Isolation and Characterization of Mosquito-Associated Spiroplasma cantharicola from Aedes japonicus Collected in Hokkaido, Japan
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Mosquito Sampling
2.2. Isolation of Filtrable Microorganisms from Field-Collected Mosquitoes
2.3. DNA Extraction
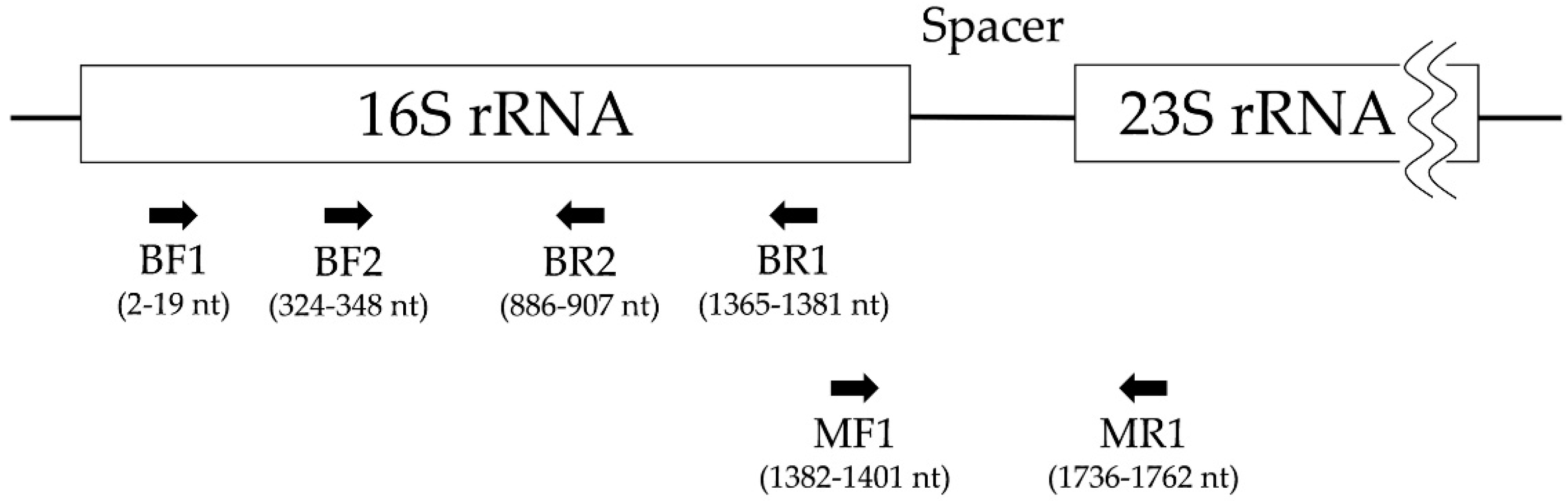
