Green, Yellow, and Red Fluorescent Proteins as Markers for Bacterial Isolates from Mosquito Midguts
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Species
2.2. Transformation of Bacterial Species
2.3. Plasmid Stability Testing
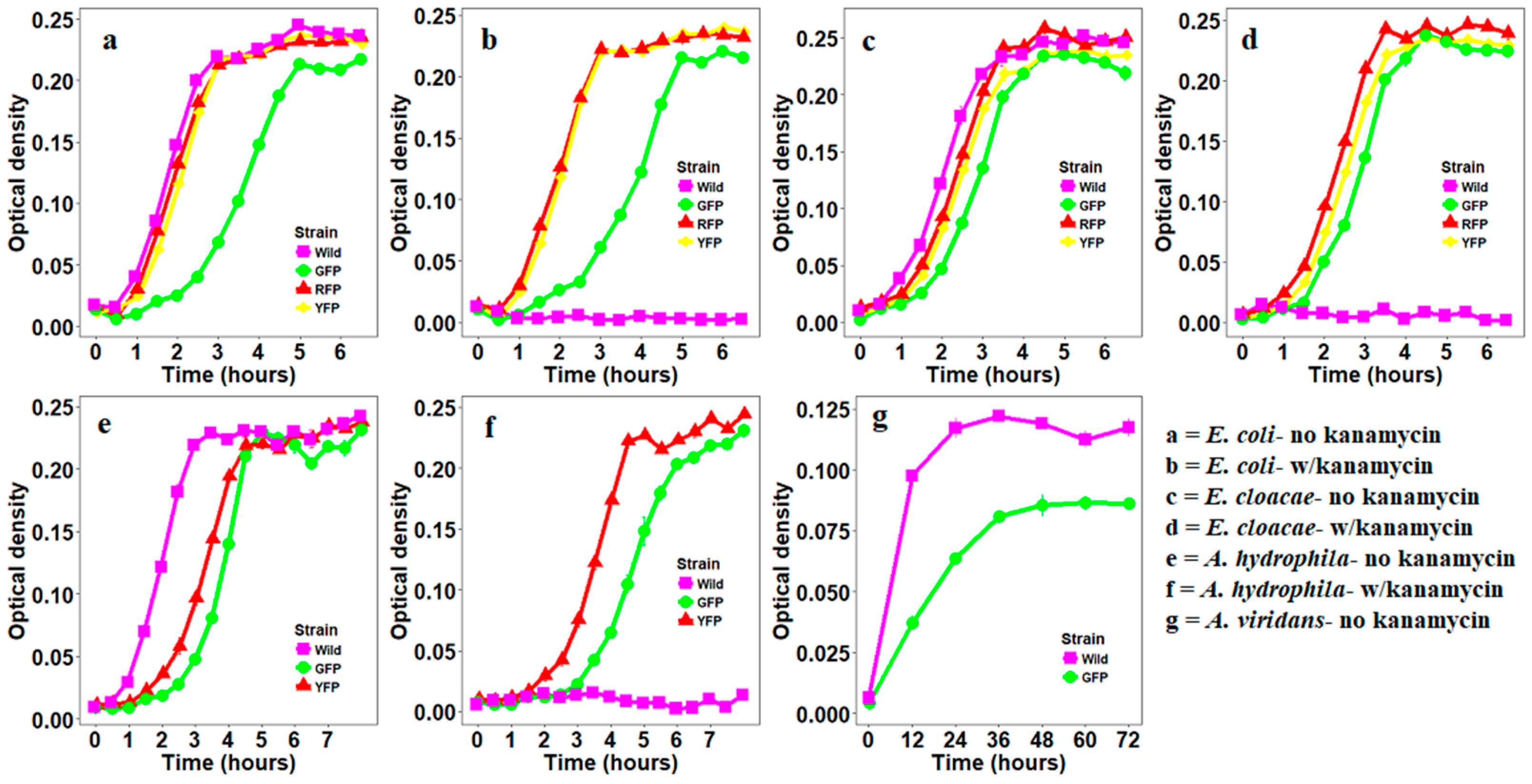
2.4. Growth Curves of Bacteria Transformed with Fluorescent Proteins
3. Results
3.1. Construction of Fluorescently Labeled Bacteria
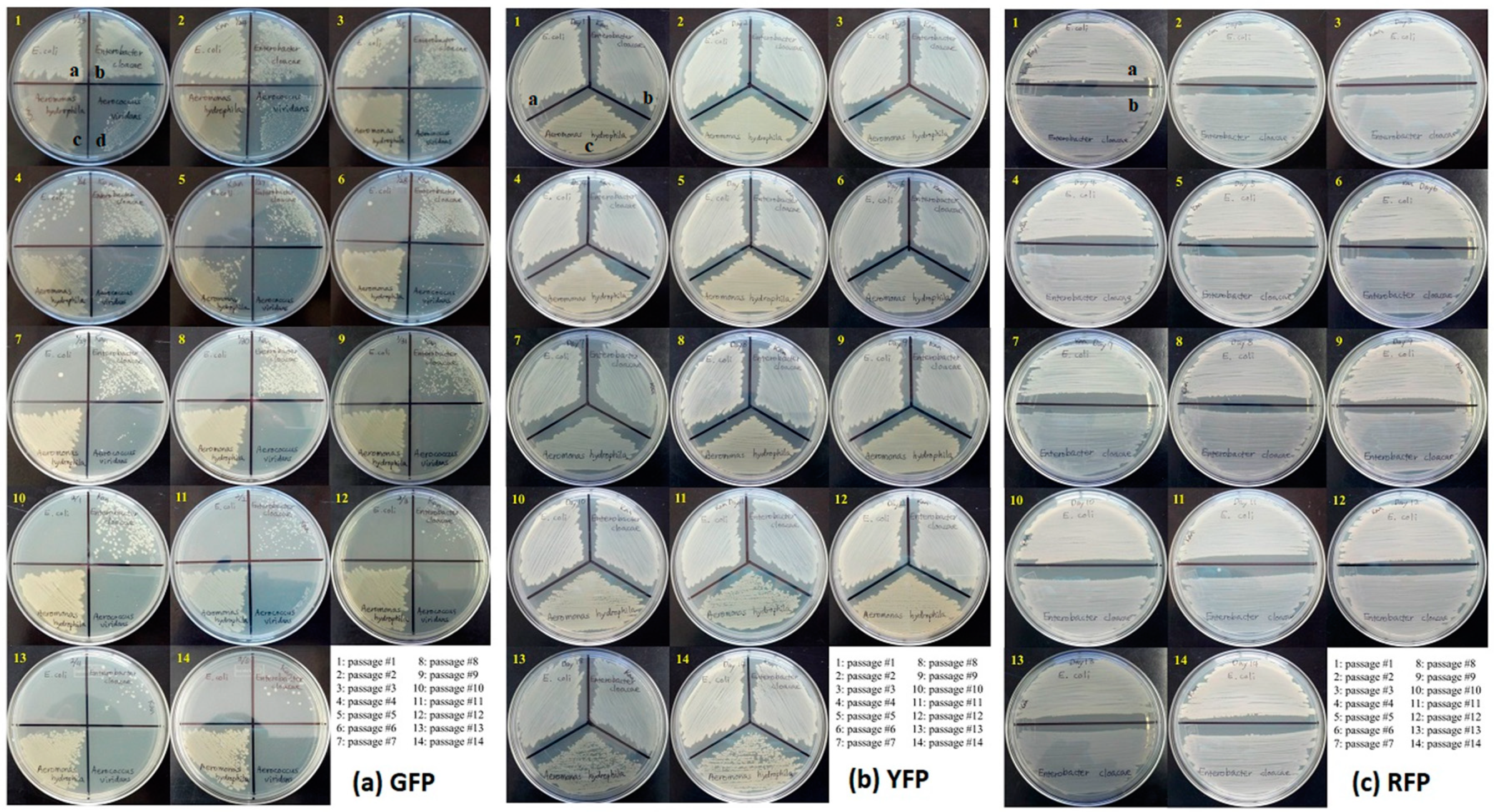
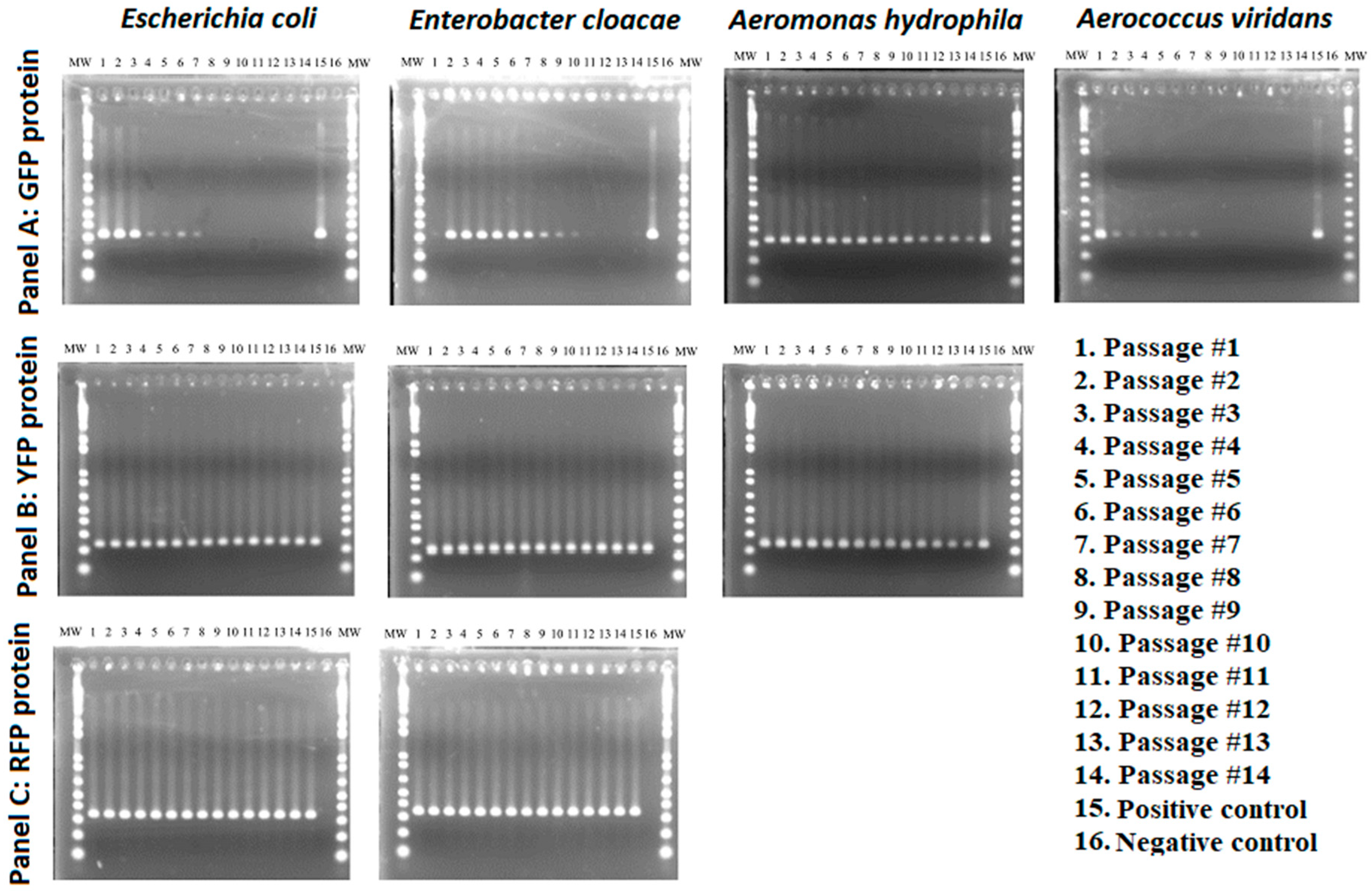
3.2. Stability of GFP Plasmid
3.3. Stability of YFP Plasmids
3.4. Stability of RFP Plasmids
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gubler, D.J. The global emergence/resurgence of arboviral diseases as public health problems. Arch. Med. Res. 2002, 33, 330–342. [Google Scholar] [CrossRef]
- Ricci, I.; Valzano, M.; Ulissi, U.; Epis, S.; Cappelli, A.; Favia, G. Symbiotic control of mosquito borne disease. Pathog.Glob. Health 2012, 106, 380–385. [Google Scholar] [CrossRef] [PubMed]
- Ramirez, J.L.; Souza-Neto, J.; Torres Cosme, R.; Rovira, J.; Ortiz, A.; Pascale, J.M.; Dimopoulos, G. Reciprocal tripartite interactions between the Aedes aegypti midgut microbiota, innate immune system and dengue virus influences vector competence. PLoS Negl. Trop. Dis. 2012, 6, e1561. [Google Scholar] [CrossRef] [PubMed]
- Apte-Deshpande, A.; Paingankar, M.; Gokhale, M.D.; Deobagkar, D.N. Serratia odorifera a midgut inhabitant of Aedes aegypti mosquito enhances its susceptibility to Dengue-2 virus. PLoS ONE 2012, 7, e40401. [Google Scholar] [CrossRef] [PubMed]
