Pilot Testing of the “Turbidimeter”, a Simple, Universal Reader Intended to Complement and Enhance Bacterial Growth Detection in Manual Blood Culture Systems in Low-Resource Settings
Abstract
1. Introduction
2. Materials and Methods
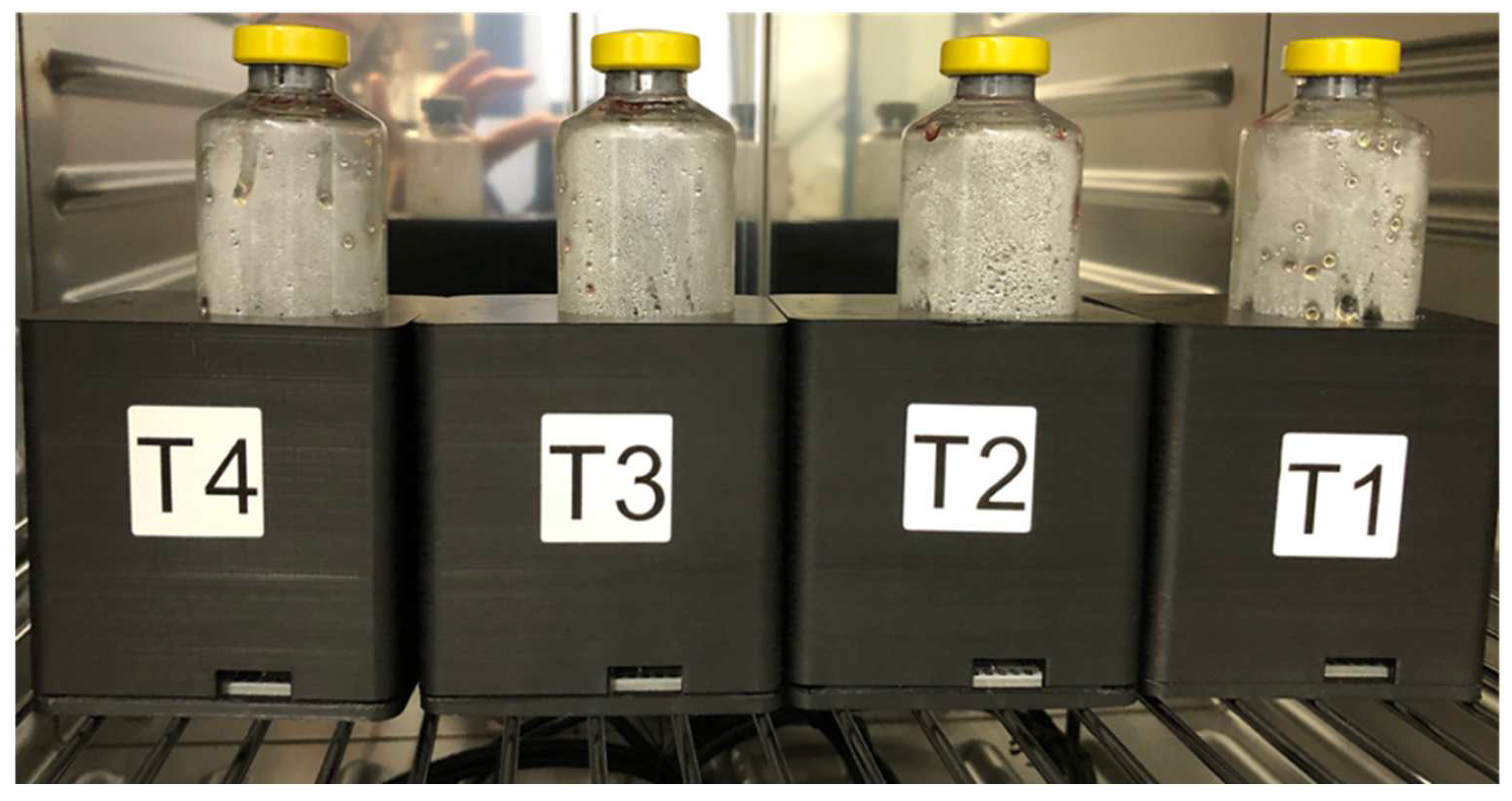
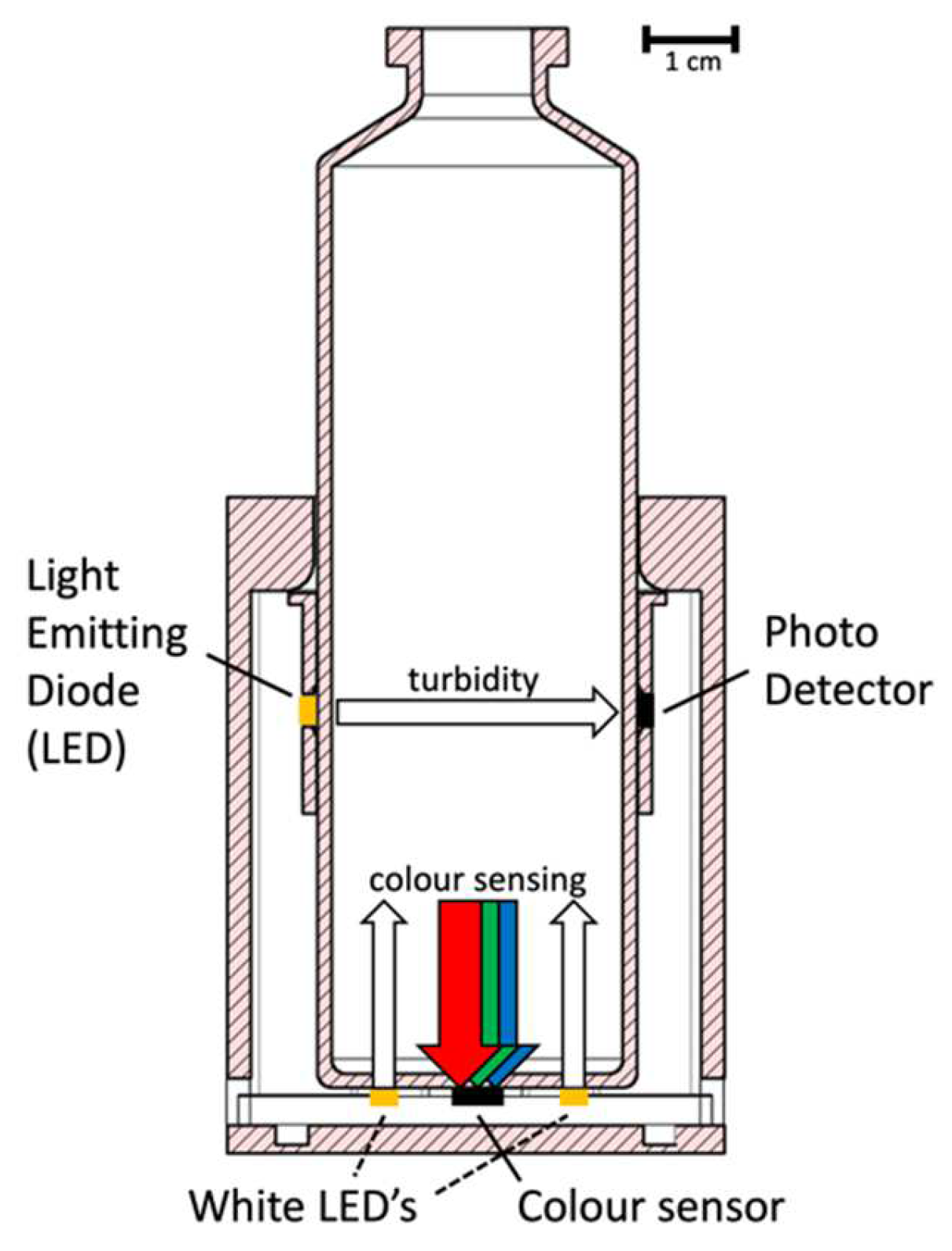
2.1. Turbidimeter Prototype: Concept and Design
2.2. Preparation of Spiked Blood Cultures
2.3. Growth Experiments with Spiked Blood Culture Bottles
2.4. Definitions
2.5. Data Analysis
3. Results
3.1. Turbidimeter Prototype: Preparing for the Pilot Testing
