1. Introduction
Molecular testing-guided targeted therapies have dramatically transformed the prognostic landscape for patients with non-small cell lung carcinoma (NSCLC). With the rapid discovery of novel oncogenic drivers, clinical guidelines for newly diagnosed advanced NSCLC continue to expand the requisite repertoire of therapeutic biomarkers [
1,
2,
3]. Although advancements in minimally invasive techniques have facilitated specimen collection with reduced morbidity, the resulting samples are frequently limited in volume. These scant specimens must support a multifaceted diagnostic workflow, including definitive histological diagnosis, ancillary immunohistochemical (IHC) staining, and comprehensive molecular profiling. This competing demand for limited material presents a formidable challenge in clinical practice. Indeed, studies indicate that 10% to 30% of small biopsies are inadequate for downstream molecular analysis due to low tumor cellularity or insufficient DNA yield [
4,
5,
6,
7,
8,
9,
10]. Consequently, a significant subset of patients either loses the opportunity for targeted intervention or must undergo the inherent risks of repeat biopsy. While liquid biopsies utilizing circulating tumor DNA (ctDNA) provide an alternative for tissue-limited cases, their clinical utility is often hindered by suboptimal sensitivity compared to tissue-based assays [
11,
12,
13].
The most prevalent minimally invasive techniques in lung cancer diagnostics include Transbronchial forceps lung biopsy (TBLB) with or without concurrent endobronchial ultrasound-guided transbronchial needle aspiration (EBUS-TBNA), and CT- or ultrasound-guided transthoracic core needle biopsy (CNB) with or without concurrent fine-needle aspiration (FNA). Pathologically, TBLB and CNB yield histological small specimens, whereas EBUS-TBNA and FNA provide cytological aspirates.
Data from a 2021 Canadian study indicated that 10% of NSCLC samples were insufficient for molecular testing, and even among tissue-adequate samples, 16% yielded insufficient DNA [
4]. Similarly, a retrospective analysis at MD Anderson Cancer Center found aggregate molecular adequacy rates of 84%, with FNA and EBUS-TBNA performing lower (72% and 77%, respectively) than CNB (90%) [
5]. Conversely, Hendry et al. reported an 89.5% adequacy rate when integrating rapid on-site evaluation (ROSE), though success rates still varied by modality: 96.8% for CNB, 85.8% for FNA, and 88.8% for EBUS-TBNA [
6].
Preliminary histological evaluation of hematoxylin and eosin (H&E) stained formalin-fixed paraffin-embedded (FFPE) tissue sections indicates that lung cancer specimens obtained via small biopsy are inherently fragile, frequently shedding microscopic tissue fragments or cell clusters into the fixative medium [
14,
15]. This phenomenon is particularly pronounced in poorly differentiated or extensively necrotic tumors. Unlike resilient normal stroma, neoplastic tissue is highly susceptible to fragmentation under the mechanical compression of biopsy forceps and the needle aspiration process. Furthermore, mechanical agitation during transport facilitates the shedding of poorly adherent neoplastic cells into the fixative solution, while stromal components remain structurally intact. At present, pathological workflows frequently neglect the diagnostic potential of tumor cells suspended in specimen fixatives. Routine processing focuses exclusively on the retrieval of macroscopically visible tissue for paraffin embedding, whereas microscopic clusters or fragments—too small for manual manipulation—remain unrecovered. As a result, this residual fixative medium, despite its potential cellular richness, is typically disposed of as medical waste.
Our previous research established a robust methodology for salvaging cell pellets from this residual fixative for molecular profiling [
14,
15]. A critical finding of our study is that the majority of lung biopsies yield high-quality tumor cell pellets within the residual medium. By integrating these pellets with traditional formalin-fixed paraffin-embedded (FFPE) tissue, we significantly improved the diagnostic adequacy rate. Our preliminary single-center study demonstrated that integrating a specialized collection workflow to salvage shed tumor cells from the fixative significantly improves the molecular success rate for histological specimens. The present project aims to validate the efficacy of these refined methods in improving molecular adequacy for lung cancer small biopsies across multiple centers.
2. Materials and Methods
2.1. Study Design and Patient Selection
This multicenter study enrolled patients diagnosed with non-small cell lung cancer (NSCLC) via conventional histopathological examination across three institutions: the First Affiliated Hospital of Guangzhou Medical University (Center A), Affiliated Cancer Hospital & Institute of Guangzhou Medical University (Center B), and Sun Yat-sen University Cancer Center (Center C). The study protocol was approved by the Institutional Review Board of the First Affiliated Hospital of Guangzhou Medical University (Ref: 2021-70; 16 August 2021).
Eligible patients underwent minimally invasive diagnostic procedures—including TBLB, CNB, and EBUS-TBNA—between January and June 2024. We prospectively and retrospectively monitored the handling workflows for small biopsy and cytological specimens at each center. The evaluation focused on the macroscopic state of specimens upon arrival at the pathology department, technician grossing methodologies, the presence of residual tissue in the fixative post-processing, and the overall pre-analytic adequacy of samples for downstream molecular analysis. Comprehensive clinicopathological data, including age, sex, specimen source, sampling technique, and IHC profiles, were retrieved from each institution’s clinical and pathological information systems.