- Boissiere, A.; Tchioffo, M.T.; Bachar, D.; Abate, L.; Marie, A.; Nsango, S.E.; Shahbazkia, H.R.; Awono-Ambene, P.H.; Levashina, E.A.; Christen, R.; et al. Midgut microbiota of the malaria mosquito vector Anopheles gambiae and interactions with Plasmodium falciparum infection. PLoS Pathog. 2012, 8, e1002742. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Manfredini, F.; Dimopoulos, G. Implication of the mosquito midgut microbiota in the defense against malaria parasites. PLoS Pathog. 2009, 5, e1000423. [Google Scholar] [CrossRef] [PubMed]
- Cirimotich, C.M.; Dong, Y.; Clayton, A.M.; Sandiford, S.L.; Souza-Neto, J.A.; Mulenga, M.; Dimopoulos, G. Natural microbe-mediated refractoriness to Plasmodium infection in Anopheles gambiae. Science 2011, 332, 855–858. [Google Scholar] [CrossRef]
- Frentiu, F.D.; Zakir, T.; Walker, T.; Popovici, J.; Pyke, A.T.; van den Hurk, A.; McGraw, E.A.; O’Neill, S.L. Limited dengue virus replication in field-collected Aedes aegypti mosquitoes infected with Wolbachia. PLoS Negl. Trop. Dis. 2014, 8, e2688. [Google Scholar] [CrossRef]
- Wang, S.; Ghosh, A.K.; Bongio, N.; Stebbings, K.A.; Lampe, D.J.; Jacobs-Lorena, M. Fighting malaria with engineered symbiotic bacteria from vector mosquitoes. Proc. Natl. Acad. Sci. USA 2012, 109, 12734–12739. [Google Scholar] [CrossRef]
- Riehle, M.A.; Moreira, C.K.; Lampe, D.; Lauzon, C.; Jacobs-Lorena, M. Using bacteria to express and display anti-Plasmodium molecules in the mosquito midgut. Int. J. Parasitol. 2007, 37, 595–603. [Google Scholar] [CrossRef]
- Yoshida, S.; Ioka, D.; Matsuoka, H.; Endo, H.; Ishii, A. Bacteria expressing single-chain immunotoxin inhibit malaria parasite development in mosquitoes. Mol. Biochem. Parasitol. 2001, 113, 89–96. [Google Scholar] [CrossRef]
