Invasion Dynamics of A Termite, Reticulitermes flavipes, at Different Spatial Scales in France
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.1.1. Sampling at the National Scale
2.1.2. Sampling at the Regional Scale
2.1.3. Sampling at the Local Scale
2.2. Molecular Procedures
2.3. Genetic Data Analyses at the National Scale
2.3.1. Phylogeographical Analyses
2.3.2. Microsatellite and Mitochondrial Data Analyses
2.3.3. Detection of Bottlenecks
2.4. Genetic Diversity Measures
2.5. Genetic Data Analyses at the Local Scale
2.5.1. Colony Assignment
2.5.2. Colony Breeding Structure
2.5.3. Isolation by Distance
3. Results
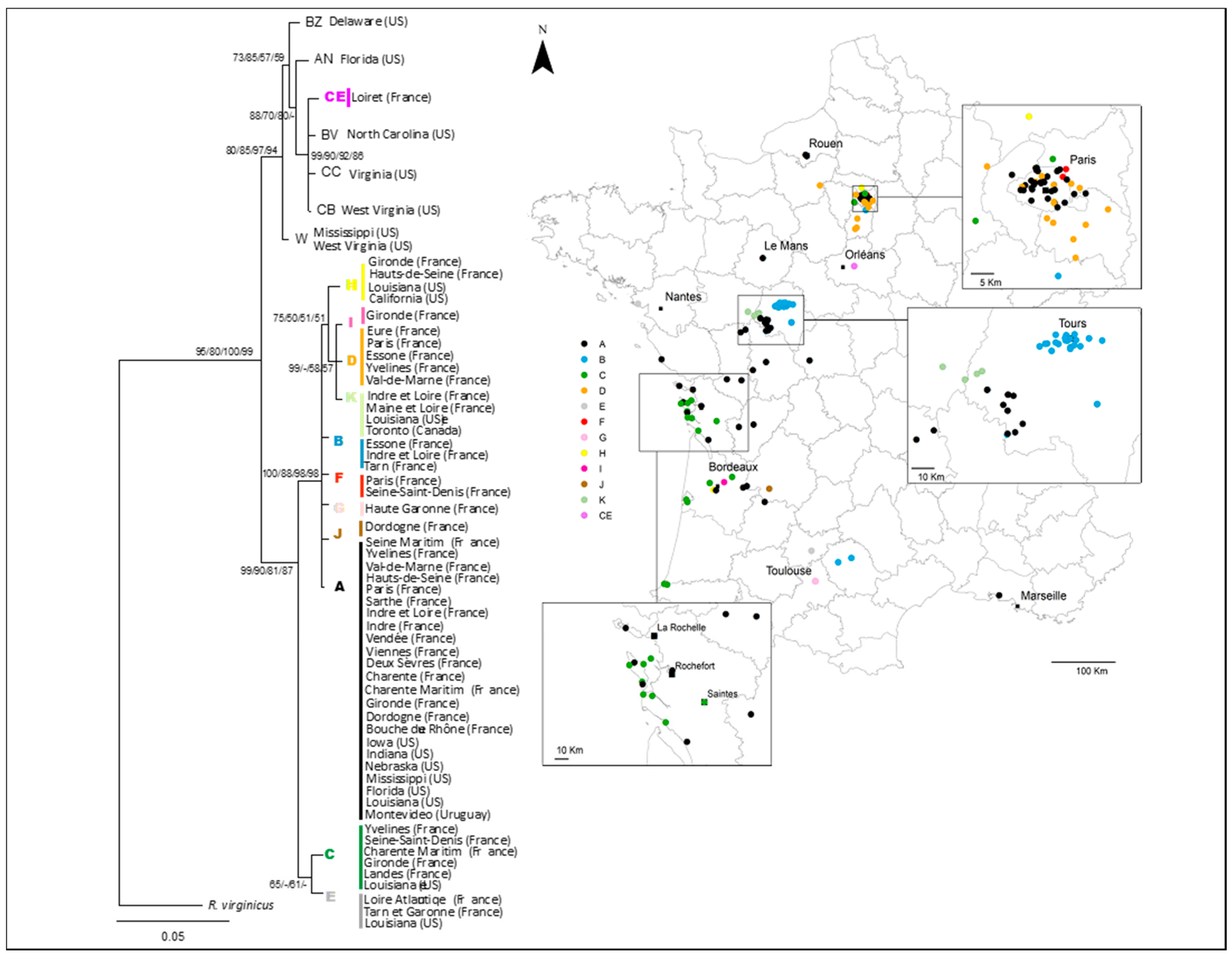
3.1. National and Regional Invasion Patterns
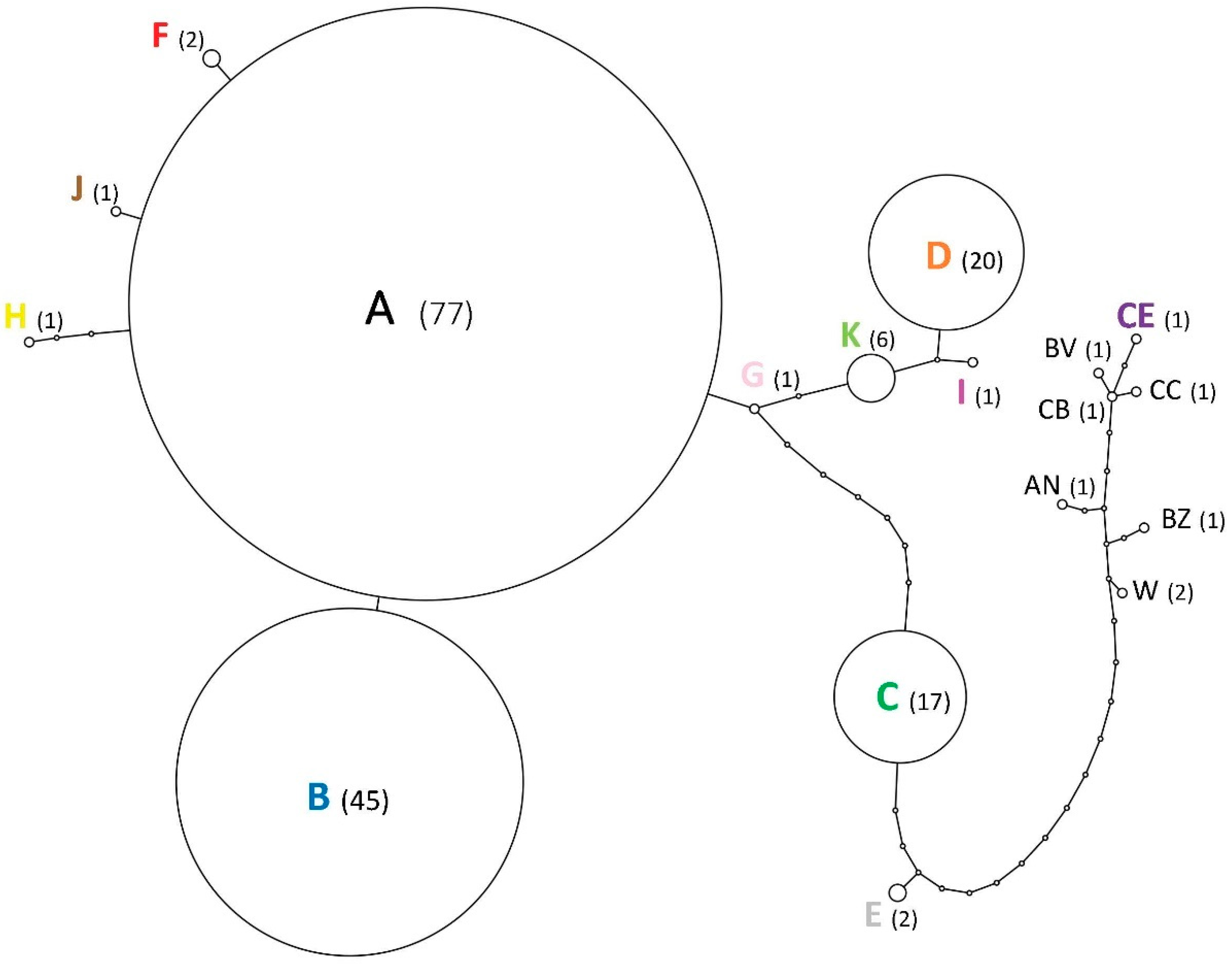
3.2. Relationships among Infestations
3.3. Genetic Diversity and Bottlenecks
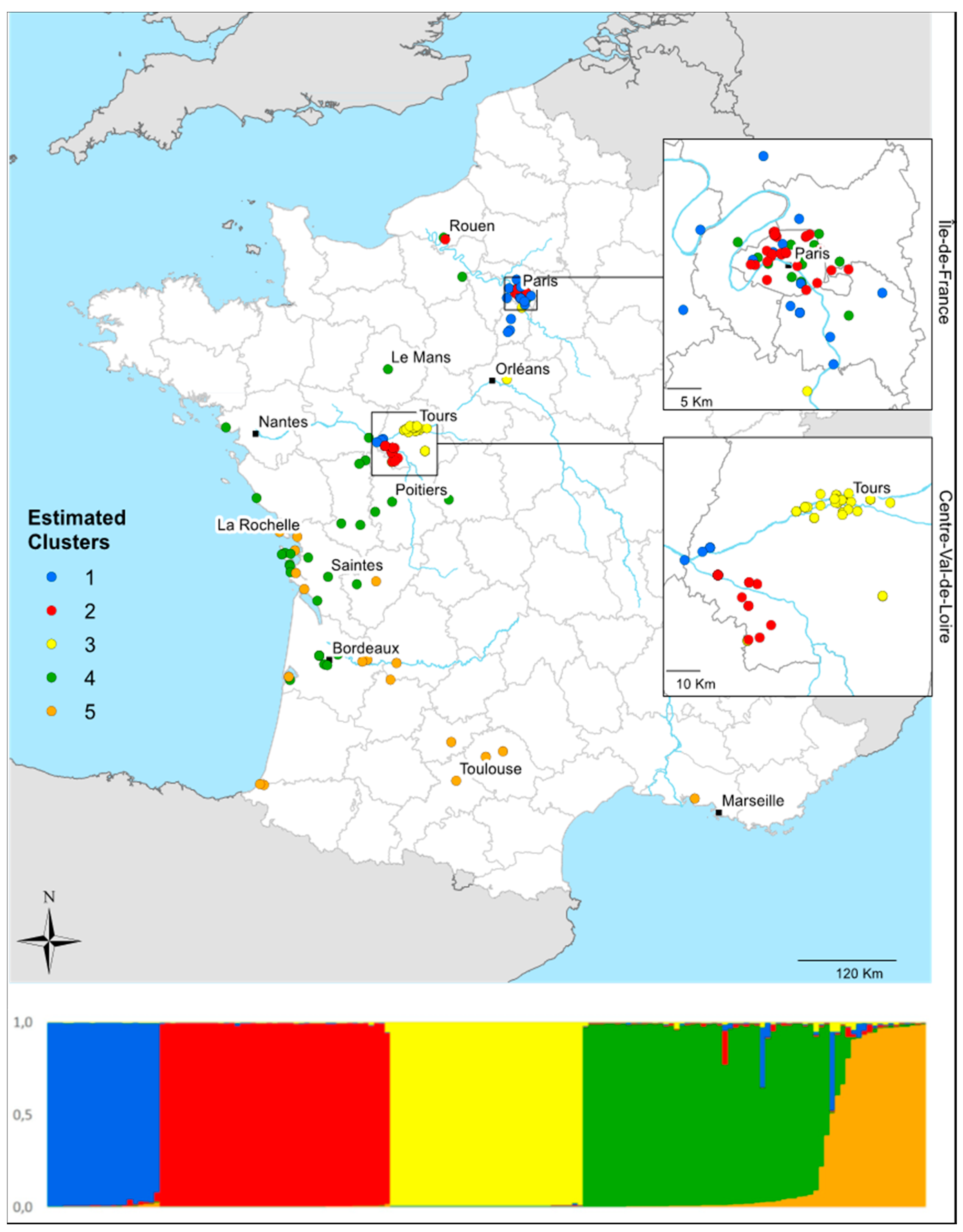
3.4. Local Invasion Patterns in Centre-Val de Loire
4. Discussion
4.1. Spread of Reticulitermes flavipes Across France
4.2. Evidence of Population Bottlenecks
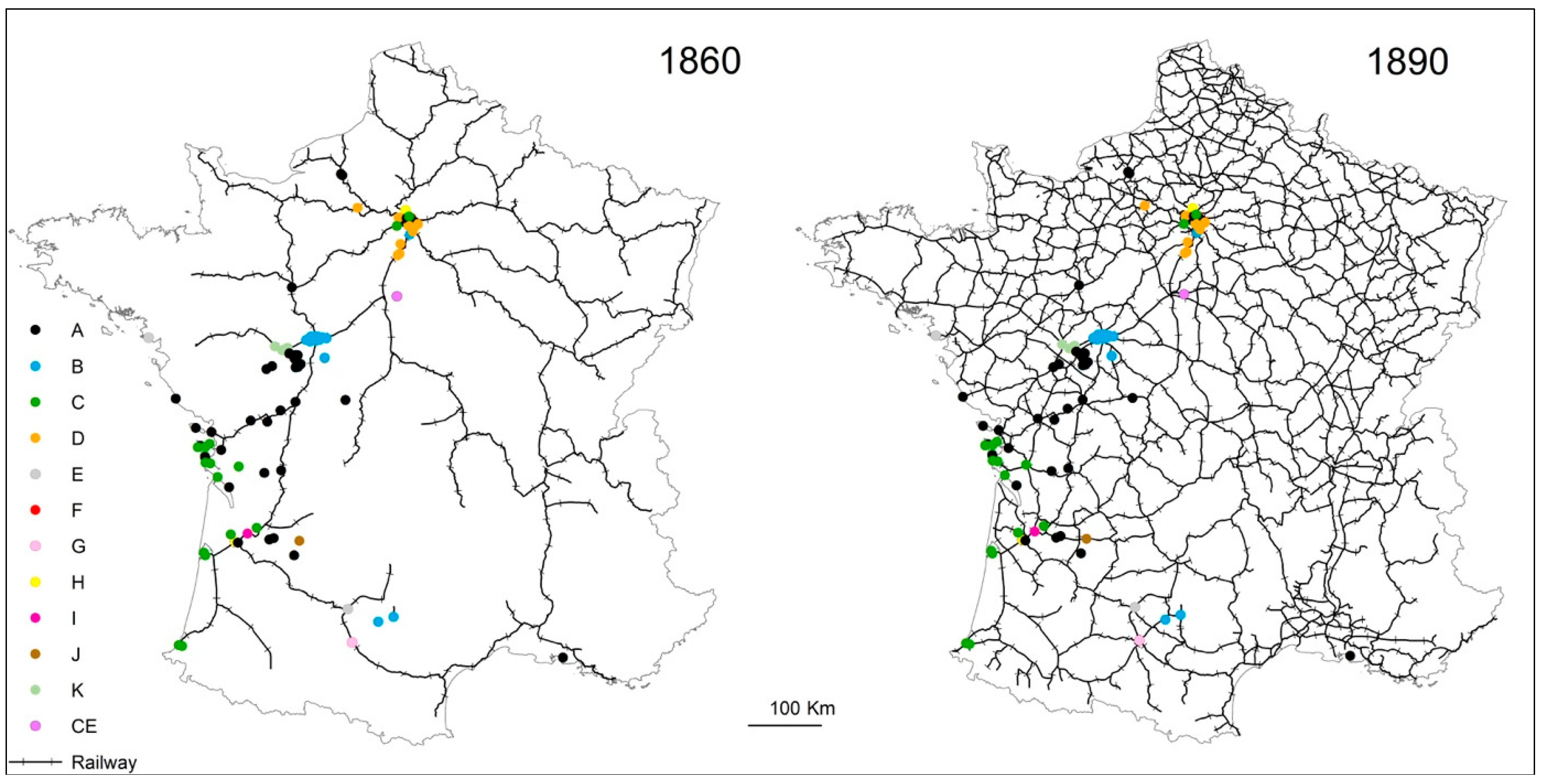
4.3. Termite Spread Driven by the Railway Network
4.4. Regional-Scale Invasion Patterns
4.5. Local-Scale Invasion Patterns
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Blackburn, T.M.; Pyšek, P.; Bacher, S.; Carlton, J.T.; Duncan, R.P.; Jarošík, V.; Wilson, J.R.U.; Richardson, D.M. A proposed unified framework for biological invasions. Trends Ecol. Evol. 2011, 26, 333–339. [Google Scholar] [CrossRef] [PubMed]
- Blackburn, T.M.; Lockwood, J.L.; Cassey, P. The influence of numbers on invasion success. Mol. Ecol. 2015, 24, 1942–1953. [Google Scholar] [CrossRef] [PubMed]
- Simberloff, D.; Martin, J.-L.; Genovesi, P.; Maris, V.; Wardle, D.A.; Aronson, J.; Courchamp, F.; Galil, B.; García-Berthou, E.; Pascal, M.; et al. Impacts of biological invasions: what’s what and the way forward. Trends Ecol. Evol. 2013, 28, 58–66. [Google Scholar] [CrossRef] [PubMed]
- Kolar, C.S.; Lodge, D.M. Progress in invasion biology: Predicting invaders. Trends Ecol. Evol. 2001, 16, 199–204. [Google Scholar] [CrossRef]
- Allendorf, F.W.; Lundquist, L.L. Introduction: Population biology, evolution, and control of invasive species. Conserv. Biol. 2003, 17, 24–30. [Google Scholar] [CrossRef]