2.2. TBLB Specimen Processing Methodologies
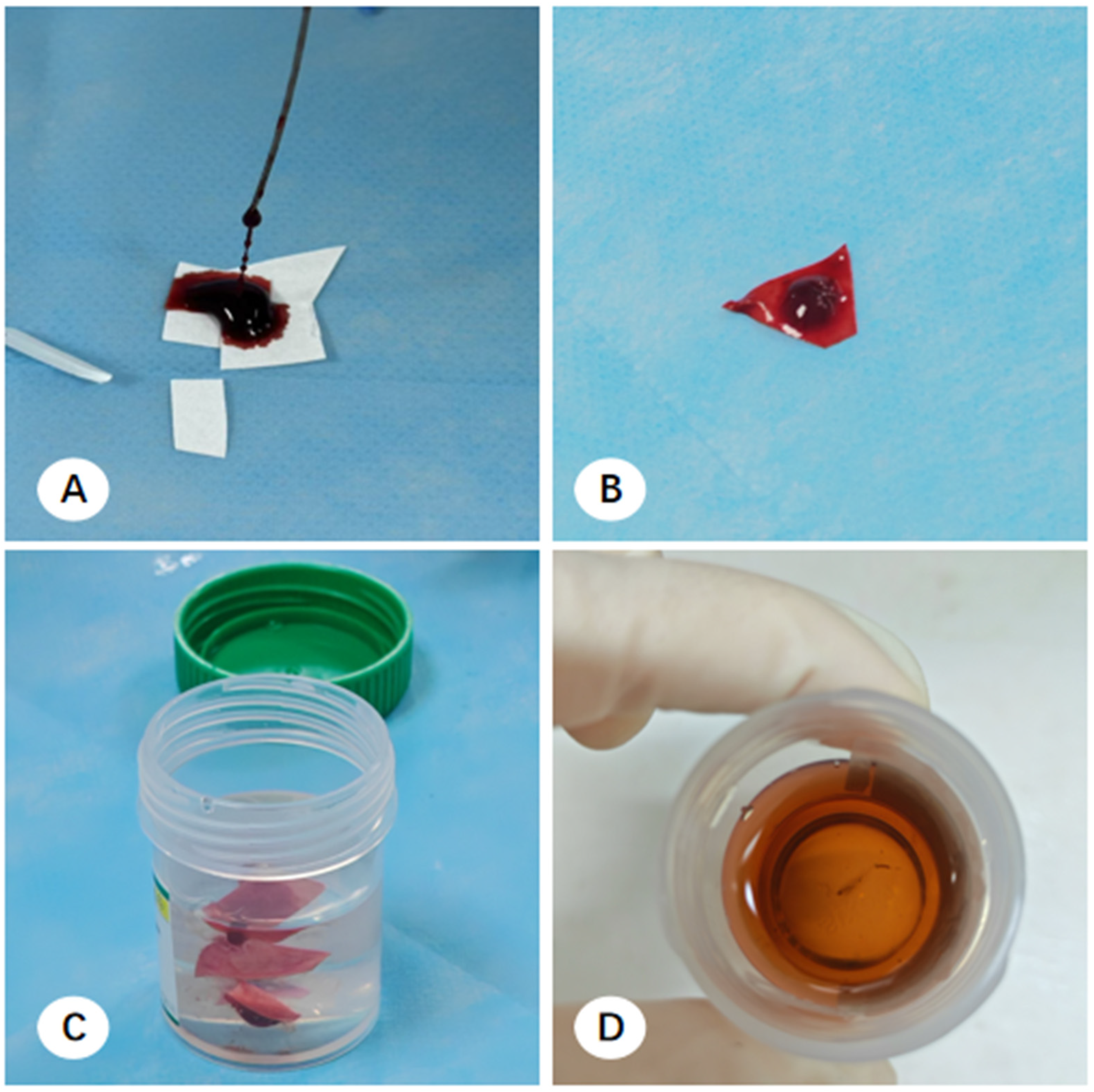
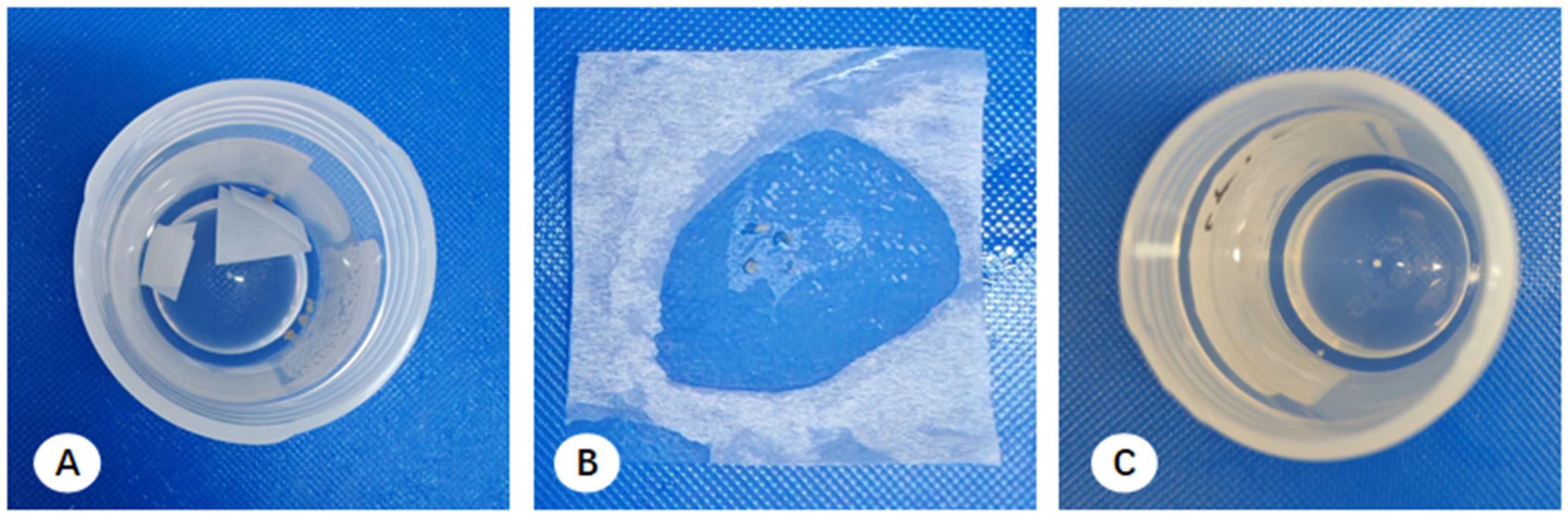
At each participating center, bronchoscopy unit support staff prepared sterilized 1 cm × 1 cm filter paper segments for specimen collection. During the TBLB procedure, two to five tissue fragments were typically collected and adhered to these segments before immersion in specimen bottles containing 10% neutral-buffered formalin.
