MEDTEC Students against Coronavirus: Investigating the Role of Hemostatic Genes in the Predisposition to COVID-19 Severity
Abstract
:1. Introduction
2. Materials and Methods
2.1. Patient Cohorts for Genetic Analyses
2.2. Imputation
2.3. SNP Selection
2.4. Statistical Analysis
2.5. Polygenic Risk Score (PRS)
2.6. Meta-Analysis
2.7. Role of MEDTEC Students
3. Results
3.1. Hemostatic Gene Variants and the Predisposition to COVID-19 Severity
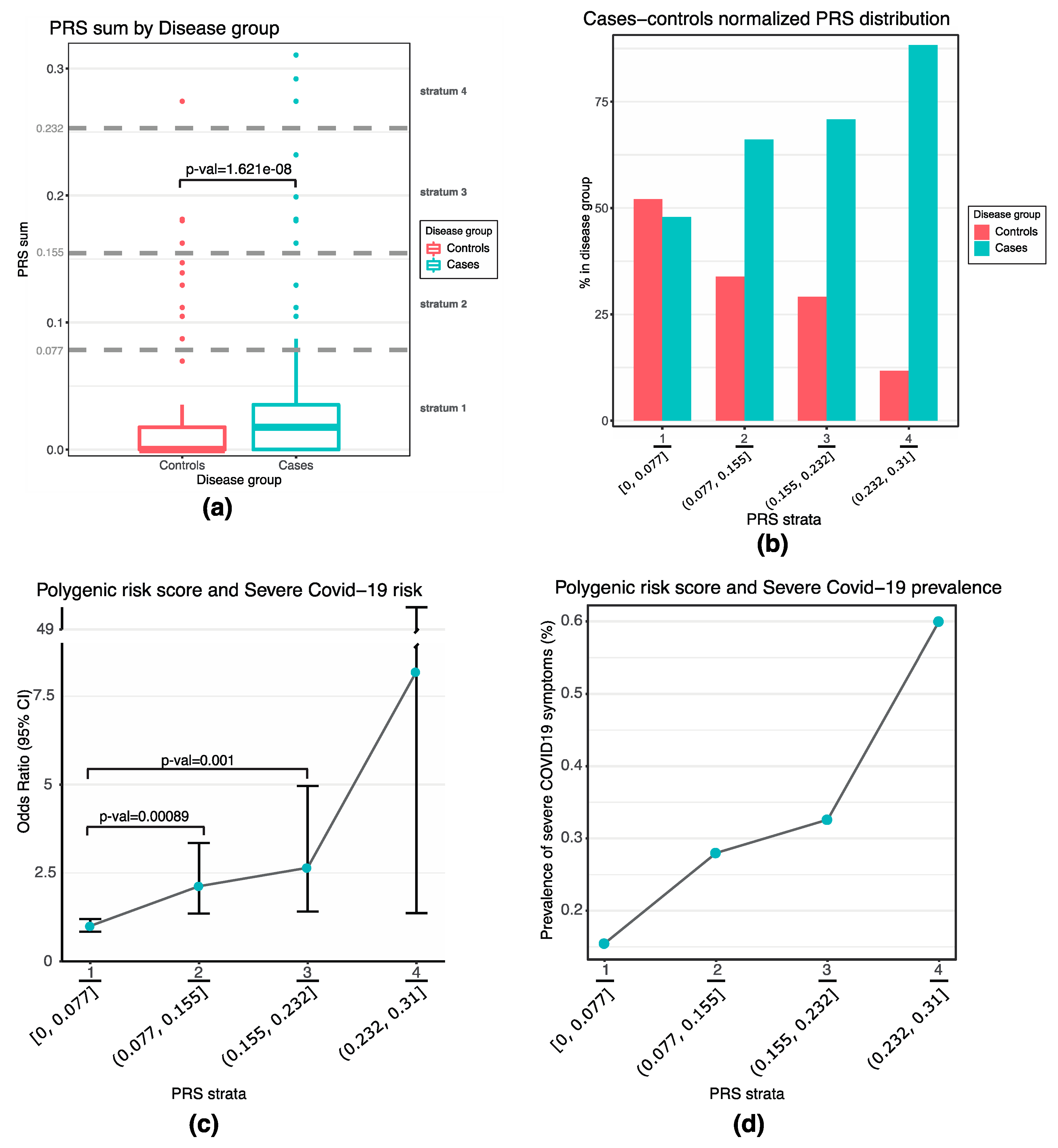
3.2. Set up of a Poligenic Risk Score Based on Hemostatic Gene Variants
3.3. Meta-Analysis Confirms the Involvement of MTHFR in Severe COVID-19 Predisposition
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Johns Hopkins Coronavirus Resource Center. Available online: https://coronavirus.jhu.edu/map.html (accessed on 8 October 2021).