- Colautti, R.I.; MacIsaac, H.J. A neutral terminology to define ‘invasive’ species. Divers. Distrib. 2004, 10, 135–141. [Google Scholar] [CrossRef]
- Frankham, R. Resolving the genetic paradox in invasive species. Heredity 2004, 94, 385. [Google Scholar] [CrossRef] [PubMed]
- Lockwood, J.L.; Cassey, P.; Blackburn, T. The role of propagule pressure in explaining species invasions. Trends Ecol. Evol. 2005, 20, 223–228. [Google Scholar] [CrossRef] [PubMed]
- Richardson, D.M.; Pyšek, P.; Rejmánek, M.; Barbour, M.G.; Panetta, F.D.; West, C.J. Naturalization and invasion of alien plants: Concepts and definitions. Divers. Distrib. 2000, 6, 93–107. [Google Scholar] [CrossRef]
- Suarez, A.V.; Holway, D.A.; Case, T.J. Patterns of spread in biological invasions dominated by long-distance jump dispersal: Insights from Argentine ants. Proc. Natl. Acad. Sci. USA 2001, 98, 1095–1100. [Google Scholar] [CrossRef]
- Tepolt, C.K.; Darling, J.A.; Bagley, M.J.; Geller, J.B.; Blum, M.J.; Grosholz, E.D. European green crabs (Carcinus maenas) in the northeastern Pacific: Genetic evidence for high population connectivity and current-mediated expansion from a single introduced source population. Divers. Distrib. 2009, 15, 997–1009. [Google Scholar] [CrossRef]
- Ascunce, M.S.; Yang, C.C.; Oakey, J.; Calcaterra, L.; Wu, W.J.; Shih, C.J.; Goudet, J.; Ross, K.G.; Shoemaker, D.W. Global invasion history of the fire ant Solenopsis invicta. Science 2011, 331, 1066–1068. [Google Scholar] [CrossRef] [PubMed]
- Eritja, R.; Palmer, J.R.B.; Roiz, D.; Sanpera-Calbet, I.; Bartumeus, F. Direct evidence of adult Aedes albopictus dispersal by car. Sci. Rep. 2017, 7, 14399. [Google Scholar] [CrossRef] [PubMed]
- Roques, A.; Auger-Rozenberg, M.-A.; Blackburn, T.M.; Garnas, J.; Pyšek, P.; Rabitsch, W.; Richardson, D.M.; Wingfield, M.J.; Liebhold, A.M.; Duncan, R.P. Temporal and interspecific variation in rates of spread for insect species invading Europe during the last 200 years. Biol. Invasions 2016, 18, 907–920. [Google Scholar] [CrossRef]
- Hastings, A.; Cuddington, K.; Davies, K.F.; Dugaw, C.J.; Elmendorf, S.; Freestone, A.; Harrison, S.; Holland, M.; Lambrinos, J.; Malvadkar, U.; et al. The spatial spread of invasions: New developments in theory and evidence. Ecol. Lett. 2005, 8, 91–101. [Google Scholar] [CrossRef]
