A Guild-Based Protocol to Target Potential Natural Enemies of Philaenus spumarius (Hemiptera: Aphrophoridae), a Vector of Xylella fastidiosa (Xanthomonadaceae): A Case Study with Spiders in the Olive Grove
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Area
2.2. Functional Response Assay
2.2.1. Selection of Predator and Prey Species
2.2.2. Laboratory Trials
2.3. Field Sampling Scheme
Sampling of Araneae
2.4. Data Analysis
2.4.1. Functional Response Assay
2.4.2. Field Sampling Scheme
3. Results
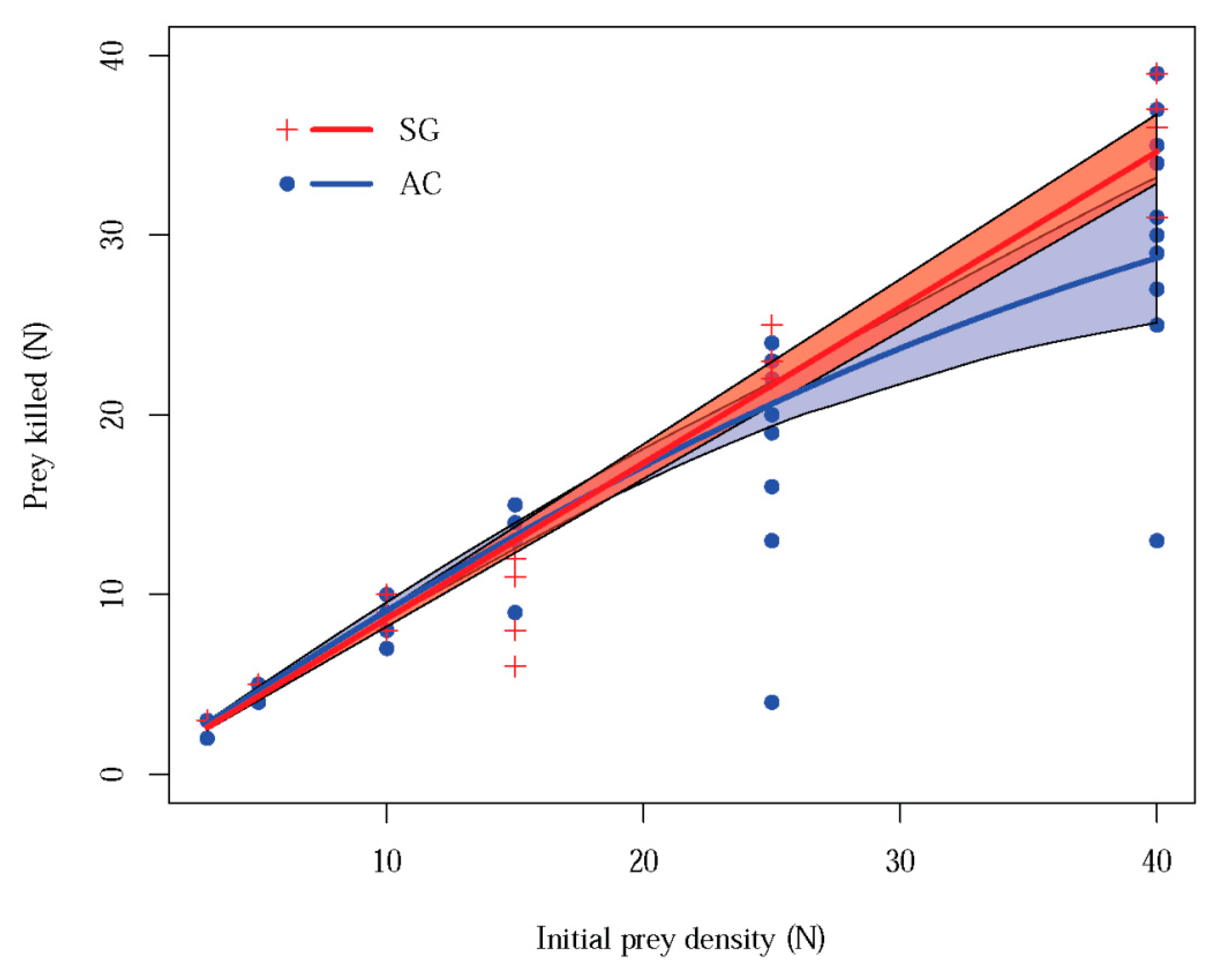
3.1. Functional Response Laboratory Assay
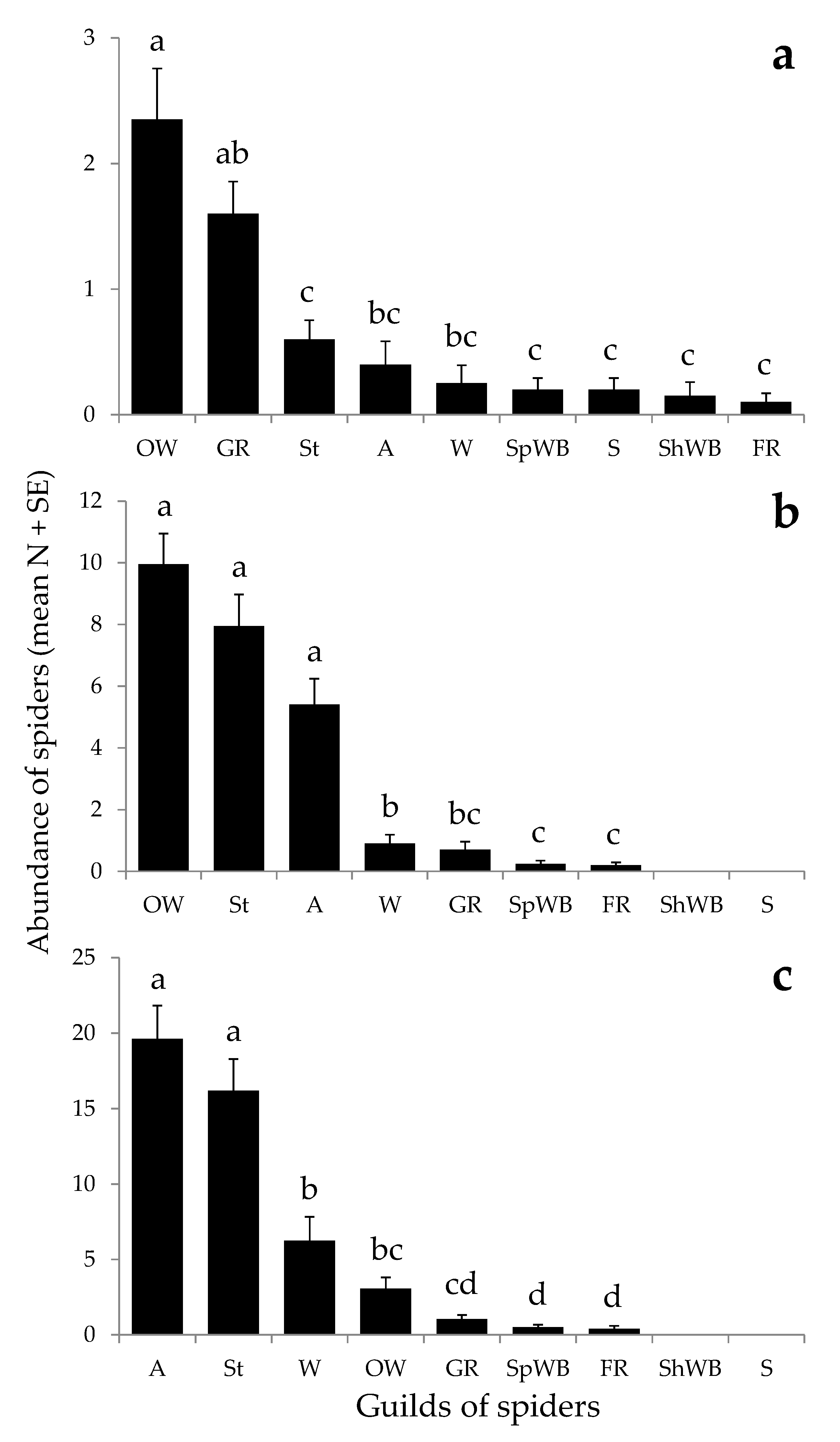
