Prioritising Data Quality Governance for AI in Prostate Cancer: A Methodological Proof-of-Concept Study Using Neural Networks for Risk Stratification
Abstract
1. Introduction
Aims of This Study
- -
- Data quality governance: to operationalise a FAIR-based “AI-readiness” protocol and test whether prioritising data quality over sample size yields a stable diagnostic signal in a small cohort.
- -
- Nomogram integration: to quantify the predictive weight gained by combining the Briganti nomogram and ISUP biopsy grade within an MLP and to compare it against traditional clinical staging.
- -
- Model stability: to conduct a sensitivity analysis across three data-partitioning schemes (20/80, 34/66, 39/61) and identify the configuration that minimises cross-entropy error.
2. Materials and Methods
2.1. Study Population and Data Collection
2.1.1. Inclusion Criteria
- Histologically confirmed diagnosis of PCa.
- Complete clinical and biochemical records required for D’Amico risk stratification [63], including PSA at diagnosis, ISUP biopsy grade, and clinical TNM staging.
- Comprehensive mpMRI findings (mrT and mrN).
2.1.2. Exclusion Criteria
- (i)
- Pre-modelling exclusion—data quality. Any patient with a missing value in any of the 11 candidate predictors was removed from the initial cohort (listwise deletion).
- (ii)
- Validation-time exclusion—software-driven. During each run of the SPSS MLP, any case in the test or hold-out sample whose categorical factor levels (e.g., a specific mrT stage or a very high PSA value) were not represented in the corresponding training sample was automatically dropped by the software, because the network cannot assign weights to factor levels it has never seen. This is a mathematical constraint of the SPSS implementation, not an investigator-driven decision.
| Quality Dimension | Metric/Target | Operational Validation Rule (Concrete Check) |
|---|---|---|
| Accuracy | 100% clinical concordance | Cross-verification of PSA values and ISUP grades between the electronic health record (EHR) and the study database. |
| Completeness | 0% missingness in predictors | Exclusion of any case with missing values in the 11 primary clinical variables (listwise deletion). |
| Validity (Range) | Biological boundary checks | PSA: (0.1 to 500 ng/mL); prostate volume: (10 to 300 cc); age: (40 to 90 years). |
| Consistency | Logical relationship | Staging consistency check: Clinical stage (cT) must not exceed pathological or imaging (mrT) findings in illogical sequences. |
| Integrity | Referential integrity | All categorical factors must map to the D’Amico classification standards (ISUP 1–5). |
| AI-Readiness | Feature scaling | Continuous variables must be normalised to a standard numerical range to prevent gradient saturation. |
Methodological Note and Limitation
2.1.3. Data Quality and Integrity
- -
- Intrinsic Accuracy and Completeness: Clinical records were verified to make sure data were accurate, reliable, and free from errors.
- -
- Consistency and Mathematical Validity: Data were represented uniformly across sources, with no residual contradictions. For the rationale, operational details and sample-size consequences of the validation-time exclusion of test cases carrying unseen factor levels, see Section 2.1.2 and Section Methodological Note and Limitation; the same considerations apply here and are not repeated.
- -
- Alignment with FAIR Principles: This study adhered to the FAIR principles in order to improve transparency and accountability. Comprehensive documentation of exclusion criteria was maintained to ensure the traceability of the datasets and the credibility of the scientific conclusions.
2.1.4. Sample Size and Statistical Power
2.2. Artificial Neural Network (ANN) Configuration
2.2.1. Network Architecture
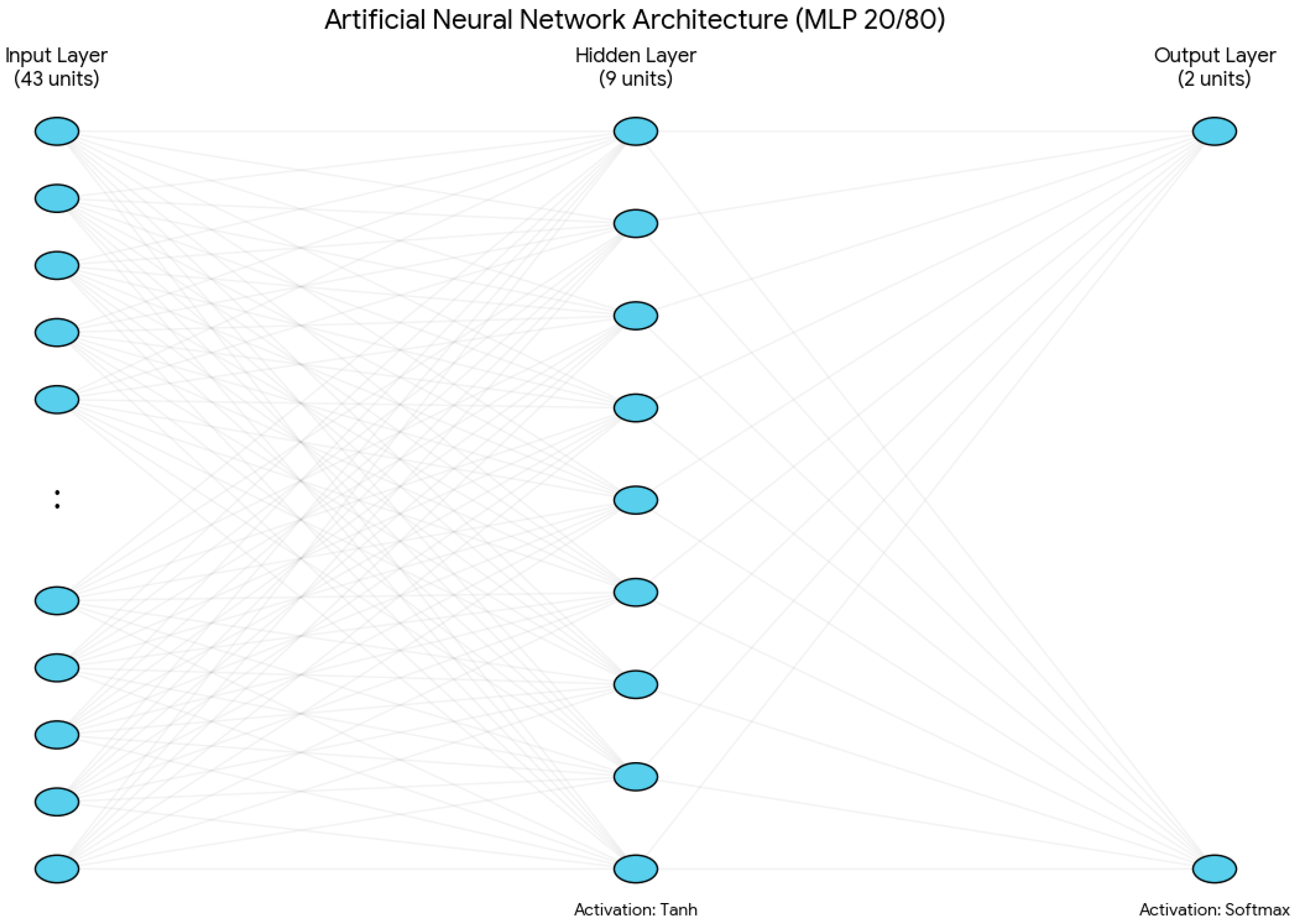
- Input Layer: The network architecture was built on 11 main clinical variables, and it has 43 input units in total for the 20/80 model. This expansion was performed automatically by the IBM SPSS MLP procedure, using 1-of-c encoding for the categorical factors. There were seven categorical variables included in this process: PSA at diagnosis, ISUP biopsy grade, biopsy laterality, clinical TNM stage, clinical nodal stage, mrT, and mrN. Under the 1-of-c scheme, these factors were transformed into 39 separate input units (one for each category level). Combined with the four units for continuous covariates, this results in a total of 43 input units.For continuous covariates, there are four variables: age, PSA density, prostate volume, and Briganti score. These continuous variables undergo a rescaling procedure by linear normalisation, which adjusts them to a standardised numerical range defined by the minimum and maximum values found in the training set. This pre-processing step is crucial, as it facilitates training convergence and prevents variables with larger numerical ranges, such as prostate volume, from disproportionately affecting the network’s weight estimations or causing “weight saturation” in the activation functions.
