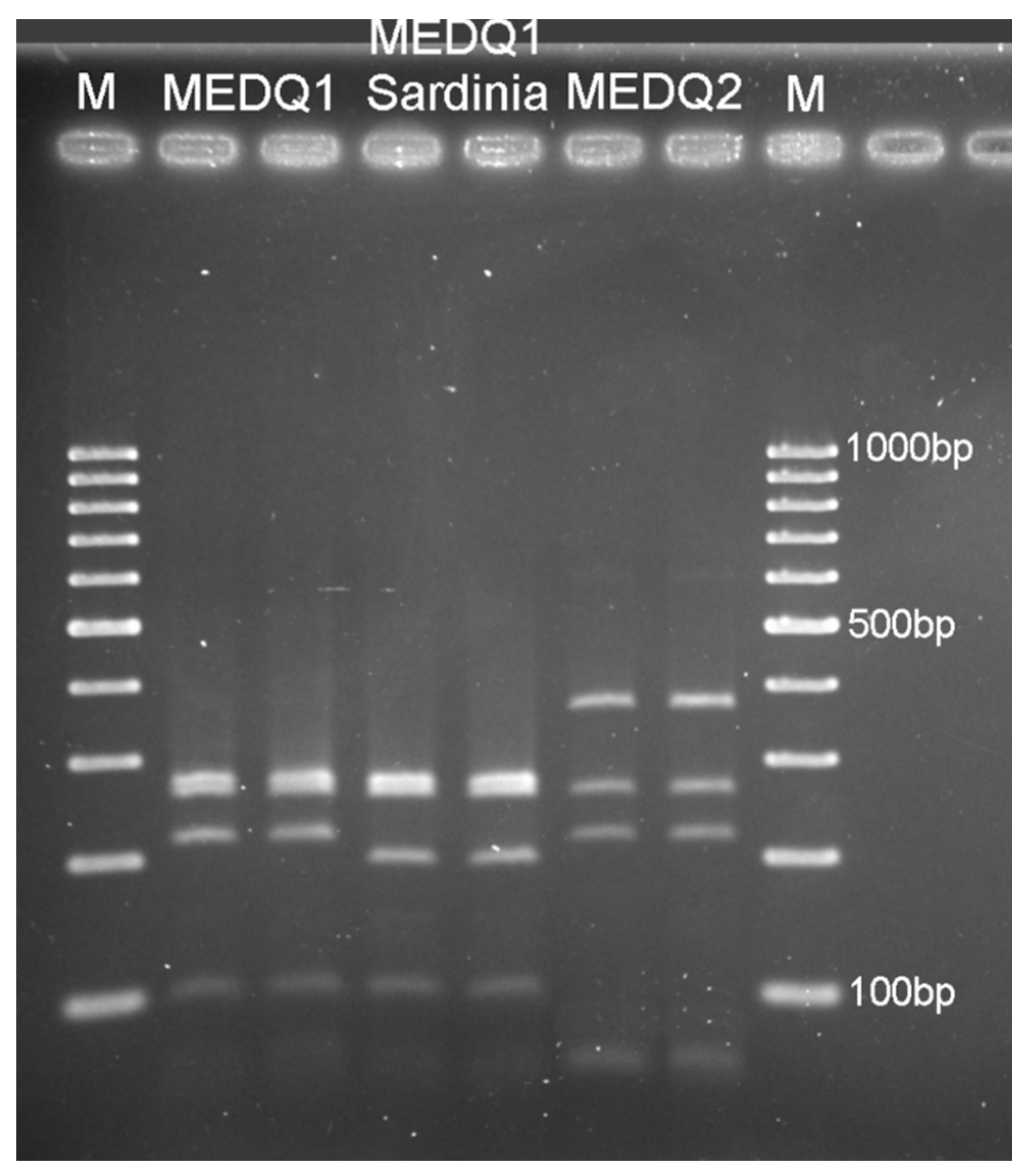
2.2. Molecular Analyses for the Identification of B. tabaci Species and Haplogroups
2.3. Sequence Analyses
3. Results
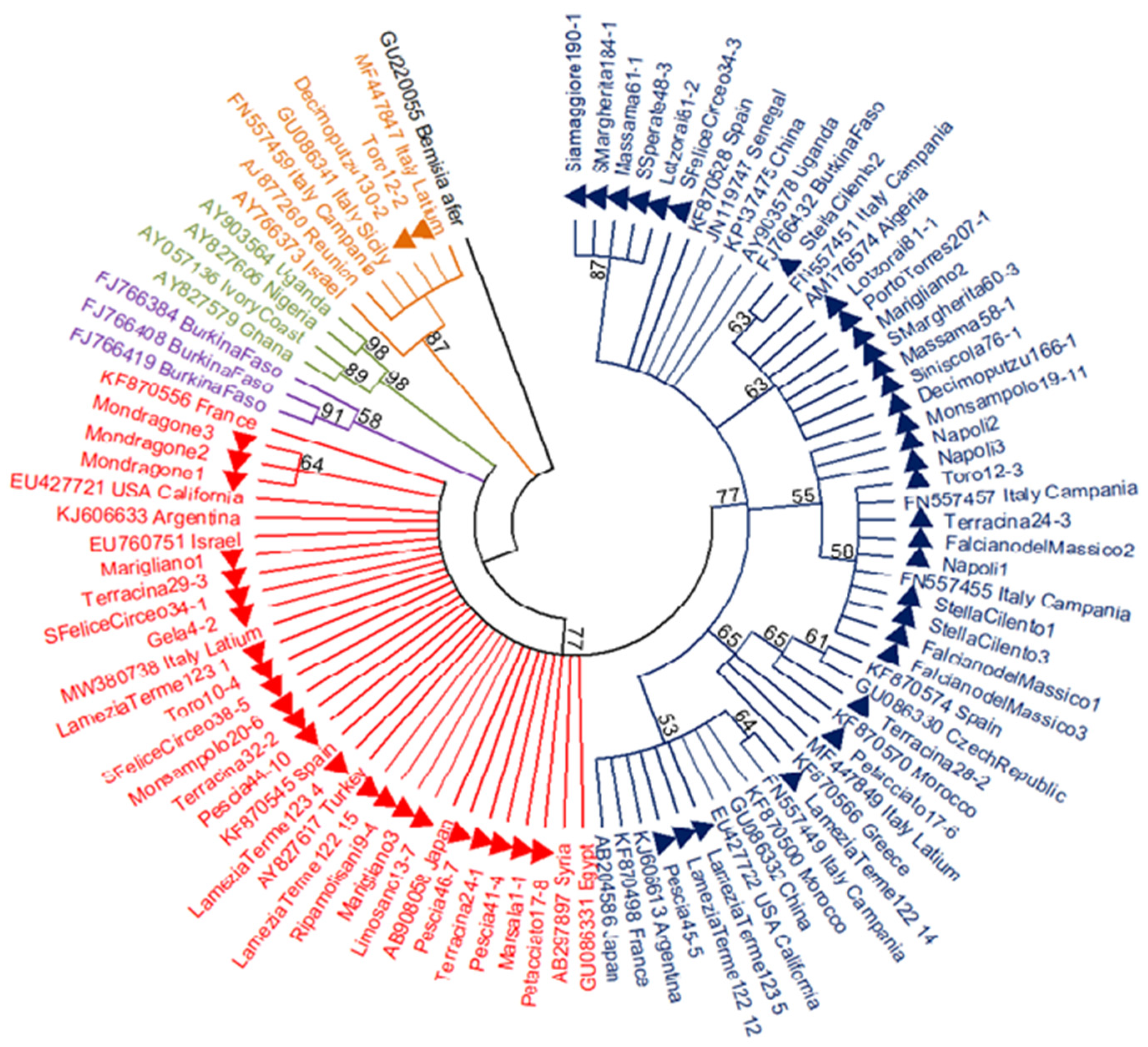
3.1. Presence and Distribution of B. tabaci Species Complex
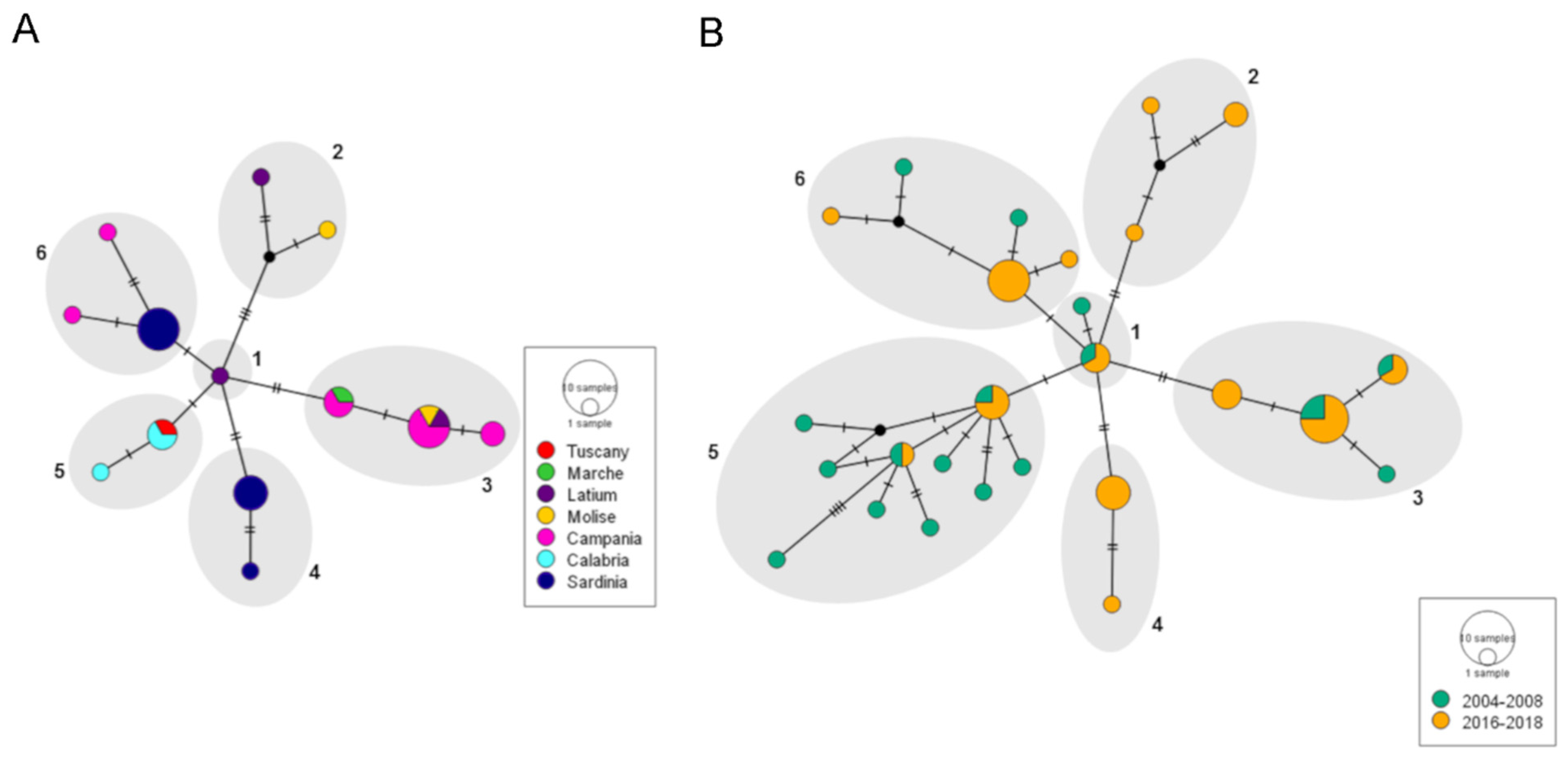
3.2. Genetic Variability within the MED Species
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Acknowledgments
Conflicts of Interest
References
- Navas-Castillo, J.; Fiallo-Olivé, E.; Sànchez-Campos, S. Emerging virus diseases transmitted by whiteflies. Annu. Rev. Phytopath. 2011, 49, 219–248. [Google Scholar] [CrossRef]
- Polston, J.E.; de Barro, P.; Boykin, L.M. Transmission specificities of plant viruses with the newly identified species of the Bemisia tabaci species complex. Pest. Manag. Sci. 2014, 70, 1547–1552. [Google Scholar] [CrossRef] [PubMed]
- De Barro, P.J.; Liu, S.S.; Boykin, L.M.; Dinsdale, A.B. Bemisia tabaci: A statement of species status. Annu. Rev. Entomol. 2011, 56, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; de Barro, P.; Zhao, H.; Wang, J.; Nardi, F.; Liu, S.S. An extensive field survey combined with a phylogenetic analysis reveals rapid and widespread invasion of two alien whiteflies in China. PLoS ONE 2011, 6, e16061. [Google Scholar] [CrossRef] [PubMed]
- Alemandri, V.; de Barro, P.; Bejerman, N.; Caro, E.B.A.; Dumón, A.D.; Mattio, M.F.; Rodriguez, S.M.; Truol, G. Species within the Bemisia tabaci (Hemiptera: Aleyrodidae) complex in soybean and bean crops in Argentina. J. Econ. Entomol. 2012, 105, 48–53. [Google Scholar] [CrossRef] [PubMed]
- Firdaus, S.; Vosman, B.; Hidayati, N.; Supena, E.D.J.; Visser, R.G.F.; van Heusden, A.W. The Bemisia tabaci species complex: Additions from different parts of the world. Insect Sci. 2013, 20, 723–733. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.; Park, J.; Lee, G.S.; Lee, S.; Akimoto, S.I. Taxonomic status of the Bemisia tabaci complex (Hemiptera: Aleyrodidae) and reassessment of the number of its constituent species. PLoS ONE 2013, 8, e63810. [Google Scholar] [CrossRef]
- Hu, J.; Zhang, X.; Jiang, Z.; Zhang, F.; Liu, Y.; Li, Z.; Zhang, Z. New putative cryptic species detection and genetic network analysis of Bemisia tabaci (Hempitera: Aleyrodidae) in China based on mitochondrial COI sequences. Mitochondrial DNA A 2017, 29, 474–484. [Google Scholar] [CrossRef]
- Mugerwa, H.; Seal, S.; Wang, H.L.; Patel, M.V.; Kabaalu, R.; Omongo, C.A.; Alicai, T.; Tairo, F.; Ndunguru, J.; Sseruwagi, P.; et al. African ancestry of New World, Bemisia tabaci-whitefly species. Sci. Rep. 2018, 8, 2734. [Google Scholar] [CrossRef]