- Demaio, J.; Pumpuni, C.B.; Kent, M.; Beier, J.C. The midgut bacterial flora of wild Aedes triseriatus, Culex pipiens, and Psorophora columbiae mosquitoes. Am. J. Trop. Med. Hyg. 1996, 54, 219–223. [Google Scholar] [CrossRef] [PubMed]
- Terenius, O.; Lindh, J.M.; Eriksson-Gonzales, K.; Bussiere, L.; Laugen, A.T.; Bergquist, H.; Titanji, K.; Faye, I. Midgut bacterial dynamics in Aedes aegypti. FEMS Microbiol. Ecol. 2012, 80, 556–565. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Gilbreath, T.M., 3rd; Kukutla, P.; Yan, G.; Xu, J. Dynamic gut microbiome across life history of the malaria mosquito Anopheles gambiae in Kenya. PLoS ONE 2011, 6, e24767. [Google Scholar] [CrossRef] [PubMed]
- Gaio Ade, O.; Gusmao, D.S.; Santos, A.V.; Berbert-Molina, M.A.; Pimenta, P.F.; Lemos, F.J. Contribution of midgut bacteria to blood digestion and egg production in Aedes aegypti (diptera: Culicidae) (L.). Parasit. Vectors 2011, 4, 105. [Google Scholar] [CrossRef] [PubMed]
- Chouaia, B.; Rossi, P.; Epis, S.; Mosca, M.; Ricci, I.; Damiani, C.; Ulissi, U.; Crotti, E.; Daffonchio, D.; Bandi, C.; et al. Delayed larval development in Anopheles mosquitoes deprived of Asaia bacterial symbionts. BMC Microbiology 2012, 12, S2. [Google Scholar] [CrossRef] [PubMed]
- Ruan, Y.M.; Xu, J.; Liu, S.S. Effects of antibiotics on fitness of the B biotype and a non-B biotype of the whitefly Bemisia tabaci. Entomol. Exp. Appl. 2006, 121, 159–166. [Google Scholar] [CrossRef]