- Hidden Layer: A single hidden layer was used, with the number of neurons determined via the IBM SPSS MLP automatic architecture selection algorithm. This procedure optimised the size of the hidden layer within a predefined range from 6 to 9 by selecting the configuration that minimised the training cross-entropy error. This architectural constraint acts as a type of structural regularisation, which creates an ‘information bottleneck’ that prevents the network from memorising the training set. By restricting the capacity of the hidden layer and combining these limits, the model is forced to prioritise the most influential predictors, such as the ISUP grade and Briganti score, over less significant categorical levels. Furthermore, to prevent ‘over-training,’ the model applied an early stopping rule that ended the iteration process at the first sign of error plateauing, where the cross-entropy error did not decrease anymore.For the 20/80 model, this led to a total of 9 hidden units. The hyperbolic tangent (tanh) activation function was applied to this layer to facilitate non-linear mapping:
- Output Layer: The target variable was the D’Amico risk group. Although the model was initialised to support three categories (high, intermediate, and low), the output neurons were dynamically reduced to two (high vs. intermediate) in the 20/80 configuration. This occurred because the low-risk group did not have enough representation in the training partition for that specific split, which did not allow for robust category initialisation (Figure 2).
2.2.2. Sensitivity Analysis and Validation
2.2.3. Model Robustness as an Extension of Data Quality Governance
- Reproducible Initialisation and Training: The framework requires the use of a fixed random seed (2,000,000) for all stochastic processes, including initial weight assignment and case selection for data partitions, to guarantee that all training runs can be precisely duplicated by independent researchers. This approach creates a repeatable baseline that can be used to compare subsequent experiments, even though it does not fully capture the range of the model’s stochastic behaviour.
- Sensitivity Analysis as a Governance Mandate: The framework requires a sensitivity analysis across several partitioning schemes rather than depending on a single data split, which could yield results that are artefacts of a particularly favourable or unfavourable partition. Three different splits (20/80, 34/66, and 39/61) were assessed for this investigation. In order to determine whether the observed performance is a feature of the architecture’s interaction with high-quality data or just a reflection of a single, lucky patient grouping, this method examines the stability of the network’s learning across various cohort compositions and sizes.
- Reporting Distributions Rather Than Point Estimates: The approach requires that findings be reported as distributions (mean ± SD) across the sensitivity analysis instead of single peak-performance percentages in order to account for the algorithmic variability inherent in small-sample machine learning. This approach gives readers a more accurate and comprehensive understanding of model stability.
- Early Stopping to Prioritise Generalisation: An aggressive early stopping rule, which stops training after one consecutive step without reducing cross-entropy error, is specified by the framework. Given the limited cohort size, it is crucial to prioritise generalisation over training-set accuracy, which is why this criterion was selected. Since this would have further lowered the already small training sample (from N = 34 to an even smaller size) and impeded the model’s ability to identify stable decision boundaries, a separate validation set for early stopping was not used.
- Accounting for Exclusion Bias: Additionally, the DQG framework requires open reporting of how the final evaluable sample is impacted by its own rules. The valid sample size varied slightly between configurations (n = 41 to n = 44) because cases with factor levels not present in a particular training split were automatically excluded to maintain mathematical validity within the SPSS MLP framework. This must be stated clearly in the framework: the analysis should be seen as a test of the architecture’s capacity to generalise from a “standardised” clinical signal rather than a straightforward comparison across identical patient subsets.
- Uncertainty Quantification: The framework requires statistical testing against a null hypothesis of a random classifier (p = 0.50) in order to determine whether classification results might be attributed to random chance. Using IBM SPSS v26, a One-Sample Binomial Test was performed for this investigation. Confidence intervals were computed using the Clopper–Pearson exact method, which is the most rigorous and conservative method for small-sample validation, especially for the N = 9 independent testing set.
- Benchmarking Against Traditional Methods: Lastly, the framework requires that the “AI-premium”, the neural network’s superior performance above conventional statistical methods, be quantified. Exact logistic regression in LogXact-11 [72], the statistical gold standard for small-sample datasets where standard maximum likelihood estimation may be incorrect, was used for this study’s baseline comparison. To enable a direct comparison with the MLP architecture, the baseline model employed the same 11 clinical predictors and the 20/80 data partitioning strategy.
3. Results
3.1. Cohort Curation and Model Performance as a Function of Data Quality
3.2. Discriminatory Capacity: Characterising the Framework’s Extracted Signal
3.3. Optimal Model Classification: A Detailed View of the Framework’s Signal
3.4. Independent Variable Importance: The Clinical Signal Preserved by the Framework
3.5. Comparative Benchmark: Evidence for Non-Linear Signal Preservation
4. Discussion
4.1. The Fragility of Perfect Metrics in a Small-Sample Context
4.2. Evidence for an “AI-Premium”: Signal Detection or Overfitting?
4.3. Clinical Significance of Predictive Variables
4.4. The Potential Added Value of the Artificial Neural Network over Existing Nomograms
Positioning of the Proposed MLP Within the Existing Risk-Stratification Landscape
4.5. Data Quality as a Strategic Imperative
Pre-Specified Sample-Size Justification and Three-Phase Multi-Institutional Validation Plan
4.6. Limitations and Future Directions
4.7. Clinical Translation: A Cautionary Framework, Not a Deployable Tool
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ANN | Artificial Neural Network |
| AUC | Area Under the Curve |
| cT | Clinical tumour stage |
| DQG | Data Quality Governance |
| DQG-AI | Data Quality Governance for AI-readiness framework |
| EHR | Electronic Health Record |
| FAIR | Findability, Accessibility, Interoperability, and Reusability |
| ISUP | International Society of Urological Pathology |
| mrN | Multiparametric imaging nodal stage (or Imaging nodal stage) |
| mrT | Multiparametric imaging tumour stage (or Imaging tumour stage) |
| MLP | Multilayer Perceptron |
| mpMRI | Multiparametric Magnetic Resonance Imaging |
| NPV | Negative Predictive Value |
| PCa | Prostate Cancer |
| PMML | Predictive Model Markup Language |
| PPV | Positive Predictive Value |
| PSA | Prostate-Specific Antigen |
| ROC | Receiver Operating Characteristic |
| SD | Standard Deviation |
| SHAPs | SHapley Additive exPlanations |
| SOP | Standard Operating Procedure |
| tanh | Hyperbolic Tangent |
| XML | Extensible Markup Language |
Appendix A. AI-Readiness and Data Quality Governance Checklist—DQG-AI Framework v1.0 (Clínica Universidad de Navarra)
- 1.
- Data Integrity and Source Governance
- 2.
- Operational Data Quality Rules (The “Technical Gatekeeper”)
- 3.
- Pre-Modelling “AI-Readiness” Transitions
- 4.