- Vargo, E.L.; Husseneder, C. Biology of subterranean termites: Insights from molecular studies of Reticulitermes and Coptotermes. Annu. Rev. Entomol. 2009, 54, 379–403. [Google Scholar] [CrossRef] [PubMed]
- Perdereau, E.; Bagnères, A.-G.; Bankhead-Dronnet, S.; Dupont, S.; Zimmermann, M.; Vargo, E.L.; Dedeine, F. Global genetic analysis reveals the putative native source of the invasive termite, Reticulitermes flavipes, in France. Mol. Ecol. 2013, 22, 1105–1119. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Teixidor, D.; Suárez, D.; García, J.; Mora, D. First report of the invasive Reticulitermes flavipes (Kollar, 1837) (Blattodea, Rhinotermitidae) in the Canary Islands. J. Appl. Entomol. 2018, 1–5. [Google Scholar] [CrossRef]
- Bagnères, A.-G.; Clément, J.L.; Blum, M.S.; Severson, R.F.; Joulie, C.; Lange, C. Cuticular hydrocarbons and defensive compounds of Reticulitermes flavipes (Kollar) and R. santonensis (feytaud): Polymorphism and chemotaxonomy. J. Chem. Ecol. 1990, 16, 3213–3244. [Google Scholar] [CrossRef]
- Bobe-Moreau, J. Mémoire sur les termites observés à Rochefort et dans divers autres lieux du département de la Charente Inférieure; Imprimerie d’Alexandre Hus: Saintes, France, 1843. [Google Scholar]
- Quatrefages, A.D. Note sur les termites de La Rochelle. Annales de la Société Zoologique 1853, 30, 16–21. [Google Scholar]
- Su, N.-Y.; Scheffrahn, R.H. Economically important termites in the United States and their control. Sociobiology 1990, 17, 77–94. [Google Scholar]
- Evans, T.A.; Forschler, B.T.; Grace, J.K. Biology of invasive termites: A Worldwide review. Annu. Rev. Entomol. 2013, 58, 455–474. [Google Scholar] [CrossRef] [PubMed]
- Buczkowski, G.; Bertelsmeier, C. Invasive termites in a changing climate: A global perspective. Ecol. Evol. 2017, 7, 974–985. [Google Scholar] [CrossRef] [PubMed]
- Thorne, B.L.; Traniello, J.F.A.; Adams, E.S.; Bulmer, M. Reproductive dynamics and colony structure of subterranean termites of the genus Reticulitermes (Isoptera Rhinotermitidae): A review of the evidence from behavioral, ecological, and genetic studies. Ethol. Ecol. Evol. 1999, 11, 149–169. [Google Scholar] [CrossRef]
- Buchli, H. L’origine des castes et les potentialités ontogéniques des termites européens du genre Reticulitermes Holmgren. Ann. Sci. Nat. Zool. 1958, 11, 263–429. [Google Scholar]