- Gendrin, M.; Rodgers, F.H.; Yerbanga, R.S.; Ouedraogo, J.B.; Basanez, M.G.; Cohuet, A.; Christophides, G.K. Antibiotics in ingested human blood affect the mosquito microbiota and capacity to transmit malaria. Nat. Commun. 2015, 6, 5921. [Google Scholar] [CrossRef]
- Nancharaiah, Y.V.; Wattiau, P.; Wuertz, S.; Bathe, S.; Mohan, S.V.; Wilderer, P.A.; Hausner, M. Dual labeling of Pseudomonas putida with fluorescent proteins for in situ monitoring of conjugal transfer of the TOL plasmid. Appl. Environ. Microbiol. 2003, 69, 4846–4852. [Google Scholar] [CrossRef]
- Ma, L.; Zhang, G.; Doyle, M.P. Green fluorescent protein labeling of Listeria, Salmonella, and Escherichia coli O157:H7 for safety-related studies. PLoS ONE 2011, 6, e18083. [Google Scholar] [CrossRef]
- Chalfie, M.; Tu, Y.; Euskirchen, G.; Ward, W.W.; Prasher, D.C. Green fluorescent protein as a marker for gene expression. Science 1994, 263, 802–805. [Google Scholar] [CrossRef] [PubMed]
- Prasher, D.C.; Eckenrode, V.K.; Ward, W.W.; Prendergast, F.G.; Cormier, M.J. Primary structure of the Aequorea victoria green-fluorescent protein. Gene 1992, 111, 229–233. [Google Scholar] [CrossRef]
- Lee, S.; Lim, W.A.; Thorn, K.S. Improved blue, green, and red fluorescent protein tagging vectors for S. cerevisiae. PLoS ONE 2013, 8, e67902. [Google Scholar] [CrossRef] [PubMed]
- Peloquin, J.J.; Lauzon, C.R.; Potter, S.; Miller, T.A. Transformed bacterial symbionts re-introduced to and detected in host gut. Curr. Microbiol. 2002, 45, 41–45. [Google Scholar] [CrossRef] [PubMed]
- Peloquin, J.J.; Kuzina, L.; Lauzon, C.R.; Miller, T.A. Transformation of internal extracellular bacteria isolated from Rhagoletis completa Cresson gut with enhanced green fluorescent protein. Curr. Microbiol. 2000, 40, 367–371. [Google Scholar] [CrossRef] [PubMed]