- Yuki, K.; Fujiogi, M.; Koutsogiannaki, S. COVID-19 pathophysiology: A review. Clin. Immunol. 2020, 215, 108427. [Google Scholar] [CrossRef] [PubMed]
- O’Driscoll, M.; Ribeiro Dos Santos, G.; Wang, L.; Cummings, D.A.T.; Azman, A.S.; Paireau, J.; Fontanet, A.; Cauchemez, S.; Salje, H. Age-specific mortality and immunity patterns of SARS-CoV-2. Nature 2021, 590, 140–145. [Google Scholar] [CrossRef] [PubMed]
- Williamson, E.J.; Walker, A.J.; Bhaskaran, K.; Bacon, S.; Bates, C.; Morton, C.E.; Curtis, H.J.; Mehrkar, A.; Evans, D.; Inglesby, P.; et al. Factors associated with COVID-19-related death using OpenSAFELY. Nature 2020, 584, 430–436. [Google Scholar] [CrossRef] [PubMed]
- Asselta, R.; Paraboschi, E.M.; Mantovani, A.; Duga, S. ACE2 and TMPRSS2 variants and expression as candidates to sex and country differences in COVID-19 severity in Italy. Aging 2020, 12, 10087–10098. [Google Scholar] [CrossRef] [PubMed]
- Severe COVID-19 GWAS Group; Ellinghaus, D.; Degenhardt, F.; Bujanda, L.; Buti, M.; Albillos, A.; Invernizzi, P.; Fernández, J.; Prati, D.; Baselli, G.; et al. Genomewide Association Study of Severe COVID-19 with Respiratory Failure. N. Engl. J. Med. 2020, 383, 1522–1534. [Google Scholar] [CrossRef] [PubMed]
- Shelton, J.F.; Shastri, A.J.; Ye, C.; Weldon, C.H.; Filshtein-Sonmez, T.; Coker, D.; Symons, A.; Esparza-Gordillo, J.; 23andMe COVID-19 Team; Aslibekyan, S.; et al. Trans-ancestry analysis reveals genetic and nongenetic associations with COVID-19 susceptibility and severity. Nat. Genet. 2021, 53, 801–808. [Google Scholar] [CrossRef] [PubMed]
- Pairo-Castineira, E.; Clohisey, S.; Klaric, L.; Bretherick, A.D.; Rawlik, K.; Pasko, D.; Walker, S.; Parkinson, N.; Fourman, M.H.; Russell, C.D.; et al. Genetic mechanisms of critical illness in COVID-19. Nature 2021, 591, 92–98. [Google Scholar] [CrossRef]
- Kosmicki, J.A.; Horowitz, J.E.; Banerjee, N.; Lanche, R.; Marcketta, A.; Maxwell, E.; Bai, X.; Sun, D.; Backman, J.; Sharma, D.; et al. Genetic association analysis of SARS-CoV-2 infection in 455,838 UK Biobank participants. medRxiv 2020. [Google Scholar] [CrossRef]
- Horowitz, J.E.; Kosmicki, J.A.; Damask, A.; Sharma, D.; Roberts, G.H.L.; Justice, A.E.; Banerjee, N.; Coignet, M.V.; Yadav, A.; Leader, J.B.; et al. Common genetic variants identify targets for COVID-19 and individuals at high risk of severe disease. medRxiv 2021. [Google Scholar] [CrossRef]
- COVID-19 Host Genetics Initiative. Mapping the human genetic architecture of COVID-19. Nature 2021. [Google Scholar] [CrossRef]
- Zhang, Q.; Bastard, P.; Liu, Z.; Le Pen, J.; Moncada-Velez, M.; Chen, J.; Ogishi, M.; Sabli, I.K.D.; Hodeib, S.; Korol, C.; et al. Inborn errors of type I IFN immunity in patients with life-threatening COVID-19. Science 2020, 370, eabd4570. [Google Scholar] [CrossRef]
- Bastard, P.; Rosen, L.B.; Zhang, Q.; Michailidis, E.; Hoffmann, H.H.; Zhang, Y.; Dorgham, K.; Philippot, Q.; Rosain, J.; Béziat, V.; et al. Autoantibodies against type I IFNs in patients with life-threatening COVID-19. Science 2020, 370, eabd4585. [Google Scholar] [CrossRef]
- Povysil, G.; Butler-Laporte, G.; Shang, N.; Wang, C.; Khan, A.; Alaamery, M.; Nakanishi, T.; Zhou, S.; Forgetta, V.; Eveleigh, R.J.M.; et al. Rare loss-of-function variants in type I IFN immunity genes are not associated with severe COVID-19. J. Clin. Investig. 2021, 147834. [Google Scholar] [CrossRef] [PubMed]
- Zhou, F.; Yu, T.; Du, R.; Fan, G.; Liu, Y.; Liu, Z.; Xiang, J.; Wang, Y.; Song, B.; Gu, X.; et al. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: A retrospective cohort study. Lancet 2020, 395, 1054–1062. [Google Scholar] [CrossRef]
- Stefely, J.A.; Christensen, B.B.; Gogakos, T.; Cone Sullivan, J.K.; Montgomery, G.G.; Barranco, J.P.; Van Cott, E.M. Marked factor V activity elevation in severe COVID-19 is associated with venous thromboembolism. Am. J. Hematol. 2020, 95, 1522–1530. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef] [Green Version]
- Kipshidze, N.; Dangas, G.; White, C.J.; Kipshidze, N.; Siddiqui, F.; Lattimer, C.R.; Carter, C.A.; Fareed, J. Viral Coagulopathy in Patients With COVID-19: Treatment and Care. Clin. Appl. Thromb. Hemost. 2020, 26, 1076029620936776. [Google Scholar] [CrossRef] [PubMed]