- DeHeer, C.J.; Vargo, E.L. Colony genetic organization and colony fusion in the termite Reticulitermes flavipes as revealed by foraging patterns over time and space. Mol. Ecol. 2004, 13, 431–441. [Google Scholar] [CrossRef]
- Perdereau, E.; Bagnères, A.-G.; Dupont, S.; Dedeine, F. High occurrence of colony fusion in a European population of the American termite Reticulitermes flavipes. Insectes Sociaux 2010, 57, 393–402. [Google Scholar] [CrossRef]
- Perdereau, E.; Bagnères, A.-G.; Vargo, E.L.; Baudouin, G.; Xu, Y.; Labadie, P.; Dupont, S.; Dedeine, F. Relationship between invasion success and colony breeding structure in a subterranean termite. Mol. Ecol. 2015, 24, 2125–2142. [Google Scholar] [CrossRef]
- Dronnet, S.; Chapuisat, M.; Vargo, E.L.; Lohou, C.; Bagnères, A.-G. Genetic analysis of the breeding system of an invasive subterranean termite, Reticulitermes santonensis, in urban and natural habitats. Mol. Ecol. 2005, 14, 1311–1320. [Google Scholar] [CrossRef]
- Baudouin, G.; Dedeine, F.; Bech, N.; Bankhead-Dronnet, S.; Dupont, S.; Bagnères, A.-G. An American termite in Paris: Temporal colony dynamics. Genetica 2017, 145, 491–502. [Google Scholar] [CrossRef]
- Suppo, C.; Robinet, C.; Perdereau, E.; Andrieu, D.; Bagnères, A.-G. Potential spread of the invasive North American termite, Reticulitermes flavipes, and the impact of climate warming. Biol. Invasions 2018, 20, 905–922. [Google Scholar] [CrossRef]
- Miura, T.; Roisin, Y.; Matsumoto, T. Molecular phylogeny and biogeography of the nasute termite genus Nasutitermes (Isoptera: Termitidae) in the Pacific Tropics. Mol. Phylogenet. Evol. 2000, 17, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Simon, C.; Frati, F.; Beckenbach, A.; Crespi, B.; Liu, H.; Flook, P. Evolution, weighting, and phylogenetic utility of mitochondrial gene sequences and a compilation of conserved polymerase chain reaction primers. Ann. Entomol. Soc. Am. 1994, 87, 651–701. [Google Scholar] [CrossRef]
- Vargo, E.L. Polymorphism at trinucleotide microsatellite loci in the subterranean termite Reticulitermes flavipes. Mol. Ecol. 2000, 9, 817–820. [Google Scholar] [CrossRef] [PubMed]
- Dronnet, S.; Bagnères, A.-G.; Juba, T.R.; Vargo, E.L. Polymorphic microsatellite loci in the European subterranean termite, Reticulitermes santonensis Feytaud. Mol. Ecol. Notes 2004, 4, 127–129. [Google Scholar] [CrossRef]