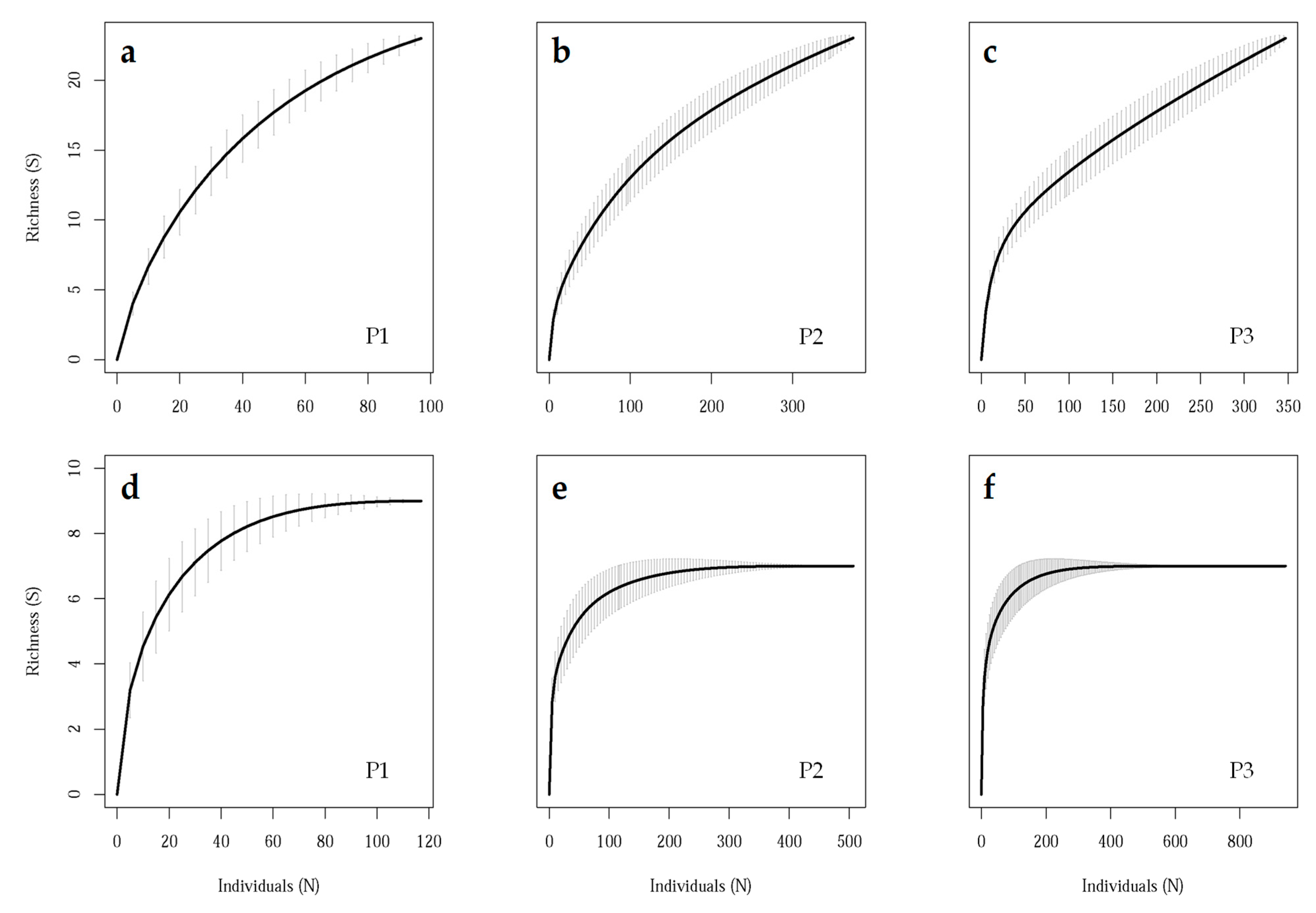
3.2. Field Sampling Scheme
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- FAOSTAT. FAOSTAT Statistics Database; Food and Agriculture Organization of the United Nations: Rome, Italy, 2018; Available online: http://www.fao.org/faostat/en/ (accessed on 18 August 2018).
