A Radiomics–Clinical Nomogram for Pre-Treatment Prediction of Neoadjuvant Chemotherapy Response in Locally Advanced Gastric Cancer
Abstract
1. Introduction
2. Materials and Methods
3. Results
3.1. Analysis of Baseline Characteristics
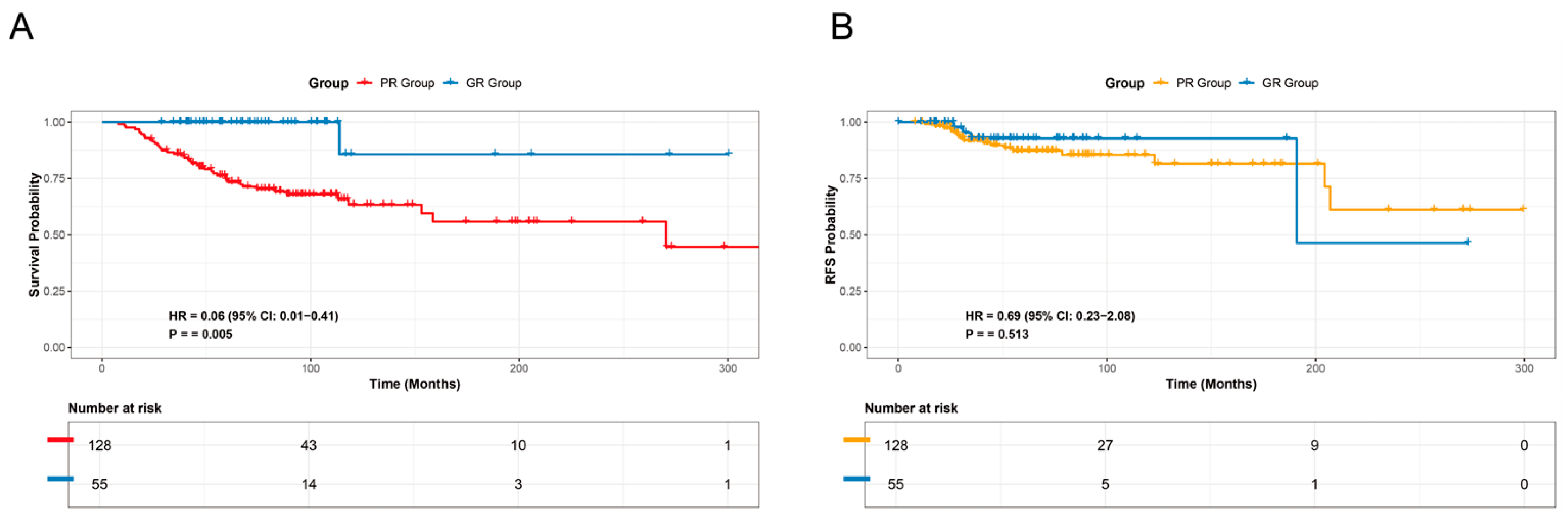
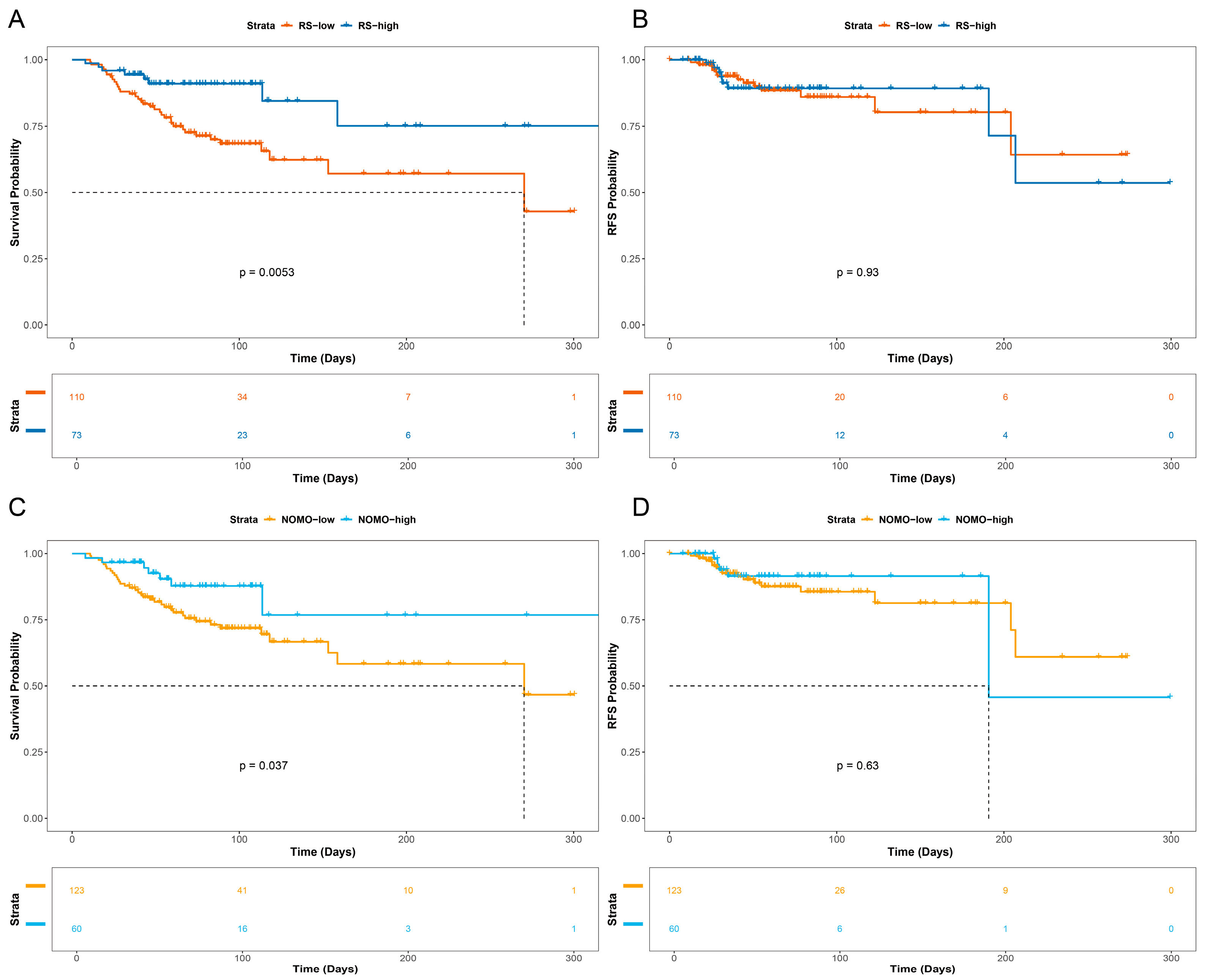
3.2. Survival Analysis of the Patient Cohort
3.3. Radiomics Feature Selection and Model Construction
3.4. Predictive Performance of Machine Learning Models
3.5. Multivariate Analysis Incorporating Clinical Data
3.6. Development of a Nomogram Model and Performance Validation and Comparison
3.7. SHAP Analysis of Radiomics Features
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sundar, R.; Nakayama, I.; Markar, S.R.; Shitara, K.; van Laarhoven, H.W.; Janjigian, Y.Y.; Smyth, E.C. Gastric cancer. Lancet 2025, 405, 2087–2102. [Google Scholar] [CrossRef] [PubMed]
- Zurlo, I.V.; Basso, M.; Strippoli, A.; Calegari, M.A.; Orlandi, A.; Cassano, A.; Di Salvatore, M.; Garufi, G.; Bria, E.; Tortora, G.; et al. Treatment of locally advanced gastric cancer (LAGC): Back to lauren’s classification in pan–cancer analysis era? Cancers 2020, 12, 1749. [Google Scholar] [CrossRef]
- Tsagkalidis, V.; Blaszczyk, M.B.; In, H. Interpretation of tumor response grade following preoperative therapy for gastric cancer: An overview. Cancers 2023, 15, 3662. [Google Scholar] [CrossRef]
- Koerner, A.S.; Moy, R.H.; Ryeom, S.W.; Yoon, S.S. The present and future of neoadjuvant and adjuvant therapy for locally advanced gastric cancer. Cancers 2023, 15, 4114. [Google Scholar] [CrossRef]
- Wang, X.Z.; Zeng, Z.Y.; Ye, X.; Sun, J.; Zhang, Z.M.; Kang, W.M. Interpretation of the development of neoadjuvant therapy for gastric cancer based on the vicissitudes of the NCCN guidelines. World J. Gastrointest. Oncol. 2020, 12, 37–53. [Google Scholar] [CrossRef]
- Oikonomopoulou, K.; Li, L.; Zheng, Y.; Simon, I.; Wolfert, R.L.; Valik, D.; Nekulova, M.; Simickova, M.; Frgala, T.; Diamandis, E.P. Prediction of ovarian cancer prognosis and response to chemotherapy by a serum-based multiparametric biomarker panel. Br. J. Cancer 2008, 99, 1103–1113. [Google Scholar] [CrossRef]