- McGonagle, D.; O’Donnell, J.S.; Sharif, K.; Emery, P.; Bridgewood, C. Immune mechanisms of pulmonary intravascular coagulopathy in COVID-19 pneumonia. Lancet Rheumatol. 2020, 2, e437–e445. [Google Scholar] [CrossRef]
- Tsivgoulis, G.; Palaiodimou, L.; Zand, R.; Lioutas, V.A.; Krogias, C.; Katsanos, A.H.; Shoamanesh, A.; Sharma, V.K.; Shahjouei, S.; Baracchini, C.; et al. COVID-19 and cerebrovascular diseases: A comprehensive overview. Ther. Adv. Neurol. Disord. 2020, 13, 1756286420978004. [Google Scholar] [CrossRef]
- Merrill, J.T.; Erkan, D.; Winakur, J.; James, J.A. Emerging evidence of a COVID-19 thrombotic syndrome has treatment implications. Nat. Rev. Rheumatol. 2020, 16, 581–589. [Google Scholar] [CrossRef] [PubMed]
- Zuo, Y.; Estes, S.K.; Ali, R.A.; Gandhi, A.A.; Yalavarthi, S.; Shi, H.; Sule, G.; Gockman, K.; Madison, J.A.; Zuo, M.; et al. Prothrombotic autoantibodies in serum from patients hospitalized with COVID-19. Sci Transl. Med. 2020, 12, eabd3876. [Google Scholar] [CrossRef]
- Myocardial Infarction Genetics Consortium; Kathiresan, S.; Voight, B.F.; Purcell, S.; Musunuru, K.; Ardissino, D.; Mannucci, P.M.; Anand, S.; Engert, J.C.; Samani, N.J.; et al. Genome-wide association of early-onset myocardial infarction with single nucleotide polymorphisms and copy number variants. Nat. Genet. 2009, 41, 334–341. [Google Scholar] [CrossRef] [Green Version]
- TOPMed Imputation Server. Available online: https://imputation.biodatacatalyst.nhlbi.nih.gov/index.html#! (accessed on 27 April 2021).
- Taliun, D.; Harris, D.N.; Kessler, M.D.; Carlson, J.; Szpiech, Z.A.; Torres, R.; Taliun, S.A.G.; Corvelo, A.; Gogarten, S.M.; Kang, H.M.; et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Nature 2021, 590, 290–299. [Google Scholar] [CrossRef]
- Chang, C.C.; Chow, C.C.; Tellier, L.C.; Vattikuti, S.; Purcell, S.M.; Lee, J.J. Second-generation PLINK: Rising to the challenge of larger and richer datasets. Gigascience 2015, 4, 7. [Google Scholar] [CrossRef]
- Stouffer, S.A.; Suchman, E.A.; Devinney, L.C.; Star, S.A.; Williams, R.M. Adjustment during Army Life; Princeton University Press: Princeton, NJ, USA, 1949. [Google Scholar]
- Gao, F.; Chang, D.; Biddanda, A.; Ma, L.; Guo, Y.; Zhou, Z.; Keinan, A. XWAS: A Software Toolset for Genetic Data Analysis and Association Studies of the X Chromosome. J. Hered 2015, 106, 666–671. [Google Scholar] [CrossRef] [Green Version]
- Lander, E.; Kruglyak, L. Genetic dissection of complex traits: Guidelines for interpreting and reporting linkage results. Nat. Genet. 1995, 11, 241–247. [Google Scholar] [CrossRef] [PubMed]
- Choi, S.W.; Mak, T.S.H.; O’Reilly, P.F. Tutorial: A guide to performing polygenic risk score analyses. Nat. Protoc. 2020, 15, 2759–2772. [Google Scholar] [CrossRef]
- The Covid19 Host Genetics Initiative. Available online: https://www.covid19hg.org/ (accessed on 20 May 2021).
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2017; Available online: https://www.R-project.org/ (accessed on 10 January 2021).
- The Regeneron–Genetic Center Database. Available online: https://rgc-covid19.regeneron.com/home (accessed on 20 July 2021).
- Mantel, N.; Haenszel, W. Statistical aspects of the analysis of data from retrospective studies of disease. J. Natl. Cancer Inst. 1959, 22, 719–748. [Google Scholar]
- Mosteller, F.; Bush, R.R. Selected quantitative techniques. In Handbook of Social Psychology; Lindzey, G., Ed.; Addison-Wesley: Cambridge, MA, USA, 1954; Volume 1, pp. 289–334. [Google Scholar]
- Liptak, T. On the combination of independent tests. Magyar. Tud Akad Mat. Kutato Int. Kozl. 1958, 3, 171–197. [Google Scholar]
- Rosendaal, F.R.; Reitsma, P.H. Genetics of venous thrombosis. J. Thromb. Haemost. 2009, 7 (Suppl. S1), 301–304. [Google Scholar] [CrossRef]
- Nyholt, D.R. A simple correction for multiple testing for single-nucleotide polymorphisms in linkage disequilibrium with each other. Am. J. Hum. Genet. 2004, 74, 765–769. [Google Scholar] [CrossRef] [Green Version]
- Nicodemus, K.K.; Liu, W.; Chase, G.A.; Tsai, Y.Y.; Fallin, M.D. Comparison of type I error for multiple test corrections in large single-nucleotide polymorphism studies using principal components versus haplotype blocking algorithms. BMC Genet. 2005, 6 (Suppl. S1), S78. [Google Scholar] [CrossRef] [Green Version]
- Genotype-Tissue Expression-GTEx Portal. Available online: https://gtexportal.org/home/locusBrowserPage/FGA (accessed on 1 September 2021).