- Saponari, M.; Boscia, D.; Nigro, F.; Martelli, G.P. Identification of DNA sequences related to Xylella fastidiosa in oleander, almond and olive trees exhibiting leaf scorch symptoms in Apulia (southern Italy). J. Plant Pathol. 2013, 95, 668. [Google Scholar] [CrossRef]
- Saponari, M.; Boscia, D.; Altamura, G.; Loconsole, G.; Zicca, S.; D’Attoma, G.; Morelli, M.; Palmisano, F.; Saponari, A.; Tavano, D.; et al. Isolation and pathogenicity of Xylella fastidiosa associated to the olive quick decline syndrome in southern Italy. Sci. Rep. 2017, 7, 17723. [Google Scholar] [CrossRef]
- Cornara, D.; Cavalieri, V.; Dongiovanni, C.; Altamura, G.; Palmisano, F.; Bosco, D.; Porcelli, F.; Almeida, R.P.P.; Saponari, M. Transmission of Xylella fastidiosa by naturally infected Philaenus spumarius (Hemiptera, Aphrophoridae) to different host plants. J. Appl. Entomol. 2017, 141, 80–87. [Google Scholar] [CrossRef]
- Marshall, A. Spittle-production and tube-building by cercopid larvae (Homoptera)–IV. Mucopolysaccharide associated with spittle-production. J. Insect Physiol. 1966, 12, 635–644. [Google Scholar] [CrossRef]
- Mello, M.L.S.; Pimentel, E.; Yamada, A.; Storopoli-Neto, A. Composition and structure of the froth of the spittlebug, Deois sp. Insect Biochem. 1987, 17, 493–502. [Google Scholar] [CrossRef]
- Weaver, C.R.; King, D.R. Meadow spittlebug Philaenus leucophthalmus (L.). Ohio. Agric. Exp. Stn. Res. Bull. 1954, 741, 1–99. [Google Scholar]
- Almeida, R.P.; Purcell, A.H. Patterns of Xylella fastidiosa colonization on the precibarium of sharpshooter vectors relative to transmission to plants. Ann. Entomol. Soc. Am. 2006, 99, 884–890. [Google Scholar] [CrossRef]
- Killiny, N.; Almeida, R.P. Xylella fastidiosa afimbrial adhesins mediate cell transmission to plants by leafhopper vectors. Appl. Environ. Microbiol. 2009, 75, 521–528. [Google Scholar] [CrossRef]
- Janse, J.D.; Obradovic, A. Xylella fastidiosa: Its biology, diagnosis, control and risks. J. Plant Pathol. 2010, 92, S1.35–S1.48. [Google Scholar]
- Whittaker, J.B. Density regulation in a population of Philaenus spumarius (L.) (Homoptera: Cercopidae). J. Anim. Ecol. 1973, 42, 163–172. [Google Scholar] [CrossRef]
- Cornara, D.; Bosco, D.; Fereres, A. Philaenus spumarius: When an old acquaintance becomes a new threat to European agriculture. J. Pest Sci. 2018, 91, 957–972. [Google Scholar] [CrossRef]
- Salerno, M.; Russo, V.; Sefa, V.; Lamaj, F.; Basher, N.; Verrastro, V.; Porcelli, F. Zelus renardii an assassin bug candidate for Philaenus spumarius biocontrol. In Proceedings of the European conference on Xylella. Finding Answer to a Global Problem, Palma del Mallorc, Spain, 13–15 November 2017; pp. 22–23. [Google Scholar]
- Osborne, L.S.; Cuda, J.P. Release of Exotic Natural Enemies for Biological Control: A Case of Damned If We Do and Damned If We Don’t? Land Use Environ. Law 2003, 18, 399–407. [Google Scholar]
- Van Driesche, R.G.; Bellows, T.S. Introduction of New Natural Enemies: Methods. In Biological Control; Van Driesche, R.G., Bellows, T.S., Eds.; Springer: Boston, MA, USA, 1996; pp. 158–177. [Google Scholar]
- Gontijo, L.M. Engineering natural enemy shelters to enhance conservation biological control in field crops. Biol. Control. 2019, 130, 155–163. [Google Scholar] [CrossRef]
- Landis, D.A.; Wratten, D.; Gurr, G.M. Habitat management to conserve natural enemies of arthropod pests in agriculture. Annu. Rev. Entomol. 2000, 45, 175–201. [Google Scholar] [CrossRef]
- Wiedenmann, R.N.; Smith, J.W., Jr. Attributes of natural enemies in ephemeral crop habitats. Biol. Control. 1997, 10, 16–22. [Google Scholar] [CrossRef]
- Dainese, M.; Schneider, G.; Krauss, J.; Steffan-Dewenter, I. Complementarity among natural enemies enhances pest suppression. Sci. Rep. 2017, 7, 8172. [Google Scholar] [CrossRef]
- Nyffeler, M.; Benz, G. Spiders in natural pest control: A review. J. Appl. Entomol. 1987, 103, 321–339. [Google Scholar] [CrossRef]
- Bogya, S.; Mols, P.J.M. The role of spiders as predators of insect pests with particular reference to orchards: A review. Acta Phytopathol. Entomol. Hung. 1996, 31, 83–159. [Google Scholar]
- Michalko, R.; Pekár, S.; Entling, M.H. An updated perspective on spiders as generalist predators in biological control. Oecologia 2019, 189, 21–36. [Google Scholar] [CrossRef]
- Uetz, G.W.; Halaj, J.; Cady, A. Guild structure of spiders in major crops. J. Arachnol. 1999, 27, 270–280. [Google Scholar]
- Michalko, R.; Pekár, S. Different hunting strategies of generalist predators result in functional differences. Oecologia 2016, 181, 1187–1197. [Google Scholar] [CrossRef]
- Greenstone, M.H. Spider Predation: How and Why We Study It. J. Arachnol. 1999, 27, 333–342. [Google Scholar]
- Fick, S.E.; Hijmans, R.J. Worldclim 2: New 1-km spatial resolution climate surfaces for global land areas. Int. J. Climatol. 2017, 37, 4302–4315. Available online: http://worldclim.org/version2 (accessed on 18 February 2019).
- Dinis, A.M.; Pereira, J.A.; Benhadi-Marín, J.; Santos, S.A.P. Feeding preferences and functional responses of Calathus granatensis and Pterostichus globosus (Coleoptera: Carabidae) on pupae of Bactrocera oleae (Diptera: Tephritidae). Bull. Entomol. Res. 2016, 106, 701–709. [Google Scholar] [CrossRef]
- Benhadi-Marín, J.; Pereira, J.A.; Barreales, D.; Sousa, J.P.; Santos, S.A.P. A simulation-based method to compare the pest suppression potential of predators: A case study with spiders. Biol. Control 2018, 123, 87–96. [Google Scholar] [CrossRef]
- Benhadi-Marín, J.; Pereira, J.A.; Sousa, J.P.; Santos, S.A.P. Functional responses of three guilds of spiders: Comparing single- and multiprey approaches. Ann. Appl. Biol. 2019, 175, 202–214. [Google Scholar] [CrossRef]
- Nentwig, W.; Blick, T.; Gloor, D.; Hänggi, A.; Kropf, C. Spiders of Europe. 2019. Available online: http://www.araneae.unibe.ch (accessed on 15 October 2019).
- Cardoso, P.; Pekár, S.; Jocqué, R.; Coddington, J.A. Global patterns of guild composition and functional diversity of spiders. PLoS ONE 2011, 6, e21710. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018; Available online: http://www.R-project.org (accessed on 1 February 2019).
- Pritchard, D. Frair: Tools for Functional Response Analysis 2017. R Package Version 0.5.100. Available online: https://CRAN.R-project.org/package=frair (accessed on 30 September 2019).
- Juliano, S.A. Nonlinear curve fitting: Predation and functional response curve. In Design and Analysis of Ecological Experiments; Scheiner, S.M., Gurevitch, J., Eds.; Oxford University Press: New York, NY, USA, 2001; pp. 178–196. [Google Scholar]
- Trexler, J.C.; McCulloch, C.E.; Travis, J. How can the functional response best be determined? Oecologia 1988, 76, 206–214. [Google Scholar] [CrossRef]
- Rogers, D. Random search and insect population models. J. Anim. Ecol. 1972, 41, 369–383. [Google Scholar] [CrossRef]
- Bolker, B. Ecological Models and Data in R; Princeton University Press: Princeton, NJ, USA, 2008; p. 408. [Google Scholar]
- Jacobs, J. Unpublished Data. Available online: https://github.com/dacb/variableO2_illumina_analyses/blob/master/rarefaction.txt (accessed on 20 December 2019).
- Oksanen, J.; Blanchet, F.G.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Vegan: Community Ecology Package. R Package Version 2.5-3 2018. Available online: https://cran.r-project.org/web/packages/vegan/index.html (accessed on 2 January 2020).
- Zuur, A.; Ieno, E.N.; Walker, N.; Saveliev, A.A.; Smith, G.M. Mixed Effects Models and Extensions in Ecology with R.; Springer: New York, NY, USA, 2009. [Google Scholar]
- Pekár, S.; Brabec, M. Generalized estimating equations: A pragmatic and flexible approach to the marginal GLM modelling of correlated data in the behavioural sciences. Ethology 2018, 124, 86–93. [Google Scholar] [CrossRef]
- Cornara, D.; Saponari, M.; Zeilinger, A.R.; de Stradis, A.; Boscia, D.; Loconsole, G.; Bosco, D.; Martelli, G.P.; Almeida, R.P.P.; Porcelli, F. Spittlebugs as vectors of Xylella fastidiosa in olive orchards in Italy. J. Pest Sci. 2017, 90, 521–530. [Google Scholar] [CrossRef]
- Cruaud, A.; Gonzalez, A.A.; Godefroid, M.; Nidelet, S.; Streito, J.C.; Thuillier, J.M.; Rossi, J.P.; Santoni, S.; Rasplus, J.Y. Using insects to detect, monitor and predict the distribution of Xylella fastidiosa: A case study in Corsica. Sci. Rep. 2018, 8, 15628. [Google Scholar] [CrossRef]
- Antonatos, S.; Papachristos, D.; Kapantaidaki, D.E.; Lytra, I.C.; Varikou, K.; Evangelou, V.I.; Milonas, P. Presence of Cicadomorpha in olive orchards of Greece with special reference to Xylella fastidiosa vectors. J. Appl. Entomol. 2020, 144, 1–11. [Google Scholar] [CrossRef]
- Morente, M.; Cornara, D.; Plaza, M.; Durán, J.M.; Capiscol, C.; Trillo, R.; Ruiz, M.; Ruz, C.; Sanjuan, S.; Pereira, J.A.; et al. Distribution and relative abundance of insect vectors of Xylella fastidiosa in olive groves of the Iberian Peninsula. Insects 2018, 9, 175. [Google Scholar] [CrossRef]
- Santoiemma, G.; Tamburini, G.; Sanna, F.; Mori, N.; Marini, L. Landscape composition predicts the distribution of Philaenus spumarius, vector of Xylella fastidiosa, in olive groves. J. Pest Sci. 2019, 92, 1101–1109. [Google Scholar] [CrossRef]
- Ben Moussa, I.E.; Mazzoni, V.; Valentini, F.; Yaseen, T.; Lorusso, D.; Speranza, S.; Digiaro, M.; Varvaro, L.; Krugner, R.; D’Onghia, A.M. Seasonal fluctuations of sap-feeding insect species infected by Xylella fastidiosa in Apulian olive groves of Southern Italy. J. Econ. Entomol. 2016, 109, 1512–1518. [Google Scholar] [CrossRef]
- Llandres, A.L.; Rodríguez-Gironés, M.A. Spider movement, UV reflectance and size, but not spider crypsis, affect the response of honeybees to Australian crab spiders. PLoS ONE 2011, 6, e17136. [Google Scholar] [CrossRef]
- Kuhn-Nentwig, L.; Stöcklin, R.; Nentwig, W. Venom composition and strategies in spiders: Is Everything Possible. In Advances in Insect Physiology; Casas, J., Ed.; Academic Press: Cambridge, MA, USA, 2011; Volume 40, pp. 1–86. [Google Scholar]

| Guild | Family | Species | Abundance (N) |
|---|---|---|---|
| Ambushers | Philodromidae | Philodromus albidus Kulczyński, 1911 | 1 |
| Philodromus sp. | 28 | ||
| Pisauridae | Pisaura mirabilis (Clerck, 1757) | 6 | |
| Thomisidae | Runcinia grammica (C. L. Koch, 1837) | 73 | |
| Synema globosum (Fabricius, 1775) | 26 | ||
| Thomisus onustus Walckenaer, 1805 | 22 | ||
| Xysticus cristatus (Clerck, 1757) | 1 | ||
| Xysticus kochi Thorell, 1872 | 12 | ||
| Immatures | 339 | ||
| Foliage runners | Cheiracanthiidae | Cheiracanthium erraticum (Walckenaer, 1802) | 5 |
| Cheiracanthium sp. | 8 | ||
| Sparassidae | Micrommata ligurina (C. L. Koch, 1845) | 1 | |
| Ground runners | Dysderidae | Dysdera crocata C. L. Koch, 1838 | 2 |
| Gnaphosidae | Haplodrassus rufipes (Lucas, 1846) | 4 | |
| Nomisia exornata (C. L. Koch, 1839) | 1 | ||
| Nomisia sp. | 4 | ||
| Zelotes sp. | 5 | ||
| Immatures | 13 | ||
| Lycosidae | Alopecosa albofasciata (Brullé, 1832) | 1 | |
| Arctosa villica (Lucas, 1846) | 2 | ||
| Pardosa proxima (C. L. Koch, 1847) | 13 | ||
| Immatures | 22 | ||
| Orb-weavers | Araneidae | Aculepeira ceropegia (Walckenaer, 1802) | 86 |
| Agalenatea redii (Scopoli, 1763) | 6 | ||
| Cyclosa algerica Simon, 1885 | 1 | ||
| Hypsosinga albovittata (Westring, 1851) | 3 | ||
| Mangora acalypha (Walckenaer, 1802) | 201 | ||
| Uloboridae | Uloborus walckenaerius Latreille, 1806 | 10 | |
| Sheet web builders | Agelenidae | Eratigena feminea (Simon, 1870) | 3 |
| Space web builders | Dictynidae | Immatures | 1 |
| Theridiidae | Asagena phalerata (Panzer, 1801) | 3 | |
| Crustulina sp. 1 | 1 | ||
| Euryopis episinoides (Walckenaer, 1847) | 1 | ||
| Phylloneta impressa (L. Koch, 1881) | 7 | ||
| Simitidion simile (C. L. Koch, 1836) | 1 | ||
| Theridiidae sp. 1 | 2 | ||
| Theridiidae sp. 2 | 1 | ||
| Immatures | 2 | ||
| Specialists | Zodariidae | Selamia reticulata (Simon, 1870) | 1 |
| Zodarion sp. | 3 | ||
| Stalkers | Oxyopidae | Oxyopes heterophthalmus (Latreille, 1804) | 105 |
| Oxyopes nigripalpis Kulczyński, 1891 | 86 | ||
| Oxyopes sp. | 276 | ||
| Salticidae | Chalcoscirtus infimus (Simon, 1868) | 2 | |
| Heliophanus cupreus (Walckenaer, 1802) | 2 | ||
| Icius hamatus (C. L. Koch, 1846) | 1 | ||
| Pellenes brevis (Simon, 1868) | 1 | ||
| Phlegra lineata (C. L. Koch, 1846) | 5 | ||
| Immatures | 17 | ||
| Wandering sheet/tangle weavers | Linyphiidae | Agyneta sp. | 7 |
| Erigoninae sp. | 62 | ||
| Linyphiidae sp. 1 | 1 | ||
| Pelecopsis inedita (O. P.-Cambridge, 1875) | 31 | ||
| Prinerigone vagans (Audouin, 1826) | 1 | ||
| Styloctetor romanus (O. P.-Cambridge, 1873) | 1 | ||
| Tenuiphantes tenuis (Blackwall, 1852) | 6 | ||
| Typhocrestus bogarti Bosmans, 1990 | 2 | ||
| Immatures | 37 | ||
| Araneae immatures (not identified) | 13 | ||
| Total | 1578 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Benhadi-Marín, J.; Villa, M.; Pereira, L.F.; Rodrigues, I.; Morente, M.; Baptista, P.; Pereira, J.A. A Guild-Based Protocol to Target Potential Natural Enemies of Philaenus spumarius (Hemiptera: Aphrophoridae), a Vector of Xylella fastidiosa (Xanthomonadaceae): A Case Study with Spiders in the Olive Grove. Insects 2020, 11, 100. https://doi.org/10.3390/insects11020100
Benhadi-Marín J, Villa M, Pereira LF, Rodrigues I, Morente M, Baptista P, Pereira JA. A Guild-Based Protocol to Target Potential Natural Enemies of Philaenus spumarius (Hemiptera: Aphrophoridae), a Vector of Xylella fastidiosa (Xanthomonadaceae): A Case Study with Spiders in the Olive Grove. Insects. 2020; 11(2):100. https://doi.org/10.3390/insects11020100
Chicago/Turabian StyleBenhadi-Marín, Jacinto, María Villa, Luís F. Pereira, Isabel Rodrigues, Marina Morente, Paula Baptista, and José Alberto Pereira. 2020. "A Guild-Based Protocol to Target Potential Natural Enemies of Philaenus spumarius (Hemiptera: Aphrophoridae), a Vector of Xylella fastidiosa (Xanthomonadaceae): A Case Study with Spiders in the Olive Grove" Insects 11, no. 2: 100. https://doi.org/10.3390/insects11020100
APA StyleBenhadi-Marín, J., Villa, M., Pereira, L. F., Rodrigues, I., Morente, M., Baptista, P., & Pereira, J. A. (2020). A Guild-Based Protocol to Target Potential Natural Enemies of Philaenus spumarius (Hemiptera: Aphrophoridae), a Vector of Xylella fastidiosa (Xanthomonadaceae): A Case Study with Spiders in the Olive Grove. Insects, 11(2), 100. https://doi.org/10.3390/insects11020100








