A DNA Barcoding Survey of an Arctic Arthropod Community: Implications for Future Monitoring
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Sites
2.2. Collecting Methods
- Malaise traps—three Townes-style Malaise traps (BugDorm, MegaView Science Co., Ltd., Taiwan) were installed at continuous monitoring sites: one near Water Lake and two near IMA. Samples were collected in 500-mL plastic bottles filled with ~350 mL of 95% ethanol, which were exchanged on a weekly basis. (NB: During this study, the polyester support strings used to anchor Malaise traps installed at the CHARS IMA site were damaged by arctic foxes, which, combined with strong wind gusts, caused the traps to collapse completely or partially between service dates. As a result, stock ropes were replaced with eight 1/16 inch galvanized “aircraft cables” (2.5 m in length). Cables were fitted with aluminum oval sleeves to form loops at the ends and a round swivel eye simplex snap was used to attach each cable to the trap. The distal end of each cable was anchored to the ground with a metal peg, reinforced as needed with heavy stones. These modifications prevented further collapsing of the traps for the remainder of the field season).
- Pan traps—ten yellow shallow plastic bowls (9” diameter) were installed at each of the monitoring sites and half-filled with soapy water. Traps were checked every other day, and the two-day catch from each pan trap site was pooled into a single bulk sample. Collected specimens were strained through a Nitex nylon fabric with a mesh size of 50 µm and fixed in 95% ethanol.
- Pitfall traps—a line of 10 translucent wide-mouth 500 mL plastic cups was installed at 3 m intervals at each monitoring site. The traps were half-filled with soapy water and capped with a 10 mm steel mesh to exclude vertebrate by-catch. The checking schedule and specimen collecting procedure was the same as for pan traps.