- Dinsdale, A.; Cook, L.; Riginos, C.; Buckley, Y.M.; de Barro, P. Refined global analysis of Bemisia tabaci (Hemiptera: Sternorrhyncha: Aleyrodoidea: Aleyrodidae) mitochondrial cytochrome oxidase 1 to identify species level genetic boundaries. Ann. Entomol. Soc. Am. 2010, 103, 196–208. [Google Scholar] [CrossRef]
- Boykin, L.M.; DeBarro, P.J. A practical guide to identifying members of the Bemisia tabaci species complex: And other morphologically identical species. Front. Ecol. Evol. 2014, 2, 1–5. [Google Scholar] [CrossRef]
- Zang, L.S.; Chen, W.Q.; Liu, S.S. Comparison of performance on different host plants between the B biotype and a non-B biotype of Bemisia tabaci from Zhejiang, China. Entomol. Exp. Appl. 2006, 121, 221–227. [Google Scholar] [CrossRef]
- Xu, J.; Lin, K.K.; Liu, S.S. Performance on different host plants of an alien and an indigenous Bemisia tabaci from China. J. Appl. Entomol 2011, 135, 771–779. [Google Scholar] [CrossRef]
- Sun, D.B.; Liu, Y.Q.; Qin, L.; Xu, J.; Li, F.F.; Liu, S.S. Competitive displacement between two invasive whiteflies: Insecticide application and host plant effects. Bull. Entomol. Res. 2013, 103, 344–353. [Google Scholar] [CrossRef]
- Brown, J.K.; Frohlich, D.R.; Rosell, R.C. The sweetpotato or silverleaf whiteflies: Biotypes of Bemisia tabaci or a species complex? Annu. Rev. Entomol. 1995, 40, 511–534. [Google Scholar] [CrossRef]
- Sseruwagi, P.; Legg, J.P.; Maruthi, M.N.; Colvin, J.; Rey, M.E.C.; Brown, J.K. Genetic diversity of Bemisia tabaci (Gennadius) (Hemiptera: Aleyrodidae) populations and presence of the B biotype and a non-B biotype that can induce silverleaf symptoms in squash, in Uganda. Ann. Appl. Biol. 2005, 147, 253–265. [Google Scholar] [CrossRef]
- Horowitz, A.R.; Kontsedalov, S.; Khasdan, V.; Ishaaya, I. Biotypes B and Q of Bemisia tabaci and their relevance to neonicotinoid and pyriproxyfen resistance. Arch. Insect Biochem. Physiol. 2005, 58, 216–225. [Google Scholar] [CrossRef] [PubMed]
- Horowitz, A.R.; Ishaaya, I. Dynamics of biotypes B and Q of the whitefly Bemisia tabaci and its impact on insecticide resistance. Pest. Manag. Sci. 2014, 70, 1568–1572. [Google Scholar] [CrossRef]
- Liu, S.-S.; de Barro, P.J.; Xu, J.; Luan, J.-B.; Zang, L.-S.; Ruan, Y.-M.; Wan, F.-H. Asymmetric mating interactions drive widespread invasion and displacement in a whitefly. Science 2007, 318, 1769–1772. [Google Scholar] [CrossRef]