Upon arrival at the pathology department, macroscopic assessment revealed multiple filter paper segments within the fixative; while some tissue remained adhered, other fragments had detached and were suspended freely. Pathologists used tweezers to scrape tissue from the paper and retrieve visible free-floating fragments, which were then wrapped in embedding paper for paraffin embedding. However, this manual recovery was limited to macroscopically visible fragments. Consequently, minute, ungraspable tissue residues consistently remained in the residual formalin across all centers, representing a systematic loss of diagnostic material (
Figure 1).
2.3. CNB Specimen Processing Methodologies
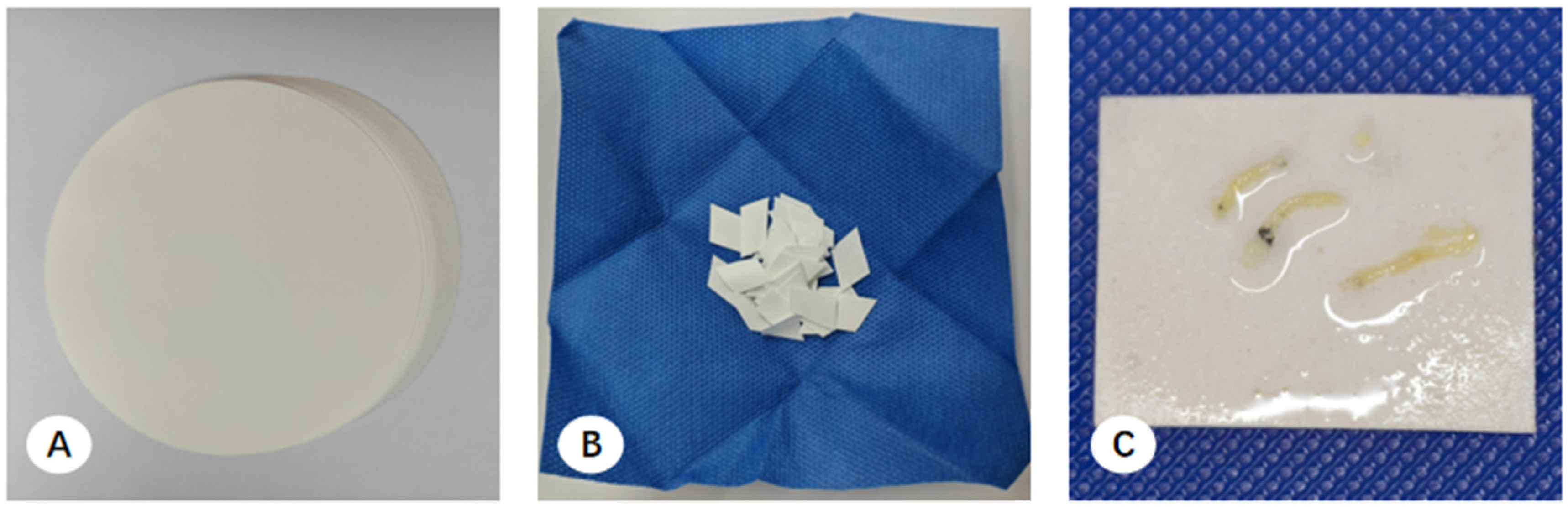
Multicenter observations indicated that, unlike the bronchoscopy units, Interventional Radiology (IR) departments generally did not include standardized adherence materials in their sterile puncture kits. Consequently, clinicians often improvised by using product qualification certificates (from ethylene oxide-sterilized kits) or 3M steam sterilization indicator cards (from autoclaved kits) to secure CNB specimens (
Figure 2). These cards—intended solely for verifying sterilization efficacy—were immersed with the tissue in 10% formalin. Gross pathological assessment revealed that these cards lost structural integrity and shed fibers upon saturation, causing them to adhere aggressively to the biopsy cores. Attempts to separate the tissue using tweezers frequently resulted in specimen fragmentation and loss of diagnostic material. Additionally, because CNB specimens often contained necrotic or friable regions. These samples frequently disintegrated into minute particles that remained suspended in the fixative, mirroring the losses observed in TBLB processing.
To address these limitations, the IR department at Center A, in collaboration with the pathology department, optimized its handling protocols by utilizing pre-cut Polypropylene Microporous Membranes (PPMM). Large PPMMs (27 cm diameter) were trimmed into 1.5 cm × 2 cm rectangles, packaged, and sterilized (
Figure 3). These membranes provided enhanced durability and minimal fiber shedding, thereby maintaining specimen integrity and significantly improving tissue recovery during the grossing process [
14].