- Coon, K.L.; Vogel, K.J.; Brown, M.R.; Strand, M.R. Mosquitoes rely on their gut microbiota for development. Mol. Ecol. 2014, 23, 2727–2739. [Google Scholar] [CrossRef] [PubMed]
- Ricci, I.; Damiani, C.; Rossi, P.; Capone, A.; Scuppa, P.; Cappelli, A.; Ulissi, U.; Mosca, M.; Valzano, M.; Epis, S.; et al. Mosquito symbioses: From basic research to the paratransgenic control of mosquito-borne diseases. J. Appl. Entomol. 2011, 135, 487–493. [Google Scholar] [CrossRef]
- Husseneder, C.; Grace, J.K. Genetically engineered termite gut bacteria (Enterobacter cloacae) deliver and spread foreign genes in termite colonies. Appl. Microbiol. Biotechnol. 2005, 68, 360–367. [Google Scholar] [CrossRef]
- Husseneder, C.; Grace, J.K.; Oishi, D.E. Use of genetically engineered Escherichia coli to monitor ingestion, loss, and transfer of bacteria in termites. Curr. Microbiol. 2005, 50, 119–123. [Google Scholar] [CrossRef]
- Zhang, Z.; Moo-Young, M.; Chisti, Y. Plasmid stability in recombinant Saccharomyces cerevisiae. Biotechnol. Adv. 1996, 14, 401–435. [Google Scholar] [CrossRef]
- Bentley, W.E.; Mirjalili, N.; Andersen, D.C.; Davis, R.H.; Kompala, D.S. Plasmid-encoded protein: The principal factor in the “metabolic burden” associated with recombinant bacteria. Biotechnol. Bioeng. 1990, 35, 668–681. [Google Scholar] [CrossRef] [PubMed]
- Fakruddin, M.; Mazumdar, R.M.; Mannan, K.S.; Chowdhury, A.; Hossain, M.N. Critical factors affecting the success of cloning, expression, and mass production of enzymes by recombinant E. coli. ISRN Biotechnol. 2013, 2013. [Google Scholar] [CrossRef] [PubMed]