- Delatte, H.; Duyck, P.F.; Triboire, A.; David, P.; Becker, N.; Bonato, O.; Reynaud, B. Differential invasion success among biotypes: Case of Bemisia tabaci. Biol. Invasions 2009, 11, 1059–1070. [Google Scholar] [CrossRef]
- Bedford, I.D.; Briddon, R.W.; Brown, J.K.; Rosell, R.C.; Markham, P.G. Geminivirus transmission and biological characterisation of Bemisia tabaci (Gennadius) biotypes from different geographic regions. Ann. Appl. Biol. 1994, 125, 311–325. [Google Scholar] [CrossRef]
- Bing, X.L.; Ruan, Y.M.; Rao, Q.; Wang, X.W.; Liu, S.S. Diversity of secondary endosymbionts among different putative species of the whitefly Bemisia tabaci. Insect Sci. 2013, 20, 194–206. [Google Scholar] [CrossRef] [PubMed]
- EFSA Panel on Plant Health. Scientific opinion on the risks to plant health posed by Bemisia tabaci species complex and viruses it transmits for the EU territory. EFSA J. 2013, 11, 3162. [Google Scholar]
- Chu, D.; Tao, Y.L.; Zhang, Y.J.; Wan, F.H.; Brown, J.K. Effects of host, temperature and relative humidity on competitive displacement of two invasive Bemisia tabaci biotypes (Q and B). Insect Sci. 2012, 19, 595–603. [Google Scholar] [CrossRef]
- Watanabe, L.F.M.; Bello, V.H.; de Marchi, B.R.; Silva, F.; Machado, L.F.; Sartori, M.M.P.; Pavan, M.A.; Krause-Sakate, R. Performance and competitive displacement of Bemisia tabaci MEAM1 and MED cryptic species on different host plants. Crop. Prot. 2019, 124, 104860. [Google Scholar] [CrossRef]
- Bonato, O.; Lurette, A.; Vidal, C.; Fargues, J. Modelling temperature-dependent bionomics of Bemisia tabaci (Q-biotype). Physiol. Entomol. 2007, 32, 50–55. [Google Scholar] [CrossRef]
- Pascual, S. Mechanisms in competition, under laboratory conditions, between Spanish biotypes B and Q of Bemisia tabaci (Gennadius). Span. J. Agric. Res. 2006, 4, 351–354. [Google Scholar] [CrossRef]
- Crowder, D.W.; Sitvarin, M.I.; Carrière, Y. Plasticity in mating behaviour drives asymmetric reproductive interference in whiteflies. Anim. Behav. 2010, 79, 579–587. [Google Scholar] [CrossRef]
- Elbaz, M.; Lahav, N.; Morin, S. Evidence for pre-zygotic reproductive barrier between the B and Q biotypes of Bemisia tabaci (Hemiptera: Aleyrodidae). Bull. Entomol. Res. 2010, 100, 581–590. [Google Scholar] [CrossRef]
- Simón, B.; Cenis, J.L.; de la Rúa, P. Distribution patterns of the Q and B biotypes of Bemisia tabaci in the Mediterranean basin based on microsatellite variation. Entomol. Exp. Appl. 2007, 124, 327–336. [Google Scholar] [CrossRef]