2.4. EBUS-TBNA Specimen Processing Methodologies
Multicenter observations identified three primary handling methods for EBUS-TBNA specimens:
- (1)
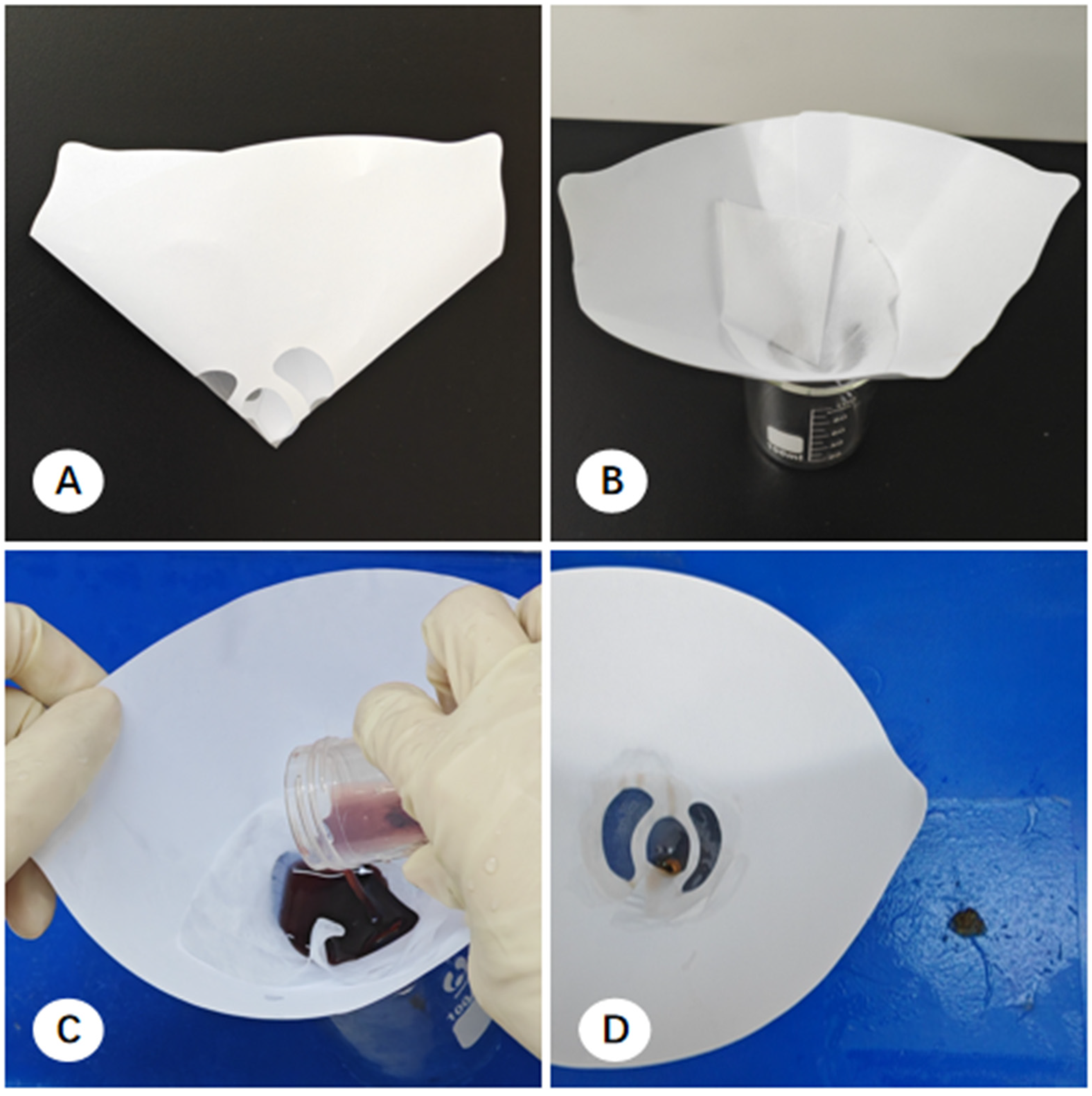
Filter Paper-Based Collection: Aspirated material was expressed from the needle via a wire stylet onto pre-cut filter paper to form a coagulated mixture of tissue and blood. The resulting clot was air-dried for 3 min to ensure adequate coagulation and prevent the rapid dispersion of cellular components upon immersion. The clot and filter paper were then submerged in 10% formalin for transport (
Figure 4) [
16].
- (2)
Direct Self-clotting in Centrifuge Tubes: Aspirated materials were expelled directly into a centrifuge tube to allow for spontaneous coagulation, followed by the addition of 10% formalin fixative (
Figure 5) [
17].
- (3)
Direct Fixation: Some pulmonologists bypassed coagulation entirely, expelling the aspirated material directly into a specimen bottle containing 10% formalin for immediate stabilization.
In the pathology laboratory, processing varied based on the collection method. For filter paper-based samples, tissue clots were scraped from the paper, and macroscopically visible flakes suspended in the fixative were retrieved. These were wrapped in embedding paper and processed into FFPE cell blocks (CBs) (
Figure 4). For direct self-clotting or direct fixation samples, technicians used tweezers to retrieve thread-like material or utilized a decanting method, where the majority of the liquid was poured out and the remaining sediment was filtered through embedding paper (
Figure 5).
To optimize sample recovery, the pathology technician at Center A implemented a Funnel Filtration Technique [
18]. A specialized funnel device was constructed using a disposable paper funnel (Beijing Ruimiao Jingyi Trading Co., Ltd., Beijing, China), which was originally de-signed for paint filtration, with a fine-mesh base. A piece of embedding paper was placed inside the funnel, which was positioned over a beaker. The solid–liquid mixture from the centrifuge tube was poured through this device, allowing the fixative to drain rapidly while ensuring that all tissue clots and dispersed cellular material were captured on the lens paper. This collected material was then wrapped and processed into CBs using the standard FFPE protocol (
Figure 6).
2.5. FFPE Specimen Processing and Pre-Analytic Assessment
All small biopsy specimens were processed on the day of laboratory receipt. Following gross assessment, formalin-fixed biopsy fragments were retrieved, wrapped in embedding paper, and placed in cassettes for paraffin embedding. The next day, FFPE blocks were sectioned at 4 μm and stained with H&E. Upon histopathological confirmation of NSCLC, the total tumor cell count and the neoplastic cell fraction were quantified for each section. Samples were classified as adequate for molecular testing if they contained more than 200 tumor cells per section and exhibited a tumor cell fraction above the established thresholds (typically >10% or >5%, depending on the specific molecular assay requirements). A specimen was deemed adequate only if it simultaneously met both criteria: having at least 200 tumor cells per section and a tumor cell fraction at or above the aforementioned thresholds.
2.6. Cell Pellet Collection and Cellularity Evaluation
During the final two months of the study, a modified protocol was implemented to salvage residual cellular material from the fixative solution. On the day of gross processing, following primary tissue retrieval, pathology technicians retained the residual fixative while discarding the adherence paper. For EBUS-TBNA specimens, the supernatant was preserved in a secondary centrifuge tube during decanting. Once the primary tissue was harvested, the supernatant was returned to the original tube for storage. The next day, in cases where NSCLC was confirmed but the FFPE tissue was deemed inadequate for molecular profiling through pre-analytic assessment, the residual fixative was processed to generate a cell pellet. The recovery technique for EBUS-TBNA samples varied by site according to local harvesting practices: Center A: Utilized a funnel filtration method; because this process sequestered all viable cellular material, the filtered residual solution was devoid of diagnostic cells and was not collected. Centers B and C: The supernatant was preserved and the residual fixative was centrifuged to harvest a cell pellet for further evaluation.
The residual medium was transferred to a 15 mL tube and centrifuged at 1000×
g for 5 min. The resulting pellets were resuspended in 1 mL of phosphate-buffered saline (PBS), transferred to a 1.5 mL microcentrifuge tube, and centrifuged at 3000×
g for 5 min. To mitigate nucleic acid crosslinking caused by prolonged formalin exposure, the pellets underwent two additional PBS wash cycles. Following the final wash and supernatant removal, the pellets were resuspended in 250 μL of PBS. A 50 μL aliquot was used to prepare a direct smear, which was air-dried and H&E-stained. The remaining 200 μL suspension was centrifuged at 3000×
g for 5 min; the supernatant was then discarded, and the resulting cell pellet was stored at −80 °C for DNA extraction [
14,
15,
18]. For cases requiring molecular testing of residual medium, cell pellet harvesting is typically completed within 48 h of sample receipt by the laboratory.
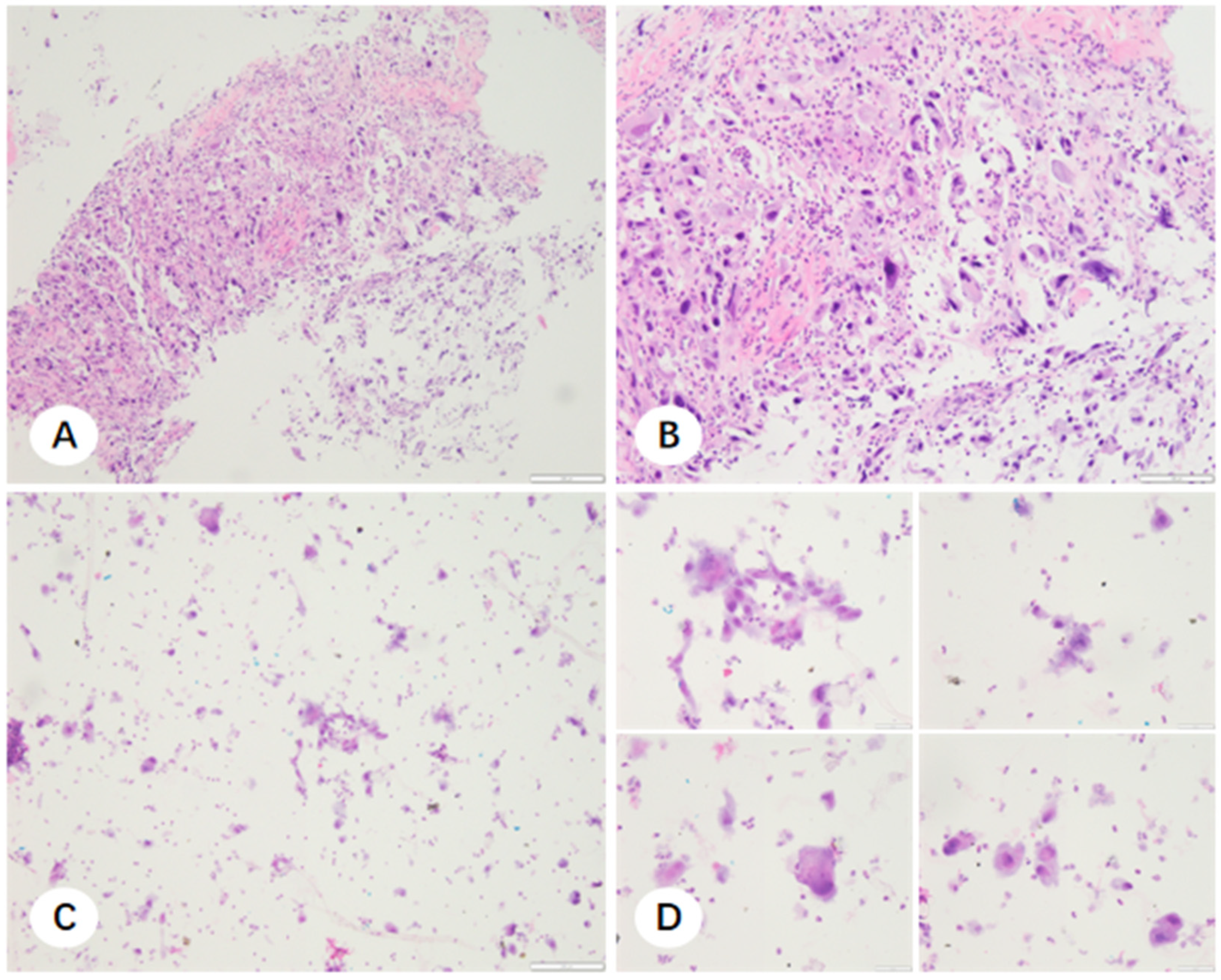
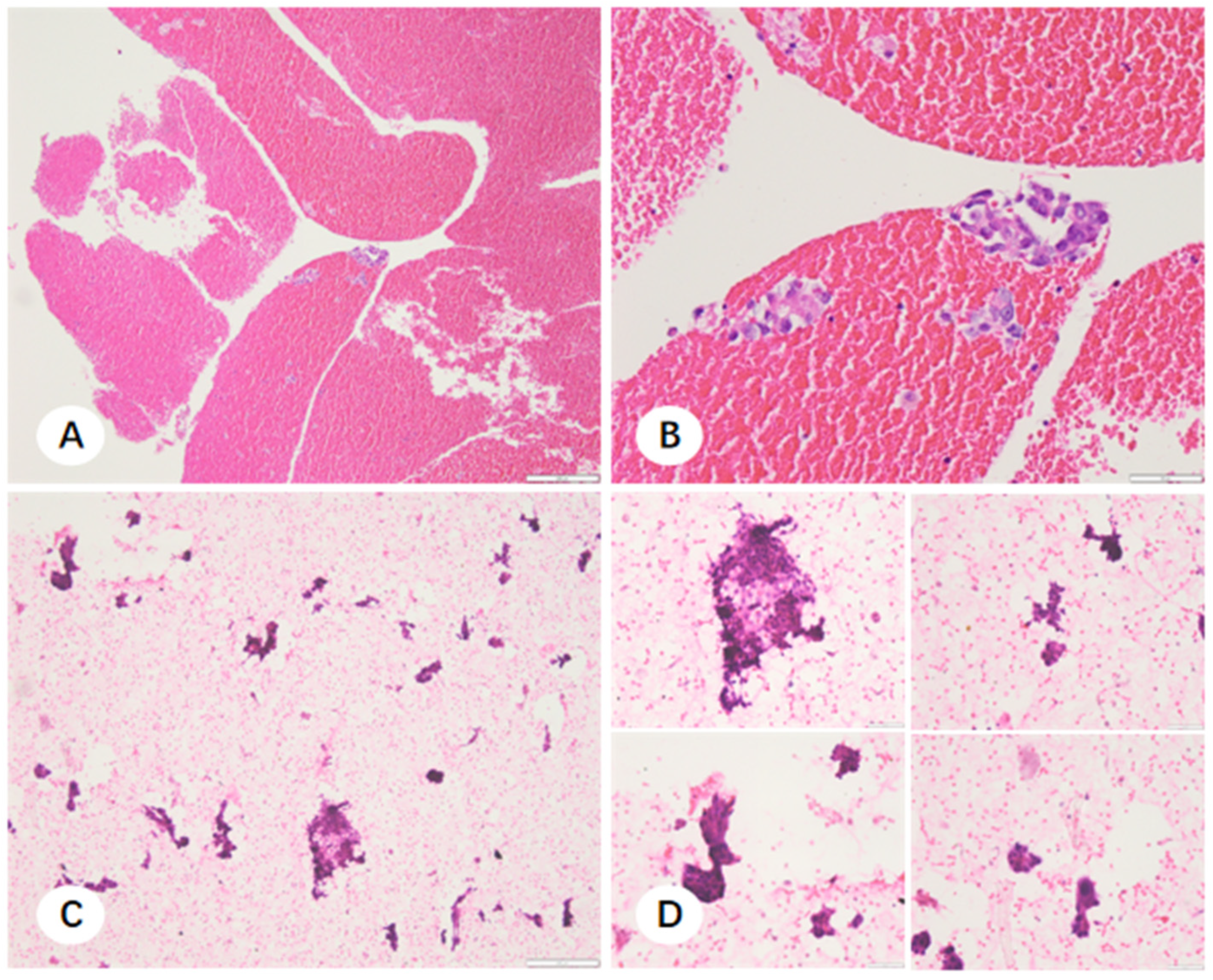
Neoplastic cellularity was quantified by scanning the index smear via light microscopy in a systematic raster pattern. Because the final stored pellet represented four times the volume of the 50 μL aliquot, molecular adequacy was extrapolated from the smear. Samples were classified as adequate if the index smear contained more than 50 tumor cells (corresponding to >200 cells in the final pellet) and a tumor fraction of >10%.
For cell pellet samples deemed adequate by cell counting evaluation, DNA was extracted using the QIAGEN DNeasy Blood and Tissue Kit (Qiagen, Hilden, Germany) following the manufacturer’s protocol. The extracted DNA was quantified with a Qubit 2.0 fluorometer (Invitrogen, Carlsbad, CA, USA). Samples were required to contain >10 ng of DNA for PCR analysis and >30 ng for NGS testing.
2.7. Statistical Analysis
Statistical analyses were performed using SPSS version 26.0 (IBM Corp., Armonk, NY, USA). Descriptive statistics were used to summarize specimen characteristics and baseline adequacy rates. To evaluate the baseline consistency across the three participating centers, Pearson’s chi-squared test (with 2 degrees of freedom, X2critical value = 5.991) was employed for inter-institutional comparisons of each biopsy modality (TBLB, CNB, and EBUS-TBNA). To control for the potential inflation of Type I error due to multiple comparisons between the three centers (Center A vs. B, A vs. C, and B vs. C), the Bonferroni correction was pre-specified, resulting in an adjusted significance threshold of α’ = (0.05/3).