2.4. Polymerase Chain Reaction and Electrophoresis
2.5. Sequencing and Genetic Analysis
2.6. Culturing of S. cantharicola
2.7. Statistical Analyses
3. Results
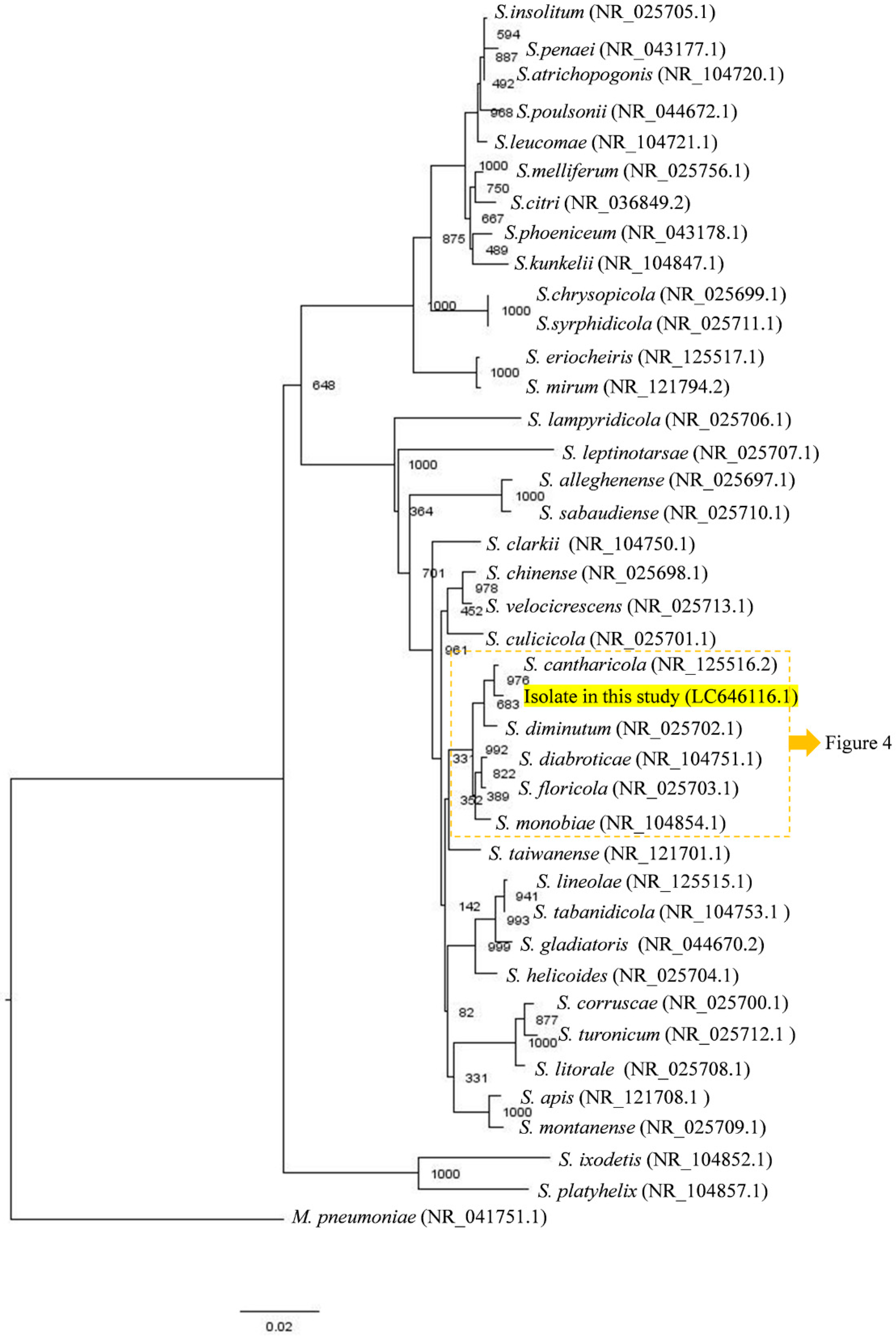
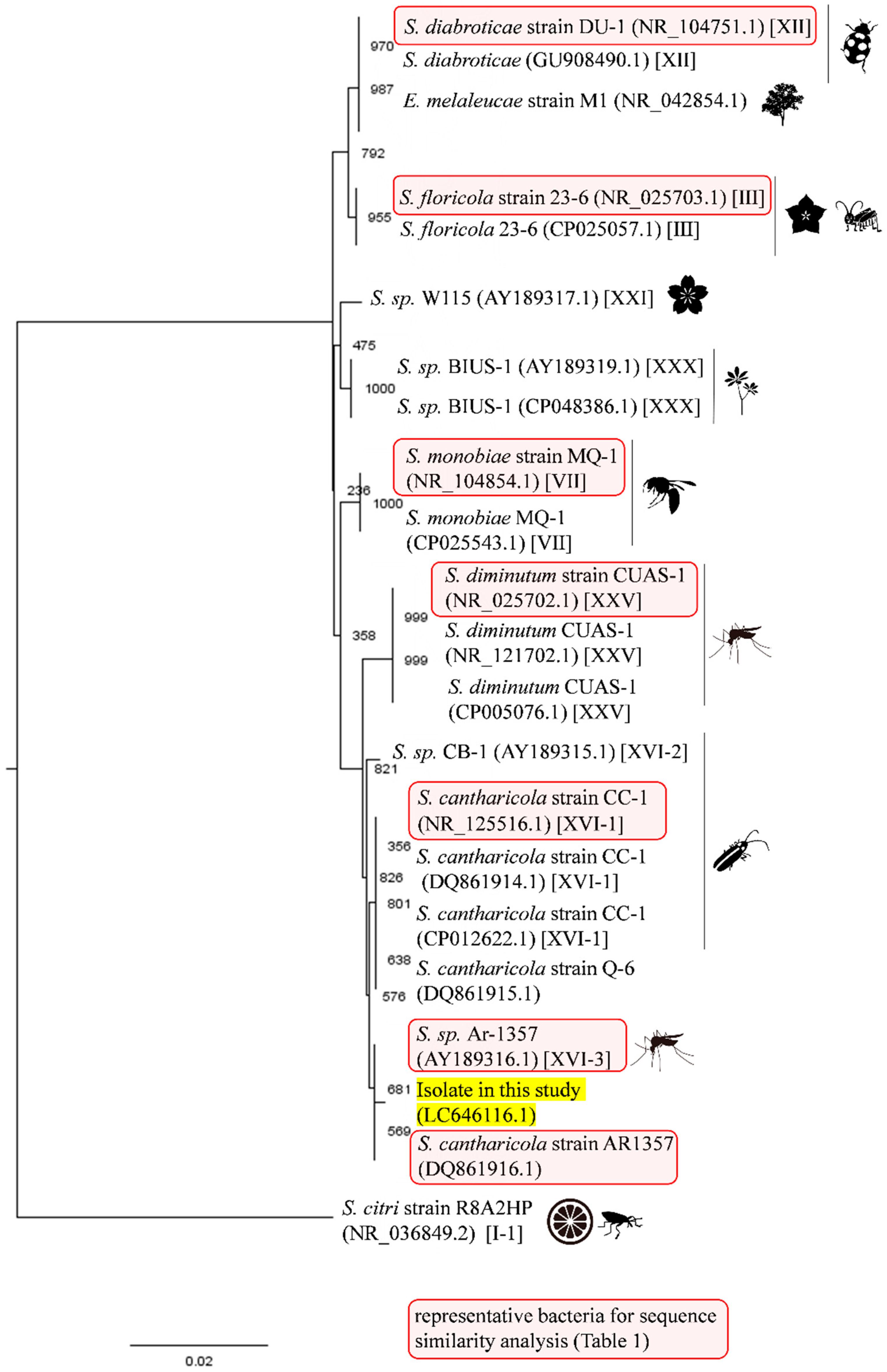
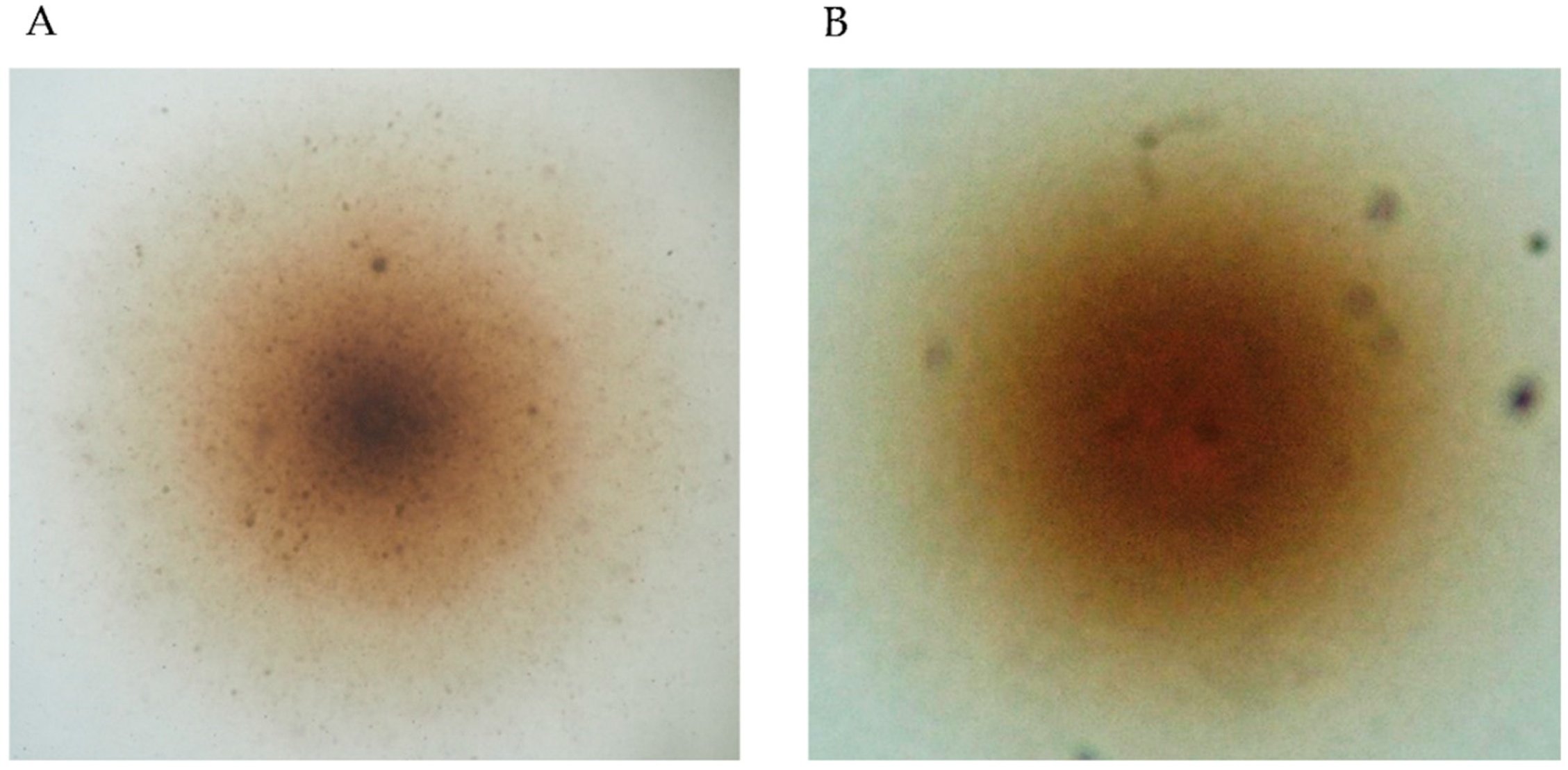