- Tsagkarakou, A.; Tsigenopoulos, C.S.; Gorman, K.; Lagnel, J.; Bedford, I.D. Biotype status and genetic polymorphism of the whitefly Bemisia tabaci (Hemiptera: Aleyrodidae) in Greece: Mitochondrial DNA and microsatellites. Bull. Entomol. Res. 2007, 97, 29–40. [Google Scholar] [CrossRef]
- Parrella, G.; Scassillo, L.; Giorgini, M. Evidence for a new genetic variant in the Bemisia tabaci species complex and the prevalence of the biotype Q in southern Italy. J. Pest. Sci. 2012, 85, 227–238. [Google Scholar] [CrossRef]
- Barboza, N.M.; Esker, P.; Inoue-Nagata, A.K.; Moriones, E. Genetic diversity and geographic distribution of Bemisia tabaci and Trialeurodes vaporariorum in Costa Rica. Ann. Appl. Biol. 2019, 174, 248–261. [Google Scholar] [CrossRef]
- Tang, X.-T.; Cai, L.; Shen, Y.; Xu, L.-L.; Du, Y.-Z. Competitive displacement between Bemisia tabaci MEAM1 and MED and evidence for multiple invasions of MED. Insects 2020, 11, 35. [Google Scholar] [CrossRef] [PubMed]
- De Barro, P.; Ahmed, M.Z. Genetic networking of the Bemisia tabaci cryptic species complex reveals pattern of biological invasions. PLoS ONE 2011, 6, e25579. [Google Scholar] [CrossRef]
- Mckenzie, C.L.; Hodges, G.; Osborne, L.S.; Byrne, F.J.; Shatters, R.G. Distribution of Bemisia tabaci (Hemiptera: Aleyrodidae) biotypes in Florida investigating the Q invasion. J. Econ. Entomol. 2009, 102, 670–676. [Google Scholar] [CrossRef]
- McKenzie, C.L.; Bethke, J.A.; Byrne, F.J.; Chamberlin, J.R.; Dennehy, T.J.; Dickey, A.M.; Gilrein, D.; Hall, P.M.; Ludwig, S.; Oetting, R.D.; et al. Distribution of Bemisia tabaci (Hemiptera: Aleyrodidae) biotypes in North America after the Q invasion. J. Econ. Entomol. 2012, 105, 753–766. [Google Scholar] [CrossRef]
- Chu, D.; Zhang, Y.J.; Brown, J.K.; Cong, B.; Xu, B.Y.; Wu, Q.J.; Zhu, G.R. The introduction of the exotic Q biotype of Bemisia tabaci from the Mediterranean region into China on ornamental crops. Fla. Entomol. 2006, 89, 168–174. [Google Scholar] [CrossRef]
- Chu, D.; Wan, F.H.; Tao, Y.L.; Liu, G.X.; Fan, Z.X.; Bi, Y.P. Genetic differentiation of Bemisia tabaci (Hemiptera: Aleyrodidae) biotype Q based on mitochondrial DNA markers. Insect Sci. 2008, 15, 115–123. [Google Scholar] [CrossRef]
- Alemandri, V.; Vaghi, C.G.; Dumón, A.D.; Argüello, E.B.; Mattio, M.F.; García, S.; Lambertini, P.M.L.; Truol, G. Three members of the Bemisia tabaci (Hemiptera: Aleyrodidae) cryptic species complex occur sympatrically in Argentina horticultural crops. J. Econ. Entomol. 2015, 18, 405–413. [Google Scholar] [CrossRef]