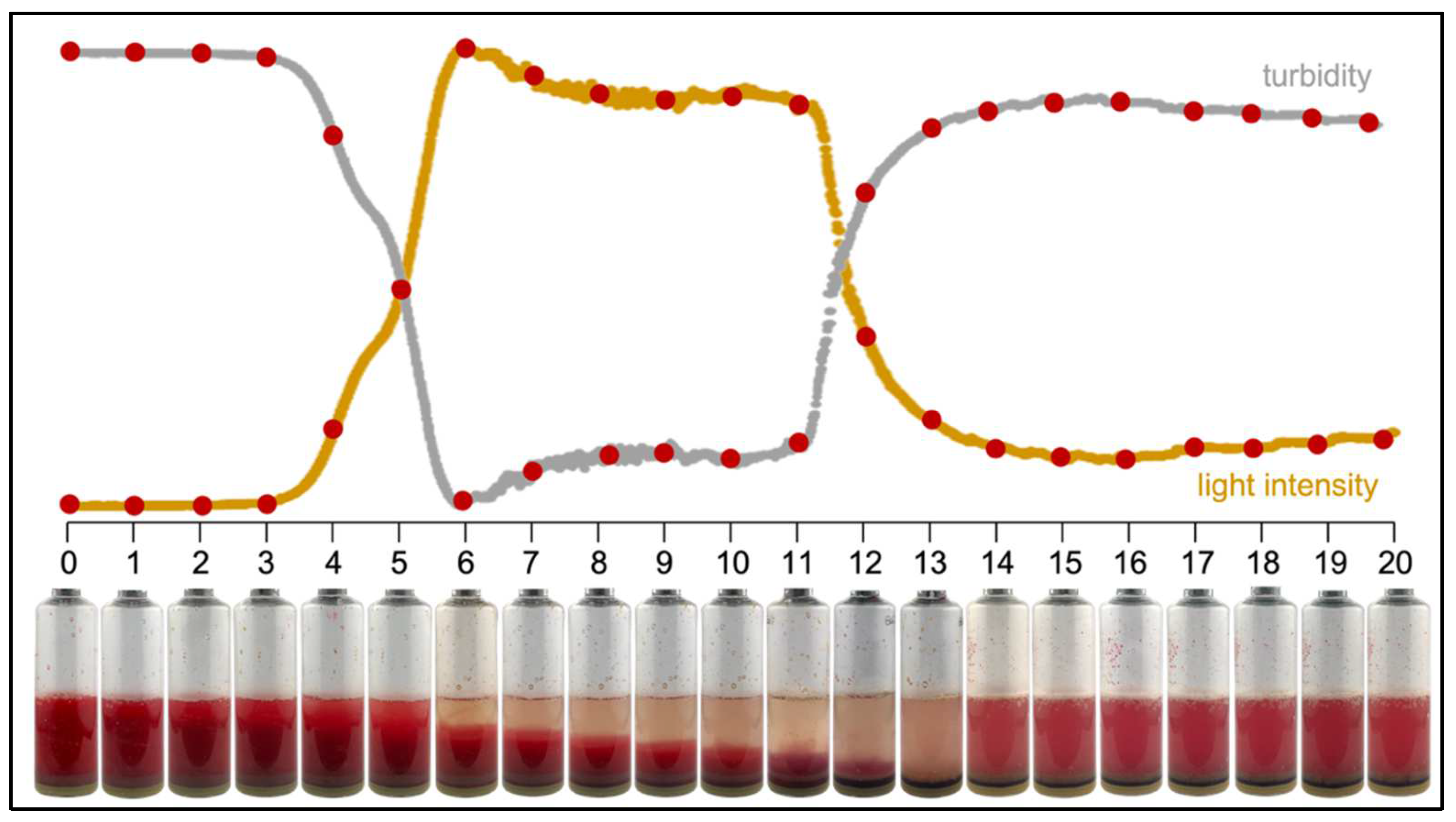
3.1.1. Sedimentation of Blood
3.1.2. Measurement Data Collection
3.2. Detection of Growth Using the Turbidimeter
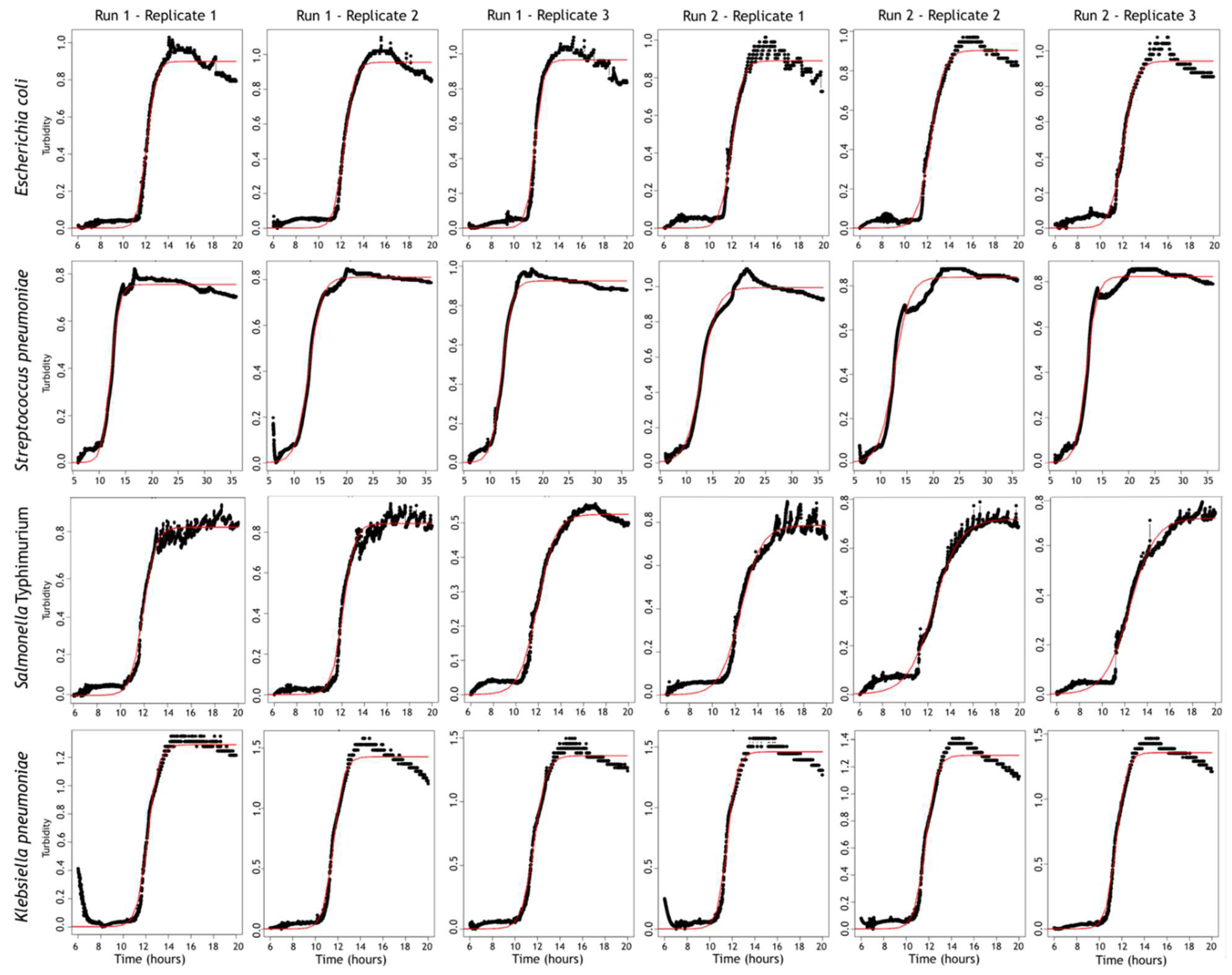
3.2.1. Detection of Turbidity
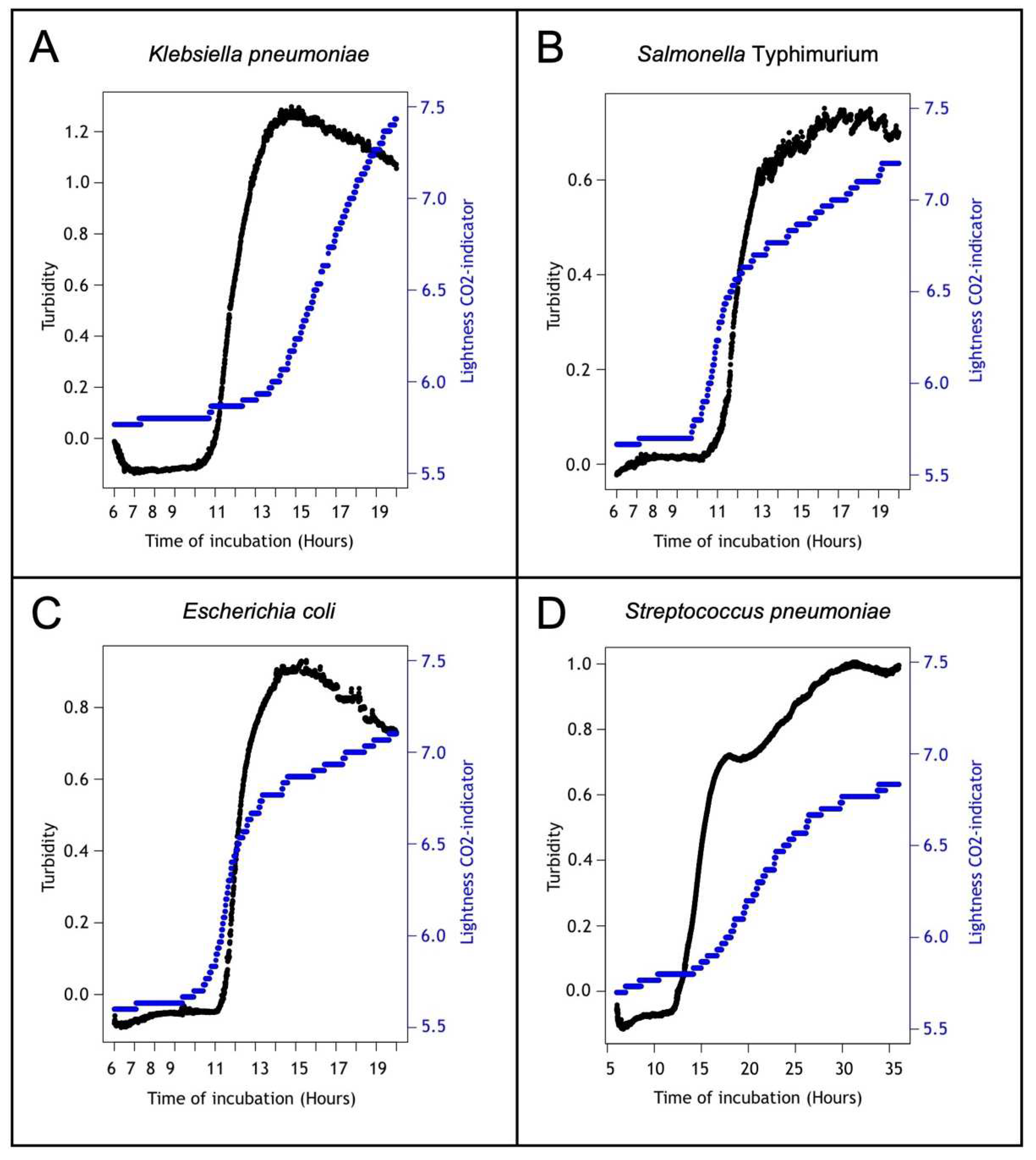
3.2.2. Detection of Color Change in the CO2 Indicator
3.2.3. Turbidimeter Performance in General
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rudd, K.E.; Johnson, S.C.; Agesa, K.M.; Shackelford, K.A.; Tsoi, D.; Kievlan, D.R.; Colombara, D.V.; Ikuta, K.S.; Kissoon, N.; Finfer, S.; et al. Global, regional, and