3.1. Isolation and Identification of Filtrable Microorganism from a Field-Collected Mosquito
3.2. Prevalence of S. cantharicola in the Field-Collected Mosquitoes and Laboratory Colonies
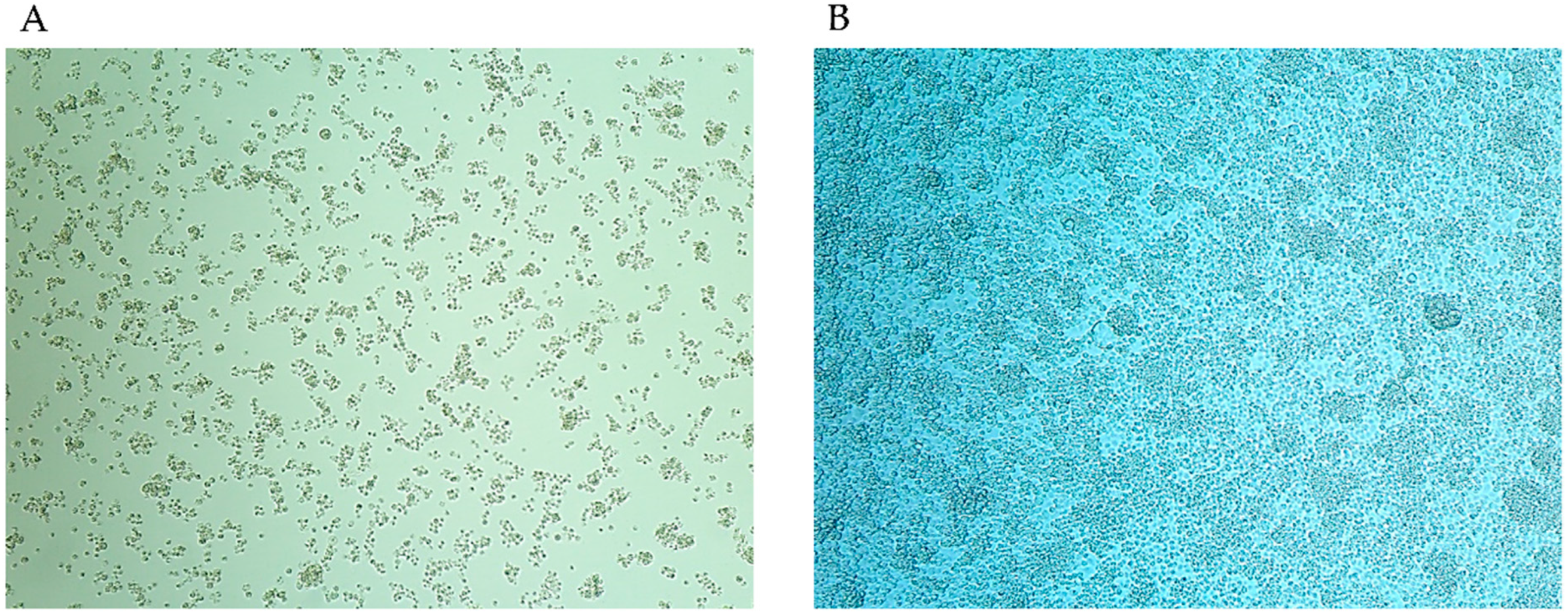
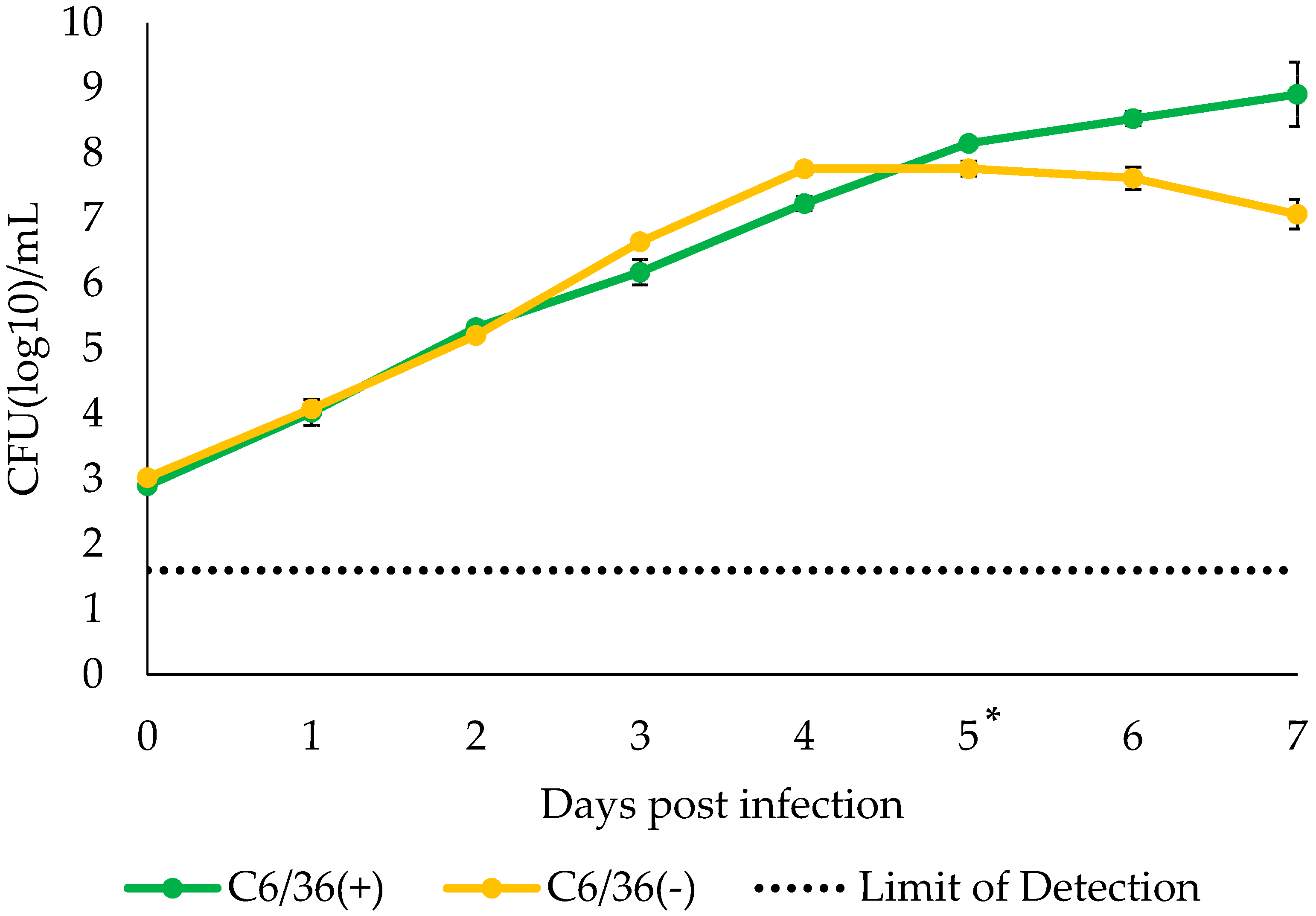
3.3. Growth Kinetics of S. cantharicola
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cisak, E.; Wójcik-Fatla, A.; Zając, V.; Sawczyn, A.; Sroka, J.; Dutkiewicz, J. Spiroplasma-an emerging arthropod-borne pathogen? Ann. Agric. Environ. Med. 2015, 22, 589–593. [Google Scholar] [CrossRef]
- Regassa, L.B.; Gasparich, G.E. Spiroplasmas: Evolutionary relationships and biodiversity. Front. Biosci. 2006, 11, 2983–3002. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Halverson, S.; Ezinwa, N. Mosquito-borne diseases. Prim. Care 2018, 45, 393–407. [Google Scholar] [CrossRef]
- AMCA. Mosquito-Borne Diseases. Available online: https://www.mosquito.org/page/diseases (accessed on 14 October 2021).