- De Moraes, L.A.; Marubayashi, J.M.; Yuki, V.A.; Ghanim, M.; Bello, V.H.; de Marchi, B.R.; da Fonseca, L.B.; Boykin, L.M.; Krause-Sakate, R.; Pavan, M.A. New invasion of Bemisia tabaci Mediterranean species in Brazil associated to ornamental plants. Phytoparasitica 2017, 45, 1–9. [Google Scholar] [CrossRef]
- Gueguen, G.; Vavre, F.; Gnankine, O.; Peterschmitt, M.; Charif, D.; Chiel, E.; Gottlieb, Y.; Ghanim, M.; Zchori-Fein, E.; Fleury, F. Endosymbiont metacommunities, mtDNA diversity and the evolution of the Bemisia tabaci (Hemiptera: Aleyrodidae) species complex. Mol. Ecol. 2010, 19, 4365–4378. [Google Scholar] [CrossRef] [PubMed]
- Boykin, L.M.; Shatters, R.G.; Rosell, R.C.; Mckenzie, C.L.; Bagnall, R.A.; de Barro, P.; Frohlich, D.R. Global relationships of Bemisia tabaci (Hemiptera: Aleyrodidae) revealed using Bayesian analysis of mitochondrial COI DNA sequences. Mol. Phylogenet. Evol. 2007, 44, 1306–1319. [Google Scholar] [CrossRef] [PubMed]
- Chu, D.; Gao, C.S.; de Barro, P.; Wan, F.H.; Zhang, Y.J.; Khan, I.A. Further insights into the strange role of bacterial endosymbionts in whitefly, Bemisia tabaci: Comparison of secondary symbionts from biotypes B and Q in China. Bull. Entomol. Res. 2011, 101, 477–486. [Google Scholar] [CrossRef]
- Vyskočilová, S.; Seal, S.; Colvin, J. Relative polyphagy of “Mediterranean” cryptic Bemisia tabaci whitefly species and global pest status implications. J. Pest. Sci. 2019, 92, 1071–1088. [Google Scholar] [CrossRef]
- Mouton, L.; Gnankiné, O.; Henri, H.; Terraz, G.; Ketoh, G.; Martin, T.; Fleury, F.; Vavre, F. Detection of genetically isolated entities within the Mediterranean species of Bemisia tabaci: New insights into the systematics of this worldwide pest. Pest. Manag. Sci. 2015, 71, 452–458. [Google Scholar] [CrossRef]
- Vyskočilová, S.; Tay, W.T.; van Brunschot, S.; Seal, S.; Colvin, J. An integrative approach to discovering cryptic species within the Bemisia tabaci whitefly species complex. Sci. Rep. 2018, 8, 10886. [Google Scholar] [CrossRef] [PubMed]
- Bosco, D.; Caciagli, P. Bionomics and ecology of Bemisia tabaci (Sternorrhyncha: Aleyrodidae) in Italy. Eur. J. Entomol. 1998, 95, 519–527. [Google Scholar]
- Rapisarda, C.; Garzia, G.T. Tomato yellow leaf curl Sardinia virus and its vector Bemisia tabaci in Sicilia (Italy): Present status and control possibilities. Bull. OEPP/EPPO 2002, 32, 25–29. [Google Scholar] [CrossRef]