- Evaluation and Stability Governance
References
- Schafer, E.J.; Laversanne, M.; Sung, H.; Soerjomataram, I.; Briganti, A.; Dahut, W.; Bray, F.; Jemal, A. Recent Patterns and Trends in Global Prostate Cancer Incidence and Mortality: An Update. Eur. Urol. 2025, 87, 302–313. [Google Scholar] [CrossRef]
- Wang, L.; Lu, B.; He, M.; Wang, Y.; Wang, Z.; Du, L. Prostate Cancer Incidence and Mortality: Global Status and Temporal Trends in 89 Countries from 2000 to 2019. Front. Public Health 2022, 10, 811044. [Google Scholar] [CrossRef]
- Chu, F.; Chen, L.; Guan, Q.; Chen, Z.; Ji, Q.; Ma, Y.; Ji, J.; Sun, M.; Huang, T.; Song, H.; et al. Global Burden of Prostate Cancer: Age-Period-Cohort Analysis from 1990 to 2021 and Projections until 2040. World J. Surg. Oncol. 2025, 23, 98. [Google Scholar] [CrossRef] [PubMed]
- Kratzer, T.B.; Mazzitelli, N.; Star, J.; Dahut, W.L.; Jemal, A.; Siegel, R.L. Prostate Cancer Statistics, 2025. CA Cancer J. Clin. 2025, 75, 485–497. [Google Scholar] [CrossRef]
- Hassanipour-Azgomi, S.; Mohammadian-Hafshejani, A.; Ghoncheh, M.; Towhidi, F.; Jamehshorani, S.; Salehiniya, H. Incidence and Mortality of Prostate Cancer and Their Relationship with the Human Development Index Worldwide. Prostate Int. 2016, 4, 118–124. [Google Scholar] [CrossRef]
- Dudipala, H.; Jani, C.T.; Gurnani, S.D.; Morgenstern-Kaplan, D.; Tran, E.; Edwards, K.; Shalhoub, J.; McKay, R.R. Evolving Global Trends in Prostate Cancer: Disparities Across Income Levels and Geographic Regions (1990–2019). JCO Glob. Oncol. 2025, 11, e2500249. [Google Scholar] [CrossRef]
- Bray, F.; Laversanne, M.; Sung, H.; Ferlay, J.; Siegel, R.L.; Soerjomataram, I.; Jemal, A. Global Cancer Statistics 2022: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2024, 74, 229–263. [Google Scholar] [CrossRef]
- Cornford, P.; Tilki, D.; van den Berghn, R.; Eberlin, D. Prostate Cancer. In EAU Guidelines; EAU Guidelines Office: Arnhem, The Netherlands, 2025. [Google Scholar]
- Olah, C.; Mairinger, F.; Wessolly, M.; Joniau, S.; Spahn, M.; Kruithof-de Julio, M.; Hadaschik, B.; Soós, A.; Nyirády, P.; Győrffy, B.; et al. Enhancing Risk Stratification Models in Localized Prostate Cancer by Novel Validated Tissue Biomarkers. Prostate Cancer Prostatic Dis. 2025, 28, 773–781. [Google Scholar] [CrossRef]
- D’Amico, A.V.; Whittington, R.; Malkowicz, S.; Schulz, D.; Blank, K.; Broderick, G.; Tomaszewski, J.; Renshaw, A.; Kaplan, I.; Beard, C.; et al. D’Amico Risk Classification for Prostate Cancer (Version: 1.28)—Evidencio. Available online: https://www.evidencio.com/models/show/300 (accessed on 10 February 2026).
- Hernandez, D.J.; Nielsen, M.E.; Han, M.; Partin, A.W. Contemporary Evaluation of the D’Amico Risk Classification of Prostate Cancer. Urology 2007, 70, 931–935. [Google Scholar] [CrossRef] [PubMed]
- Cooperberg, M.R. Clinical Risk Stratification for Prostate Cancer: Where Are We, and Where Do We Need to Go? Can. Urol. Assoc. J. 2017, 11, 101. [Google Scholar] [CrossRef] [PubMed]
- Rastinehad, A.R.; Baccala, A.A.; Chung, P.H.; Proano, J.M.; Kruecker, J.; Xu, S.; Locklin, J.K.; Turkbey, B.; Shih, J.; Bratslavsky, G.; et al. D’Amico Risk Stratification Correlates with Degree of Suspicion of Prostate Cancer on Multiparametric Magnetic Resonance Imaging. J. Urol. 2011, 185, 815–820. [Google Scholar] [CrossRef]
- Chierigo, F.; Flammia, R.S.; Sorce, G.; Hoeh, B.; Hohenhorst, L.; Tian, Z.; Saad, F.; Gallucci, M.; Briganti, A.; Montorsi, F.; et al. The Association of the Type and Number of D’Amico High-Risk Criteria with Rates of Pathologically Non-Organ-Confined Prostate Cancer. Cent. Eur. J. Urol. 2023, 76, 104–108. [Google Scholar] [CrossRef]
- Huang, J.; Ssentongo, P.; Sharma, R. Editorial: Cancer Burden, Prevention and Treatment in Developing Countries. Front. Public Health 2023, 10, 1124473. [Google Scholar] [CrossRef] [PubMed]
- Brand, N.R.; Qu, L.G.; Chao, A.; Ilbawi, A.M. Delays and Barriers to Cancer Care in Low- and Middle-Income Countries: A Systematic Review. Oncologist 2019, 24, e1371–e1380. [Google Scholar] [CrossRef]
- Daniels, J.; Mosadi, L.E.; Nyantakyi, A.Y.; Ayabilah, E.A.; Tackie, J.N.O.; Kyei, K.A. Metastatic Breast Cancer in Resource-Limited Settings: Insights from a Retrospective Cross-Sectional Study at a Radiotherapy Centre in Sub-Saharan Africa. Ecancermedicalscience 2025, 19, 1955. [Google Scholar] [CrossRef]
- Cazap, E.; Magrath, I.; Kingham, T.P.; Elzawawy, A. Structural Barriers to Diagnosis and Treatment of Cancer in Low- and Middle-Income Countries: The Urgent Need for Scaling Up. J. Clin. Oncol. 2016, 34, 14–19. [Google Scholar] [CrossRef]
- Morsi, M.H.; Elawfi, B.; ALsaad, S.A.; Nazar, A.; Mostafa, H.A.; Awwad, S.A.; Abdelwahab, M.M.; Tarakhan, H.; Baghagho, E. Unveiling the Disparities in the Field of Precision Medicine: A Perspective. Health Sci. Rep. 2025, 8, e71102. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z. Evaluation of ChatGPT’s Capabilities in Medical Report Generation. Cureus 2023, 15, e37589. [Google Scholar] [CrossRef]
- Fu, E.; Yager, P.; Floriano, P.N.; Christodoulides, N.; McDevitt, J.T. Perspective on Diagnostics for Global Health. IEEE Pulse 2011, 2, 40–50. [Google Scholar] [CrossRef]
- Hubley, J.H. Barriers to Health Education in Developing Countries. Health Educ. Res. 1986, 1, 233–245. [Google Scholar] [CrossRef]
- Coetzer, M. How AI Ready Data Strengthens Clinical Analytics and Outcomes Research. Available online: https://imatsolutions.com/index.php/2025/12/the-role-of-ai-ready-data-in-strengthening-clinical-research-predictive-modeling-and-care-quality/ (accessed on 10 February 2026).
- Domagalski, M.J.; Lu, Y.; Pilozzi, A.; Williamson, A.; Chilappagari, P.; Luker, E.; Shelley, C.D.; Dabic, A.; Keller, M.A.; Rodriguez, R.M.; et al. Preparing Clinical Research Data for Artificial Intelligence Readiness: Insights from the National Institute of Diabetes and Digestive and Kidney Diseases Data Centric Challenge. J. Am. Med. Inform. Assoc. 2025, 32, 1609–1616. [Google Scholar] [CrossRef]
- Bönisch, C.; Schmidt, C.; Kesztyüs, D.; Kestler, H.A.; Kesztyüs, T. Proposal for Using AI to Assess Clinical Data Integrity and Generate Metadata: Algorithm Development and Validation. JMIR Med. Inform. 2025, 13, e60204. [Google Scholar] [CrossRef] [PubMed]
- Wichard, J.D.; Cammann, H.; Stephan, C.; Tolxdorff, T. Classification Models for Early Detection of Prostate Cancer. BioMed Res. Int. 2008, 2008, 218097. [Google Scholar] [CrossRef]
- Chen, S.; Jian, T.; Chi, C.; Liang, Y.; Liang, X.; Yu, Y.; Jiang, F.; Lu, J. Machine Learning-Based Models Enhance the Prediction of Prostate Cancer. Front. Oncol. 2022, 12, 941349. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.; Feng, X.; Hu, Y.; Yang, J.; Chen, Y.; Bastow, J.; Zheng, Z.-J.; Xu, M. Factors Associated with the Utilization of Diagnostic Tools among Countries with Different Income Levels during the COVID-19 Pandemic. Glob. Health Res. Policy 2023, 8, 45. [Google Scholar] [CrossRef]
- Elmarakeby, H.A.; Hwang, J.; Arafeh, R.; Crowdis, J.; Gang, S.; Liu, D.; AlDubayan, S.H.; Salari, K.; Kregel, S.; Richter, C.; et al. Biologically Informed Deep Neural Network for Prostate Cancer Discovery. Nature 2021, 598, 348–352. [Google Scholar] [CrossRef]
- Shanei, A.; Etehadtavakol, M.; Azizian, M.; Ng, E.Y.K. Comparison of Different Kernels in a Support Vector Machine to Classify Prostate Cancerous Tissues in T2-Weighted Magnetic Resonance Imaging. Explor. Res. Hypothesis Med. 2022, 8, 25–35. [Google Scholar] [CrossRef]
- Chiu, P.K.-F.; Shen, X.; Wang, G.; Ho, C.-L.; Leung, C.-H.; Ng, C.-F.; Choi, K.-S.; Teoh, J.Y.-C. Enhancement of Prostate Cancer Diagnosis by Machine Learning Techniques: An Algorithm Development and Validation Study. Prostate Cancer Prostatic Dis. 2022, 25, 672–676. [Google Scholar] [CrossRef]
- Singh, S.; Pathak, A.K.; Kural, S.; Kumar, L.; Bhardwaj, M.G.; Yadav, M.; Trivedi, S.; Das, P.; Gupta, M.; Jain, G. Integrating MiRNA Profiling and Machine Learning for Improved Prostate Cancer Diagnosis. Sci. Rep. 2025, 15, 30477. [Google Scholar] [CrossRef]
- Jiang, M.; Miao, Z.; Xu, R.; Guo, M.; Li, X.; Li, G.; Luo, P.; Hu, S. Clinical-Radiomics Hybrid Modeling Outperforms Conventional Models: Machine Learning Enhances Stratification of Adverse Prognostic Features in Prostate Cancer. Front. Oncol. 2025, 15, 1625158. [Google Scholar] [CrossRef]
- Zamo, F.C.D.; Ebongue, A.N.; Bongue, D.; Ndontchueng, M.M.; Njeh, C.F. Classification of PSQA Outcomes in Prostate VMAT Treatments: A Comparative Study of Machine Learning Models. Biomed. Eng. Adv. 2026, 11, 100206. [Google Scholar] [CrossRef]
- Nieboer, D.; Vergouwe, Y.; Roobol, M.J.; Ankerst, D.P.; Kattan, M.W.; Vickers, A.J.; Steyerberg, E.W. Nonlinear Modeling Was Applied Thoughtfully for Risk Prediction: The Prostate Biopsy Collaborative Group. J. Clin. Epidemiol. 2015, 68, 426–434. [Google Scholar] [CrossRef]
- Zhao, Y.; Zhang, L.; Zhang, S.; Li, J.; Shi, K.; Yao, D.; Li, Q.; Zhang, T.; Xu, L.; Geng, L.; et al. Machine Learning-Based MRI Imaging for Prostate Cancer Diagnosis: Systematic Review and Meta-Analysis. Prostate Cancer Prostatic Dis. 2025, 29, 426–434. [Google Scholar] [CrossRef] [PubMed]
- Morote, J.; Miró, B.; Hernando, P.; Paesano, N.; Picola, N.; Muñoz-Rodriguez, J.; Ruiz-Plazas, X.; Muñoz-Rivero, M.V.; Celma, A.; García-de Manuel, G.; et al. Developing a Predictive Model for Significant Prostate Cancer Detection in Prostatic Biopsies from Seven Clinical Variables: Is Machine Learning Superior to Logistic Regression? Cancers 2025, 17, 1101. [Google Scholar] [CrossRef]
- Zhang, Y.-D.; Wang, J.; Wu, C.-J.; Bao, M.-L.; Li, H.; Wang, X.-N.; Tao, J.; Shi, H.-B. An Imaging-Based Approach Predicts Clinical Outcomes in Prostate Cancer through a Novel Support Vector Machine Classification. Oncotarget 2016, 7, 78140–78151. [Google Scholar] [CrossRef]
- Siech, C.; Wenzel, M.; Grosshans, N.; Cano Garcia, C.; Humke, C.; Koll, F.J.; Tian, Z.; Karakiewicz, P.I.; Kluth, L.A.; Chun, F.K.H.; et al. The Association Between Lymphovascular or Perineural Invasion in Radical Prostatectomy Specimen and Biochemical Recurrence. Cancers 2024, 16, 3648. [Google Scholar] [CrossRef]
- Chung, D.H.; Han, J.H.; Jeong, S.-H.; Yuk, H.D.; Jeong, C.W.; Ku, J.H.; Kwak, C. Role of Lymphatic Invasion in Predicting Biochemical Recurrence after Radical Prostatectomy. Front. Oncol. 2023, 13, 1226366. [Google Scholar] [CrossRef]
- Fajkovic, H.; Mathieu, R.; Lucca, I.; Hiess, M.; Hübner, N.; Al Awamlh, B.A.H.; Lee, R.; Briganti, A.; Karakiewicz, P.; Lotan, Y.; et al. Validation of Lymphovascular Invasion Is an Independent Prognostic Factor for Biochemical Recurrence after Radical Prostatectomy. Urol. Oncol. Semin. Orig. Investig. 2016, 34, 233.e1–233.e6. [Google Scholar] [CrossRef]
- Karwacki, J.; Stodolak, M.; Dłubak, A.; Nowak, Ł.; Gurwin, A.; Kowalczyk, K.; Kiełb, P.; Holdun, N.; Szlasa, W.; Krajewski, W.; et al. Association of Lymphovascular Invasion with Biochemical Recurrence and Adverse Pathological Characteristics of Prostate Cancer: A Systematic Review and Meta-Analysis. Eur. Urol. Open Sci. 2024, 69, 112–126. [Google Scholar] [CrossRef]
- Schmidhuber, J. Deep Learning in Neural Networks: An Overview. Neural Netw. 2015, 61, 85–117. [Google Scholar] [CrossRef] [PubMed]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; van der Laak, J.A.W.M.; van Ginneken, B.; Sánchez, C.I. A Survey on Deep Learning in Medical Image Analysis. Med. Image Anal. 2017, 42, 60–88. [Google Scholar] [CrossRef]
- Aggarwal, R.; Sounderajah, V.; Martin, G.; Ting, D.S.W.; Karthikesalingam, A.; King, D.; Ashrafian, H.; Darzi, A. Diagnostic Accuracy of Deep Learning in Medical Imaging: A Systematic Review and Meta-Analysis. npj Digit. Med. 2021, 4, 65. [Google Scholar] [CrossRef]
- Erdem, E.; Bozkurt, F. A Comparison of Various Supervised Machine Learning Techniques for Prostate Cancer Prediction. Eur. J. Sci. Technol. 2021, 9, 610–620. [Google Scholar] [CrossRef]