- Atlas of GWAS Summary Statistics. Available online: https://atlas.ctglab.nl/ (accessed on 1 September 2021).
- Zhu, Z.; Wang, X.; Li, X.; Lin, Y.; Shen, S.; Liu, C.L.; Hobbs, B.D.; Hasegawa, K.; Liang, L.; International COPD Genetics Consortium; et al. Genetic overlap of chronic obstructive pulmonary disease and cardiovascular disease-related traits: A large-scale genome-wide cross-trait analysis. Respir Res. 2019, 20, 64. [Google Scholar] [CrossRef]
- Watanabe, K.; Stringer, S.; Frei, O.; Umićević Mirkov, M.; de Leeuw, C.; Polderman, T.J.C.; van der Sluis, S.; Andreassen, O.A.; Neale, B.M.; Posthuma, D. A global overview of pleiotropy and genetic architecture in complex traits. Nat. Genet. 2019, 51, 1339–1348. [Google Scholar] [CrossRef]
- Wu, Y.; Byrne, E.M.; Zheng, Z.; Kemper, K.E.; Yengo, L.; Mallett, A.J.; Yang, J.; Visscher, P.M.; Wray, N.R. Genome-wide association study of medication-use and associated disease in the UK Biobank. Nat. Commun. 2019, 10, 1891. [Google Scholar] [CrossRef] [Green Version]
- Astle, W.J.; Elding, H.; Jiang, T.; Allen, D.; Ruklisa, D.; Mann, A.L.; Mead, D.; Bouman, H.; Riveros-Mckay, F.; Kostadima, M.A.; et al. The Allelic Landscape of Human Blood Cell Trait Variation and Links to Common Complex Disease. Cell 2016, 167, 1415–1429.e19. [Google Scholar] [CrossRef] [Green Version]
- International Consortium for Blood Pressure Genome-Wide Association Studies. Genetic variants in novel pathways influence blood pressure and cardiovascular disease risk. Nature 2011, 478, 103–109. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Guo, R.; Kim, S.H.; Shah, H.; Zhang, S.; Liang, J.H.; Fang, Y.; Gentili, M.; Leary, C.N.O.; Elledge, S.J.; et al. SARS-CoV-2 hijacks folate and one-carbon metabolism for viral replication. Nat. Commun. 2021, 12, 1676. [Google Scholar] [CrossRef]
- Sharma, P.; Senthilkumar, R.D.; Brahmachari, V.; Sundaramoorthy, E.; Mahajan, A.; Sharma, A.; Sengupta, S. Mining literature for a comprehensive pathway analysis: A case study for retrieval of homocysteine related genes for genetic and epigenetic studies. Lipids Health Dis. 2006, 5, 1. [Google Scholar] [CrossRef] [Green Version]
- Karst, M.; Hollenhorst, J.; Achenbach, J. Life-threatening course in coronavirus disease 2019 (COVID-19): Is there a link to methylenetetrahydrofolic acid reductase (MTHFR) polymorphism and hyperhomocysteinemia? Med. Hypotheses 2020, 144, 110234. [Google Scholar] [CrossRef]
- Ponti, G.; Pastorino, L.; Manfredini, M.; Ozben, T.; Oliva, G.; Kaleci, S.; Iannella, R.; Tomasi, A. COVID-19 spreading across world correlates with C677T allele of the methylenetetrahydrofolate reductase (MTHFR) gene prevalence. J. Clin. Lab. Anal. 2021, 35, e23798. [Google Scholar] [CrossRef] [PubMed]
- Ponti, G.; Roli, L.; Oliva, G.; Manfredini, M.; Trenti, T.; Kaleci, S.; Iannella, R.; Balzano, B.; Coppola, A.; Fiorentino, G.; et al. Homocysteine (Hcy) assessment to predict outcomes of ho-spitalized COVID-19 patients: A multicenter study on 313 Covid-19 patients. Clin. Chem. Lab. Med. 2021, 59, e354–e357. [Google Scholar] [CrossRef]
- Durand, P.; Lussier-Cacan, S.; Blache, D. Acute methionine load-induced hyperhomocysteinemia enhances platelet aggregation, thromboxane biosynthesis, and macrophage-derived tissue factor activity in rats. FASEB J. 1997, 11, 1157–1168. [Google Scholar] [CrossRef] [PubMed]