- Sweep-netting—arthropods were collected from air and from vegetation with a standard insect sweep-net, then hand-sorted or aspirated, followed by placement in a kill jar or fixation with 95% ethanol. At each of the two monitoring sites, sweep-netting was performed weekly for five continuous minutes (six times at WL site and five times at IMA site).
- Soil and litter sifting—three 30-m soil sifting transects were established parallel to the pitfall trap line at each of the two monitoring sites, with the middle transect adjacent to the pitfall traps, and one transect on either side of the pitfall line at a distance of 10 m. Ten ca. 1-L samples of litter and surface soil were taken with a trowel at 3-m intervals, sifted through a 5-mm mesh to remove larger debris, and pooled into a single bulk sample. The resulting soil and litter samples (ca. 3 L in volume) were placed in Berlese funnels for one week to extract specimens, which were collected in vials with 95% ethanol. The soil transects were sampled four times during the study period for each continuous monitoring site. In addition to the transects, samples of varying size were taken opportunistically by hand or with a trowel from soil surface, moss, debris, remains of dry vertebrate carcasses, or leaf litter in various habitats across the study area. Each sample was sifted on site through a 5 mm mesh, followed by hand sorting in the lab and fixation of all extracted organisms in 95% ethanol.
- Aquatic sampling—arthropods were collected from the water column, submerged vegetation, and bottom sediments of tundra ponds and streams using a hand-held plankton net, D-net or small hand net, then strained from water and fixed with 95% ethanol.
- Freehand collecting—individual arthropods were opportunistically picked by hand, with forceps, or hand-netted, followed by fixation with 95% ethanol.
2.3. Sample Processing and DNA Barcode Sequencing