- Shu, X.; Liu, Y.; Qiao, X.; Ai, G.; Liu, L.; Liao, J.; Deng, Z.; He, X. Radiomic-Based Machine Learning Model for the Accurate Prediction of Prostate Cancer Risk Stratification. Br. J. Radiol. 2023, 96, 20220238. [Google Scholar] [CrossRef]
- Gupta, S.; Kumar, M. Prostate Cancer Prognosis Using Multi-Layer Perceptron and Class Balancing Techniques. In 2021 Thirteenth International Conference on Contemporary Computing (IC3-2021); ACM: New York, NY, USA, 2021; pp. 1–6. [Google Scholar] [CrossRef]
- Yang, T.; Zhang, H.; Peng, H.; Niu, X.; Yang, F.; Zhang, J.; Wang, Q.; Fan, J.; Song, Y.; Tao, W. PSMA PET/MRI-based Swin Transformer Architecture for Gleason Score Prediction in Prostate Cancer. Med. Phys. 2026, 53, e70274. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, C.; Hulsey, G.; James, K.; James, T.; Carlson, J.M. Deep Learning Classification of Prostate Cancer Using MRI Histopathologic Data. Radiol. Imaging Cancer 2025, 7, e240381. [Google Scholar] [CrossRef]
- Li, D.; Han, X.; Gao, J.; Zhang, Q.; Yang, H.; Liao, S.; Guo, H.; Zhang, B. Deep Learning in Prostate Cancer Diagnosis Using Multiparametric Magnetic Resonance Imaging With Whole-Mount Histopathology Referenced Delineations. Front. Med. 2022, 8, 810995. [Google Scholar] [CrossRef]
- He, M.; Cao, Y.; Chi, C.; Yang, X.; Ramin, R.; Wang, S.; Yang, G.; Mukhtorov, O.; Zhang, L.; Kazantsev, A.; et al. Research Progress on Deep Learning in Magnetic Resonance Imaging–Based Diagnosis and Treatment of Prostate Cancer: A Review on the Current Status and Perspectives. Front. Oncol. 2023, 13, 1189370. [Google Scholar] [CrossRef] [PubMed]
- Quarta, L.; Stabile, A.; Pellegrino, F.; Scilipoti, P.; Longoni, M.; Cannoletta, D.; Zaurito, P.; Santangelo, A.; Viti, A.; Barletta, F.; et al. Tailored Use of PSA Density According to Multiparametric MRI Index Lesion Location: Results of a Large, Multi-Institutional Series. Prostate Cancer Prostatic Dis. 2025, 29, 111–115. [Google Scholar] [CrossRef]
- Wang, X.; Zhong, S.; Fang, K.; Du, Y.; Huang, J. Application and Prospect of Artificial Intelligence in Diagnostic Imaging of Prostate Cancer. npj Digit. Med. 2026, 9, 168. [Google Scholar] [CrossRef]
- Zhao, K.; Hung, A.L.Y.; Pang, K.; Hajipour, P.; Wu, H.; Raman, S.; Sung, K. PCa-Mamba: Spatiotemporal State Space Models for Prostate Cancer Detection in Multi-Parametric MRI. Med. Image Anal. 2026, 111, 104033. [Google Scholar] [CrossRef]
- Alzate-Grisales, J.A.; Mora-Rubio, A.; Peán-Teruel, M.; Beltrán, A.N.; Torres, C.R.; García, J.M.O.; García-García, F.; Tabares-Soto, R.; García-Gómez, J.M.; de la Iglesia-Vayá, M. Clinically Significant Prostate Cancer Detection with Deep Learning in a Multi-Center Magnetic Resonance Imaging Study. Sci. Rep. 2026, 16, 10976. [Google Scholar] [CrossRef]
- Rodrigues, N.M.; de Almeida, J.G.; Verde, A.S.C.; Gaivão, A.M.; Bireiro, C.; Santiago, I.; Ip, J.; Belião, S.; Matos, C.; Vanneschi, L.; et al. Effective Reduction of Unnecessary Biopsies through a Deep-Learning-Assisted Aggressive Prostate Cancer Detector. Sci. Rep. 2025, 15, 15211. [Google Scholar] [CrossRef]
- Ciobanu-Caraus, O.; Aicher, A.; Kernbach, J.M.; Regli, L.; Serra, C.; Staartjes, V.E. A Critical Moment in Machine Learning in Medicine: On Reproducible and Interpretable Learning. Acta Neurochir. 2024, 166, 14. [Google Scholar] [CrossRef] [PubMed]
- McDermott, M.B.A.; Wang, S.; Marinsek, N.; Ranganath, R.; Foschini, L.; Ghassemi, M. Reproducibility in Machine Learning for Health Research: Still a Ways to Go. Sci. Transl. Med. 2021, 13, eabb1655. [Google Scholar] [CrossRef]
- Guillen-Aguinaga, M.; Aguinaga-Ontoso, E.; Guillen-Aguinaga, L.; Guillen-Grima, F.; Aguinaga-Ontoso, I. Data Quality in the Age of AI: A Review of Governance, Ethics, and the FAIR Principles. Data 2025, 10, 201. [Google Scholar] [CrossRef]
- Clark, T.; Caufield, H.; Parker, J.A.; Al Manir, S.; Amorim, E.; Eddy, J.; Gim, N.; Gow, B.; Goar, W.; Haendel, M.; et al. AI-Readiness for Biomedical Data: Bridge2AI Recommendations. bioRxiv 2024. [Google Scholar] [CrossRef]
- Aksenova, A.; Johny, A.; Adams, T.; Gribbon, P.; Jacobs, M.; Hofmann-Apitius, M. Current State of Data Stewardship Tools in Life Science. Front. Big Data 2024, 7, 1428568. [Google Scholar] [CrossRef]
- D’Amico, A.V. Biochemical Outcome After Radical Prostatectomy, External Beam Radiation Therapy, or Interstitial Radiation Therapy for Clinically Localized Prostate Cancer. JAMA 1998, 280, 969. [Google Scholar] [CrossRef]
- Diamand, R.; Oderda, M.; Albisinni, S.; Fourcade, A.; Fournier, G.; Benamran, D.; Iselin, C.; Fiard, G.; Descotes, J.-L.; Assenmacher, G.; et al. External Validation of the Briganti Nomogram Predicting Lymph Node Invasion in Patients with Intermediate and High-Risk Prostate Cancer Diagnosed with Magnetic Resonance Imaging-Targeted and Systematic Biopsies: A European Multicenter Study. Urol. Oncol. Semin. Orig. Investig. 2020, 38, 847.e9–847.e16. [Google Scholar] [CrossRef]
- Hansen, J.; Rink, M.; Bianchi, M.; Kluth, L.A.; Tian, Z.; Ahyai, S.A.; Shariat, S.F.; Briganti, A.; Steuber, T.; Fisch, M.; et al. External Validation of the Updated Briganti Nomogram to Predict Lymph Node Invasion in Prostate Cancer Patients Undergoing Extended Lymph Node Dissection. Prostate 2013, 73, 211–218. [Google Scholar] [CrossRef]
- Małkiewicz, B.; Ptaszkowski, K.; Knecht, K.; Gurwin, A.; Wilk, K.; Kiełb, P.; Dudek, K.; Zdrojowy, R. External Validation of the Briganti Nomogram to Predict Lymph Node Invasion in Prostate Cancer—Setting a New Threshold Value. Life 2021, 11, 479. [Google Scholar] [CrossRef]
- ISO/IEC 42001:2023; Information Technology—Artificial Intelligence—Management System. International Organization for Standardization: Geneva, Switzerland, 2023.
- Wang, R.Y.; Strong, D.M. Beyond Accuracy: What Data Quality Means to Data Consumers. J. Manag. Inf. Syst. 1996, 12, 5–33. [Google Scholar] [CrossRef]
- Faul, F.; Erdfelder, E.; Buchner, A.; Lang, A.-G. Statistical Power Analyses Using G*Power 3.1: Tests for Correlation and Regression Analyses. Behav. Res. Methods 2009, 41, 1149–1160. [Google Scholar] [CrossRef]
- Faul, F.; Erdfelder, E.; Lang, A.-G.; Buchner, A. G*Power 3: A Flexible Statistical Power Analysis Program for the Social, Behavioral, and Biomedical Sciences. Behav. Res. Methods 2007, 39, 175–191. [Google Scholar] [CrossRef] [PubMed]
- Dean, A.G.; Sullivan, K.M.; Soe, M.M.; Sullivan, K.M. OpenEpi: Open Source Epidemiologic Statistics for Public Health, Versión 3.01. 2013. Available online: https://openepi.com/ (accessed on 14 March 2026).
- Cytel Inc. LogXact-11: Software for Exact Logistic Regression, A, Version; Cytel Inc.: Waltham, MA, USA, 2015. [Google Scholar]
- Sur, P.; Candès, E.J. A Modern Maximum-Likelihood Theory for High-Dimensional Logistic Regression. Proc. Natl. Acad. Sci. USA 2019, 116, 14516–14525. [Google Scholar] [CrossRef]
- Yee, T.W. On the Hauck-Donner Effect in Wald Tests: Detection, Tipping Points, and Parameter Space Characterization. J. Am. Stat. Assoc. 2022, 117, 1763–1774. [Google Scholar] [CrossRef]
- Hauck, W.W.; Donner, A. Wald’s Test as Applied to Hypotheses in Logit Analysis. J. Am. Stat. Assoc. 1977, 72, 851–853. [Google Scholar] [CrossRef]
- Van Calster, B.; Collins, G.S.; Vickers, A.J.; Wynants, L.; Kerr, K.F.; Barreñada, L.; Varoquaux, G.; Singh, K.; Moons, K.G.; Hernandez-Boussard, T.; et al. Evaluation of Performance Measures in Predictive Artificial Intelligence Models to Support Medical Decisions: Overview and Guidance. Lancet Digit. Health 2025, 7, 100916. [Google Scholar] [CrossRef]
- Roberts, M.; Driggs, D.; Thorpe, M.; Gilbey, J.; Yeung, M.; Ursprung, S.; Aviles-Rivero, A.I.; Etmann, C.; McCague, C.; Beer, L.; et al. Common Pitfalls and Recommendations for Using Machine Learning to Detect and Prognosticate for COVID-19 Using Chest Radiographs and CT Scans. Nat. Mach. Intell. 2021, 3, 199–217. [Google Scholar] [CrossRef]
- Varoquaux, G.; Cheplygina, V. Machine Learning for Medical Imaging: Methodological Failures and Recommendations for the Future. npj Digit. Med. 2022, 5, 48. [Google Scholar] [CrossRef]
- Yu, A.C.; Mohajer, B.; Eng, J. External Validation of Deep Learning Algorithms for Radiologic Diagnosis: A Systematic Review. Radiol. Artif. Intell. 2022, 4, e210064. [Google Scholar] [CrossRef]
- Arun, S.; Grosheva, M.; Kosenko, M.; Robertus, J.L.; Blyuss, O.; Gabe, R.; Munblit, D.; Offman, J. Systematic Scoping Review of External Validation Studies of AI Pathology Models for Lung Cancer Diagnosis. npj Precis. Oncol. 2025, 9, 166. [Google Scholar] [CrossRef]
- Tseng, A.S.; Shelly-Cohen, M.; Attia, I.Z.; Noseworthy, P.A.; Friedman, P.A.; Oh, J.K.; Lopez-Jimenez, F. Spectrum Bias in Algorithms Derived by Artificial Intelligence: A Case Study in Detecting Aortic Stenosis Using Electrocardiograms. Eur. Heart J. Digit. Health 2021, 2, 561–567. [Google Scholar] [CrossRef]
- Yu, L.; Gao, X.-S.; Zhang, L.; Miao, Y. Generalizability of Memorization Neural Networks. In 38th Conference on Neural Information Processing Systems (NeurIPS 2024); Globerson, A., Mackey, L., Belgrave, D., Fan, A., Paquet, U., Tomczak, J., Zhang, C., Eds.; Neural Information Processing Systems Foundation, Inc.: Vancouver, BC, USA, 2024; pp. 1–46. [Google Scholar]
- Zhang, C.; Bengio, S.; Hardt, M.; Mozer, M.C.; Singer, Y. Identity Crisis: Memorization and Generalization under Extreme Overparameterization. arXiv 2020, arXiv:1902.04698. [Google Scholar] [CrossRef]
- Belkin, M.; Hsu, D.; Ma, S.; Mandal, S. Reconciling Modern Machine-Learning Practice and the Classical Bias–Variance Trade-Off. Proc. Natl. Acad. Sci. USA 2019, 116, 15849–15854. [Google Scholar] [CrossRef]
- Hastie, T.; Tibshiran, R.; Friedman, J. The Elements of Statistical Learning: Data Mining, Inference, and Prediction, 2nd ed.; Springer: New York, NY, USA, 2017. [Google Scholar]
- Crespo Márquez, A. The Curse of Dimensionality. In Digital Maintenance Management; Springer Science and Business Media Deutschland GmbH: New York, NY, USA, 2022; pp. 67–86. [Google Scholar] [CrossRef]
- Tishby, N.; Zaslavsky, N. Deep Learning and the Information Bottleneck Principle. In 2015 IEEE Information Theory Workshop (ITW); IEEE: New York, NY, USA, 2015; pp. 1–5. [Google Scholar] [CrossRef]
- Koch, R.D.M.; Ghosh, A. Two-Phase Perspective on Deep Learning Dynamics. Phys. Rev. E 2025, 112, 025307. [Google Scholar] [CrossRef]
- Chapman, B.P.; Weiss, A.; Duberstein, P.R. Statistical Learning Theory for High Dimensional Prediction: Application to Criterion-Keyed Scale Development. Psychol. Methods 2016, 21, 603–620. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Rosasco, L.; Caponnetto, A. On Early Stopping in Gradient Descent Learning. Constr. Approx. 2007, 26, 289–315. [Google Scholar] [CrossRef]
- Lee, F. What Is Bias-Variance Tradeoff? Available online: https://www.ibm.com/think/topics/bias-variance-tradeoff (accessed on 15 February 2026).
- Hu, Y.; Zhang, X.; Slavin, V.; Belsti, Y.; Tiruneh, S.A.; Callander, E.; Enticott, J. Beyond Comparing Machine Learning and Logistic Regression in Clinical Prediction Modelling: Shifting from Model Debate to Data Quality. J. Med. Internet Res. 2025, 27, e77721. [Google Scholar] [CrossRef]
- Condon, D. Performance of Artificial Neural Networks on Small Structured Datasets; AMSI Vacation Research Scholarships: Victoria, Australia, 2019. [Google Scholar]
- Bailly, A.; Blanc, C.; Francis, É.; Guillotin, T.; Jamal, F.; Wakim, B.; Roy, P. Effects of Dataset Size and Interactions on the Prediction Performance of Logistic Regression and Deep Learning Models. Comput. Methods Programs Biomed. 2022, 213, 106504. [Google Scholar] [CrossRef]
- Issitt, R.W.; Cortina-Borja, M.; Bryant, W.; Bowyer, S.; Taylor, A.M.; Sebire, N. Classification Performance of Neural Networks Versus Logistic Regression Models: Evidence from Healthcare Practice. Cureus 2022, 14, e22443. [Google Scholar] [CrossRef]
- Nguyen, T.; Nguyen, H.-T.; Nguyen-Hoang, T.-A. Data Quality Management in Big Data: Strategies, Tools, and Educational Implications. J. Parallel Distrib. Comput. 2025, 200, 105067. [Google Scholar] [CrossRef]
- Liu, J.; Zhao, J.; Zhang, M.; Chen, N.; Sun, G.; Yang, Y.; Zhang, X.; Chen, J.; Shen, P.; Shi, M.; et al. The Validation of the 2014 International Society of Urological Pathology (ISUP) Grading System for Patients with High-Risk Prostate Cancer: A Single-Center Retrospective Study. Cancer Manag. Res. 2019, 11, 6521–6529. [Google Scholar] [CrossRef]
- Offermann, A.; Hupe, M.C.; Sailer, V.; Merseburger, A.S.; Perner, S. The New ISUP 2014/WHO 2016 Prostate Cancer Grade Group System: First Résumé 5 Years after Introduction and Systemic Review of the Literature. World J. Urol. 2020, 38, 657–662. [Google Scholar] [CrossRef]
- Epstein, J.I.; Egevad, L.; Amin, M.B.; Delahunt, B.; Srigley, J.R.; Humphrey, P.A. The 2014 International Society of Urological Pathology (ISUP) Consensus Conference on Gleason Grading of Prostatic Carcinoma. Am. J. Surg. Pathol. 2016, 40, 244–252. [Google Scholar] [CrossRef]
- Persily, J.; Chandarana, H.; Tong, A.; Ranganath, R.; Taneja, S.; Nayan, M. Development and Deployment of a Machine Learning Model to Triage the Use of Prostate MRI (ProMT-ML) in Patients With Suspected Prostate Cancer. J. Magn. Reson. Imaging 2026, 63, 698–708. [Google Scholar] [CrossRef]
- Giganti, F.; Moreira da Silva, N.; Yeung, M.; Davies, L.; Frary, A.; Ferrer Rodriguez, M.; Sushentsev, N.; Ashley, N.; Andreou, A.; Bradley, A.; et al. AI-Powered Prostate Cancer Detection: A Multi-Centre, Multi-Scanner Validation Study. Eur. Radiol. 2025, 35, 4915–4924. [Google Scholar] [CrossRef]
- Barberis, A.; Aerts, H.J.W.L.; Buffa, F.M. Robustness and Reproducibility for AI Learning in Biomedical Sciences: RENOIR. Sci. Rep. 2024, 14, 1933. [Google Scholar] [CrossRef]
- Colliot, O.; Thibeau-Sutre, E.; Burgos, N. Reproducibility in Machine Learning for Medical Imaging. In Machine Learning for Brain Disorders; Colliot, O., Ed.; Humana: Lexington, KY, USA, 2023; Volume 197, pp. 631–653. [Google Scholar] [CrossRef]
- Beam, A.L.; Manrai, A.K.; Ghassemi, M. Challenges to the Reproducibility of Machine Learning Models in Health Care. JAMA 2020, 323, 305. [Google Scholar] [CrossRef]
- Yuan, M.; Chang, D.; Lu, W.; Ma, K.; Gu, Y.; Xia, T.; Peng, J.; Zhang, Y.; Fu, L.; Zhao, B. Interpretable Habitat and Peritumoral Radiomics from Multiparametric MRI for Preoperative High-Risk Prostate Cancer Prediction: A Multi-Institutional Study. J. Transl. Med. 2026, 24, 399. [Google Scholar] [CrossRef]
- de Kanter, E.; Kaul, T.; Heus, P.; de Groot, T.M.; Kuijten, R.H.; Reitsma, J.B.; Collins, G.S.; Hooft, L.; Moons, K.G.M.; Damen, J.A.A. Adherence to TRIPOD+AI Guideline: An Updated Reporting Assessment Tool. J. Clin. Epidemiol. 2026, 191, 112118. [Google Scholar] [CrossRef] [PubMed]
- Martin, G.P.; Riley, R.D.; Ensor, J.; Grant, S.W. Statistical Primer: Sample Size Considerations for Developing and Validating Clinical Prediction Models. Eur. J. Cardio Thorac. Surg. 2025, 67, ezaf142. [Google Scholar] [CrossRef] [PubMed]
- Alwosheel, A.; van Cranenburgh, S.; Chorus, C.G. Is Your Dataset Big Enough? Sample Size Requirements When Using Artificial Neural Networks for Discrete Choice Analysis. J. Choice Model. 2018, 28, 167–182. [Google Scholar] [CrossRef]
- Harrell, F.E. Regression Modeling Strategies, 2nd ed.; Springer: New York, NY, USA, 2015. [Google Scholar]
- Peduzzi, P.; Concato, J.; Kemper, E.; Holford, T.R.; Feinstein, A.R. A Simulation Study of the Number of Events per Variable in Logistic Regression Analysis. J. Clin. Epidemiol. 1996, 49, 1373–1379. [Google Scholar] [CrossRef]
- Tsegaye, B.; Snell, K.I.E.; Archer, L.; Kirtley, S.; Riley, R.D.; Sperrin, M.; Van Calster, B.; Collins, G.S.; Dhiman, P. Larger Sample Sizes Are Needed When Developing a Clinical Prediction Model Using Machine Learning in Oncology: Methodological Systematic Review. J. Clin. Epidemiol. 2025, 180, 111675. [Google Scholar] [CrossRef]
- Kapoor, S.; Narayanan, A. Leakage and the Reproducibility Crisis in Machine-Learning-Based Science. Patterns 2023, 4, 100804. [Google Scholar] [CrossRef]
- Young, V.M.; Gates, S.; Garcia, L.Y.; Salardini, A. Data Leakage in Deep Learning for Alzheimer’s Disease Diagnosis: A Scoping Review of Methodological Rigor and Performance Inflation. Diagnostics 2025, 15, 2348. [Google Scholar] [CrossRef]
- Starcke, J.; Spadafora, J.; Spadafora, J.; Spadafora, P.; Toma, M. The Effect of Data Leakage and Feature Selection on Machine Learning Performance for Early Parkinson’s Disease Detection. Bioengineering 2025, 12, 845. [Google Scholar] [CrossRef] [PubMed]
- Brierley, J.D.; Giuliani, M.; O’Sullivan, B.; Rous, B.; Van Eycken, E. TNM Classification of Malignant Tumours, 9th ed.; John Wiley & Sons: Hoboken, NJ, USA, 2025. [Google Scholar]
- Epstein, J.I.; Egevad, L.; Amin, M.B.; Delahunt, B.; Srigley, J.R.; Humphrey, P.A. The 2014 International Society of Urological Pathology (ISUP) Consensus Conference on Gleason Grading of Prostatic Carcinoma. Am. J. Surg. Pathol. 2016, 40, 244–252. [Google Scholar] [CrossRef]



| Partitioning Scheme | Total Sample | Valid Cases (n) | Excluded Cases (n) | Exclusion Rate (%) | Primary Reason for Exclusion |
|---|---|---|---|---|---|
| Model 20/80 | 49 | 43 | 6 | 12.20% | Factor levels (e.g., PSA values or mrT stages) not present in the training set. |
| Model 34/66 | 49 | 44 | 5 | 10.20% | Factor levels or dependent variable values (Low-risk strata) not present in training. |
| Model 39/61 | 49 | 41 | 8 | 16.30% | Factor levels (clinical outliers) not represented in the training sample. |
| Metric | Model 20/80 | Model 34/66 | Model 39/61 |
|---|---|---|---|
| D’Amico Strata Evaluated | Binary (high/int) | Ternary (high/int/low) | Binary (high/int) |
| Training Sample n (%) | 34 (79.10%) | 29 (65.90%) | 25 (61.00%) |
| Testing Sample n (%) | 9 (20.9%) | 15 (34.1%) | 16 (39.0%) |
| Training Cross-Entropy Error | 0.161 | 3.842 | 0.209 |
| Testing Cross-Entropy Error | 0.001 | 4.227 | 4.636 |
| Training Incorrect Predictions (%) | 0.00% | 6.90% | 0.00% |
| Testing Incorrect Predictions (%) | 0.00% | 13.30% | 6.30% |
| Overall Training Accuracy (%) | 100% | 93.10% | 100.00% |
| Overall Testing Accuracy (%) | 100% (95% CI: 66.4–100) † | 86.70% (95% CI: 62.1–96.3) | 93.80% (95% CI: 71.7–98.9) |
| Correct Classifications (n/N) | (9/9) | (13/15) | (15/16) |
| Sensitivity (High Risk) | 100% (95% CI: 56.5–100) | 85.7% (95% CI: 48.7–97.4) | 87.5% (95% CI: 52.9–97.8) |
| Specificity (Int. Risk) | 100% (95% CI: 51.0–100) | 87.5% (95% CI: 52.9–97.8) | 100% (95% CI: 67.6–100) |
| PPV (Positive Predictive Value) | 100% (95% CI: 56.5–100) | 85.7% (95% CI: 48.7–97.4) | 100% (95% CI: 64.6–100) |
| NPV (Negative Predictive Value) | 100% (95% CI: 51.0–100) | 87.5% (95% CI: 52.9–97.8) | 88.9% (95% CI: 56.5–98.0) |
| Risk Group | AUC Model 20/80 | AUC Model 34/66 | AUC Model 39/61 |
|---|---|---|---|
| High | 1 | 0.996 | 0.99 |
| Intermediate | 1 | 0.991 | 0.99 |
| Low | N/A | 1 | N/A |
| Sample | Observed Risk Group | Predicted: High | Predicted: Intermediate | Percent Correct |
|---|---|---|---|---|
| Training | High | 22 | 0 | 100% |
| Intermediate | 0 | 12 | 100% | |
| Testing | High | 5 | 0 | 100% |
| Intermediate | 0 | 4 | 100% | |
| Global Percentage | 55.60% | 44.40% | 100% |
| Variable | Importance | Normalized Importance |
|---|---|---|
| ISUP BX | 0.181 | 100.00% |
| BRIGANTI | 0.180 | 99.30% |
| PSA DENSITY | 0.127 | 70.20% |
| PSA at DX | 0.098 | 54.20% |
| AGE | 0.098 | 54.00% |
| mrT | 0.071 | 39.10% |
| PROSTATE VOLUME c.c. | 0.069 | 38.20% |
| BX LATERALITY | 0.067 | 37.00% |
| Clinical TNM Stage | 0.053 | 29.10% |
| mrN | 0.032 | 17.90% |
| Nodal Stage | 0.025 | 14.10% |
| Feature/Metric | Exact Logistic Regression (Baseline) | Multilayer Perceptron (MLP 20/80) |
|---|---|---|
| Statistical Engine | Exact Likelihood Estimation (LogXact). | Backpropagation/Softmax (SPSS). |
| Model Type | Linear/parametric. | Non-linear/connectionist + 1. |
| Overall Significance | p < 0.001 (likelihood ratio). | p = 0.002 (Exact Binomial Test). |
| Predictor: ISUP Grade | Significant (p = 0.005). | Dominant (100% importance). |
| Predictor: Briganti Nomogram | Not significant (p = 0.800). | Critical (99.3% importance). |
| Predictor: PSA Density | Not significant (p = 0.500). | High impact (70.2% importance). |
| Predictor: Age | Degenerate estimate (DEGEN). † | Moderate impact (54.0% importance). |
| Testing Accuracy | Outperformed by MLP | 100.00%. |
| Testing AUC | <1.000. | 1. |
| Clinical Interpretation | Limited to linear histological signal. | Captures non-linear feature synergies. |
| Tool/ Approach | Input Data | Primary Clinical Task | Approximate Development Sample | Data-Quality Governance | Principal Claim |
|---|---|---|---|---|---|
| D’Amico classification [63] | 3 clinical/biochemical variables | Pre-treatment risk stratification | Multi-cohort historical | Implicit (clinical staging conventions) | Validated three-tier risk groups |
| Briganti nomogram [64,65,66] | 4–5 clinical/biochemical/pathological variables | Probability of lymph node invasion | >1000 patients (multi-cohort) | Standardised pathology review | Calibrated probabilistic estimate |
| Exact logistic regression (Table 7) | 5 top MLP predictors | Linear risk discrimination | N = 43 valid cases (this study) | Same DQG protocol as MLP | Confirms ISUP signal; cannot integrate non-linear features |
| Image-based deep learning [101] (e.g., PCa-Mamba [55] multi-centre 3D EfficientNet [56]) | mpMRI sequences (T2W, DWI, and DCE) | Lesion detection/csPCa | Thousands of MRI sessions, multi-centre | Variable; PI-CAI provides a curated benchmark | Lesion-level AUC ≈ 0.80–0.90 |
| Tabular ML triage (e.g., ProMT-ML [100] | 4 routinely available variables | Pre-MRI triage | >11,000 retrospective + 4500 prospective | Routine-care data; deployed pipeline | Outperforms PSA-based triage |
| Proposed MLP (this work) | 11 clinical/biochemical/imaging-derived variables | Post-detection D’Amico risk stratification | N = 49 (proof-of-concept) | Explicit FAIR-based DQG with ISO/IEC 25012 alignment, dual blinded extraction, AI-readiness SOPs (Appendix A) | Methodological framework—not a deployable model |
| Phase | Site (s) | Target Sample Size | Primary Objective | Pre-Specified Endpoints |
|---|---|---|---|---|
| A | CUN Pamplona (development site). | ≥220 evaluable patients (≥110 high-risk events; replenished low-risk stratum). | Cohort expansion to enable three-class classifier and formal internal cross-validation. | Discrimination (AUC); calibration; bootstrap-based variable-importance CIs; SHAP attribution; documented anti-leakage safeguards (training-only fitting of scalers and imputers; patient-level partition integrity). |
| B | CUN Madrid (second site, same institutional system). | Independent cohort ≥150 evaluable patients. | Within-system external validation; isolation of population vs. curation heterogeneity. | AUC; calibration intercept/slope; Brier score; decision-curve analysis; TRIPOD + AI subgroup reporting; verification that anti-leakage safeguards from Phase A are preserved across sites. |
| C | Spanish multi-centre network (additional centres). | Multi-centre cohort ≥500 evaluable patients. | True multi-institutional external validation; test of framework portability across heterogeneous EHR/imaging/pathology ecosystems. | Site-stratified AUC and calibration; data-drift monitoring; net clinical benefit vs. standard nomograms; pre-registered analysis plan. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Talavera-Cobo, V.; Robles-Garcia, J.E.; Guillen-Grima, F.; Calva-Lopez, A.; Tapia-Tapia, M.; Labairu-Huerta, L.; Ancizu-Marckert, F.J.; Guillen-Aguinaga, L.; Sanchez-Zalabardo, D.; Miñana-Lopez, B. Prioritising Data Quality Governance for AI in Prostate Cancer: A Methodological Proof-of-Concept Study Using Neural Networks for Risk Stratification. Diagnostics 2026, 16, 1454. https://doi.org/10.3390/diagnostics16101454
Talavera-Cobo V, Robles-Garcia JE, Guillen-Grima F, Calva-Lopez A, Tapia-Tapia M, Labairu-Huerta L, Ancizu-Marckert FJ, Guillen-Aguinaga L, Sanchez-Zalabardo D, Miñana-Lopez B. Prioritising Data Quality Governance for AI in Prostate Cancer: A Methodological Proof-of-Concept Study Using Neural Networks for Risk Stratification. Diagnostics. 2026; 16(10):1454. https://doi.org/10.3390/diagnostics16101454
Chicago/Turabian StyleTalavera-Cobo, Vanessa, Jose Enrique Robles-Garcia, Francisco Guillen-Grima, Andres Calva-Lopez, Mario Tapia-Tapia, Luis Labairu-Huerta, Francisco Javier Ancizu-Marckert, Laura Guillen-Aguinaga, Daniel Sanchez-Zalabardo, and Bernardino Miñana-Lopez. 2026. "Prioritising Data Quality Governance for AI in Prostate Cancer: A Methodological Proof-of-Concept Study Using Neural Networks for Risk Stratification" Diagnostics 16, no. 10: 1454. https://doi.org/10.3390/diagnostics16101454
APA StyleTalavera-Cobo, V., Robles-Garcia, J. E., Guillen-Grima, F., Calva-Lopez, A., Tapia-Tapia, M., Labairu-Huerta, L., Ancizu-Marckert, F. J., Guillen-Aguinaga, L., Sanchez-Zalabardo, D., & Miñana-Lopez, B. (2026). Prioritising Data Quality Governance for AI in Prostate Cancer: A Methodological Proof-of-Concept Study Using Neural Networks for Risk Stratification. Diagnostics, 16(10), 1454. https://doi.org/10.3390/diagnostics16101454