- Fryer, R.H.; Wilson, B.D.; Gubler, D.B.; Fitzgerald, L.A.; Rodgers, G.M. Homocysteine, a risk factor for premature vascular disease and thrombosis, induces tissue factor activity in endothelial cells. Arterioscler. Thromb. 1993, 13, 1327–1333. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Cao, W.; Jiang, W.; Xiao, M.; Li, Y.; Tang, N.; Liu, Z.; Yan, X.; Zhao, Y.; Li, T.; et al. Profile of natural anticoagulant, coagulant factor and anti-phospholipid antibody in critically ill COVID-19 patients. J. Thromb. Thrombolysis 2020, 50, 580–586. [Google Scholar] [CrossRef]
- Stanne, T.M.; Pedersen, A.; Gisslén, M.; Jern, C. Low admission protein C levels are a risk factor for disease worsening and mortality in hospitalized patients with COVID-19. Thromb. Res. 2021, 204, 13–15. [Google Scholar] [CrossRef]
- Corrêa, T.D.; Cordioli, R.L.; Campos Guerra, J.C.; Caldin da Silva, B.; Dos Reis Rodrigues, R.; de Souza, G.M.; Midega, T.D.; Campos, N.S.; Carneiro, B.V.; Campos, F.N.D.; et al. Coagulation profile of COVID-19 patients admitted to the ICU: An exploratory study. PLoS ONE 2020, 15, e0243604. [Google Scholar] [CrossRef]
- Panigada, M.; Bottino, N.; Tagliabue, P.; Grasselli, G.; Novembrino, C.; Chantarangkul, V.; Pesenti, A.; Peyvandi, F.; Tripodi, A. Hypercoagulability of COVID-19 patients in intensive care unit: A report of thromboelastography findings and other parameters of hemostasis. J. Thromb. Haemost. 2020, 18, 1738–1742. [Google Scholar] [CrossRef]
- Hardy, M.; Michaux, I.; Lessire, S.; Douxfils, J.; Dogné, J.M.; Bareille, M.; Horlait, G.; Bulpa, P.; Chapelle, C.; Laporte, S.; et al. Prothrombotic hemostasis disturbances in patients with severe COVID-19: Individual daily data. Data Brief. 2020, 33, 106519. [Google Scholar] [CrossRef]
- Cao, W.J.; Niiya, M.; Zheng, X.W.; Shang, D.Z.; Zheng, X.L. Inflammatory cytokines inhibit ADAMTS13 synthesis in hepatic stellate cells and endothelial cells. J. Thromb. Haemost. 2008, 6, 1233–1235. [Google Scholar] [CrossRef] [Green Version]
- Bernardo, A.; Ball, C.; Nolasco, L.; Moake, J.F.; Dong, J.F. Effects of inflammatory cytokines on the release and cleavage of the endothelial cell-derived ultralarge von Willebrand factor multimers under flow. Blood 2004, 104, 100–106. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goshua, G.; Pine, A.B.; Meizlish, M.L.; Chang, C.H.; Zhang, H.; Bahel, P.; Baluha, A.; Bar, N.; Bona, R.D.; Burns, A.J.; et al. Endotheliopathy in COVID-19-associated coagulopathy: Evidence from a single-centre, cross-sectional study. Lancet Haematol. 2020, 7, e575–e582. [Google Scholar] [CrossRef]
- Ward, S.E.; Curley, G.F.; Lavin, M.; Fogarty, H.; Karampini, E.; McEvoy, N.L.; Clarke, J.; Boylan, M.; Alalqam, R.; Worrall, A.P.; et al. Von Willebrand factor propeptide in severe coronavirus disease 2019 (COVID-19): Evidence of acute and sustained endothelial cell activation. Br. J. Haematol. 2021, 192, 714–719. [Google Scholar] [CrossRef] [PubMed]
- Helms, J.; Tacquard, C.; Severac, F.; Leonard-Lorant, I.; Ohana, M.; Delabranche, X.; Merdji, H.; Clere-Jehl, R.; Schenck, M.; Fagot Gandet, F.; et al. High risk of thrombosis in patients with severe SARS-CoV-2 infection: A multicenter prospective cohort study. Intensive Care Med. 2020, 46, 1089–1098. [Google Scholar] [CrossRef] [PubMed]
- Peyvandi, F.; Artoni, A.; Novembrino, C.; Aliberti, S.; Panigada, M.; Boscarino, M.; Gualtierotti, R.; Rossi, F.; Palla, R.; Martinelli, I.; et al. Hemostatic alterations in COVID-19. Haematologica 2021, 106, 1472–1475. [Google Scholar] [CrossRef] [PubMed]