- Gurakan, G.C.; Bayindirli, A.; Ogel, Z.B.; Tanboga, P. Stability of a recombinant plasmid carrying alpha-amylase gene in Bacillus subtilis. World J. Microbiol. Biotechnol. 1998, 14, 293–295. [Google Scholar] [CrossRef]

| Target | Primer Name | Sequence | Amplicon |
|---|---|---|---|
| All | UniversalF UniversalR | 5-CGA AAC CCG ACA GGA CTA TAA A-3 5-CTA CAT ACC TCG CTC TGC TAA TC-3 | 328 bp |
| RFP | RFP1F RFP1R | 5-CGA CAT CAA ACT GGA CAT CAC C -3 5-CAC ATG TTC TTT CCT GCG TTA TC-3 | 344 bp |
| YFP | YFP1F YFP1R | 5-CTG CTT GTC GGC CAT GAT ATA G-3 5-CAT GAA GCA GCA CGA CTT CT-3 | 241 bp |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Muturi, E.J.; Ramirez, J.L.; Kim, C.-H. Green, Yellow, and Red Fluorescent Proteins as Markers for Bacterial Isolates from Mosquito Midguts. Insects 2019, 10, 49. https://doi.org/10.3390/insects10020049
Muturi EJ, Ramirez JL, Kim C-H. Green, Yellow, and Red Fluorescent Proteins as Markers for Bacterial Isolates from Mosquito Midguts. Insects. 2019; 10(2):49. https://doi.org/10.3390/insects10020049
Chicago/Turabian StyleMuturi, Ephantus J., Jose L. Ramirez, and Chang-Hyun Kim. 2019. "Green, Yellow, and Red Fluorescent Proteins as Markers for Bacterial Isolates from Mosquito Midguts" Insects 10, no. 2: 49. https://doi.org/10.3390/insects10020049
APA StyleMuturi, E. J., Ramirez, J. L., & Kim, C.-H. (2019). Green, Yellow, and Red Fluorescent Proteins as Markers for Bacterial Isolates from Mosquito Midguts. Insects, 10(2), 49. https://doi.org/10.3390/insects10020049