- Simón, B.; Cenis, J.L.; Demichelis, S.; Rapisarda, C.; Caciagli, P.; Bosco, D. Survey of Bemisia tabaci (Hemiptera: Aleyrodidae) biotypes in Italy with the description of a new biotype (T) from Euphorbia characias. Bull. Entomol. Res. 2003, 93, 259–264. [Google Scholar] [CrossRef]
- Parrella, G.; Cozzolino, A.; Manco, E.; Stinca, A.; Formisano, G.; Giorgini, M. Identificata in campania la specie Italy (=biotipo T) del complesso di specie Bemisia tabaci. Protezione Colture 2016, 4, 28–30. [Google Scholar]
- Bosco, D.; Loria, A.; Sartor, C.; Cenis, J.L. PCR-RFLP identification of Bemisia tabaci biotypes in the Mediterranean basin. Phytoparasitica 2006, 34, 243–251. [Google Scholar] [CrossRef]
- Cavalieri, V.; Rapisarda, C. Indagini molecolari sui biotipi di Bemisia tabaci in Sicilia (Hemiptera: Aleyrodidae). Boll. Zool. Agr. Bachic. 2008, 40, 145–154. [Google Scholar]
- Parrella, G.; Nappo, A.G.; Manco, E.; Greco, B.; Giorgini, M. Invasion of the Q2 mitochondrial variant of Mediterranean Bemisia tabaci in Southern Italy: Possible role of bacterial endosymbionts. Pest. Manag. Sci. 2014, 70, 1514–1523. [Google Scholar] [CrossRef] [PubMed]
- Mouton, L.; Thierry, M.; Henri, H.; Baudin, R.; Gnankin’e, O.; Reynaud, B.; Zchori-Fein, E.; Becker, N.; Fleury, F.; Delatte, H. Evidence of diversity and recombination in Arsenophonus symbionts of the Bemisia tabaci species complex. BMC Microbiol. 2012, 12, S10. [Google Scholar] [CrossRef]
- Bertin, S.; Luigi, M.; Parrella, G.; Giorgini, M.; Davino, S.; Tomassoli, L. Survey of the distribution of Bemisia tabaci (Hemiptera: Aleyrodidae) in Lazio region (Central Italy): A threat for the northward expansion of tomato leaf curl New Delhi virus (Begomovirus: Geminiviridae) infection. Phytoparasitica 2018, 46, 171–182. [Google Scholar] [CrossRef]
- Simon, C.; Frati, F.; Beckenbach, A.; Crespi, B.; Liu, H.; Flook, P. Evolution, weighting and phylogenetic utility of mitochondrial gene sequences and a compilation of conserved polymerase chain reaction primers. Ann. Entomol. Soc. Am. 1994, 87, 651–701. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Rozas, J.; Ferrer-Mata, A.; Sánchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sánchez-Gracia, A. DnaSP v6: DNA sequence polymorphism analysis of large datasets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef] [PubMed]
- Bandelt, H.; Forster, P.; Röhl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 1999, 16, 37–48. [Google Scholar] [CrossRef]
- Leigh, J.W.; Bryant, D. POPART: Full-feature software for haplotype network construction. Methods Ecol. Evol. 2015, 6, 1110–1116. [Google Scholar] [CrossRef]