- Gould, E.A.; Solomon, T. Pathogenic flaviviruses. Lancet 2008, 371, 500–509. [Google Scholar] [CrossRef]
- White, N.J. Antimalarial drug resistance. J. Clin. Investig. 2004, 113, 1084–1092. [Google Scholar] [CrossRef] [PubMed]
- Ranson, H.; N’Guessan, R.; Lines, J.; Moiroux, N.; Nkuni, Z.; Corbel, V. Pyrethroid resistance in african anopheline mosquitoes: What are the implications for malaria control? Trends Parasitol. 2011, 27, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, A.A.; Montgomery, B.L.; Popovici, J.; Iturbe-Ormaetxe, I.; Johnson, P.H.; Muzzi, F.; Greenfield, M.; Durkan, M.; Leong, Y.S.; Dong, Y.; et al. Successful establishment of wolbachia in aedes populations to suppress dengue transmission. Nature 2011, 476, 454–457. [Google Scholar] [CrossRef]
- Moreira, L.A.; Iturbe-Ormaetxe, I.; Jeffery, J.A.; Lu, G.; Pyke, A.T.; Hedges, L.M.; Rocha, B.C.; Hall-Mendelin, S.; Day, A.; Riegler, M.; et al. A wolbachia symbiont in aedes aegypti limits infection with dengue, chikungunya, and plasmodium. Cell 2009, 139, 1268–1278. [Google Scholar] [CrossRef] [Green Version]
- Humphery-Smith, I.; Grulet, O.; Chastel, C. Pathogenicity of spiroplasma taiwanense for larval aedes aegypti mosquitoes. Med. Vet. Entomol. 1991, 5, 229–232. [Google Scholar] [CrossRef]
- Humphery-Smith, I.; Grulet, O.; Le Goff, F.; Chastel, C. Spiroplasma (mollicutes: Spiroplasmataceae) pathogenic for aedes aegypti and anopheles stephensi (diptera: Culicidae). J. Med. Entomol. 1991, 28, 219–222. [Google Scholar] [CrossRef]
- Jupatanakul, N.; Sim, S.; Dimopoulos, G. The insect microbiome modulates vector competence for arboviruses. Viruses 2014, 6, 4294–4313. [Google Scholar] [CrossRef]
- Chang, T.H.; Lo, W.S.; Ku, C.; Chen, L.L.; Kuo, C.H. Molecular evolution of the substrate utilization strategies and putative virulence factors in mosquito-associated spiroplasma species. Genome Biol. Evol. 2014, 6, 500–509. [Google Scholar] [CrossRef] [PubMed]
- Chepkemoi, S.T.; Mararo, E.; Butungi, H.; Paredes, J.; Masiga, D.; Sinkins, S.P.; Herren, J.K. Identification of spiroplasmainsolitum symbionts in anopheles gambiae. Wellcome Open Res. 2017, 2, 90. [Google Scholar] [CrossRef] [PubMed]
- Vazeille-Falcoz, M.; Perchec-Merien, A.M.; Rodhain, F. Experimental infection of aedes aegypti mosquitoes, suckling mice, and rats with four mosquito spiroplasmas. J. Invertebr. Pathol. 1994, 63, 37–42. [Google Scholar] [CrossRef] [PubMed]
- Utarini, A.; Indriani, C.; Ahmad, R.A.; Tantowijoyo, W.; Arguni, E.; Ansari, M.R.; Supriyati, E.; Wardana, D.S.; Meitika, Y.; Ernesia, I.; et al. Efficacy of wolbachia-infected mosquito deployments for the control of dengue. N. Engl. J. Med. 2021, 384, 2177–2186. [Google Scholar] [CrossRef]
- Chastel, C.; Devau, B.; Le Goff, F.; Simitzis-Le Flohic, A.M.; Gruffaz, R.; Kerdraon, G.; Gilot, B. Mosquito spiroplasmas from france and their ecology. Isr. J. Med. Sci. 1987, 23, 683–686. [Google Scholar]
- Whitcomb, R.F.; Chastel, C.; Abalain-Colloc, M.; Stevens, C.; Tully, J.G.; Rose, D.L.; Carle, P.; Bové, J.M.; Henegar, R.B.; Hackett, K.J.; et al. Spiroplasma cantharicola sp. Nov., from cantharid beetles (coleoptera: Cantharidae). Int. J. Syst. Evol. Microbiol. 1993, 43, 421–424. [Google Scholar] [CrossRef]
- Tanaka, K.; Mizusawa, K.; Saugstad, E.S. A revision of the adult and larval mosquitoes of japan (including the ryukyu archipelago and the ogasawara islands) and korea (diptera: Culicidae). Contrib. Am. Entomol. Inst. 1979, 16, 1–987. [Google Scholar]