For the assessment of adequacy improvement following the addition of cell pellets, each specimen served as its own internal control. The significance of the incremental adequacy was analyzed using McNemar’s test for paired categorical data. Specifically, the McNemar chi-square asymptotic formula was used when the number of discordant pairs (b + c) was ≥25; for instances where b + c < 25, the Binomial exact test was applied to ensure statistical accuracy. p-value < 0.05 was considered statistically significant.
4. Discussion
In the current landscape of precision oncology, the management of lung cancer relies heavily on obtaining sufficient neoplastic material for both definitive histological subtyping and comprehensive molecular profiling. As targeted therapies and immunotherapies expand, the “tissue is issue” challenge has intensified; small biopsy specimens obtained via minimally invasive techniques—such as image-guided CNB or EBUS-TBNA—frequently face the risk of tissue exhaustion. Consequently, optimizing the processing of these limited specimens has become a critical priority to ensure that every patient receives a complete molecular diagnosis from a single invasive procedure [
4,
5,
6,
7,
8,
9,
10].
Strategic optimization involves a multi-modal approach to specimen handling that shifts away from traditional, tissue-depleting workflows. A cornerstone of this modern strategy is the salvage of “medical waste” substrates, including rinsing the coaxial or aspiration needle and harvesting cell pellets from the residual transport fixative [
14,
15,
18,
19,
20,
21,
22]. These fluids often contain high-quality exfoliated cells and tissue fragments that, when processed into cell pellet, cell blocks or liquid-based cytology (LBC) slides, provide an essential secondary reservoir of tumor cells. These adjuncts not only increase diagnostic sensitivity—particularly in cases where the primary solid core is necrotic or scant—but also offer a high-yield source of high-quality nucleic acids for downstream Next-Generation Sequencing (NGS) [
14,
15].
Enriched tumor cells are frequently found in ancillary fluids—such as bronchial biopsy rinses, core needle rinses, and residual fixative media—due to a combination of the biological characteristics of lung cancer and the mechanical trauma of the biopsy procedure. Lung cancer tissues are often characterized by reduced cell-to-cell adhesion and structural fragility. Unlike healthy, organized tissue, malignant cells are easily dislodged from the main tumor mass. The act of inserting a needle or using biopsy forceps acts as a “mechanical scraping”. This physical disruption breaks off small clusters of tumor cells and microscopic tissue fragments from the lesion. As the needle or forceps transgresses the tumor and is withdrawn, cells often migrate or “spill” into the lumen of the needle or the surrounding fluid track. Once the solid core biopsy is placed into a transport fixative (like formalin), the mechanical trauma already sustained causes tumor cells to continue shedding into the surrounding liquid.
A study by Rosell et al. demonstrated that cytological examination of rinse fluid from bronchial biopsies increased the diagnostic yield for endobronchial malignancies by 4.8%. This technique maximizes the use of materials often discarded during bronchoscopy and adds no significant time or expense to the procedure [
19]. Research by Mojica et al. suggests that salvaging cells from CNB washes can significantly extend the utility of small specimens without compromising the integrity of the primary tissue core [
20]. Lan et al. (2021) further validated this approach, showing that LBC performed on core needle rinses (CNR) significantly improves diagnostic sensitivity for peripheral lung nodules [
21]. By analyzing 406 patients, they found that combining CNR with traditional CNB increased sensitivity from 92.5% to 96.4%, effectively reducing false-negative rates [
21].
Two additional studies explored using these alternative sample sources for genetic testing in NSCLC. Sakairi et al. (2014) investigated using saline rinse solutions from bronchoscopic biopsy needles—termed ultra-microsamples—for comprehensive biomarker testing [
22]. Their analysis of 1474 molecular tests showed a 99.3% concordance rate with results from conventional histological samples [
22]. A 2022 study in thoracic cancer evaluated the feasibility of using supernatants from core-needle biopsy samples as a liquid biopsy source for 48 patients with lung adenocarcinoma. The study reported a 95.8% concordance rate for DNA-level mutation detection compared to tissue samples. While the results confirm that supernatants are a valuable source for genotyping, the researchers noted that methods for preserving and extracting RNA from these specimens require further improvement [
23].
Our preliminary single-center studies demonstrated the significant potential of collecting cell pellets from the residual fixative medium of both TBLB and CNB samples. Conventionally, this medium, containing tiny tissue fragments and cell clusters, is discarded. Our study found that the fixative medium was frequently enriched with tumor cells. Among 324 TBLB samples, 21.6% had inadequate FFPE tissue sections for molecular analysis; however, 75.7% of these cases yielded adequate cell pellet samples. Adequacy rates for molecular testing of CNB sample were 86.4% in FFPE samples and 92.3% in cell pellet samples. Incorporating cell pellet analysis increased overall molecular testing adequacy to 95.2%.
In this study, we investigated the adequacy rates of molecular testing for small lung cancer biopsy specimens obtained via minimally invasive procedures at our hospital and two other high-volume oncology centers in the same city. We evaluated a total of 3705 NSCLC cases diagnosed through minimally invasive techniques across these three institutions during a six-month period in 2024. Using a conventional threshold for adequacy—defined as cases containing >200 tumor cells per section and a tumor cell fraction > 10%—the overall adequacy rate was 80.6% (2986/3705). Under this adjusted standard of adequacy criteria to >200 tumor cells per section and a tumor cell fraction > 5%. the overall adequacy rate in our cohort increased to 88.9% (3293/3705). Nevertheless, a subset of specimens remains insufficient for molecular analysis, suggesting that there is still significant room for improvement in specimen handling protocols and laboratory workflows.
During the final two months of the study, we applied our previously established single-center optimization strategy for small specimens to those that remained inadequate for molecular testing. Specifically, we collected cell sediment from the residual fixative after tissue sampling to prepare cell smears, which were then evaluated for tumor cell quantity and adequacy. Among the 154 cases collected, 86 met the requirements for molecular testing. By combining paraffin sections with cell sediment analysis, the overall adequacy rate for molecular testing was successfully improved from 81.1% to 91.7%.
Our preliminary observations identified specific nuances in specimen collection techniques within the bronchoscopy and interventional radiology (IR) departments that adversely affect downstream processing in pathology. These issues were prevalent across all three participating hospitals. In most IR departments, sterile puncture kits lack pre-cut filter paper for tissue adhesion. Consequently, clinicians often improvise by using product qualification certificates (from ethylene oxide sterilization) or 3M steam sterilization indicator cards (from autoclaves) to secure CNB specimens.
Gross pathological assessment revealed that these indicator cards soften and lose structural integrity—frequently shedding fibers—once saturated with fixative. This practice complicates the routine collection of small tissue samples for paraffin sectioning and hinders the recovery of residual cell sediments. To address this, our hospital’s IR and pathology departments collaborated to optimize handling by using sterilized, pre-cut PPMM segments. This transition has improved tissue recovery and specimen integrity while minimizing fiber contamination.
Aspirated materials from EBUS-TBNA can be processed as liquid-based cytology, direct smears, or cell blocks (CBs) [
2]. CB preparation is particularly critical, as it allows for superior morphological evaluation, IHC staining, and molecular testing. However, unlike surgical histology, CB preparation lacks a standardized protocol; a 2014 survey identified over ten different methods, each with distinct advantages. This lack of standardization contributes to practical challenges, as evidenced by the 44% of cytopathologists who express dissatisfaction with the quality of CB specimens [
24]. The “tissue coagulum clot” (TCC) and “self-clotting” (SC) techniques are alternatives designed to improve cellular yield over conventional saline rinsing. In the TCC method, aspirated material is expelled onto filter paper [
16], a technique that has demonstrated a diagnostic yield of 88.7% compared to 56.4% for saline rinsing. In the SC method, material is expelled directly into a centrifuge tube to air-dry before formalin fixation [
17]. While filter paper-based TCC improves EBUS-TBNA yields, it can impede downstream pathology workflows. Consequently, our bronchoscopy unit has transitioned to “direct self-clotting” in centrifuge tubes without filter paper. Simultaneously, the pathology department optimized collection using a funnel filtration method to ensure that all tissue clots and dispersed cellular materials are harvested [
18].
Observations at the other two participating hospitals revealed inconsistent collection methods, with some utilizing filter-paper-based TCC and others using the SC technique. During harvesting, technicians typically used tweezers for large fragments or decanting and filtration for smaller sediments. These manual processes are highly susceptible to specimen loss; tweezers often leave residual tumor cells in the vial, while decanting risks the accidental disposal of suspended material. Notably, during the final two months of this study, we confirmed that in cases where initial CB sections had insufficient cellularity, recovering the residual fixative or decanted supernatant often yielded enough tumor cells for molecular testing.
The persistence of these suboptimal practices at Centers B and C—specifically the use of non-validated adherence materials (such as 3M sterilization indicator cards) for CNB specimens and decanting-based workflows for EBUS-TBNA samples—constitutes a widespread, systemic issue in the pre-analytical phase of diagnostics. We strongly advocate for a coordinated effort between the Society of Interventional Radiology (SIR), the College of American Pathologists (CAP), and the International Association for the Study of Lung Cancer (IASLC) to establish and enforce standardized technical protocols. Specifically, clinical societies should issue standards mandating that sterile core needle biopsy (CNB) kits incorporate validated, medical-grade, and fixative-resistant materials—such as the PPMM segments successfully piloted in our study. Integrating these materials into the manufacturing of pre-packaged kits would shift the responsibility for specimen integrity from ad hoc clinical improvisation to a standardized, manufacturer-validated process. This transition is essential to eliminate fiber contamination, preserve tissue architecture during fixation, and ultimately ensure that the limited tissue obtained via CNB is fully optimized for downstream genomic analyses. For EBUS-TBNA and other fine needle aspiration (FNA) procedures, we advocate for the development and adoption of specialized commercial products. These products should be designed to efficiently filter out excess fixative media while ensuring maximal retention of cellular material for diagnostic and molecular studies.
In a previous study, we compared sequencing data from 190 NSCLC patients who underwent both tissue-based next-generation sequencing (tissue-NGS) and plasma-based NGS (plasma-NGS). The overall concordance rate between tissue and plasma samples was 78.9% (150/190). The sensitivity of tissue-NGS and plasma-NGS was 95.0% and 71.9%, respectively. The suboptimal sensitivity of plasma-NGS was attributed to the fact that some tumors do not shed ctDNA into the bloodstream or shed it at levels too low to be detected [
11]. We also validated the concordance of variant detection between FFPE tissue sections and matched cell pellet samples, confirming that somatic mutations identified in FFPE sections were consistently detected in corresponding cell pellets [
14,
15]. Furthermore, we compared the molecular testing performance of plasma and cell pellet samples. In 45 cases with insufficient FFPE tissue, both plasma-based and cell pellet-based NGS were performed. All samples were successfully sequenced, and targetable mutations found in plasma were also identified in matched cell pellets. Notably, in four cases, targetable mutations were detected only in cell pellets—highlighting the superior sensitivity of cell pellets over plasma for molecular testing [
14].
Given that collecting cell pellets is neither time-consuming nor costly, and imposes no additional burden or risk on patients, we recommend the following clinical workflow: in cases with insufficient FFPE tissue, cell pellet samples should be prioritized for sequencing when available. Plasma should be used only if adequate cell pellet material cannot be obtained. Moreover, any negative plasma result should be followed by a repeat tissue biopsy to avoid false negatives and ensure that actionable driver mutations are not missed.
In the clinical workflow at Center A, tumor cellularity in NSCLC small biopsy FFPE sections is routinely evaluated and documented in the diagnostic report. For cases where NSCLC is confirmed, but the FFPE tissue is deemed inadequate for molecular profiling upon pre-analytical review, the residual specimen fixative is subsequently processed to generate both an index smear and a cell pellet. A cytology diagnosis is issued based on the index smear, stating the tumor cellularity and its adequacy for molecular analysis. The matching cell pellet sample that is determined to be adequate then serves as a substitute for sequencing.
This study has several limitations. Factors such as operator experience, number of needle passes, and needle size were not standardized across participating institutions, which may have affected the cellular yield of biopsy specimens. These are major confounding variables that could largely account for the observed inter-center differences. Nevertheless, the primary objective of this study was to identify potential areas for improving molecular adequacy in NSCLC diagnostic samples under such real-world variables, and to evaluate the value of implementing residual fixative-derived cell pellets as a supplementary source for molecular profiling.
While pathology laboratories can control the post-receipt processing timeline—typically ensuring cell pellet harvesting within 48 h—the total formalin fixation time is variable, depending on the time elapsed from biopsy acquisition to laboratory arrival. This pre-analytical variability poses a risk to DNA quality in the resulting cell pellets. Therefore, to maintain nucleic acid integrity and ensure reliable downstream results, clinical workflows should adhere to appropriate fixation times, ideally between 6 and 72 h, for small biopsy specimens intended for molecular testing.
Additionally, only 10 eligible cell pellet cases from Center A were selected for sequencing, based on DNA yield thresholds and clinical demand; therefore, the reproducibility of these DNA yield results and sequencing success rates at Centers B and C was not confirmed. In those centers, suboptimal adherence materials (3M sterilization indicator cards) and decanting-based workflows were still in use. The potential impact of this inter-center variability in pre-analytical methods represents a recognized constraint of the current molecular dataset, underscoring the need for standardized protocol implementation and subsequent multi-center validation.