2.4. BINs and Taxonomic Assignment
2.5. Archiving and Imaging
2.6. Data Analyses
2.7. Data Availability
3. Results
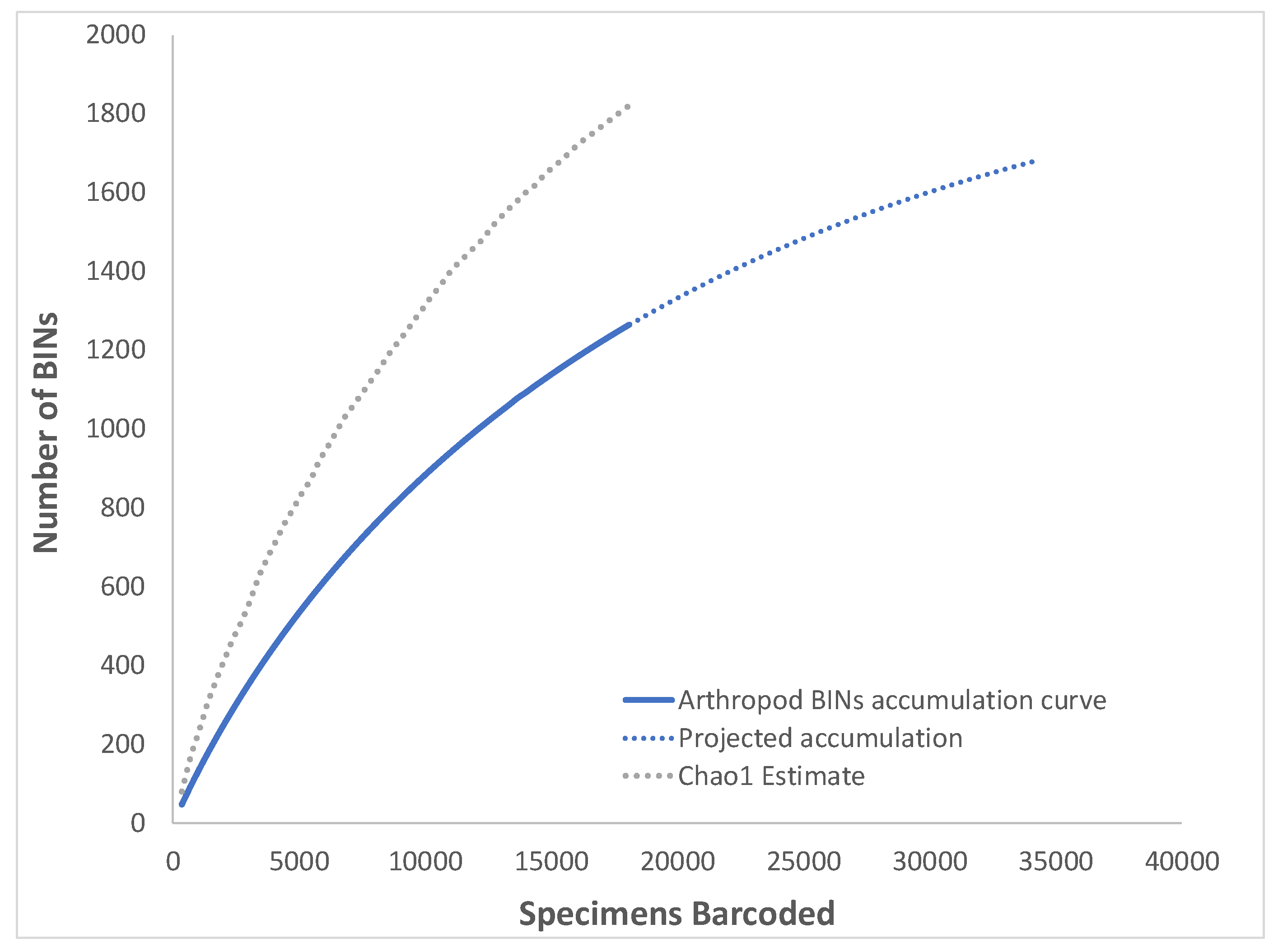
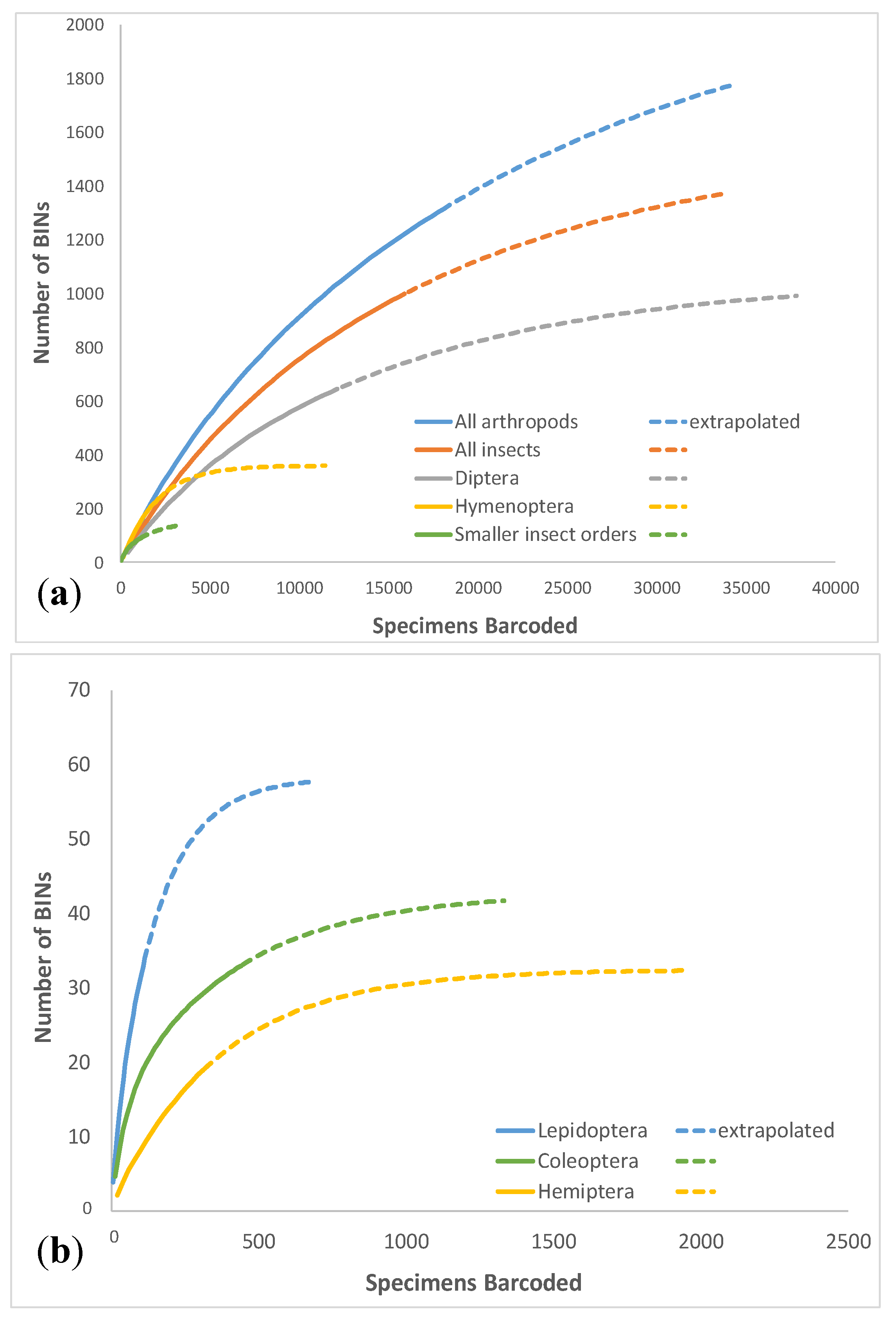
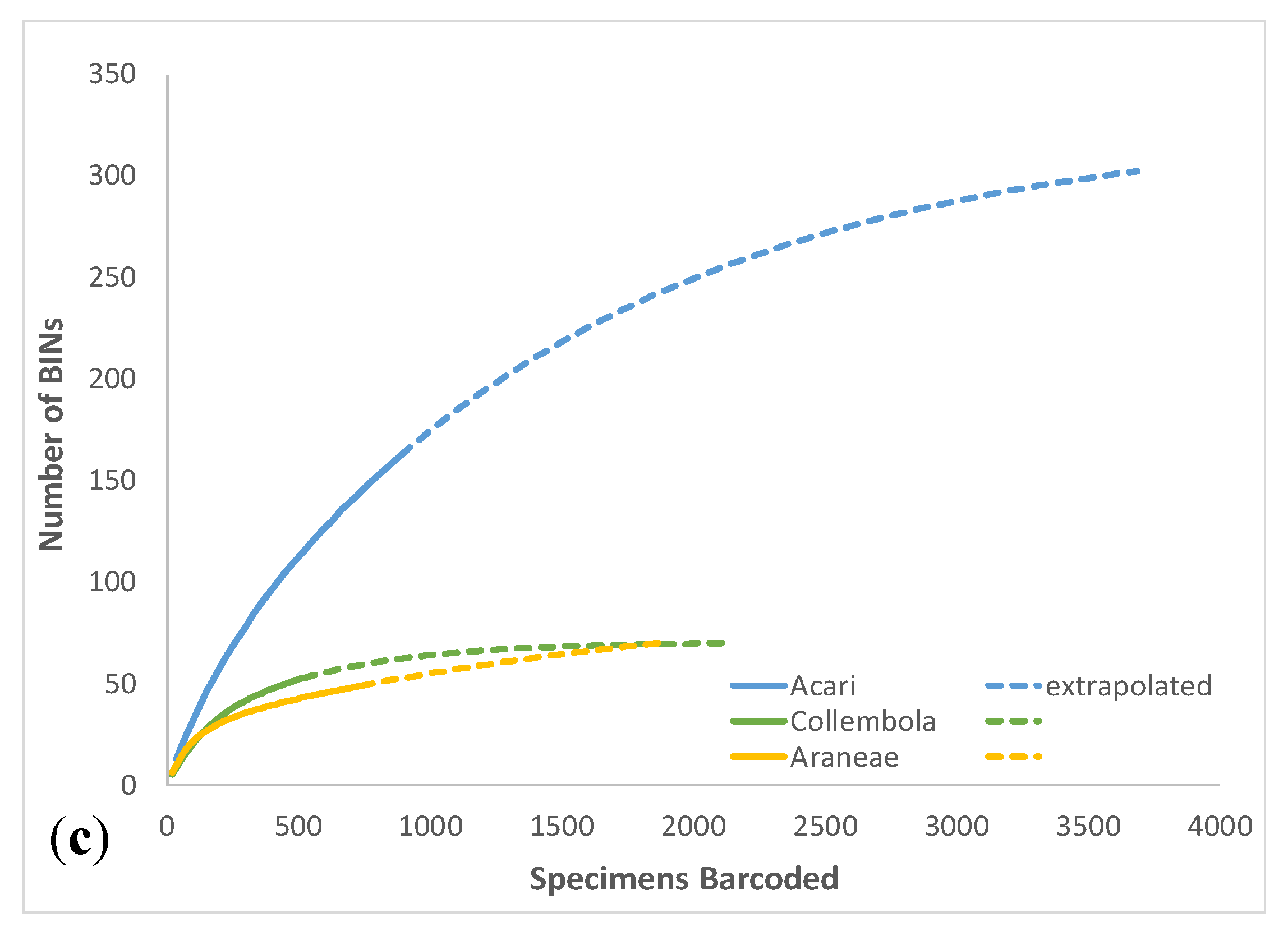
3.1. Overall Arthropod Diversity and Sequencing Success
3.2. Diversity at Continuously Monitored Sites
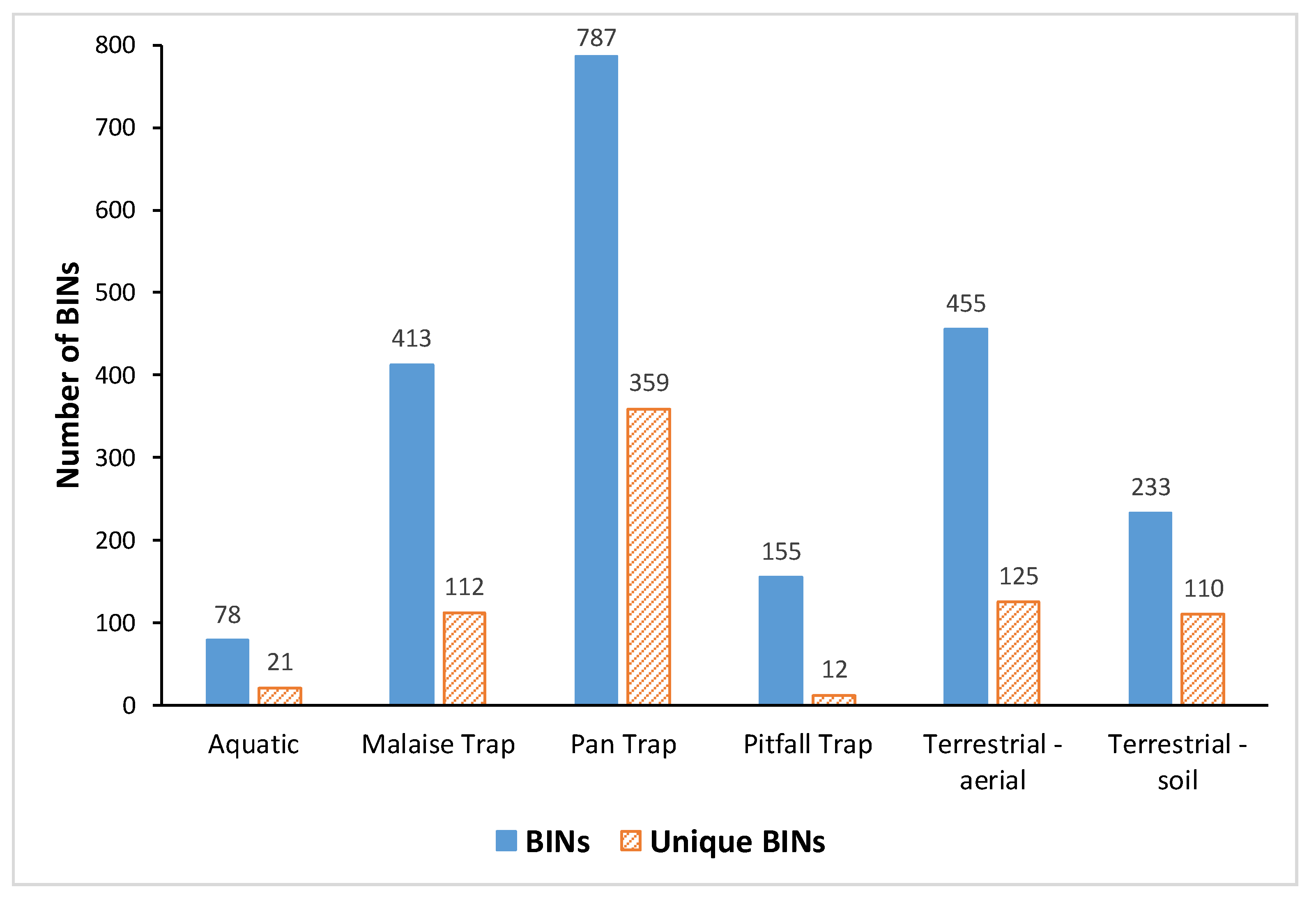
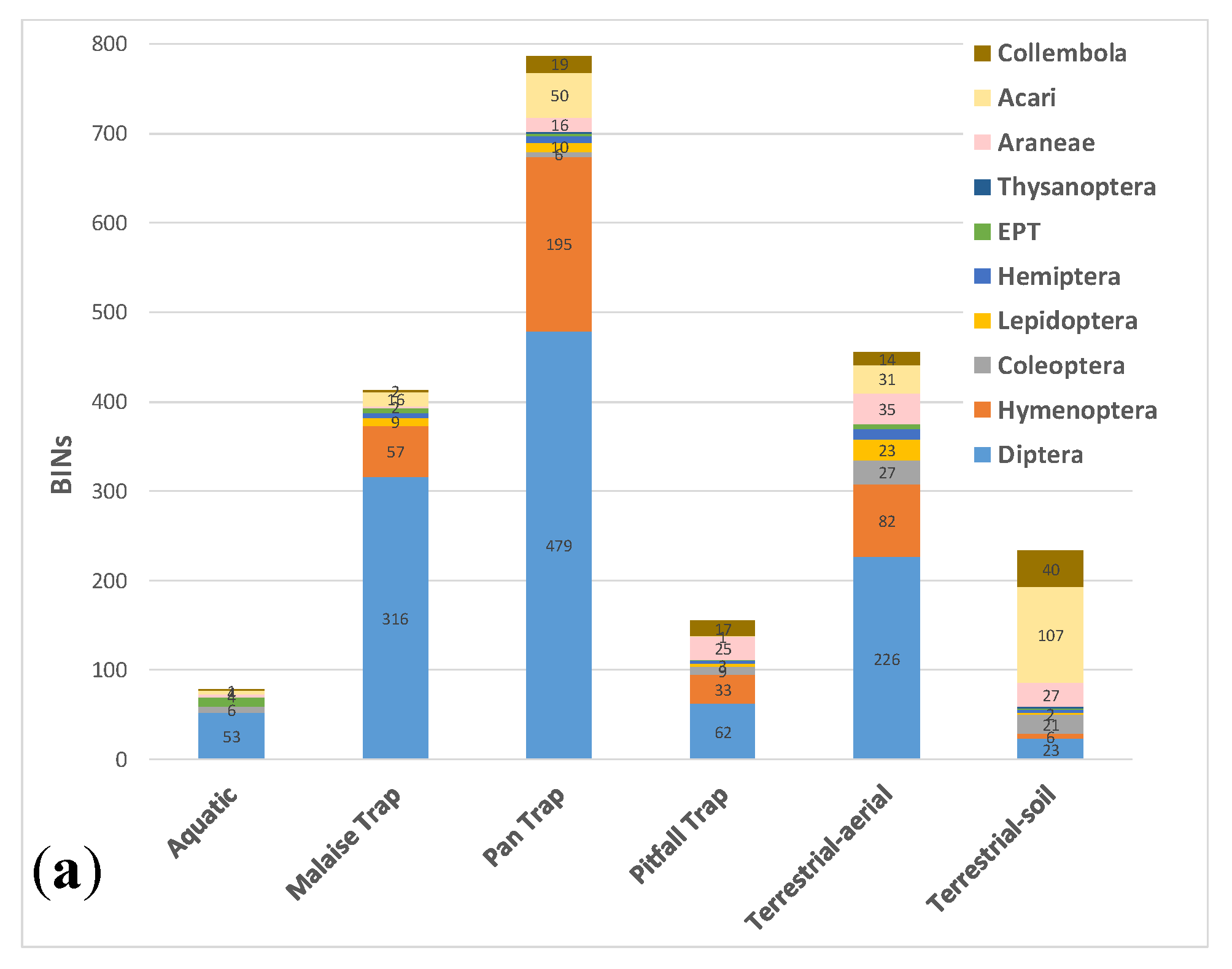
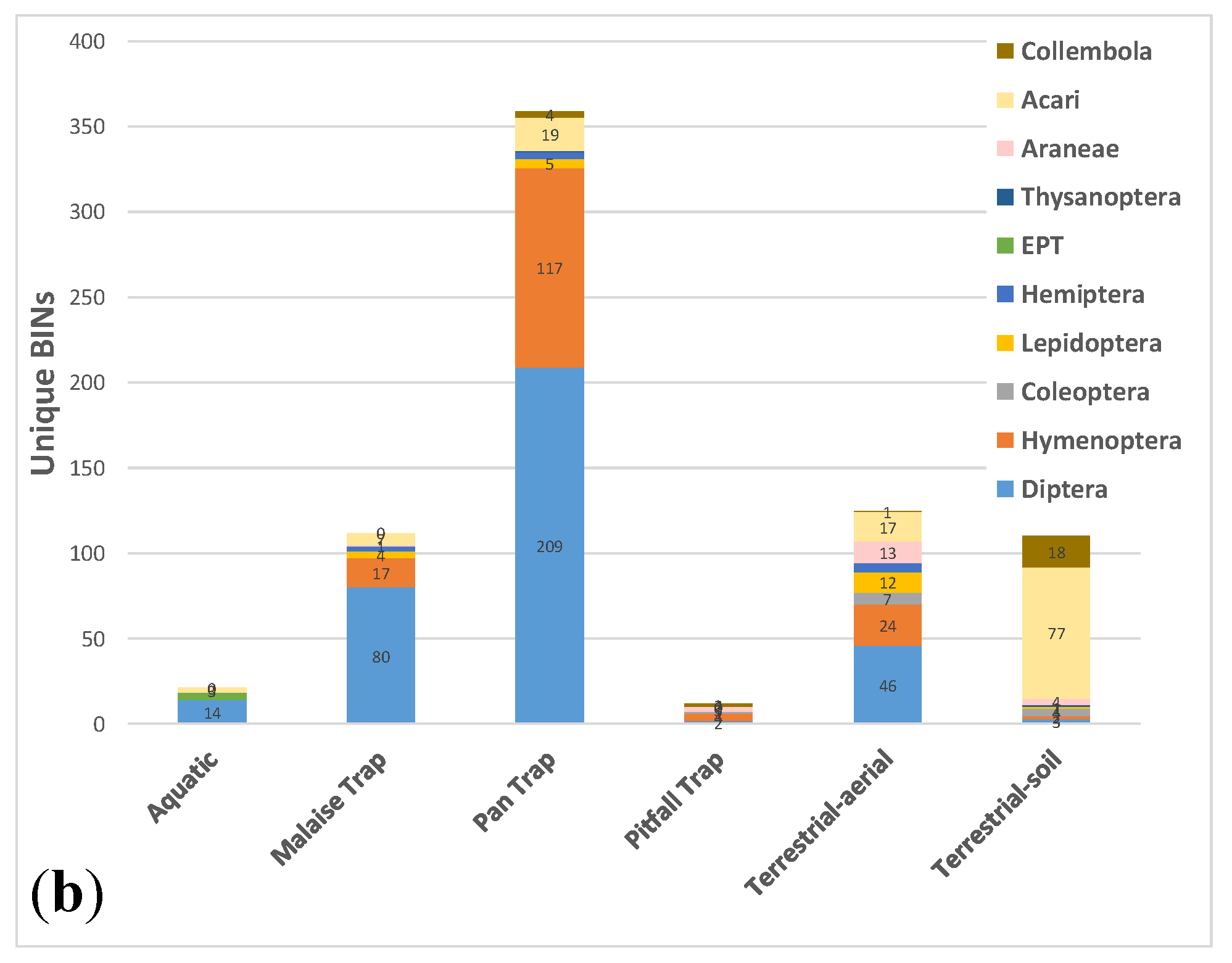
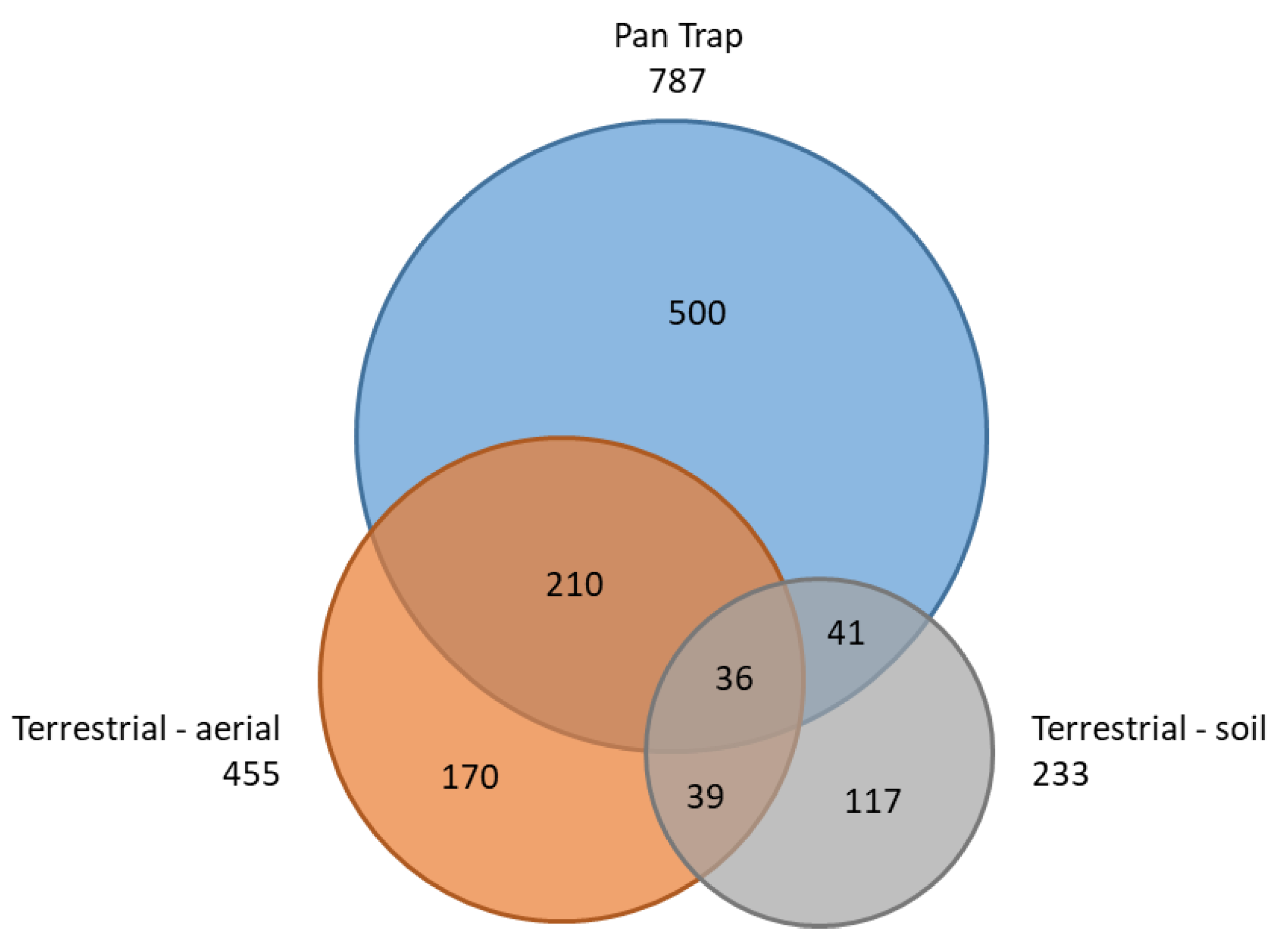
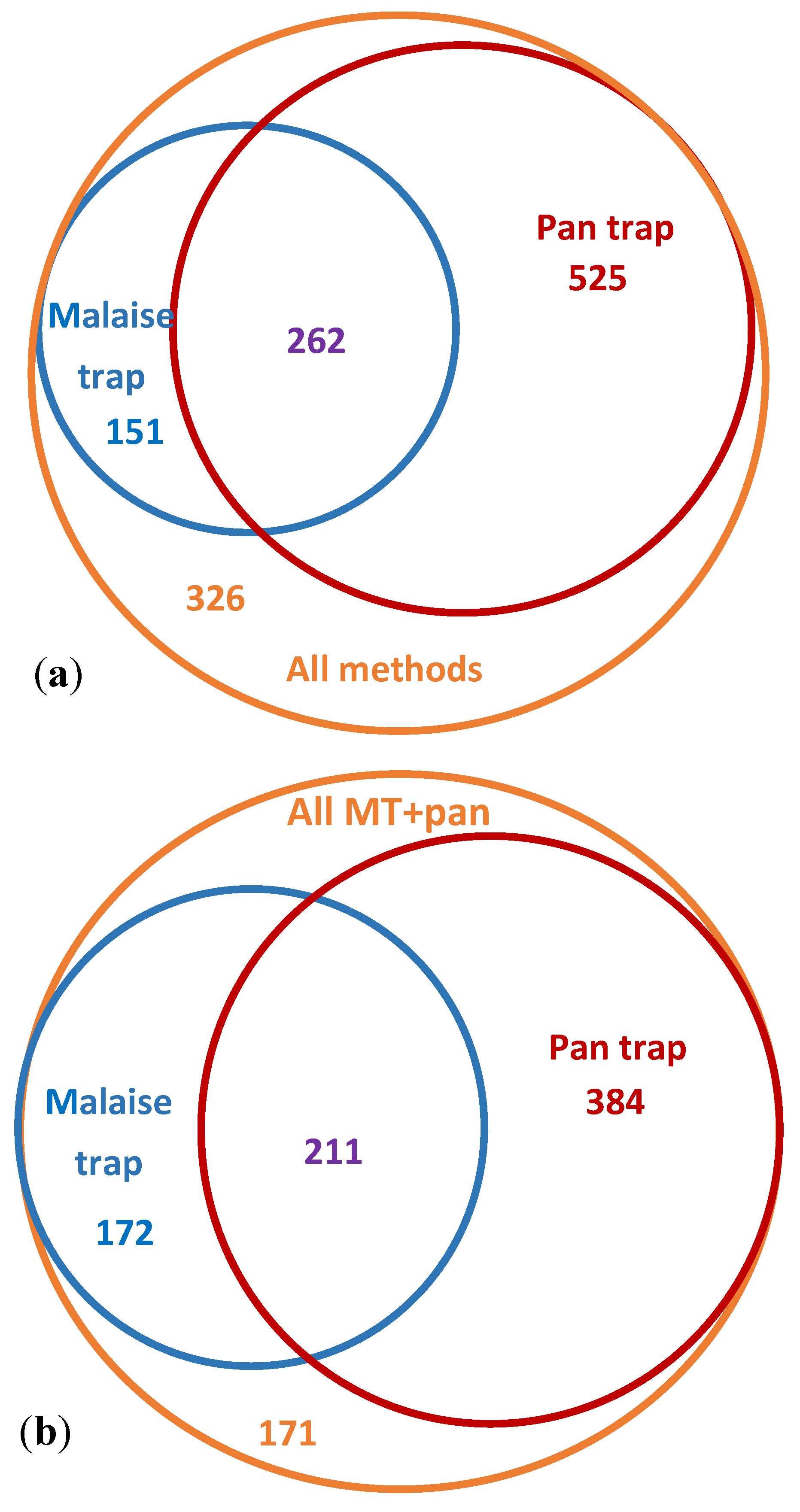
3.3. Comparative Performance of Sampling Methods
4. Discussion
4.1. Arthropod Diversity at Ikaluktutiak (Cambridge Bay)
4.2. Comparative Assessment of Collecting Methods
4.3. Implications for Community-Based Monitoring
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Robinet, C.; Roques, A. Direct impacts of recent climate warming on insect populations. Integr. Zool. 2010, 5, 132–142. [Google Scholar] [CrossRef] [PubMed]
- Bale, J.S.; Masters, G.J.; Hodkinson, I.D.; Awmack, C.; Bezemer, T.M.; Brown, V.K.; Butterfield, J.; Buse, A.; Coulson, J.C.; Farrar, J.; et al. Herbivory in global climate change research: Direct effects of rising temperature on insect herbivores. Glob. Chang. Biol. 2002, 8, 1–16. [Google Scholar] [CrossRef]
- Parmesan, C.; Ryrholm, N.; Stefanescu, C.; Hill, J.K.; Thomas, C.D.; Descimon, H.; Huntley, B.; Kaila, L.; Kullberg, J.; Tammaru, T.; et al. Poleward shifts in geographical ranges of butterfly species associated with regional warming. Nature 1999, 399, 579–583. [Google Scholar] [CrossRef]
- Fernandez-Triana, J.; Smith, M.A.; Boudreault, C.; Goulet, H.; Hebert, P.D.N.; Smith, A.C.; Roughley, R. A Poorly Known High-Latitude Parasitoid Wasp Community: Unexpected Diversity and Dramatic Changes through Time. PLoS ONE 2011, 6, e23719. [Google Scholar] [CrossRef] [PubMed]
- Koltz, A.M.; Schmidt, N.M.; Høye, T.T. Differential arthropod responses to warming are altering the structure of Arctic communities. R. Soc. Open Sci. 2018, 5, 171503. [Google Scholar] [CrossRef]
- Høye, T.T.; Culler, L.E. Tundra arthropods provide key insights into ecological responses to environmental change. Polar Biol. 2018, 41, 1523–1529. [Google Scholar] [CrossRef]
- Høye, T.T.; Forchhammer, M.C. Phenology of High-Arctic Arthropods: Effects of Climate on Spatial, Seasonal, and Inter-Annual Variation. In Advances in Ecological Research: High-Arctic Ecosystem Dynamics in a Changing Climate; Meltofte, H., Christensen, T.R., Elberling, B., Forchhammer, M.C., Rasch, M., Eds.; Academic Press: Cambridge, MA, USA, 2008; Volume 40, pp. 299–324. ISBN 978-0-12-373665-9. [Google Scholar]
- Myers-Smith, I.H.; Grabowski, M.M.; Thomas, H.J.D.; Angers-Blondin, S.; Daskalova, G.N.; Bjorkman, A.D.; Cunliffe, A.M.; Assmann, J.J.; Boyle, J.S.; McLeod, E.; et al. Eighteen years of ecological monitoring reveals multiple lines of evidence for tundra vegetation change. Ecol. Monogr. 2019, 89, e01351. [Google Scholar] [CrossRef]
- Gill, H.; Lantz, T. A Community-Based Approach to Mapping Gwich’in Observations of Environmental Changes in the Lower Peel River Watershed, NT. J. Ethnobiol. 2014, 34, 294–315. [Google Scholar] [CrossRef]
- Johnson, N.; Alessa, L.; Behe, C.; Danielsen, F.; Gearheard, S.; Gofman-Wallingford, V.; Kliskey, A.; Krümmel, E.-M.; Lynch, A.; Mustonen, T.; et al. The contributions of community-based monitoring and traditional knowledge to Arctic Observing Networks: Reflections on the state of the field. Arctic 2015, 68, 28–40. [Google Scholar] [CrossRef]
- Schmeller, D.S.; Böhm, M.; Arvanitidis, C.; Barber-Meyer, S.; Brummitt, N.; Chandler, M.; Chatzinikolaou, E.; Costello, M.J.; Ding, H.; García-Moreno, J.; et al. Building capacity in biodiversity monitoring at the global scale. Biodivers. Conserv. 2017, 26, 2765–2790. [Google Scholar] [CrossRef]
- Ji, Y.; Ashton, L.; Pedley, S.M.; Edwards, D.P.; Tang, Y.; Nakamura, A.; Kitching, R.; Dolman, P.M.; Woodcock, P.; Edwards, F.A.; et al. Reliable, verifiable and efficient monitoring of biodiversity via metabarcoding. Ecol. Lett. 2013, 16, 1245–1257. [Google Scholar] [CrossRef] [PubMed]
- Elbrecht, V.; Vamos, E.E.; Meissner, K.; Aroviita, J.; Leese, F. Assessing strengths and weaknesses of DNA metabarcoding-based macroinvertebrate identification for routine stream monitoring. Methods Ecol. Evol. 2017, 8, 1265–1275. [Google Scholar] [CrossRef]
- Young, M.R.; Proctor, H.C.; deWaard, J.R.; Hebert, P.D.N. DNA barcodes expose unexpected diversity in Canadian mites. Mol. Ecol. 2019, 28, 5347–5359. [Google Scholar] [CrossRef] [PubMed]
- Hebert, P.D.N.; Ratnasingham, S.; Zakharov, E.V.; Telfer, A.C.; Levesque-Beaudin, V.; Milton, M.A.; Pedersen, S.; Jannetta, P.; deWaard, J.R. Counting animal species with DNA barcodes: Canadian insects. Philos Trans. R Soc. 2016, 371, 20150333. [Google Scholar] [CrossRef] [PubMed]
- Cordero, R.D.; Sanchez-Ramirez, S.; Currie, D.C. DNA barcoding of aquatic insects reveals unforeseen diversity and recurrent population divergence patterns through broad-scale sampling in northern Canada. Polar Biol. 2017, 40, 1687–1695. [Google Scholar] [CrossRef]
- Decaëns, T.; Porco, D.; James, S.W.; Brown, G.G.; Chassany, V.; Dubs, F.; Dupont, L.; Lapied, E.; Rougerie, R.; Rossi, J.-P.; et al. DNA barcoding reveals diversity patterns of earthworm communities in remote tropical forests of French Guiana. Soil Biol. Biochem. 2016, 92, 171–183. [Google Scholar] [CrossRef]
- Leray, M.; Knowlton, N. DNA barcoding and metabarcoding of standardized samples reveal patterns of marine benthic diversity. Proc. Natl. Acad. Sci. USA 2015, 112, 2076–2081. [Google Scholar] [CrossRef]
- Wirta, H.K.; Hebert, P.D.N.; Kaartinen, R.; Prosser, S.W.; Várkonyi, G.; Roslin, T. Complementary molecular information changes our perception of food web structure. Proc. Natl. Acad. Sci. USA 2014, 111, 1885–1890. [Google Scholar] [CrossRef]
- Kaartinen, R.; Stone, G.N.; Hearn, J.; Lohse, K.; Roslin, T. Revealing secret liaisons: DNA barcoding changes our understanding of food webs. Ecol. Entomol. 2010, 35, 623–638. [Google Scholar] [CrossRef]
- Ratnasingham, S.; Hebert, P.D.N. A DNA-based registry for all animal species: The Barcode Index Number (BIN) system. PLoS ONE 2013, 8, e66213. [Google Scholar] [CrossRef]
- Ratnasingham, S.; Hebert, P.D.N. BOLD: The Barcode of Life Data System (www.barcodinglife.org). Mol. Ecol. Notes 2007, 7, 355–364. [Google Scholar] [CrossRef] [PubMed]
- Pentinsaari, M.; Vos, R.; Mutanen, M. Algorithmic single-locus species delimitation: Effects of sampling effort, variation and nonmonophyly in four methods and 1870 species of beetles. Mol. Ecol. Resour. 2017, 17, 393–404. [Google Scholar] [CrossRef] [PubMed]
- Huemer, P.; Mutanen, M.; Sefc, K.M.; Hebert, P.D.N. Testing DNA Barcode Performance in 1000 Species of European Lepidoptera: Large Geographic Distances Have Small Genetic Impacts. PLoS ONE 2014, 9, e115774. [Google Scholar] [CrossRef] [PubMed]
- Wirta, H.; Várkonyi, G.; Rasmussen, C.; Kaartinen, R.; Schmidt, N.M.; Hebert, P.D.N.; Barták, M.; Blagoev, G.; Disney, H.; Ertl, S.; et al. Establishing a community-wide DNA barcode library as a new tool for arctic research. Mol. Ecol. Resour. 2016, 16, 809–822. [Google Scholar] [CrossRef]
- Cameron, E.R.; Buddle, C.M. Seasonal change and microhabitat association of Arctic spider assemblages (Arachnida: Araneae) on Victoria Island (Nunavut, Canada). Can. Entomol. 2017, 149, 357–371. [Google Scholar] [CrossRef]
- Loboda, S.; Buddle, C.M. Small- to large-scale patterns of ground-dwelling spider (Araneae) diversity across northern Canada. Facets 2018, 3, 880–895. [Google Scholar] [CrossRef]
- Rogy, P.; Wheeler, T.A.; Solecki, A.M. Spatial distribution of acalyptrate fly (Diptera) assemblages in Northern Canada. Polar Biol. 2019, 42, 1489–1500. [Google Scholar] [CrossRef]
- Loboda, S.; Savage, J.; Buddle, C.M.; Schmidt, N.M.; Høye, T.T. Declining diversity and abundance of High Arctic fly assemblages over two decades of rapid climate warming. Ecography 2018, 41, 265–277. [Google Scholar] [CrossRef]
- Walker, D.A.; Raynolds, M.K.; Daniëls, F.J.A.; Einarsson, E.; Elvebakk, A.; Gould, W.A.; Katenin, A.E.; Kholod, S.S.; Markon, C.J.; Melnikov, E.S.; et al. The Circumpolar Arctic vegetation map. J. Veg. Sci. 2005, 16, 267–282. [Google Scholar] [CrossRef]
- McLennan, D.S.; MacKenzie, W.H.; Meidinger, D.; Wagner, J.; Arko, C. A Standardized Ecosystem Classification for the Coordination and Design of Long-term Terrestrial Ecosystem Monitoring in Arctic-Subarctic Biomes. Arctic 2018, 71, 1–15. [Google Scholar] [CrossRef]
- Polar Data Catalogue Geospatial Search. Available online: https://www.polardata.ca/pdcsearch/PDCSearch.jsp?doi_id=13060 (accessed on 26 September 2019).
- deWaard, J.R.; Levesque-Beaudin, V.; deWaard, S.L.; Ivanova, N.V.; McKeown, J.T.A.; Miskie, R.; Naik, S.; Perez, K.H.J.; Ratnasingham, S.; Sobel, C.N.; et al. Expedited assessment of terrestrial arthropod diversity by coupling Malaise traps with DNA barcoding. Genome 2019, 62, 85–95. [Google Scholar] [CrossRef] [PubMed]
- Hebert, P.D.N.; Braukmann, T.W.A.; Prosser, S.W.J.; Ratnasingham, S.; Dewaard, J.R.; Ivanova, N.V.; Janzen, D.H.; Hallwachs, W.; Naik, S.; Sones, J.E.; et al. A Sequel to Sanger: Amplicon sequencing that scales. BMC Genom. 2018, 19, 219. [Google Scholar] [CrossRef] [PubMed]
- Colwell, R.K. EstimateS, Version 9.1: Statistical Estimation of Species Richness and Shared Species from Samples. Available online: http://viceroy.eeb.uconn.edu/estimates (accessed on 27 November 2019).
- Danks, H.V. Arctic Arthropods: A Review of Systematics and Ecology with Particular Reference to the North. American Fauna; Entomological Society of Canada: Ottawa, ON, Canada, 1981; ISBN 0-9690829-0-8. [Google Scholar]
- Ernst, C.M.; Loboda, S.; Buddle, C.M. Capturing northern biodiversity: Diversity of arctic, subarctic and north boreal beetles and spiders are affected by trap type and habitat. Insect Conserv. Divers. 2016, 9, 63–73. [Google Scholar] [CrossRef]
- Oliverio, A.M.; Gan, H.; Wickings, K.; Fierer, N. A DNA metabarcoding approach to characterize soil arthropod communities. Soil Biol. Biochem. 2018, 125, 37–43. [Google Scholar] [CrossRef]
- Borrell, Y.J.; Miralles, L.; Huu, H.D.; Mohammed-Geba, K.; Garcia-Vazquez, E. DNA in a bottle—Rapid metabarcoding survey for early alerts of invasive species in ports. PLoS ONE 2017, 12, e0183347. [Google Scholar] [CrossRef] [PubMed]
- Kouril, D.; Furgal, C.; Whillans, T. Trends and key elements in community-based monitoring: A systematic review of the literature with an emphasis on Arctic and Subarctic regions. Environ. Rev. 2015, 24, 151–163. [Google Scholar] [CrossRef]
- Wheeler, H.C.; Berteaux, D.; Furgal, C.; Cazelles, K.; Yoccoz, N.G.; Grémillet, D. Identifying key needs for the integration of social–ecological outcomes in arctic wildlife monitoring. Conserv. Biol. 2019, 33, 861–872. [Google Scholar] [CrossRef]
- Chandler, M.; See, L.; Copas, K.; Bonde, A.M.Z.; López, B.C.; Danielsen, F.; Legind, J.K.; Masinde, S.; Miller-Rushing, A.J.; Newman, G.; et al. Contribution of citizen science towards international biodiversity monitoring. Biol. Conserv. 2017, 213, 280–294. [Google Scholar] [CrossRef]
- Hoagland, K.E. The taxonomic impediment and the convention on biodiversity. ASC Newsl. 1996, 24, 61–62. [Google Scholar]

| Habitat Category: | Active Collecting | Stationary Traps |
|---|---|---|
| Aerial | Sweep netting | Malaise trap, pan trap |
| Vegetation | Sweep netting, freehand collecting | |
| Soil–surface | Freehand collecting | Pitfall trap |
| Soil–interstitial | Sifting | |
| Freshwater–aquatic | Plankton net, D-net, aquatic hand-net |
| Class | Taxon | Specimens | BINs | Singletons | Doubletons | Chao1 | Chao1 95% Cl Lower | Chao1 95% Cl Upper | Chao1 SD | % BINS Sampled | % Singletons | Upper-Lower Discrepancy |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Insecta | Coleoptera | 440 | 33 | 7 | 3 | 41.2 | 34.6 | 75.3 | 8.3 | 80.2% | 21.2% | 2.18 |
| Diptera | 12,113 | 644 | 212 | 82 | 918.0 | 834.8 | 1037.6 | 51.1 | 70.2% | 32.9% | 1.24 | |
| Ephemeroptera | 78 | 3 | 0 | 0 | not calculated | |||||||
| Hemiptera | 312 | 19 | 10 | 2 | 43.9 | 24.1 | 141.9 | 24.2 | 43.3% | 52.6% | 5.90 | |
| Hymenoptera | 2402 | 255 | 91 | 39 | 361.1 | 316.8 | 437.2 | 29.8 | 70.6% | 35.7% | 1.38 | |
| Lepidoptera | 114 | 34 | 13 | 5 | 50.8 | 38.5 | 96.3 | 12.6 | 67.0% | 38.2% | 2.50 | |
| Plecoptera | 8 | 2 | 0 | 0 | ||||||||
| Thysanoptera | 19 | 3 | 1 | 0 | not calculated | |||||||
| Trichoptera | 416 | 6 | 0 | 0 | not calculated | |||||||
| Total Insecta | 15,902 | 999 | 334 | 131 | 1424.8 | 1317.9 | 1567.4 | 63.1 | 70.1% | 33.4% | 1.19 | |
| Arachnida | Araneae | 781 | 50 | 12 | 4 | 68.0 | 54.5 | 121.3 | 29.8 | 73.6% | 24.0% | 2.22 |
| Mesostigmata | 74 | 20 | 7 | 4 | not calculated | |||||||
| Sarcoptiformes | 356 | 48 | 17 | 9 | not calculated | |||||||
| Trombidiformes | 415 | 74 | 32 | 13 | not calculated | |||||||
| Unidentified Acari | 40 | 20 | not calculated | |||||||||
| Total Acari | 885 | 162 | 68 | 30 | 239.0 | 203.4 | 305.1 | 25.0 | 67.8% | 42.0% | 1.50 | |
| Total Arachnida | 1666 | 212 | 136 | 60 | not calculated | |||||||
| Collembola | Entomobryomorpha | 106 | 26 | 8 | 1 | not calculated | ||||||
| Poduromorpha | 150 | 13 | 3 | 0 | not calculated | |||||||
| Symphypleona | 272 | 14 | 5 | 0 | not calculated | |||||||
| Total Collembola | 528 | 53 | 16 | 1 | 180.8 | 74.8 | 802.1 | 143.3 | 29.3% | 30.2% | 10.73 | |
| Total | Total Arthropoda | 18,096 | 1264 | 430 | 166 | 1820.9 | 1695.4 | 1982.9 | 72.9 | 69.4% | 34.0% | 1.17 |
| BIN Count | BIN Overlap | ||||||
|---|---|---|---|---|---|---|---|
| Method | BINs | Unique BINs | Aquatic | Malaise Trap | Pan Trap | Pitfall Trap | Sweep net |
| Aquatic | 78 | 21 | |||||
| Malaise Trap | 413 | 112 | 40 | ||||
| Pan Trap | 787 | 359 | 38 | 262 | |||
| Pitfall Trap | 155 | 12 | 10 | 50 | 121 | ||
| Terrestrial-aerial | 455 | 125 | 42 | 186 | 246 | 82 | |
| Terrestrial-soil | 233 | 110 | 7 | 20 | 77 | 46 | 75 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pentinsaari, M.; Blagoev, G.A.; Hogg, I.D.; Levesque-Beaudin, V.; Perez, K.; Sobel, C.N.; Vandenbrink, B.; Borisenko, A. A DNA Barcoding Survey of an Arctic Arthropod Community: Implications for Future Monitoring. Insects 2020, 11, 46. https://doi.org/10.3390/insects11010046
Pentinsaari M, Blagoev GA, Hogg ID, Levesque-Beaudin V, Perez K, Sobel CN, Vandenbrink B, Borisenko A. A DNA Barcoding Survey of an Arctic Arthropod Community: Implications for Future Monitoring. Insects. 2020; 11(1):46. https://doi.org/10.3390/insects11010046
Chicago/Turabian StylePentinsaari, Mikko, Gergin A. Blagoev, Ian D. Hogg, Valerie Levesque-Beaudin, Kate Perez, Crystal N. Sobel, Bryan Vandenbrink, and Alex Borisenko. 2020. "A DNA Barcoding Survey of an Arctic Arthropod Community: Implications for Future Monitoring" Insects 11, no. 1: 46. https://doi.org/10.3390/insects11010046
APA StylePentinsaari, M., Blagoev, G. A., Hogg, I. D., Levesque-Beaudin, V., Perez, K., Sobel, C. N., Vandenbrink, B., & Borisenko, A. (2020). A DNA Barcoding Survey of an Arctic Arthropod Community: Implications for Future Monitoring. Insects, 11(1), 46. https://doi.org/10.3390/insects11010046