- Yin, H.; Sun, L.; Yuan, Y.; Zhu, Y. PPIC-labeled CAFs: Key players in neoadjuvant chemotherapy resistance for gastric cancer. Transl. Oncol. 2024, 48, 102080. [Google Scholar] [CrossRef]
- Abboretti, F.; Lambert, C.; Schäfer, M.; Pereira, B.; Le Roy, B.; Mège, D.; AFC-French Association of Surgery. Neoadjuvant chemotherapy does not improve survival in cT2N0M0 gastric adenocarcinoma patients: A multicenter propensity score analysis. Ann. Surg. Oncol. 2024, 31, 5273–5282. [Google Scholar] [CrossRef] [PubMed]
- Fan, L.; Li, J.; Zhang, H.; Yin, H.; Zhang, R.; Zhang, J.; Chen, X. Machine learning analysis for the noninvasive prediction of lymphovascular invasion in gastric cancer using PET/CT and enhanced CT-based radiomics and clinical variables. Abdom. Radiol. 2022, 47, 1209–1222. [Google Scholar] [CrossRef]
- Ge, H.T.; Chen, J.W.; Wang, L.L.; Zou, T.X.; Zheng, B.; Liu, Y.F.; Lin, W.W. Preoperative prediction of lymphovascular and perineural invasion in gastric cancer using spectral computed tomography imaging and machine learning. World J. Gastroenterol. 2024, 30, 542–555. [Google Scholar] [CrossRef] [PubMed]
- Van Griethuysen, J.J.; Lambregts, D.M.; Trebeschi, S.; Lahaye, M.J.; Bakers, F.C.; Vliegen, R.F.; Beets-Tan, R.G. Radiomics performs comparable to morphologic assessment by expert radiologists for prediction of response to neoadjuvant chemoradiotherapy on baseline staging MRI in rectal cancer. Abdom. Radiol. 2020, 45, 632–643. [Google Scholar] [CrossRef]
- Tong, Y.; Zhu, Y.; Zhao, Y.; Shan, Z.; Liu, D.; Zhang, J. Evaluation and comparison of predictive value of tumor regression grades according to mandard and Becker in locally advanced gastric adenocarcinoma. Cancer Res. Treat. 2021, 53, 112–122. [Google Scholar] [CrossRef]
- Hoshi, H. Management of gastric adenocarcinoma for general surgeons. Surg. Clin. 2020, 100, 523–534. [Google Scholar] [CrossRef] [PubMed]
- Biondi, A.; Lirosi, M.C.; D’uGo, D.; Fico, V.; Ricci, R.; Santullo, F.; Rizzuto, A.; Cananzi, F.C.; Persiani, R. Neo-adjuvant chemo(radio)therapy in gastric cancer: Current status and future perspectives. World J. Gastrointest. Oncol. 2015, 7, 389–400. [Google Scholar] [CrossRef] [PubMed]
- Su, P.F.; Yu, J.C. Progress in neoadjuvant therapy for gastric cancer. Oncol. Lett. 2022, 23, 172. [Google Scholar] [CrossRef]
- Sah, B.R.; Owczarczyk, K.; Siddique, M. Radiomics in esophageal and gastric cancer. Abdom. Radiol. 2019, 44, 2048–2058. [Google Scholar] [CrossRef]
- Li, Q.; Feng, Q.X.; Qi, L.; Liu, C.; Zhang, J.; Yang, G.; Liu, X.S. Prognostic aspects of lymphovascular invasion in localized gastric cancer: New insights into the radiomics and deep transfer learning from contrast-enhanced CT imaging. Abdom. Radiol. 2022, 47, 496–507. [Google Scholar] [CrossRef]
- Huang, J.; Chen, Y.; Zhang, Y.; Xie, J.; Liang, Y.; Yuan, W.; Zhou, T.; Gao, R.; Wen, R.; Xia, Y.; et al. Comparison of clinical-computed tomography model with 2D and 3D radiomics models to predict occult peritoneal metastases in advanced gastric cancer. Abdom. Radiol. 2021, 47, 66–75. [Google Scholar] [CrossRef]
- Wang, J.; Liu, X.; Hu, B.; Gao, Y.; Chen, J.; Li, J. Development and validation of an MRI-based radiomic nomogram to distinguish between good and poor responders in patients with locally advanced rectal cancer undergoing neoadjuvant chemoradiotherapy. Abdom. Radiol. 2021, 46, 1805–1815. [Google Scholar] [CrossRef]
- Zhang, J.; Zhang, Q.; Zhao, B.; Shi, G. Deep learning nomogram for predicting neoadjuvant chemotherapy response in locally advanced gastric cancer patients. Abdom. Radiol. 2024, 49, 3780–3796. [Google Scholar] [CrossRef] [PubMed]
- Wen, L.; Liu, J.; Hu, P.; Bi, F.; Liu, S.; Jian, L.; Zhu, S.; Nie, S.; Cao, F.; Lu, Q.; et al. MRI-based radiomic models outperform radiologists in predicting pathological complete response to neoadjuvant chemoradiotherapy in locally advanced rectal cancer. Acad. Radiol. 2023, 30, S176–S184. [Google Scholar] [CrossRef]