- DeHeer, C.J.; Kutnik, M.; Vargo, E.L.; Bagnères, A.-G. The breeding system and population structure of the termite Reticulitermes grassei in Southwestern France. Heredity 2005, 95, 408–415. [Google Scholar] [CrossRef] [PubMed]
- Galtier, N.; Gouy, M.; Gautier, C. SEAVIEW and PHYLO_WIN: Two graphic tools for sequence alignment and molecular phylogeny. Bioinformatics 1996, 12, 543–548. [Google Scholar] [CrossRef]
- Guindon, S.; Gascuel, O. A Simple, fast, and accurate algorithm to estimate large phylogenies by Maximum Likelihood. Syst. Biol. 2003, 52, 696–704. [Google Scholar] [CrossRef] [PubMed]
- Nylander, J.A.A. MrAIC.pl. Program Distributed by the Author; Evolutionary Biology Centre, Uppsala University: Uppsala, Sweden, 2004. [Google Scholar]
- Huelsenbeck, J.P.; Ronquist, F.; Nielsen, R.; Bollback, J.P. Bayesian inference of phylogeny and its impact on evolutionary biology. Science 2001, 294, 2310. [Google Scholar] [CrossRef]
- Clement, M.; Posada, D.; Crandall, K.A. TCS: A computer program to estimate gene genealogies. Mol. Ecol. 2000, 9, 1657–1659. [Google Scholar] [CrossRef] [PubMed]
- Múrias dos Santos, A.; Cabezas, M.P.; Tavares, A.I.; Xavier, R.; Branco, M. tcsBU: A tool to extend TCS network layout and visualization. Bioinformatics 2016, 32, 627–628. [Google Scholar] [CrossRef] [PubMed]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [CrossRef] [PubMed]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef]
- Earl, D.A.; VonHoldt, B.M. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]
- Piry, S.; Luikart, G.; Cornuet, J.-M. Computer note. BOTTLENECK: A computer program for detecting recent reductions in the effective size using allele frequency data. J. Hered. 1999, 90, 502–503. [Google Scholar] [CrossRef]
- Excoffier, L.; Laval, G.; Schneider, S. Arlequin (version 3.0): An integrated software package for population genetics data analysis. Evol. Bioinform. 2005, 1. [Google Scholar] [CrossRef]
- Goudet, J. FSTAT (version 1.2): A computer program to calculate F-Statistics. J. Hered. 1995, 86, 485–486. [Google Scholar] [CrossRef]
- Raymond, M.; Rousset, F. GENEPOP (Version 1.2): Population genetics software for exact tests and ecumenicism. J. Hered. 1995, 86, 9–10. [Google Scholar] [CrossRef]
- Vargo, E.L. Hierarchical analysis of colony and population genetic structure of the eastern subterranean termite, Reticulitermes flavipes, using two classes of molecular markers. Evolution 2003, 57, 2805–2818. [Google Scholar] [CrossRef]
- Bulmer, M.S.; Adams, E.S.; Traniello, J.F.A. Variation in colony structure in the subterranean termite Reticulitermes flavipes. Behav. Ecol. Sociobiol. 2001, 49, 236–243. [Google Scholar] [CrossRef]
- Slatkin, M. A Measure of population subdivision based on microsatellite allele frequencies. Genetics 1995, 139, 457–462. [Google Scholar]
- Dlugosch, K.M.; Parker, I.M. Founding events in species invasions: Genetic variation, adaptive evolution, and the role of multiple introductions. Mol. Ecol. 2008, 17, 431–449. [Google Scholar] [CrossRef]
- Andrieu, D.; Perdereau, E.; Robinet, C.; Suppo, C.; Dupont, S.; Cornillon, M.; Bagnères, A.-G. Géographie des termites souterrains en région CentreVal de Loire: Le risque d’une espèce invasive. CyberGeo 2017, 2017. [Google Scholar] [CrossRef]
- Baudouin, G.; Bech, N.; Bagnères, A.-G.; Dedeine, F. Spatial and genetic distribution of a north American termite, Reticulitermes flavipes, across the landscape of Paris. Urban Ecosyst. 2018, 21, 751–764. [Google Scholar] [CrossRef]
- Schwartz, R.; Gregory, I.; Thévenin, T. Spatial History: Railways, uneven development, and population change in France and Great-Britain, 1850–1914. J. Interdiscip. History 2011, 42, 53–88. [Google Scholar] [CrossRef]
- Lesne, P. Une station nouvelle du Termite lucifuge. Acad. Sci. 1923, 1507–1508. [Google Scholar] [CrossRef]
- Scheffrahn, R.H.; Su, N.-Y. Distribution of the termite genus Coptotermes (Isoptera: Rhinotermitidae) in Florida. Fla. Entomol. 2005, 88, 201–204. [Google Scholar] [CrossRef]
- Simms, M.D.; Riegel, C.; Husseneder, C. Genetic analysis of Coptotermes formosanus infestations along the Riverfront Railroad in New Orleans, LA before and after treatment. In Proceedings of the ESA Annual Meeting and Exhibition, Ft. Lauderdale, FL, USA, 15–18 December 2005. [Google Scholar]
- Lee, K.C.; Sun, J.-Z.; Zhu, Y.; Mallette, E.J. A case study of the formosan subterranean termite, Coptotermes formosanus (Isoptera: Rhinotermitidae) transported with a non-cellulosic commercial carrier in South Mississippi. Sociobiology 2009, 53, 619–630. [Google Scholar]
- Austin, J.W.; Glenn, G.J.; Gold, R.E. Protecting urban infrastructure from Formosan termite (Isoptera: Rhinotermitidae) attack: A case study for United States railroads. Sociobiology 2008, 51, 231–247. [Google Scholar]
- Lawson Handley, L.J.; Estoup, A.; Evans, D.M.; Thomas, C.E.; Lombaert, E.; Facon, B.; Aebi, A.; Roy, H.E. Ecological genetics of invasive alien species. BioControl 2011, 56, 409–428. [Google Scholar] [CrossRef]
- Lombaert, E.; Guillemaud, T.; Deleury, E.; Lombaert, E. A simulation-based evaluation of STRUCTURE software for exploring the introduction routes of invasive species. BioRxiv 2016, 1–28. [Google Scholar] [CrossRef]
| Loci | The Number of Alleles | |||||
|---|---|---|---|---|---|---|
| Na | ||||||
| Locus | S1 | S2 | S3 | S4 | S5 | All |
| Rf11-1 | 4 | 6 | 4 | 6 | 7 | 10 |
| Rf6-1 | 8 | 7 | 4 | 11 | 11 | 20 |
| Rs1 | 5 | 5 | 2 | 15 | 10 | 19 |
| Rf21-1 | 7 | 6 | 2 | 24 | 18 | 32 |
| Rs43 | 3 | 2 | 1 | 4 | 4 | 5 |
| Rf15-2 | 3 | 3 | 1 | 4 | 5 | 5 |
| Rs15 | 3 | 4 | 3 | 19 | 7 | 21 |
| Rf1-3 | 6 | 2 | 1 | 10 | 4 | 10 |
| Mean | 4.88 | 4.38 | 2.25 | 11.63 | 8.25 | 15.25 |
| ± SD | ±1.96 (a) | ±1.92 (a) | ±1.28 (b) | ±7.27 (c) | ±4.71 (a,c) | ±9.38 |
| Rs | ||||||
| Locus | S1 | S2 | S3 | S4 | S5 | All |
| Rf11-1 | 4 | 4.67 | 3.94 | 4.45 | 7 | 7.44 |
| Rf6-1 | 7.98 | 5.31 | 3.50 | 8.98 | 11 | 13.06 |
| Rs1 | 4.84 | 4.81 | 1.50 | 11.80 | 10 | 10.79 |
| Rf21-1 | 6.89 | 5.05 | 2 | 17.29 | 18 | 16.62 |
| Rs43 | 2.8§ | 2 | 1 | 2.68 | 4 | 3.83 |
| Rf15-2 | 3 | 2.96 | 1 | 3.43 | 5 | 4.90 |
| Rs15 | 2.98 | 3.90 | 2.55 | 14.17 | 7 | 10.86 |
| Rf1-3 | 5.97 | 2 | 1 | 8.95 | 4 | 6.95 |
| Mean | 4.81 | 3.84 | 2.06 | 8.97 | 8.25 | 9.31 |
| ± SD | ±1.96 (a,c) | ±1.35 (a) | ±1.17 (b) | ±5.28 (a,c) | ±4.71 (c) | ±4.32 |
| Hs | ||||||
| Locus | S1 | S2 | S3 | S4 | S5 | All |
| Rf11-1 | 0.727 | 0.587 | 0.612 | 0.591 | 0.837 | 0.670 |
| Rf6-1 | 0.867 | 0.711 | 0.671 | 0.852 | 0.895 | 0.799 |
| Rs1 | 0.564 | 0.767 | 0.028 | 0.884 | 0.828 | 0.614 |
| Rf21-1 | 0.782 | 0.648 | 0.44 | 0.944 | 0.951 | 0.752 |
| Rs43 | 0.331 | 0.506 | 0 | 0.115 | 0.413 | 0.272 |
| Rf15-2 | 0.612 | 0.542 | 0 | 0.525 | 0.716 | 0.477 |
| Rs15 | 0.423 | 0.654 | 0.307 | 0.914 | 0.704 | 0.601 |
| Rf1-3 | 0.669 | 0.307 | 0 | 0.835 | 0.503 | 0.463 |
| Mean | 0.622 (a) | 0.590 (a) | 0.257 (b) | 0.707 (a) | 0.731 (a) | 0.581 |
| Sign Test | Wilcoxon Test | Mode-Shift | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| One Tail | Two Tails | |||||||||