- Ward, S.E.; Fogarty, H.; Karampini, E.; Lavin, M.; Schneppenheim, S.; Dittmer, R.; Morrin, H.; Glavey, S.; Ni Cheallaigh, C.; Bergin, C.; et al. ADAMTS13 regulation of VWF multimer distribution in severe COVID-19. J. Thromb. Haemost. 2021, 19, 1914–1921. [Google Scholar] [CrossRef] [PubMed]
- Martinelli, N.; Montagnana, M.; Pizzolo, F.; Friso, S.; Salvagno, G.L.; Forni, G.L.; Gianesin, B.; Morandi, M.; Lunardi, C.; Lippi, G.; et al. A relative ADAMTS13 deficiency supports the presence of a secondary microangiopathy in COVID 19. Thromb. Res. 2020, 193, 170–172. [Google Scholar] [CrossRef]
- Philippe, A.; Gendron, N.; Bory, O.; Beauvais, A.; Mirault, T.; Planquette, B.; Sanchez, O.; Diehl, J.L.; Chocron, R.; Smadja, D.M. Von Willebrand factor collagen-binding capacity predicts in-hospital mortality in COVID-19 patients: Insight from VWF/ADAMTS13 ratio imbalance. Angiogenesis 2021, 24, 407–411. [Google Scholar] [CrossRef]
- Marco, A.; Marco, P. Von Willebrand factor and ADAMTS13 activity as clinical severity markers in patients with COVID-19. J. Thromb. Thrombolysis 2021, 1–7. [Google Scholar] [CrossRef]
- Sweeney, J.M.; Barouqa, M.; Krause, G.J.; Gonzalez-Lugo, J.D.; Rahman, S.; Gil, M.R. Low ADAMTS13 Activity Correlates with Increased Mortality in COVID-19 Patients. TH Open 2021, 5, e89–e103. [Google Scholar] [CrossRef] [PubMed]
- Pascreau, T.; Zia-Chahabi, S.; Zuber, B.; Tcherakian, C.; Farfour, E.; Vasse, M. ADAMTS 13 deficiency is associated with abnormal distribution of von Willebrand factor multimers in patients with COVID-19. Thromb. Res. 2021, 204, 138–140. [Google Scholar] [CrossRef] [PubMed]
- De Bruijn, S.; Maes, M.B.; De Waele, L.; Vanhoorelbeke, K.; Gadisseur, A. First report of a de novo iTTP episode associated with an mRNA-based anti-COVID-19 vaccination. J. Thromb. Haemost. 2021, 19, 2014–2018. [Google Scholar] [CrossRef] [PubMed]
- Yocum, A.; Simon, E.L. Thrombotic Thrombocytopenic Purpura after Ad26.COV2-S Vaccination. Am. J. Emerg Med. 2021, 49, 441.e3–441.e4. [Google Scholar] [CrossRef]
- Albánez, S.; Ogiwara, K.; Michels, A.; Hopman, W.; Grabell, J.; James, P.; Lillicrap, D. Aging and ABO blood type influence von Willebrand factor and factor VIII levels through interrelated mechanisms. J. Thromb. Haemost. 2016, 14, 953–963. [Google Scholar] [CrossRef] [PubMed] [Green Version]

| Chr | Gene | Protein | Gene Size (kb) | Analyzed Region (kb) | Coordinates of the Analyzed Region (hg38) | Analyzed SNPs (n) |
|---|---|---|---|---|---|---|
| 1 | MTHFR | MTHR | 20.3 | 520.3 | 1:11535729-12056103 | 1821 |
| 1 | F5 | FA5 (FV) | 74.5 | 574.5 | 1:169261953-169836531 | 2276 |
| 1 | F13B | F13B (FXIII) | 28.0 | 528 | 1:196789190-197317267 | 956 |
| 1 | MTR | METH | 108.7 | 608.7 | 1:236545280-237153981 | 2270 |
| 2 | TFPI | TFPI | 90.2 | 590.2 | 2:187214230-187804492 | 1655 |
| 2 | PROC | PROC | 10.8 | 510.8 | 2:127168419-127679246 | 1492 |
| 3 | PROS1 | PROS | 101.0 | 601 | 3:93623036-94224090 | 592 |
| 4 | FGB | FIBB | 49.7 | 549.7 | 4:154312979-154862750 | 1606 |
| 4 | FGA | |||||
| 4 | FGG | |||||
| 4 | F11 | FA11 (FXI) | 23.7 | 523.7 | 4:186015963-186539681 | 2251 |
| 5 | ITGA2 | ITA2 (GPIa) | 105.4 | 605.4 | 5:52739325-53344779 | 2469 |
| 5 | THBS4 | TSP4 | 47.9 | 547.9 | 5:79785348-80333284 | 2047 |
| 5 | F12 | FA12 (FXII) | 7.4 | 507.4 | 5:177152137-177659576 | 890 |
| 6 | F13A1 | F13A (FXIII) | 176.6 | 676.6 | 6:5894077-6570691 | 2895 |
| 6 | THBS2 | TSP2 | 38.3 | 538.3 | 6:168965779-169504114 | 2080 |
| 7 | CD36 | CD36 (GPIIIb) | 304.8 | 804.8 | 7:80119575-80924418 | 2110 |
| 7 | SERPINE1 | PAI-1 | 12.1 | 512.1 | 7:100877088-101389266 | 1929 |