- Folmer, O.; Black, M.; Hoeh, W.; Lutz, R.; Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit i from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 1994, 3, 294–299. [Google Scholar]
- Uchida, L.; Shibuya, M.; Morales-Vargas, R.E.; Hagiwara, K.; Muramatsu, Y. Zika virus potential vectors among aedes mosquitoes from hokkaido, northern japan: Implications for potential emergence of zika disease. Pathogens 2021, 10, 938. [Google Scholar] [CrossRef]
- Igarashi, A. Isolation of a singh’s aedes albopictus cell clone sensitive to dengue and chikungunya viruses. J. Gen. Virol. 1978, 40, 531–544. [Google Scholar] [CrossRef] [PubMed]
- Fredriksson, N.J.; Hermansson, M.; Wilen, B.M. The choice of pcr primers has great impact on assessments of bacterial community diversity and dynamics in a wastewater treatment plant. PLoS ONE 2013, 8, e76431. [Google Scholar] [CrossRef] [PubMed]
- Beckers, B.; Op De Beeck, M.; Thijs, S.; Truyens, S.; Weyens, N.; Boerjan, W.; Vangronsveld, J. Performance of 16s rdna primer pairs in the study of rhizosphere and endosphere bacterial microbiomes in metabarcoding studies. Front. Microbiol. 2016, 7, 650. [Google Scholar] [CrossRef] [Green Version]
- Uemori, T.; Asada, K.; Kato, I.; Harasawa, R. Amplification of the 16s-23s spacer region in rrna operons of mycoplasmas by the polymerase chain reaction. Syst. Appl. Microbiol. 1992, 15, 181–186. [Google Scholar] [CrossRef]
- Hatamoto, M.; Imachi, H.; Ohashi, A.; Harada, H. Identification and cultivation of anaerobic, syntrophic long-chain fatty acid-degrading microbes from mesophilic and thermophilic methanogenic sludges. Appl. Environ. Microbiol 2007, 73, 1332–1340. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lazarevic, V.; Gaia, N.; Girard, M.; Schrenzel, J. Decontamination of 16s rrna gene amplicon sequence datasets based on bacterial load assessment by qpcr. BMC Microbiol. 2016, 16, 73. [Google Scholar] [CrossRef] [Green Version]
- Moulder, R.W.; French, F.E.; Chang, C.J. Simplified media for spiroplasmas associated with tabanid flies. Can. J. Microbiol. 2002, 48, 1–6. [Google Scholar] [CrossRef]
- Abalain-Colloc, M.L.; Williamson, D.L.; Carle, P.; Abalain, J.H.; Bonnet, F.; Tully, J.G.; Konai, M.; Whitcomb, R.F.; Bové, J.M.; Chastel, C. Division of group xvi spiroplasmas into subgroups. Int. J. Syst. Evol. Microbiol. 1993, 43, 342–346. [Google Scholar] [CrossRef] [Green Version]
- Koban, M.B.; Kampen, H.; Scheuch, D.E.; Frueh, L.; Kuhlisch, C.; Janssen, N.; Steidle, J.L.M.; Schaub, G.A.; Werner, D. The asian bush mosquito aedes japonicus japonicus (diptera: Culicidae) in europe, 17 years after its first detection, with a focus on monitoring methods. Parasit. Vectors 2019, 12, 109. [Google Scholar] [CrossRef] [Green Version]
- Peach, D.A.H.; Almond, M.; Pol, J.C. Modeled distributions of aedes japonicus japonicus and aedes togoi (diptera: Culicidae) in the united states, canada, and northern latin america. J. Vector Ecol. 2019, 44, 119–129. [Google Scholar] [CrossRef] [Green Version]
- Konai, M.; Clark, E.A.; Camp, M.; Koeh, A.L.; Whitcomb, R.F. Temperature ranges, growth optima, and growth rates of spiroplasma (spiroplasmataceae, class mollicutes) species. Curr. Microbiol. 1996, 32, 314–319. [Google Scholar] [CrossRef] [PubMed]
- Duret, S.; Batailler, B.; Dubrana, M.P.; Saillard, C.; Renaudin, J.; Béven, L.; Arricau-Bouvery, N. Invasion of insect cells by spiroplasma citri involves spiralin relocalization and lectin/glycoconjugate-type interactions. Cell Microbiol. 2014, 16, 1119–1132. [Google Scholar] [CrossRef] [PubMed]