| IAM | TPM | SMM | IAM | TPM | SMM | IAM | TPM | SMM | ||
| S1 | NS | NS | NS | p < 0.05 | NS | NS | NS | NS | NS | Normal |
| S2 | p < 0.01 | NS | NS | p < 0.01 | p < 0.05 | NS | p < 0.01 | NS | NS | Normal |
| S3 | NS | NS | NS | NS | NS | NS | NS | NS | NS | Normal |
| S4 | NS | NS | p < 0.001 | NS | NS | NS | NS | NS | p < 0.01 | Normal |
| S5 | NS | NS | NS | NS | NS | NS | NS | NS | NS | Normal |
| Tours | Joué-Les-Tours | La Riche | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Locus | Na | Rs | Hs | Rs | Hs | Rs | Hs | |||
| Rf6-1 | 3 | 2.55 | 0.561 | 2.60 | 0.590 | 2.88 | 0.643 | |||
| RS1 | 2 | 1.42 | 0.149 | 1.20 | 0.062 | 1.90 | 0.335 | |||
| Rf11-1 | 3 | 2.41 | 0.557 | 2.72 | 0.683 | 2.58 | 0.502 | |||
| Rf21-1 | 2 | 1.85 | 0.380 | 1.77 | 0.349 | 1.86 | 0.306 | |||
| Rf24-2 | 2 | 1.93 | 0.474 | 1.94 | 0.469 | 1.99 | 0.472 | |||
| RS15 | 2 | 1.59 | 0.242 | 1.70 | 0.241 | 1.94 | 0.359 | |||
| Mean | 2.33 | 1.96 | 0.394 | 1.99 | 0.399 | 2.19 | 0.436 | |||
| ± SD | ±0.52 | ±0.44 (a) | ±0.58 (a) | ±0.43 (a) | ||||||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Perdereau, E.; Baudouin, G.; Bankhead-Dronnet, S.; Chevalier, Z.; Zimmermann, M.; Dupont, S.; Dedeine, F.; Bagnères, A.-G. Invasion Dynamics of A Termite, Reticulitermes flavipes, at Different Spatial Scales in France. Insects 2019, 10, 30. https://doi.org/10.3390/insects10010030
Perdereau E, Baudouin G, Bankhead-Dronnet S, Chevalier Z, Zimmermann M, Dupont S, Dedeine F, Bagnères A-G. Invasion Dynamics of A Termite, Reticulitermes flavipes, at Different Spatial Scales in France. Insects. 2019; 10(1):30. https://doi.org/10.3390/insects10010030
Chicago/Turabian StylePerdereau, Elfie, Guillaume Baudouin, Stéphanie Bankhead-Dronnet, Zoé Chevalier, Marie Zimmermann, Simon Dupont, Franck Dedeine, and Anne-Geneviève Bagnères. 2019. "Invasion Dynamics of A Termite, Reticulitermes flavipes, at Different Spatial Scales in France" Insects 10, no. 1: 30. https://doi.org/10.3390/insects10010030
APA StylePerdereau, E., Baudouin, G., Bankhead-Dronnet, S., Chevalier, Z., Zimmermann, M., Dupont, S., Dedeine, F., & Bagnères, A.-G. (2019). Invasion Dynamics of A Termite, Reticulitermes flavipes, at Different Spatial Scales in France. Insects, 10(1), 30. https://doi.org/10.3390/insects10010030