national sepsis incidence and mortality, 1990–2017: Analysis for the Global Burden of Disease Study. Lancet 2020, 395, 200–211. [Google Scholar] [CrossRef]
- Bebell, L.M.; Muiru, A.N. Antibiotic Use and Emerging Resistance: How Can Resource-Limited Countries Turn the Tide? Glob. Heart 2014, 9, 347. [Google Scholar] [CrossRef] [PubMed]
- Founou, R.C.; Founou, L.L.; Essack, S.Y. Clinical and economic impact of antibiotic resistance in developing countries: A systematic review and meta-analysis. PLoS ONE 2017, 12, e0189621. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, J.; Hardy, L.; Semret, M.; Lunguya, O.; Phe, T.; Affolabi, D.; Yansouni, C.; Vandenberg, O. Diagnostic Bacteriology in District Hospitals in Sub-Saharan Africa: At the Forefront of the Containment of Antimicrobial Resistance. Front. Med. 2019, 6, 205. [Google Scholar] [CrossRef]
- Murray, C.J.; Ikuta, K.S.; Sharara, F.; Swetschinski, L.; Robles Aguilar, G.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E.; et al. Global burden of bacterial antimicrobial resistance in 2019: A systematic analysis. Lancet 2022, 399, 629–655. [Google Scholar] [CrossRef]