- Fletcher, J.; Wayadande, A.; Melcher, U.; Ye, F. The phytopathogenic mollicute-insect vector interface: A closer look. Phytopathology 1998, 88, 1351–1358. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kwon, M.O.; Wayadande, A.C.; Fletcher, J. Spiroplasma citri movement into the intestines and salivary glands of its leafhopper vector, circulifer tenellus. Phytopathology 1999, 89, 1144–1151. [Google Scholar] [CrossRef] [Green Version]
- Humphery-Smith, I.; Grulet, O.; Le Lay, G.; Chastel, C. Pathogenicity of spiroplasma sabaudiense (mollicute) for the cells (c6/36) of aedes albopictus (insecta: Diptera) in vitro. Bull. Soc. Pathol. Exot. Fil. 1988, 81, 752–765. [Google Scholar]
- Steiner, T.; McGarrity, G.J.; Phillips, D.M. Cultivation and partial characterization of spiroplasmas in cell cultures. Infect. Immun. 1982, 35, 296–304. [Google Scholar] [CrossRef] [Green Version]
- Abalain-Colloc, M.L.; Rosen, L.; Tully, J.G.; Bovë, J.M.; Chastel, C.; Williamson, D.L. Spiroplasma taiwanense sp.Nov. From culex tritaeniorhynchus mosquitoes collected in taiwan. Int. J. Syst. Evol. Microbiol. 1988, 38, 103–107. [Google Scholar]

| Isolate in This Study (LC646116.1) | S. cantharicola Strain AR1357 (DQ861916.1) | Spiroplasma sp. Ar-1357 (AY189316.1) | S. cantharicola Strain CC-1 (NR_125516.1) | S. diminutum Strain CUAS-1 (NR_025702.1) | S. monobiae Strain MQ-1 (NR_104854.1) | S. floricola Strain 23-6 (NR_025703.1) | |
|---|---|---|---|---|---|---|---|
| S. cantharicola strain AR1357 (DQ861916.1) | 99.8% (1281/1283) | ||||||
| Spiroplasma sp. Ar-1357 (AY189316.1) | 99.8% (1281/1283) | 99.8% (1502/1505) | |||||
| S. cantharicola strain CC-1 (NR_125516.1) | 99.7% (1279/1283) | 99.9% (1503/1505) | 99.7% (1500/1504) | ||||
| S. diminutum strain CUAS-1 (NR_025702.1) | 99.2% (1273/1283) | 98.9% (1351/1366) | 99.4% (1507/1516) | 99.3% (1493/1504) | |||
| S. monobiae strain MQ-1 (NR_104854.1) | 99.1% (1272/1283) | 99.3% (1456/1467) | 99.3% (1457/1467) | 99.2% (1455/1467) | 98.9% (1451/1467) | ||
| S. floricola strain 23-6 (NR_025703.1) | 98.8% (1268/1283) | 98.9% (1351/1366) | 99.0% (1364/1378) | 98.9% (1351/1366) | 99.0% (1364/1378) | 99.4% (1355/1363) | |
| S. diabroticae strain DU-1 (NR_104751.1) | 98.8% (1267/1283) | 98.9% (1487/1504) | 99.0% (1493/1508) | 98.9% (1487/1504) | 99.2% (1496/1508) | 99.3% (1457/1467) | 99.7% (1366/1370) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shimooka, M.; Sakurai, Y.; Muramatsu, Y.; Uchida, L. Isolation and Characterization of Mosquito-Associated Spiroplasma cantharicola from Aedes japonicus Collected in Hokkaido, Japan. Insects 2021, 12, 1056. https://doi.org/10.3390/insects12121056
Shimooka M, Sakurai Y, Muramatsu Y, Uchida L. Isolation and Characterization of Mosquito-Associated Spiroplasma cantharicola from Aedes japonicus Collected in Hokkaido, Japan. Insects. 2021; 12(12):1056. https://doi.org/10.3390/insects12121056
Chicago/Turabian StyleShimooka, Makoto, Yoshimi Sakurai, Yasukazu Muramatsu, and Leo Uchida. 2021. "Isolation and Characterization of Mosquito-Associated Spiroplasma cantharicola from Aedes japonicus Collected in Hokkaido, Japan" Insects 12, no. 12: 1056. https://doi.org/10.3390/insects12121056
APA StyleShimooka, M., Sakurai, Y., Muramatsu, Y., & Uchida, L. (2021). Isolation and Characterization of Mosquito-Associated Spiroplasma cantharicola from Aedes japonicus Collected in Hokkaido, Japan. Insects, 12(12), 1056. https://doi.org/10.3390/insects12121056