| 8 | PLAT | TPA | 32.9 | 532.9 | 8:41924717-42457676 | 927 |
| 9 | ADAMTS13 | ATS13 (ADAMTS13) | 37.4 | 537.4 | 9:133171999-133709403 | 2195 |
| 11 | F2 | THRB | 20.3 | 520.3 | 11:46469192-46989506 | 697 |
| 12 | VWF | VWF | 175.7 | 675.7 | 12:5698873-6374670 | 2122 |
| 13 | CPB2 | CBPB2 (TAFI) | 51.8 | 551.8 | 13:45803186-46355076 | 1811 |
| 13 | F7 | FA7 (FVII) | 14.8 | 514.8 | 13:112855787-113370681 | 1411 |
| 13 | F10 | FA10 (FX) | 26.7 | 526.7 | 13:112872798-113399529 | 1540 |
| 15 | THBS1 | TSP1 | 17.8 | 517.8 | 15:39331078-39848921 | 1443 |
| 17 | GPIBA | GPIBA | 2.7 | 502.7 | 17:4682274-5185030 | 1502 |
| 17 | ITGB3 | ITB3 (GPIIIa) | 58.8 | 558.8 | 17:47003841-47562711 | 1695 |
| 18 | LMAN1 | LMAN1 | 31.4 | 531.4 | 18:59077823-59609276 | 2327 |
| 20 | THBD | TRBM | 4.0 | 504 | 20:22795632-23299664 | 1488 |
| X | F8 | FA8 (FVIII) | 186.9 | 686.9 | 23:154585788-155272723 | 570 |
| X | F9 | FA9 (FIX) | 32.7 | 532.7 | 23:139280735-139813458 | 778 |
| Total | 32 genes | - | 1893 | 16,893 | - | 49,845 |
| (a) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Uncorrected Analyses | Corrected Analyses | ||||||||
| Chr | Gene | SNP | MAF Cases | MAF Controls | p Value | OR (95%CI) | p Value | OR (95%CI) | |
| 2 | PROC | chr2:127192625:G:A | 0.065 | 0.037 | 8.51 × 10−4 | 1.82 (1.28–2.61) | 8.77 × 10−5 | 2.23 (1.50–3.34) | |
| 1 | MTHFR | chr1:11753033:G:A | 0.247 | 0.183 | 1.33 × 10−4 | 1.47 (1.20–1.79) | 1.08 × 10−4 | 1.56 (1.25–1.95) | |
| 1 | MTR | chr1:237145686:A:G | 0.023 | 0.010 | 4.53 × 10−3 | 2.39 (1.29–4.43) | 3.14 × 10−4 | 3.49 (1.77–6.88) | |
| 9 | ADAMTS13 | chr9:133179750:G:C | 0.030 | 0.011 | 1.58 × 10−4 | 2.77 (1.60–4.80) | 4.26 × 10−4 | 2.90 (1.60–5.24) | |
| 6 | THBS2 | chr6:169195156:A:T | 0.039 | 0.021 | 6.25 × 10−3 | 1.87 (1.19–2.96) | 7.84 × 10−4 | 2.35 (1.43–3.87) | |
| 17 | ITGB3 | chr17:47019591:G:C | 0.321 | 0.270 | 7.05 × 10−3 | 1.28 (1.07–1.53) | 1.41 × 10−3 | 1.39 (1.14–1.71) | |
| 7 | SERPINE1 | chr7:101070945:A:G | 0.324 | 0.267 | 2.55 × 10−3 | 1.32 (1.10–1.58) | 1.73 × 10−3 | 1.38 (1.13–1.69) | |
| 6 | F13A1 | chr6:6163858:G:C | 0.108 | 0.070 | 6.99 × 10−4 | 1.61 (1.22–2.13) | 1.80 × 10−3 | 1.67 (1.21–2.31) | |
| 18 | LMAN1 | chr18:59208206:A:G | 0.401 | 0.337 | 1.57 × 10−3 | 1.32 (1.11–1.56) | 1.81 × 10−3 | 1.36 (1.12–1.65) | |
| 4 | F11 | chr4:186056516:A:G | 0.065 | 0.106 | 1.31 × 10−3 | 0.59 (0.42–0.82) | 1.84 × 10−3 | 0.56 (0.39–0.81) | |
| 7 | CD36 | chr7:80591832:AAATCAGC:A | 0.039 | 0.021 | 6.25 × 10−3 | 1.87 (1.19–2.96) | 1.98 × 10−3 | 2.15 (1.33–3.50) | |
| 15 | THBS1 | chr15:39455553:G:C | 0.178 | 0.133 | 2.17 × 10−3 | 1.42 (1.13–1.77) | 2.50 × 10−3 | 1.46 (1.14–1.86) | |
| 12 | VWF | chr12:6068637:T:C | 0.054 | 0.031 | 3.71 × 10−3 | 1.76 (1.20–2.60) | 2.50 × 10−3 | 1.95 (1.27–3.01) | |
| 13 | F7–F10 | chr13:113257337:C:T | 0.288 | 0.234 | 3.50 × 10−3 | 1.32 (1.10–1.59) | 3.13 × 10−3 | 1.37 (1.11–1.69) | |
| 20 | THBD | chr20:22949512:A:G | 0.048 | 0.025 | 1.51 × 10−3 | 1.94 (1.28–2.93) | 3.18 × 10−3 | 2.01 (1.26–3.19) | |
| 4 | FG_genes | chr4:154774926:G:A | 0.054 | 0.033 | 8.81 × 10−3 | 1.67 (1.13–2.45) | 4.45 × 10−3 | 1.88 (1.22–2.91) | |
| 1 | F13B | chr1:196913749:A:AT | 0.036 | 0.017 | 1.84 × 10−3 | 2.12 (1.31–3.44) | 5.04 × 10−3 | 2.12 (1.25–3.59) | |
| 13 | CPB2 | chr13:45931697:A:G | 0.008 | 0.026 | 3.59 × 10−3 | 0.28 (0.12–0.70) | 5.29 × 10−3 | 0.18 (0.056–0.61) | |
| 8 | PLAT | chr8:42278531:C:T | 0.017 | 0.007 | 1.79 × 10−2 | 2.33 (1.13–4.77) | 5.69 × 10−3 | 2.88 (1.36–6.08) | |