- Ayukekbong, J.A.; Ntemgwa, M.; Atabe, A.N. The threat of antimicrobial resistance in developing countries: Causes and control strategies. Antimicrob. Resist. Infect. Control 2017, 6, 47. [Google Scholar] [CrossRef]
- Ombelet, S.; Barbé, B.; Affolabi, D.; Ronat, J.-B.; Lompo, P.; Lunguya, O.; Jacobs, J.; Hardy, L. Best Practices of Blood Cultures in Low- and Middle-Income Countries. Front. Med. 2019, 6, 131. [Google Scholar] [CrossRef]
- Okeke, I.N. Diagnostic Insufficiency in Africa. Clin. Infect. Dis. 2006, 42, 1501–1503. [Google Scholar] [CrossRef]
- Petti, C.A.; Polage, C.R.; Quinn, T.C.; Ronald, A.R.; Sande, M.A. Laboratory Medicine in Africa: A Barrier to Effective Health Care. Clin. Infect. Dis. 2006, 42, 377–382. [Google Scholar] [CrossRef]
- Otu, A.; Elston, J.; Nsutebu, E. Sepsis in Africa: Practical steps to stem the tide. Pan Afr. Med. J. 2015, 21, 323. [Google Scholar] [CrossRef]
- Dunne, W.M.; Burnham, C.-A.D. (Eds.) The Dark Art of Blood Cultures; ASM Press: Washington, DC, USA, 2018. [Google Scholar]
- Weinstein, M.P. Current Blood Culture Methods and Systems: Clinical Concepts, Technology, and Interpretation of Results. Clin. