The Pathogen Profile of a Honey Bee Queen Does Not Reflect That of Her Workers
Abstract
1. Introduction
2. Methods
2.1. Queen Exchange and Sample Collection
2.2. Sample Processing
2.3. Molecular Analysis
2.4. Analysis
3. Results
3.1. Pathogen Prevalence
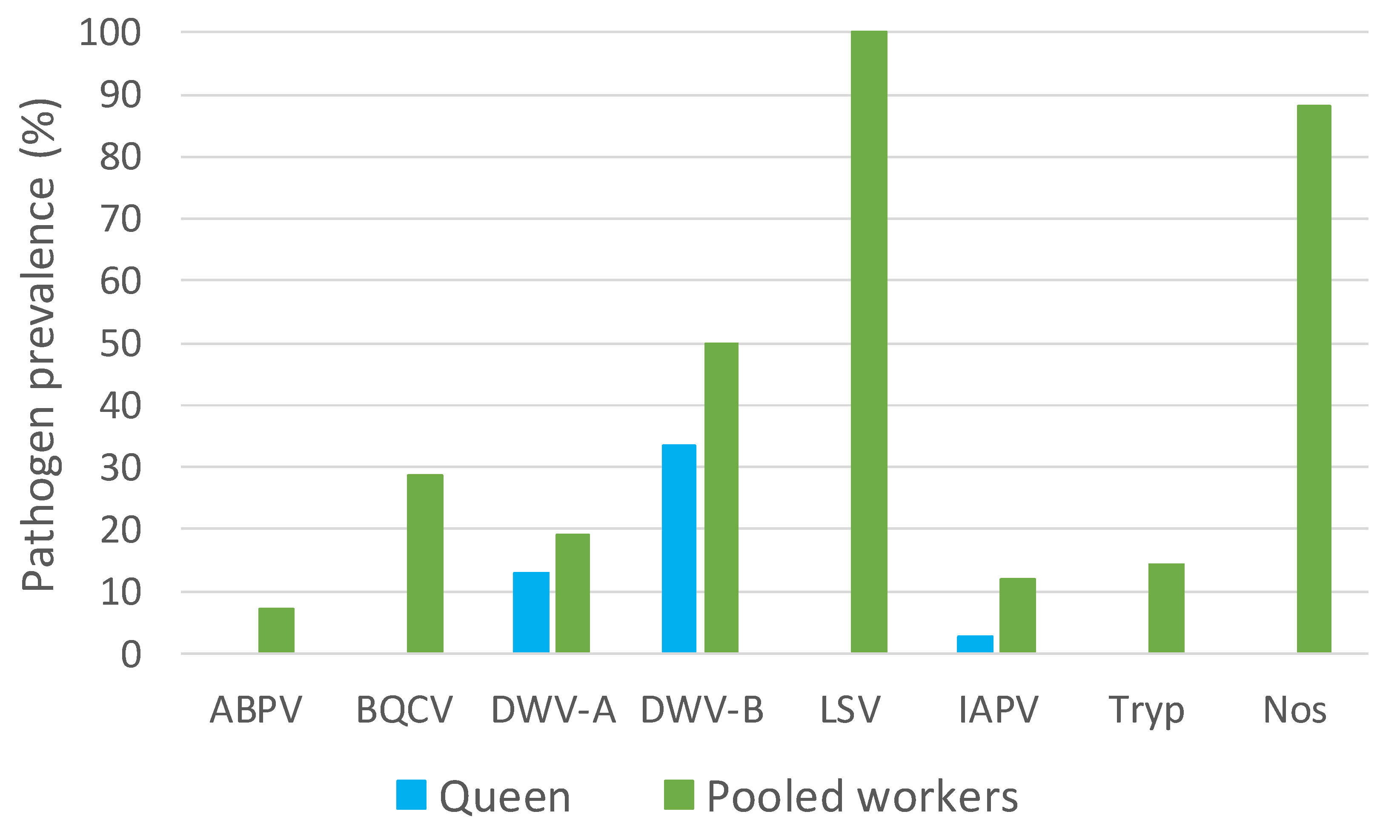
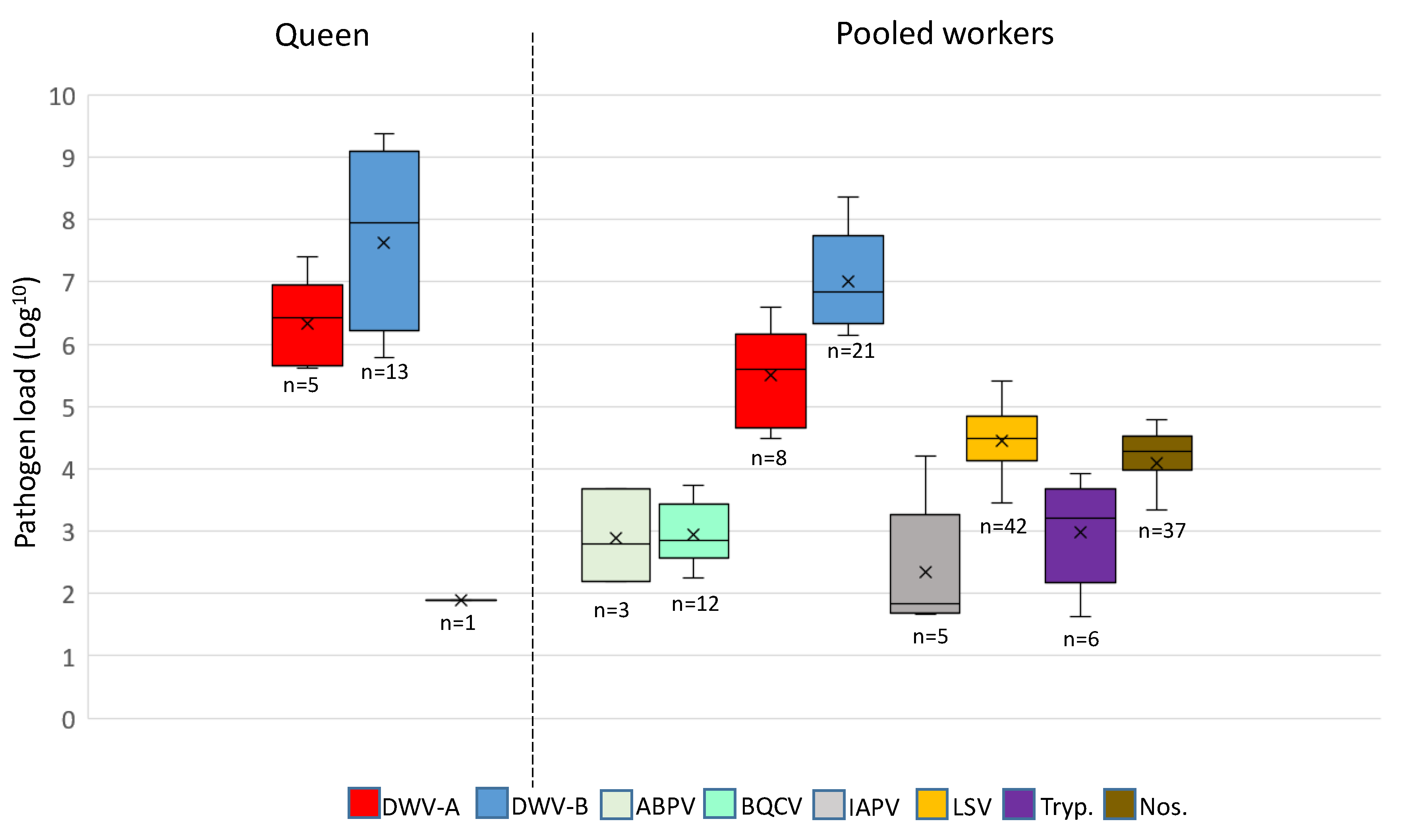
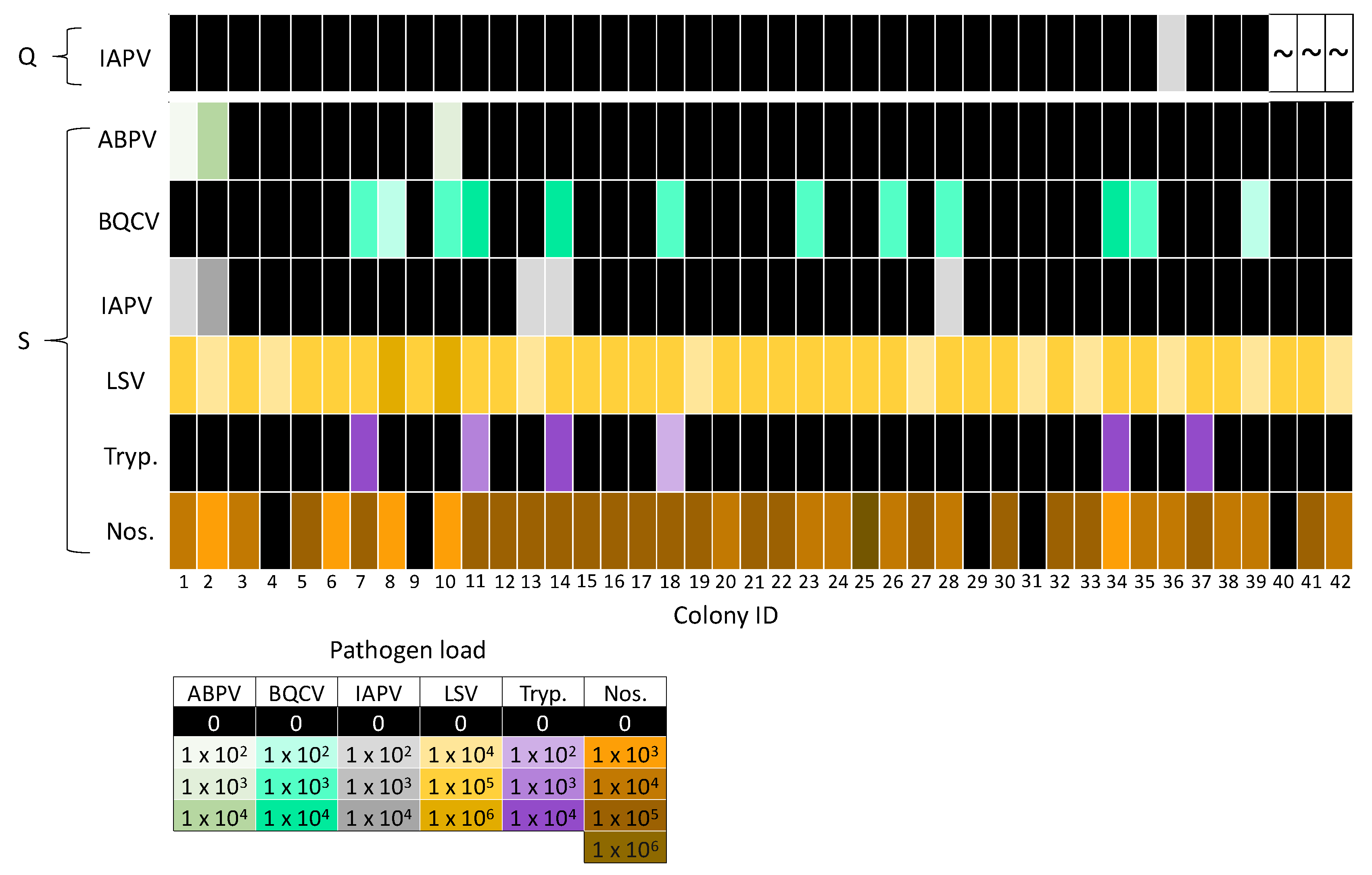
3.2. A Comparison of Pathogens in Pooled Workers and Queens
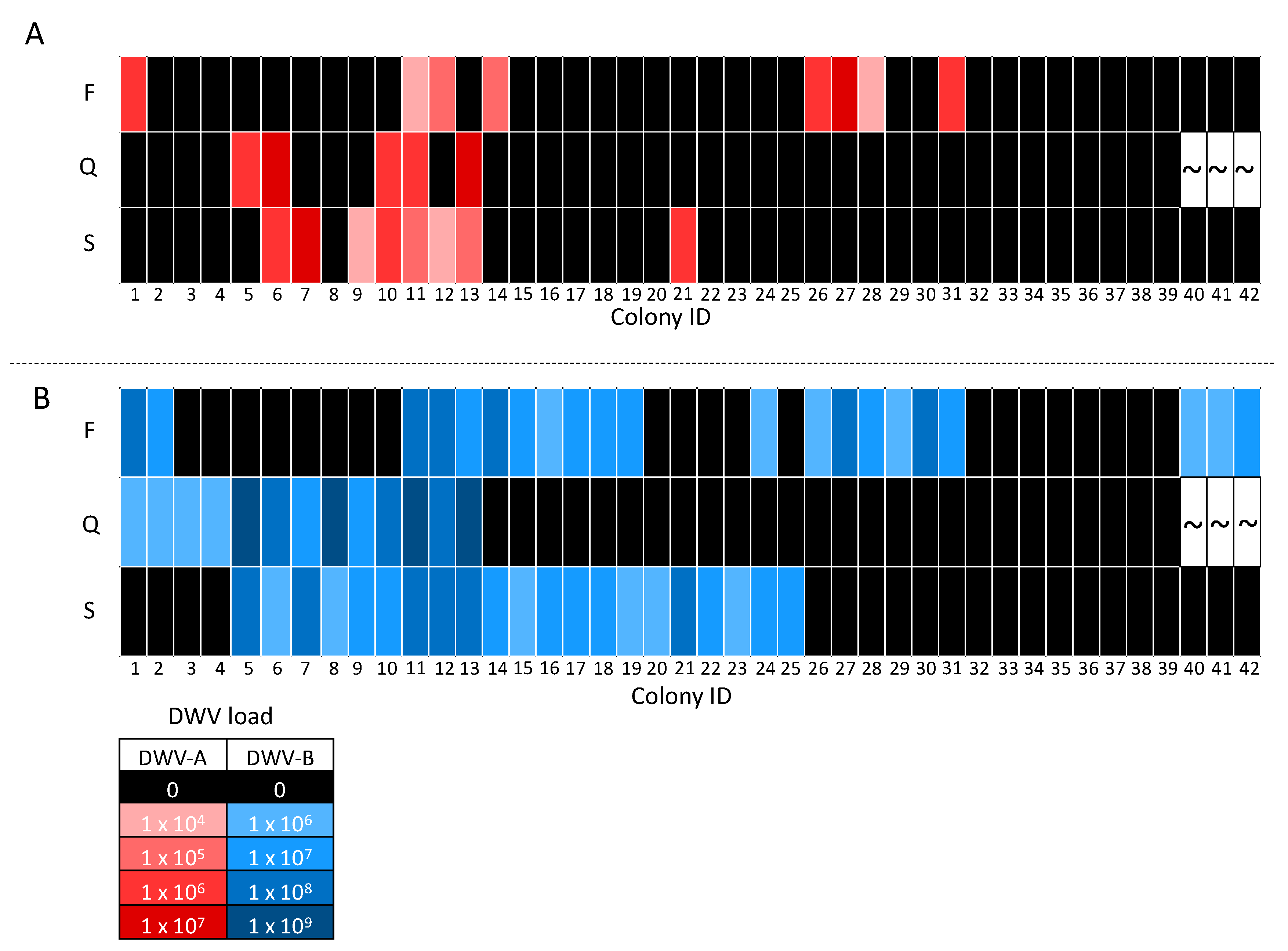
3.3. DWV Transmission between Pooled Workers and Queens
4. Discussion
Author Contributions
Funding
Conflicts of Interest
References
- Winston, M.L. The Biology of the Honey Bee; Harvard University Press: Cambridge, MA, USA, 1987; ISBN 9780674074095. [Google Scholar]
- Allen, M.D. The honeybee queen and her attendants. Anim. Behav. 1960, 8, 201–208. [Google Scholar] [CrossRef]
- Schmid-Hempel, P. Parasites in Social Insects; Princeton University Press: Princeton, NJ, USA, 1998; ISBN 9780691059242. [Google Scholar]
- Day, T. Virulence evolution and the timing of disease life-history events. Trends Ecol. Evol. 2003, 18, 113–118. [Google Scholar] [CrossRef]
- Ebert, D.; Bull, J.J. Challenging the trade-off model for the evolution of virulence: Is virulence management feasible? Trends Microbiol. 2003, 11, 15–20. [Google Scholar] [CrossRef]
- Fries, I.; Camazine, S. Implications of horizontal and vertical pathogen transmission for honey bee epidemiology. Apidologie 2001, 32, 199–214. [Google Scholar] [CrossRef]
- Yue, C.; Schroder, M.; Gisder, S.; Genersch, E. Vertical-transmission routes for deformed wing virus of honeybees (Apis mellifera). J. Gen. Virol. 2007, 88, 2329–2336. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Evans, J.; Feldlaufer, M. Horizontal and vertical transmission of viruses in the honey bee, Apis mellifera. J. Invertebr. Pathol. 2006, 92, 152–159. [Google Scholar] [CrossRef] [PubMed]
- de Miranda, J.R.; Genersch, E. Deformed wing virus. J. Invertebr. Pathol. 2010, 103, S48–S61. [Google Scholar] [CrossRef] [PubMed]
- Ball, B.V. Varroa Jacobsoni as a Virus Vector. In Present Status of Varroatosis in Europe and Progress in the Varroa Mite Control; Cavallero, R., Ed.; ECSC-EEC-EAEC: Luxembourg, 1989. [Google Scholar]
- Bowen-Walker, P.; Martin, S.; Gunn, A. The Transmission of Deformed Wing Virus between Honeybees (Apis mellifera L.) by the Ectoparasitic MiteVarroa jacobsoniOud. J. Invertebr. Pathol. 1999, 73, 101–106. [Google Scholar] [CrossRef]
- Highfield, A.C.; El Nagar, A.; Mackinder, L.C.M.; Noël, L.M.L.J.; Hall, M.J.; Martin, S.J.; Schroeder, D.C. Deformed wing virus implicated in overwintering honeybee colony losses. Appl. Environ. Microbiol. 2009, 75, 7212–7220. [Google Scholar] [CrossRef]
- Martin, S.J.; Ball, B.V.; Carreck, N.L. Prevalence and persistence of deformed wing virus (DWV) in untreated or acaricide-treated Varroa destructor infested honey bee (Apis mellifera) colonies. J. Apic. Res. 2010, 49, 72–79. [Google Scholar] [CrossRef]
- Martin, S.J.; Highfield, A.C.; Brettell, L.; Villalobos, E.M.; Budge, G.E.; Powell, M.; Nikaido, S.; Schroeder, D.C. Global honey bee viral landscape altered by a parasitic mite. Science 2012, 336, 1304–1306. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, D.C.; Martin, S.J. Deformed wing virus. Virulence 2012, 3, 589–591. [Google Scholar] [CrossRef] [PubMed]
- Bailey, L.; Ball, B.V. Honey Bee Pathology; Academic Press: Cambridge, UK, 1991; ISBN 9781483288093. [Google Scholar]
- Tentcheva, D.; Gauthier, L.; Zappulla, N.; Dainat, B.; Cousserans, F.; Colin, M.E.; Bergoin, M. Prevalence and Seasonal Variations of Six Bee Viruses in Apis mellifera L. and Varroa destructor Mite Populations in France. Appl. Environ. Microbiol. 2004, 70, 7185–7191. [Google Scholar] [CrossRef]
- Chen, Y.P.; Siede, R. Honey Bee Viruses. Adv. Virus Res. 2007, 70, 33–80. [Google Scholar] [CrossRef]
- Runckel, C.; Flenniken, M.L.; Engel, J.C.; Ruby, J.G.; Ganem, D.; Andino, R.; DeRisi, J.L. Temporal Analysis of the Honey Bee Microbiome Reveals Four Novel Viruses and Seasonal Prevalence of Known Viruses, Nosema, and Crithidia. PLoS ONE 2011, 6, e20656. [Google Scholar] [CrossRef]
- Li, J.L.; Cornman, R.S.; Evans, J.D.; Pettis, J.S.; Zhao, Y.; Murphy, C.; Peng, W.J.; Wu, J.; Hamilton, M.; Boncristiani, H.F.; et al. Systemic spread and propagation of a plant-pathogenic virus in European honeybees, Apis mellifera. mBio 2014, 5, e00898-13. [Google Scholar] [CrossRef]
- Glenny, W.; Cavigli, I.; Daughenbaugh, K.F.; Radford, R.; Kegley, S.E.; Flenniken, M.L. Honey bee (Apis mellifera) colony health and pathogen composition in migratory beekeeping operations involved in California almond pollination. PLoS ONE 2017, 12, e0182814. [Google Scholar] [CrossRef] [PubMed]
- Traynor, K.S.; Rennich, K.; Forsgren, E.; Rose, R.; Pettis, J.; Kunkel, G.; Madella, S.; Evans, J.; Lopez, D.; vanEngelsdorp, D. Multiyear survey targeting disease incidence in US honey bees. Apidologie 2016, 47, 325–347. [Google Scholar] [CrossRef]
- McMenamin, A.J.; Flenniken, M.L. Recently identified bee viruses and their impact on bee pollinators. Curr. Opin. Insect Sci. 2018, 26, 120–129. [Google Scholar] [CrossRef]
- Yue, C.; Genersch, E. RT-PCR analysis of Deformed wing virus in honeybees (Apis mellifera) and mites (Varroa destructor). J. Gen. Virol. 2005, 86, 3419–3424. [Google Scholar] [CrossRef]
- Mockel, N.; Gisder, S.; Genersch, E. Horizontal transmission of deformed wing virus: Pathological consequences in adult bees (Apis mellifera) depend on the transmission route. J. Gen. Virol. 2011, 92, 370–377. [Google Scholar] [CrossRef] [PubMed]
- Locke, B.; Semberg, E.; Forsgren, E.; de Miranda, J.R. Persistence of subclinical deformed wing virus infections in honeybees following Varroa mite removal and a bee population turnover. PLoS ONE 2017, 12, e0180910. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Levitt, A.L.; Rajotte, E.G.; Holmes, E.C.; Ostiguy, N.; vanEngelsdorp, D.; Lipkin, W.I.; dePamphilis, C.W.; Toth, A.L.; Cox-Foster, D.L. RNA Viruses in Hymenopteran Pollinators: Evidence of Inter-Taxa Virus Transmission via Pollen and Potential Impact on Non-Apis Hymenopteran Species. PLoS ONE 2010, 5, e14357. [Google Scholar] [CrossRef]
- Shen, M.; Cui, L.; Ostiguy, N.; Cox-Foster, D. Intricate transmission routes and interactions between picorna-like viruses (Kashmir bee virus and sacbrood virus) with the honeybee host and the parasitic varroa mite. J. Gen. Virol. 2005, 86, 2281–2289. [Google Scholar] [CrossRef] [PubMed]
- Amiri, E.; Meixner, M.; Büchler, R.; Kryger, P. Chronic Bee Paralysis Virus in Honeybee Queens: Evaluating Susceptibility and Infection Routes. Viruses 2014, 6, 1188–1201. [Google Scholar] [CrossRef]
- de Miranda, J.R.; Cordoni, G.; Budge, G. The Acute bee paralysis virus–Kashmir bee virus–Israeli acute paralysis virus complex. J. Invertebr. Pathol. 2010, 103, S30–S47. [Google Scholar] [CrossRef]
- Koziy, R.V.; Wood, S.C.; Kozii, I.V.; van Rensburg, C.J.; Moshynskyy, I.; Dvylyuk, I.; Simko, E. Deformed Wing Virus Infection in Honey Bees (Apis mellifera L.). Vet. Pathol. 2019, 56, 636–641. [Google Scholar] [CrossRef]
- Hung, A.C.F. PCR detection of Kashmir bee virus in honey bee excreta. J. Apic. Res. 2000, 39, 103–106. [Google Scholar] [CrossRef]
- Chen, Y.P.; Pettis, J.S.; Collins, A.; Feldlaufer, M.F. Prevalence and transmission of honeybee viruses. Appl. Environ. Microbiol. 2006, 72, 606–611. [Google Scholar] [CrossRef]
- Forsgren, E. European foulbrood in honey bees. J. Invertebr. Pathol. 2010, 103, S5–S9. [Google Scholar] [CrossRef]
- White, G.F. The Cause of European Foul Brood—Gershom Franklin White—Google Books. Available online: https://books.google.co.uk/books?hl=en&lr=&id=hW4uAAAAYAAJ&oi=fnd&pg=PA1&dq=White,+G.+F.+(1912).+The+cause+of+European+Foul+Brood.+Government+Printing+Office:+Washington+D.+C.,+USA.&ots=lYQ2UQQmUG&sig=ObfSrPZAvKuoBqpdWFRU79O_sbo&redir_esc=y#v=onepage&q=Wh (accessed on 13 April 2020).
- Genersch, E.; Forsgren, E.; Pentikäinen, J.; Ashiralieva, A.; Rauch, S.; Kilwinski, J.; Fries, I. Reclassification of Paenibacillus larvae subsp. pulvifaciens and Paenibacillus larvae subsp. larvae as Paenibacillus larvae without subspecies differentiation. Int. J. Syst. Evol. Microbiol. 2006, 56, 501–511. [Google Scholar] [CrossRef] [PubMed]
- White, G.F. Bacteria of the Apiary; Technical series; Government Printing Office: Washington, DC, USA, 1906. [Google Scholar]
- Bailey, L. Honey Bee Pathology. Annu. Rev. Entomol. 1968, 1, 191–212. [Google Scholar] [CrossRef]
- Roberts, K.E.; Evison, S.E.F.; Baer, B.; Hughes, W.O.H. The cost of promiscuity: Sexual transmission of Nosema microsporidian parasites in polyandrous honey bees. Sci. Rep. 2015, 5, 10982. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Evans, J.D.; Smith, I.B.; Pettis, J.S. Nosema ceranae is a long-present and wide-spread microsporidian infection of the European honey bee (Apis mellifera) in the United States. J. Invertebr. Pathol. 2008, 97, 186–188. [Google Scholar] [CrossRef]
- Higes, M.; Martín-Hernández, R.; Botías, C.; Bailón, E.G.; González-Porto, A.V.; Barrios, L.; del Nozal, M.J.; Bernal, J.L.; Jiménez, J.J.; Palencia, P.G.; et al. How natural infection by Nosema ceranae causes honeybee colony collapse. Environ. Microbiol. 2008, 10, 2659–2669. [Google Scholar] [CrossRef]
- Higes, M.; Martín-Hernández, R.; Garrido-Bailón, E.; González-Porto, A.V.; García-Palencia, P.; Meana, A.; del Nozal, M.J.; Mayo, R.; Bernal, J.L. Honeybee colony collapse due to Nosema ceranae in professional apiaries. Environ. Microbiol. Rep. 2009, 1, 110–113. [Google Scholar] [CrossRef]
- Williams, M.-K.F.; Tripodi, A.D.; Szalanski, A.L. Molecular survey for the honey bee (Apis mellifera L.) trypanosome parasites Crithidia mellificae and Lotmaria passim. J. Apic. Res. 2019, 58, 553–558. [Google Scholar] [CrossRef]
- Rueppell, O.; Aumer, D.; Moritz, R.F. Ties between ageing plasticity and reproductive physiology in honey bees (Apis mellifera) reveal a positive relation between fecundity and longevity as consequence of advanced social evolution. Curr. Opin. Insect Sci. 2016, 16, 64–68. [Google Scholar] [CrossRef]
- Page, R.E.; Peng, C.Y.-S. Aging and development in social insects with emphasis on the honey bee, Apis mellifera L. Exp. Gerontol. 2001, 36, 695–711. [Google Scholar] [CrossRef]
- Amiri, E.; Strand, M.K.; Rueppell, O.; Tarpy, D.R. Queen Quality and the Impact of Honey Bee Diseases on Queen Health: Potential for Interactions between Two Major Threats to Colony Health. Insects 2017, 8, 48. [Google Scholar] [CrossRef]
- Rueppell, O.; Christine, S.; Mulcrone, C.; Groves, L. Aging without functional senescence in honey bee workers. Curr. Biol. 2007, 17, R274–R275. [Google Scholar] [CrossRef] [PubMed]
- Amdam, G.V.; Hartfelder, K.; Norberg, K.; Hagen, A.; Omholt, S.W. Altered Physiology in Worker Honey Bees (Hymenoptera: Apidae) Infested with the Mite Varroa destructor (Acari: Varroidae): A Factor in Colony Loss During Overwintering? J. Econ. Entomol. 2004, 97, 741–747. [Google Scholar] [CrossRef]
- Lee, K.V.; Goblirsch, M.; Mcdermott, E.; Tarpy, D.R.; Spivak, M. Is the Brood Pattern within a Honey Bee Colony a Reliable Indicator of Queen Quality? Insects 2019, 10, 12. [Google Scholar] [CrossRef] [PubMed]
- Evans, J.D.; Schwarz, R.S.; Chen, Y.P.; Budge, G.; Cornman, R.S.; De la Rua, P.; de Miranda, J.R.; Foret, S.; Foster, L.; Gauthier, L.; et al. Standard methods for molecular research in Apis mellifera. J. Apic. Res. 2013, 52, 1–54. [Google Scholar] [CrossRef]
- De Jong, D.; De ANDREA Roma, D.; Goncalves, L.S. A comparative analysis of shaking solutions for the detection of Varroa jacobsoni on adult honey bees. Apidologie 1982, 13, 297–306. [Google Scholar] [CrossRef]
- Invitrogen. TRIzol Reagent Manual; Life Technologies Cooperation: Carlsbad, CA, USA, 2016. [Google Scholar]
- Alburaki, M.; Chen, D.; Skinner, J.A.; Meikle, W.G.; Tarpy, D.R.; Adamczyk, J.; Stewart, S.D. Honey bee survival and pathogen prevalence: From the perspective of landscape and exposure to pesticides. Insects 2018, 9, 1–15. [Google Scholar] [CrossRef]
- Kevill, J.L.; Highfield, A.; Mordecai, G.J.; Martin, S.J.; Schroeder, D.C. ABC assay: Method development and application to quantify the role of three DWV master variants in overwinter colony losses of European honey bees. Viruses 2017, 9, 314. [Google Scholar] [CrossRef]
- Dunn, O.J. Multiple Comparisons among Means. J. Am. Stat. Assoc. 1961, 56, 52–64. [Google Scholar] [CrossRef]
- Chaimanee, V.; Chantawannakul, P.; Chen, Y.; Evans, J.D.; Pettis, J.S. Effects of host age on susceptibility to infection and immune gene expression in honey bee queens (Apis mellifera) inoculated with Nosema ceranae. Apidologie 2014, 45, 451–463. [Google Scholar] [CrossRef]
- Chen, Y.; Pettis, J.S.; Feldlaufer, M.F. Detection of multiple viruses in queens of the honey bee Apis mellifera L. J. Invertebr. Pathol. 2005, 90, 118–121. [Google Scholar] [CrossRef]
- Amiri, E.; Kryger, P.; Meixner, M.D.; Strand, M.K.; Tarpy, D.R.; Rueppell, O. Quantitative patterns of vertical transmission of deformed wing virus in honey bees. PLoS ONE 2018, 13, e0195283. [Google Scholar] [CrossRef] [PubMed]
- Daughenbaugh, K.F.; Martin, M.; Brutscher, L.M.; Cavigli, I.; Garcia, E.; Lavin, M.; Flenniken, M.L. Honey Bee Infecting Lake Sinai Viruses. Viruses 2015, 7, 3285–3309. [Google Scholar] [CrossRef] [PubMed]
- Ravoet, J.; De Smet, L.; Wenseleers, T.; de Graaf, D.C. Genome sequence heterogeneity of Lake Sinai Virus found in honey bees and Orf1/RdRP-based polymorphisms in a single host. Virus Res. 2015, 201, 67–72. [Google Scholar] [CrossRef] [PubMed]
- FRIES, I. Comb replacement and Nosema dieseas (Nosema apiz Z.)in honey bee colonies. Apidologie 1988, 19, 343–354. [Google Scholar] [CrossRef]
- Higes, M.; Martín-Hernández, R.; García-Palencia, P.; Marín, P.; Meana, A. Horizontal transmission of Nosema ceranae (Microsporidia) from worker honeybees to queens (Apis mellifera). Environ. Microbiol. Rep. 2009, 1, 495–498. [Google Scholar] [CrossRef]
- Cox-Foster, D.L.; Conlan, S.; Holmes, E.C.; Palacios, G.; Evans, J.D.; Moran, N.A.; Quan, P.L.; Briese, T.; Hornig, M.; Geiser, D.M.; et al. A Metagenomic Survey of Microbes in Honey Bee Colony Collapse Disorder. Science 2007, 318, 283–287. [Google Scholar] [CrossRef]
- Maori, E.; Lavi, S.; Mozes-Koch, R.; Gantman, Y.; Peretz, Y.; Edelbaum, O.; Tanne, E.; Sela, I. Isolation and characterization of Israeli acute paralysis virus, a dicistrovirus affecting honeybees in Israel: Evidence for diversity due to intra- and inter-species recombination. J. Gen. Virol. 2007, 88, 3428–3438. [Google Scholar] [CrossRef]
- Amiri, E.; Seddon, G.; Zuluaga Smith, W.; Strand, M.K.; Tarpy, D.R.; Rueppell, O. Israeli Acute Paralysis Virus: Honey Bee Queen–Worker Interaction and Potential Virus Transmission Pathways. Insects 2019, 10, 9. [Google Scholar] [CrossRef]
- Fievet, J.; Tentcheva, D.; Gauthier, L.; de Miranda, J.; Cousserans, F.; Colin, M.; Bergoin, M. Localization of deformed wing virus infection in queen and drone Apis mellifera L. Virol. J. 2006, 3, 16. [Google Scholar] [CrossRef]
- Evans, J.D.; Wheeler, D.E. Differential gene expression between developing queens and workers in the honey bee, Apis mellifera. Proc. Natl. Acad. Sci. USA 1999, 96, 5575–5580. [Google Scholar] [CrossRef]
- DeGrandi-Hoffman, G.; Chen, Y.; Huang, E.; Huang, M.H. The effect of diet on protein concentration, hypopharyngeal gland development and virus load in worker honey bees (Apis mellifera L.). J. Insect Physiol. 2010, 56, 1184–1191. [Google Scholar] [CrossRef] [PubMed]
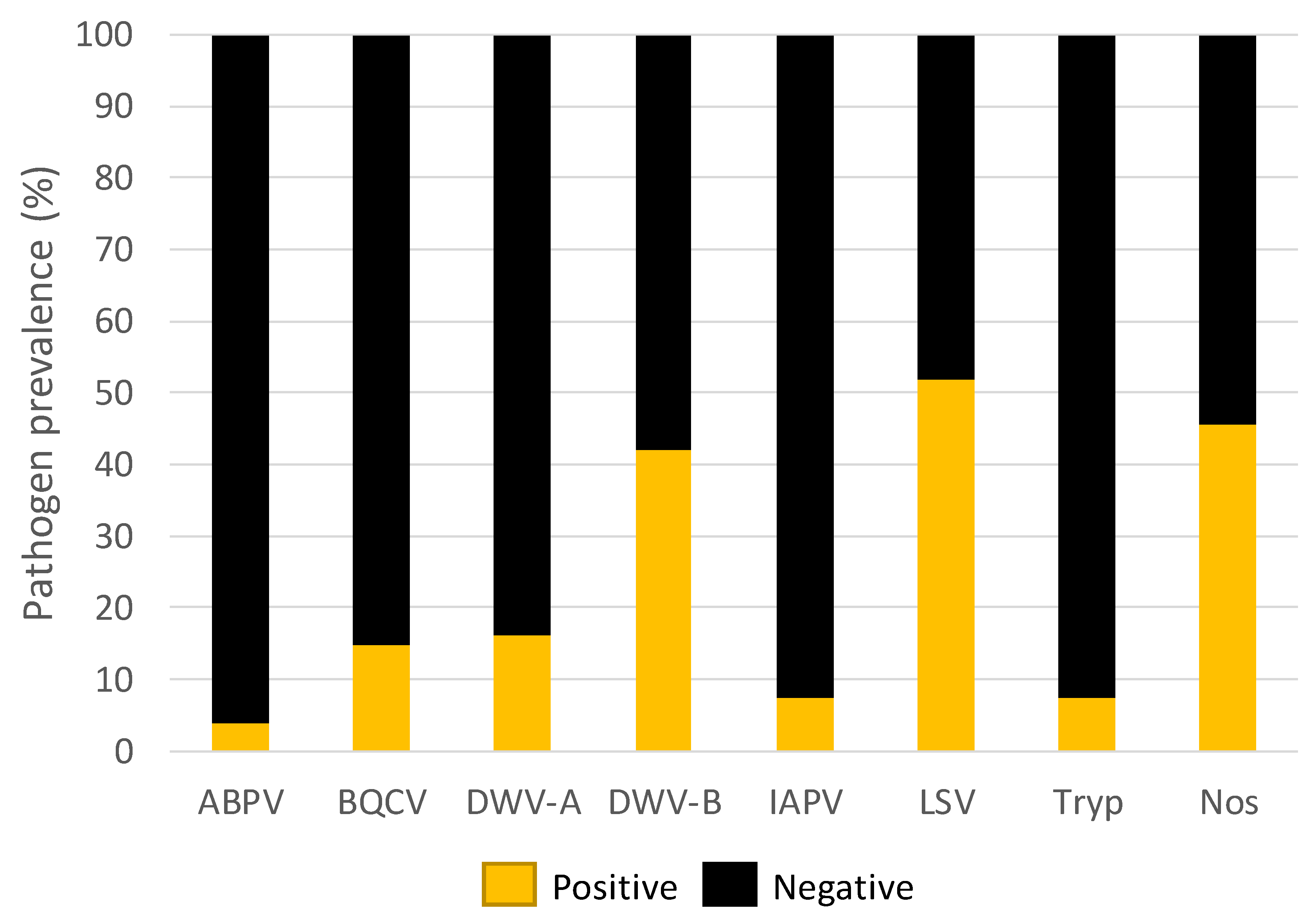
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kevill, J.L.; Lee, K.; Goblirsch, M.; McDermott, E.; Tarpy, D.R.; Spivak, M.; Schroeder, D.C. The Pathogen Profile of a Honey Bee Queen Does Not Reflect That of Her Workers. Insects 2020, 11, 382. https://doi.org/10.3390/insects11060382
Kevill JL, Lee K, Goblirsch M, McDermott E, Tarpy DR, Spivak M, Schroeder DC. The Pathogen Profile of a Honey Bee Queen Does Not Reflect That of Her Workers. Insects. 2020; 11(6):382. https://doi.org/10.3390/insects11060382
Chicago/Turabian StyleKevill, Jessica L., Katie Lee, Michael Goblirsch, Erin McDermott, David R. Tarpy, Marla Spivak, and Declan C. Schroeder. 2020. "The Pathogen Profile of a Honey Bee Queen Does Not Reflect That of Her Workers" Insects 11, no. 6: 382. https://doi.org/10.3390/insects11060382
APA StyleKevill, J. L., Lee, K., Goblirsch, M., McDermott, E., Tarpy, D. R., Spivak, M., & Schroeder, D. C. (2020). The Pathogen Profile of a Honey Bee Queen Does Not Reflect That of Her Workers. Insects, 11(6), 382. https://doi.org/10.3390/insects11060382