- Zhang, P.-F.; Zhang, W.-H.; Liu, X.-J.; He, D.; Yang, K.; Gou, H.-F.; Hu, J.-K. Chemotherapy combined with cadonilimab (AK104) as neoadjuvant treatment for locally advanced gastric/gastro-oesophageal junction adenocarcinoma: Study protocol for a single-arm, phase II clinical trial. BMJ Open 2024, 14, e081529. [Google Scholar] [CrossRef]
- Cozac-Szőke, A.-R.; Tinca, A.C.; Negovan, A.; Vilaia, A.; Cozac, D.-A.; Cocuz, I.-G.; Sabău, A.H.; Hagău, R.-D.; Chiorean, D.-M.; Lazar, A.-B.; et al. Comprehensive Analysis of SIGLEC-15 and PD-L1 Expression Identifies Distinct Prognostic Profiles in Gastric Cancer. Int. J. Mol. Sci. 2025, 26, 8637. [Google Scholar] [CrossRef] [PubMed]
- Deng, J.; Zhang, W.; Xu, M.; Zhou, J. Imaging advances in efficacy assessment of gastric cancer neoadjuvant chemotherapy. Abdom. Radiol. 2023, 48, 3661–3676. [Google Scholar] [CrossRef] [PubMed]
- Zhong, H.; Wang, T.; Hou, M.; Liu, X.; Tian, Y.; Cao, S.; Zhou, Y. Deep learning radiomics nomogram based on enhanced CT to predict the response of metastatic lymph nodes to neoadjuvant chemotherapy in locally advanced gastric cancer. Ann. Surg. Oncol. 2024, 31, 421–432. [Google Scholar] [CrossRef] [PubMed]




| GR Group | PR Group | p | |
|---|---|---|---|
| Variable | |||
| (n = 55) | (n = 128) | ||
| Sex [n (%)] | |||
| Female | 12 (21.8) | 29 (22.7) | 0.000 |
| Male | 43 (78.2) | 99 (77.3) | |
| Age [n (%)] | |||
| >60 years | 25 (45.5) | 75 (58.6) | 0.140 |
| ≤60 years | 30 (54.5) | 53 (41.4) | |
| BMI [kg/m2] | |||
| Mean ± sd | 23.3 ± 2.90 | 23.0 ± 2.69 | 0.437 |
| M (P25, P75) | 22.9 (17.6, 31.4) | 22.7 (17.6, 33.1) | |
| Treatment regimen [n (%)] | |||
| Immunotherapy | 17 (30.9) | 21 (16.4) | 0.082 |
| other | 13 (23.6) | 28 (21.9) | |
| SOX | 22 (40.0) | 62 (48.4) | |
| XELOX | 3 (5.5) | 17 (13.3) | |
| cTstage [n (%)] | |||
| 1 | 0 | 2 (1.6) | 0.014 |
| 2 | 0 | 4 (3.1) | |
| 3 | 15 (27.3) | 36 (28.1) | |
| 4a | 33 (60.0) | 84 (65.6) | |
| 4b | 7 (12.7) | 2 (1.6) | |
| cNstage [n (%)] | |||
| 0 | 0 | 3 (2.3) | 0.056 |
| 1 | 1 (1.8) | 15 (11.7) | |
| 2 | 14 (25.5) | 38 (29.7) | |
| 3 | 40 (72.7) | 72 (56.3) | |
| WBC [n×109/L] | |||
| Mean ± sd | 6.54 ± 2.26 | 6.22 ± 1.89 | 0.354 |
| M (P25, P75) | 5.89 (2.56, 13.8) | 5.83 (2.57, 14.0) | |
| RBC [n×1012/L] | |||
| Mean ± sd | 4.07± 0.801 | 4.28 ± 0.589 | 0.078 |
| M (P25, P75) | 4.03 (2.29, 5.73) | 4.32 (2.77, 6.01) | |
| PLT [n×109/L] | |||
| Mean ± sd | 239 ± 93.2 | 223 ± 70.8 | 0.262 |
| M (P25, P75) | 223 (107, 500) | 220 (61.0, 440) | |
| LYM [n×109/L] | |||
| Mean ± sd | 1.62 ± 0.565 | 1.63 ± 0.646 | 0.863 |
| M (P25, P75) | 1.54 (0.320, 3.49) | 1.54 (0.570, 4.70) | |
| NEU [n×109/L] | |||
| Mean ± sd | 4.21 ± 1.96 | 4.55 ± 6.05 | 0.559 |
| M (P25, P75) | 3.72 (1.35, 12.1) | 3.67 (1.56, 69.6) | |
| HB [g/L] | |||
| Mean ± sd | 118 ± 26.8 | 124 ± 22.6 | 0.114 |
| M (P25, P75) | 120 (58.0, 170) | 129 (51.0, 175) | |
| Ca [mmol/L] | |||
| Mean ± sd | 2.23 ± 0.110 | 2.23 ± 0.152 | 0.804 |
| M (P25, P75) | 2.21 (2.02, 2.47) | 2.24(1.07, 2.50) | |
| Mg [mmol/L] | |||
| Mean ± sd | 0.863 ± 0.0652) | 0.890 ± 0.141 | 0.071 |
| M (P25, P75) | 0.860 (0.670, 0.990) | 0.880 (0.690, 2.28) | |
| CEA [n (%)] | |||
| >5 ng/mL | 17 (30.9) | 36 (28.1) | 0.839 |
| ≤5 ng/mL | 38 (69.1) | 92 (71.9) | |
| AFP [n (%)] | |||
| >7 ng/mL | 9 (16.4) | 15 (11.7) | 0.539 |
| ≤7 ng/mL | 46 (83.6) | 113 (88.3) | |
| CA199 [n (%)] | |||
| >30 U/mL | 17 (30.9) | 27 (21.1) | 0.216 |
| ≤30 U/mL | 38 (69.1) | 101 (78.9) |
| Variable | Training Set | Validation Set | p |
|---|---|---|---|
| (n = 128) | (n = 55) | ||
| Sex [n (%)] | |||
| Female | 33 (25.8) | 8 (14.5) | 0.139 |
| Male | 95 (74.2) | 47 (85.5) | |
| AGE [n (%)] | |||
| >60 years | 69 (53.9) | 31 (56.4) | 0.885 |
| ≤60 years | 59 (46.1) | 24 (43.6) | |
| BMI [kg/m2] | |||
| Mean ± sd | 23.0 ± 2.91 | 23.2 ± 2.33 | 0.761 |
| M (P25, P75) | 22.7 (17.6, 33.1) | 22.8 (17.6, 30.7) | |
| Missing | 10 (7.8) | 7 (12.7) | |
| Treatment Regimen [n (%)] | |||
| Immunotherapy | 31 (24.2) | 7 (12.7) | 0.315 |
| other | 26 (20.3) | 15 (27.3) | |
| SOX | 58 (45.3) | 26 (47.3) | |
| XELOX | 13 (10.2) | 7 (12.7) | |
| cTstage [n (%)] | |||
| 1 | 2 (1.6) | 0 | 0.364 |
| 2 | 4 (3.1) | 0 | |
| 3 | 33 (25.8) | 18 (32.7) | |
| 4a | 84 (65.6) | 33 (60.0) | |
| 4b | 5 (3.9) | 4 (7.3) | |
| cNstage [n (%)] | |||
| 0 | 3 (2.3) | 0 | 0.249 |
| 1 | 14 (10.9) | 2 (3.6) | |
| 2 | 36 (28.1) | 16 (29.1) | |
| 3 | 75 (58.6) | 37 (67.3) | |
| WBC [n×109/L] | |||
| Mean ± sd | 6.17 ± 1.88 | 6.63 ± 2.26 | 0.190 |
| M (P25, P75) | 5.71 (2.56, 14.0) | 6.16 (2.57, 13.8) | |
| RBC [n×1012/L] | |||
| Mean ± sd | 4.19 ± 0.665) | 4.29 ± 0.665 | 0.320 |
| M (P25, P75) | 4.19(2.48, 6.01) | 4.37 (2.29, 5.73) | |
| PLT [n×109/L] | |||
| Mean ± sd | 228 ± 76.2 | 226 ± 83.5 | 0.879 |
| M (P25, P75) | 221 (105, 473) | 220 (61.0, 500) | |
| LYM [n×109/L] | |||
| Mean ± sd | 1.61 (0.635) | 1.66 (0.593) | 0.585 |
| M (P25, P75) | 1.53 (0.320, 4.70) | 1.59(0.670, 4.10) | |
| NEU [n×109/L] | |||
| Mean ± sd | 4.51 ± 6.05 | 4.31 ± 1.98 | 0.743 |
| M (P25, P75) | 3.65 (1.35, 69.6) | 3.74 (1.56, 12.1) | |
| HB [g/L] | |||
| Mean ± sd | 122 ± 25.2 | 124 ± 21.5 | 0.477 |
| M (P25, P75) | 124 (51.0, 175) | 128(67.0, 170) | |
| Ca [mmol/L] | |||
| Mean ± sd | 2.23 ± 0.152 | 2.23 ± 0.111 | 0.759 |
| M (P25, P75) | 2.24 (1.07, 2.50) | 2.22 (2.03, 2.48) | |
| Mg [mmol/L] | |||
| Mean ± sd | 0.887 ± 0.141 | 0.871 ± 0.0683 | 0.295 |
| M (P25, P75) | 0.870 (0.670, 2.28) | 0.870(0.720, 0.990) | |
| CEA [n (%)] | |||
| >5 ng/mL | 35 (27.3) | 18 (32.7) | 0.577 |
| ≤5 ng/mL | 93 (72.7) | 37 (67.3) | |
| AFP [n (%)] | |||
| >7 ng/mL | 16 (12.5) | 8 (14.5) | 0.891 |
| ≤7 ng/mL | 112 (87.5) | 47 (85.5) | |
| CA199 [n(%)] | |||
| >30 U/mL | 32 (25.0) | 12 (21.8) | 0.785 |
| ≤30 U/mL | 96 (75.0) | 43 (78.2) | |
| Response [n (%)] | |||
| GR | 37 (28.9) | 18 (32.7) | 0.733 |
| PR | 91 (71.1) | 37 (67.3) |
| Model | Random Forest | SVM | Tree | Logistic Regression |
|---|---|---|---|---|
| Random forest | NA | 0.004 | 0.001 | 0.008 |
| SVM | 0.004 | NA | 0.515 | 0.328 |
| Decision tree | 0.001 | 0.515 | NA | 0.467 |
| Logistic regression | 0.008 | 0.328 | 0.467 | NA |
| Variable | Univariate Analysis | Multivariate Analysis | |||||