- Patti, I.; Rapisarda, C. Reperti morfo-biologici sugli Aleirodidi nocivi alle piante coltivate in Italia. Bollettino Zoologia Agrarian Bachicoltura 1981, 16, 135–190. [Google Scholar]
- Gilioli, G.; Pasquali, S.; Parisi, S.; Winter, S. Modelling the potential distribution of Bemisia tabaci in Europe in light of the climate change scenario. Pest. Manag. Sci. 2014, 70, 1611–1623. [Google Scholar] [CrossRef]
- Bradshaw, C.D.; Hemming, D.; Baker, R.; Everatt, M.; Eyre, D.; Korycinska, A. A novel approach for exploring climatic factors limiting current pest distributions: A case study of Bemisia tabaci in north-west Europe and assessment of potential future establishment in the United Kingdom under climate change. PLoS ONE 2019, 14, e0221057. [Google Scholar] [CrossRef]
- Terraz, G.; Gueguen, G.; Arnó, J.; Fleurya, F.; Mouton, L. Nuclear and cytoplasmic differentiation among Mediterranean populations of Bemisia tabaci: Testing the biological relevance of cytotypes. Pest. Manag. Sci. 2014, 70, 1503–1513. [Google Scholar] [CrossRef] [PubMed]
- Gauthier, N.; Clouet, C.; Perrakis, A.; Kapantaidaki, D.; Peterschmitt, M.; Tsagkarakou, A. Genetic structure of Bemisia tabaci MED populations from home-range countries, inferred by nuclear and cytoplasmic markers: Impact on the distribution of the insecticide resistance genes. Pest. Manag. Sci. 2014, 70, 1477–1491. [Google Scholar] [CrossRef]
- Kontsedalov, S.; Zchori-Fein, E.; Chiel, E.; Gottlieb, Y.; Inbar, M.; Ghanim, M. The presence of Rickettsia is associated with increased susceptibility of Bemisia tabaci (Homoptera: Aleyrodidae) to insecticides. Pest. Manag. Sci. 2008, 64, 789–792. [Google Scholar] [CrossRef] [PubMed]
- Gottlieb, Y.; Zchori-Fein, E.; Mozes-Daube, N.; Kontsedalov, S.; Skaljac, M.; Brumin, M.; Sobol, I.; Czosnek, H.; Vavre, F.; Fleury, F.; et al. The transmission efficiency of tomato yellow leaf curl virus by the whitefly Bemisia tabaci is correlated with the presence of a specific symbiotic bacterium species. J. Virol. 2010, 84, 9310–9317. [Google Scholar] [CrossRef]
- Brumin, M.; Kontsedalov, S.; Ghanim, M. Rickettsia influences thermotolerance in the whitefly Bemisia tabaci B biotype. Insect Sci. 2011, 18, 57–66. [Google Scholar] [CrossRef]
- Sànchez-Campos, S.; Navas-Castillo, J.; Camero, R.; Soria, C.; Dìaz, J.A.; Moriones, E. Displacement of tomato yellow leaf curl virus (TYLCV)-Sr by TYLCV-Is in tomato epidemics in Spain. Phytopathology 1999, 89, 1038–1043. [Google Scholar] [CrossRef]