| 5 | ITGA2 | chr5:53162220:CAGAG:C | 0.029 | 0.013 | 4.56 × 10−3 | 2.15 (1.25–3.71) | 7.60 × 10−3 | 2.30 (1.25–4.23) | |
| 2 | TFPI | chr2:187303697:A:G | 0.030 | 0.013 | 1.13 × 10−3 | 2.38 (1.39–4.07) | 8.74 × 10−3 | 2.25 (1.23–4.14) | |
| 1 | F5 | chr1:169677905:G:A | 0.116 | 0.151 | 1.90 × 10−2 | 0.74 (0.57–0.95) | 9.74 × 10−3 | 0.68 (0.51–0.91) | |
| 5 | THBS4 | chr5:80174425:G:T | 0.203 | 0.163 | 1.23 × 10−2 | 1.31 (1.06–1.61) | 9.83 × 10−3 | 1.37 (1.08–1.73) | |
| 3 | PROS1 | chr3:94043799:A:C | 0.032 | 0.014 | 1.84 × 10−3 | 2.24 (1.33–3.76) | 1.19 × 10−2 | 2.10 (1.18–3.74) | |
| 11 | F2 | chr11:46517560:T:C | 0.331 | 0.382 | 1.39 × 10−2 | 0.80 (0.63–0.96) | 1.19 × 10−2 | 0.77 (0.63–0.95) | |
| 17 | GP1BA | chr17:4921551:T:C | 0.020 | 0.010 | 2.59 × 10−2 | 2.06 (1.08–3.95) | 1.31 × 10−2 | 2.50 (1.21–5.15) | |
| 5 | F12 | chr5:177464930:A:G | 0.026 | 0.013 | 1.71 × 10−2 | 1.97 (1.12–3.46) | 1.38 × 10−2 | 2.17 (1.17–4.03) | |
| (b) | |||||||||
| Chr | Gene | SNP | pValue_M | OR (95%CI)_M | pValue_F | OR (95%CI)_F | P_Comb_Stouffer | ||
| X | F9 | chrX:139407485:G:C | 8.5 × 10−4 | 1.27 (0.96–1.68) | 0.49 | 0.91 (0.63–1.31) | 0.0046 | ||
| Corrected Analyses | ||||||
|---|---|---|---|---|---|---|
| Chr | Gene | SNP | SNP ID | Minor Allele | p Value | OR (95%CI) |
| 1 | F5 | chr1:169549811:C:T | rs6025 | T | 0.92 | 0.96 (0.48–1.94) |
| 6 | F13A1 | chr6:6318562:C:A | rs5985 | A | 0.91 | 0.99 (0.78–1.24) |
| 11 | F2 | chr11:46739505:G:A | rs1799963 | A | 0.015 | 0.23 (0.070–0.75) |
| Italians (Our Cohort) | Regeneron Cohort | Meta-Analysis | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Gene | SNP | Analysed Individuals (n) | p Value | OR (95%CI) | Analysed Individuals (n) | p Value | OR (95%CI) | Pooled OR (95%CI) | p Value (M-H) | p Value (F Weighted) |
| PROC | chr2:127192625:G:A | 2000 | 8.51 × 10−4 | 1.82 (1.28–2.61) | 429151 | 0.026 | 0.71 (0.53–0.96) | 0.98 (0.79–1.22) | 0.87 | 0.12 |
| MTHFR | chr1:11753033:G:A | 2000 | 1.33 × 10−4 | 1.47 (1.20–1.79) | 654056 | 0.030 | 1.13 (1.01–1.28) | 1.21 (1.09–1.33) | 2.55 × 10−4 | 4.34 × 10−14 |
| MTR | chr1:237145686:A:G | 2000 | 4.53 × 10−3 | 2.39 (1.29–4.43) | 898324 | 0.98 | 1.00 (0.88–1.14) | 1.03 (0.91–1.18) | 0.61 | 0.0027 |
| ADAMTS13 | chr9:133179750:G:C | 2000 | 1.58 × 10−4 | 2.77 (1.60–4.80) | 656078 | 0.030 | 0.75 (0.58–0.97) | 0.89 (0.72–1.11) | 0.33 | 0.056 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cappadona, C.; Paraboschi, E.M.; Ziliotto, N.; Bottaro, S.; Rimoldi, V.; Gerussi, A.; Azimonti, A.; Brenna, D.; Brunati, A.; Cameroni, C.; et al. MEDTEC Students against Coronavirus: Investigating the Role of Hemostatic Genes in the Predisposition to COVID-19 Severity. J. Pers. Med. 2021, 11, 1166. https://doi.org/10.3390/jpm11111166
Cappadona C, Paraboschi EM, Ziliotto N, Bottaro S, Rimoldi V, Gerussi A, Azimonti A, Brenna D, Brunati A, Cameroni C, et al. MEDTEC Students against Coronavirus: Investigating the Role of Hemostatic Genes in the Predisposition to COVID-19 Severity. Journal of Personalized Medicine. 2021; 11(11):1166. https://doi.org/10.3390/jpm11111166
Chicago/Turabian StyleCappadona, Claudio, Elvezia Maria Paraboschi, Nicole Ziliotto, Sandro Bottaro, Valeria Rimoldi, Alessio Gerussi, Andrea Azimonti, Daniele Brenna, Andrea Brunati, Charlotte Cameroni, and et al. 2021. "MEDTEC Students against Coronavirus: Investigating the Role of Hemostatic Genes in the Predisposition to COVID-19 Severity" Journal of Personalized Medicine 11, no. 11: 1166. https://doi.org/10.3390/jpm11111166