|---|---|---|---|---|---|---|---|
| OR | 95.0% CI | p | OR | 95.0% CI | p | ||
| Sex | Male | 1.050 | 0.490~2.249 | 0.901 | |||
| Female | |||||||
| Age | >60 years | 0.589 | 0.312~1.113 | 0.103 | |||
| ≤60 years | |||||||
| Treatment regimen | SOX | 0.940 | 0.041 | ||||
| XELOX | 0.497 | 0.133~1.862 | 0.300 | 0.508 | 0.199~1.297 | 0.157 | |
| Immu | 2.281 | 1.022~5.095 | 0.044 | 0.213 | 0.047~0.971 | 0.046 | |
| other | 1.308 | 0.577~2.965 | 0.520 | 1.409 | 0.494~4.016 | 0.521 | |
| BMI | 1.051 | 0.931~1.186 | 0.420 | ||||
| CEA | 0.997 | 0.990~1.005 | 0.461 | ||||
| AFP | 1.000 | 0.998~1.002 | 0.861 | ||||
| CA199 | 1.000 | 0.998~1.002 | 0.674 | ||||
| Radiomic score | 5.253 | 2.655~10.394 | 0.000 | 7.448 | 3.466~16.004 | 0.000 | |
| cT stage | 1 | 1.466 | 0.803~2.676 | 0.213 | |||
| 2 | |||||||
| 3 | |||||||
| 4 | |||||||
| cN stage | 0 | 2.053 | 1.189~3.543 | 0.010 | 2.282 | 1.228~4.242 | 0.009 |
| 1 | |||||||
| 2 | |||||||
| 3 | |||||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Zhou, Q.; Xu, J.; Li, Q.; Li, F.; Xu, H. A Radiomics–Clinical Nomogram for Pre-Treatment Prediction of Neoadjuvant Chemotherapy Response in Locally Advanced Gastric Cancer. Diagnostics 2026, 16, 945. https://doi.org/10.3390/diagnostics16060945
Zhou Q, Xu J, Li Q, Li F, Xu H. A Radiomics–Clinical Nomogram for Pre-Treatment Prediction of Neoadjuvant Chemotherapy Response in Locally Advanced Gastric Cancer. Diagnostics. 2026; 16(6):945. https://doi.org/10.3390/diagnostics16060945
Chicago/Turabian StyleZhou, Qianzheng, Jun Xu, Qiong Li, Fengyuan Li, and Hao Xu. 2026. "A Radiomics–Clinical Nomogram for Pre-Treatment Prediction of Neoadjuvant Chemotherapy Response in Locally Advanced Gastric Cancer" Diagnostics 16, no. 6: 945. https://doi.org/10.3390/diagnostics16060945
APA StyleZhou, Q., Xu, J., Li, Q., Li, F., & Xu, H. (2026). A Radiomics–Clinical Nomogram for Pre-Treatment Prediction of Neoadjuvant Chemotherapy Response in Locally Advanced Gastric Cancer. Diagnostics, 16(6), 945. https://doi.org/10.3390/diagnostics16060945