- Pan, H.; Chu, D.; Yan, W.; Su, Q.; Liu, B.; Wang, S.; Wu, Q. Rapid spread of tomato yellow leaf curl virus in China is aided differentially by two invasive whiteflies. PLoS ONE 2012, 7, e34817. [Google Scholar] [CrossRef]
- Panno, S.; Iacono, G.; Davino, M.; Marchione, S.; Zappardo, V.; Bella, P.; Tomassoli, L.; Accotto, G.P.; Davino, S. First report of Tomato leaf curl New Delhi virus affecting zucchini squash in an important horticultural area of southern Italy. New Dis. Rep. 2016, 33, 6. [Google Scholar] [CrossRef]
- Panno, S.; Caruso, A.G.; Troiano, E.; Luigi, M.; Manglli, A.; Vatrano, T.; Iacono, G.; Marchione, S.; Bertin, S.; Tomassoli, L.; et al. Emergence of tomato leaf curl New Delhi virus in Italy: Estimation of incidence and genetic diversity. Plant. Pathol. 2019, 68, 601–608. [Google Scholar] [CrossRef]
- Monci, F.; Sánchez-Campos, S.; Navas-Castillo, J.; Moriones, E. A natural recombinant between the geminiviruses Tomato yellow leaf curl Sardinia virus and Tomato yellow leaf curl virus exhibits a novel pathogenic phenotype and is becoming prevalent in Spanish populations. Virology 2002, 303, 317–326. [Google Scholar] [CrossRef] [PubMed]
- Fiallo-Olivé, E.; Trenado, H.P.; Louro, D.; Navas-Castillo, J. Recurrent speciation of a tomato yellow leaf curl geminivirus in Portugal by recombination. Sci. Rep. 2019, 9, 1332. [Google Scholar] [CrossRef]
- Bertin, S.; (CREA-DC, Rome, Italy). Personal communication, 2020.
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bertin, S.; Parrella, G.; Nannini, M.; Guercio, G.; Troiano, E.; Tomassoli, L. Distribution and Genetic Variability of Bemisia tabaci Cryptic Species (Hemiptera: Aleyrodidae) in Italy. Insects 2021, 12, 521. https://doi.org/10.3390/insects12060521
Bertin S, Parrella G, Nannini M, Guercio G, Troiano E, Tomassoli L. Distribution and Genetic Variability of Bemisia tabaci Cryptic Species (Hemiptera: Aleyrodidae) in Italy. Insects. 2021; 12(6):521. https://doi.org/10.3390/insects12060521
Chicago/Turabian StyleBertin, Sabrina, Giuseppe Parrella, Mauro Nannini, Giorgia Guercio, Elisa Troiano, and Laura Tomassoli. 2021. "Distribution and Genetic Variability of Bemisia tabaci Cryptic Species (Hemiptera: Aleyrodidae) in Italy" Insects 12, no. 6: 521. https://doi.org/10.3390/insects12060521
APA StyleBertin, S., Parrella, G., Nannini, M., Guercio, G., Troiano, E., & Tomassoli, L. (2021). Distribution and Genetic Variability of Bemisia tabaci Cryptic Species (Hemiptera: Aleyrodidae) in Italy. Insects, 12(6), 521. https://doi.org/10.3390/insects12060521








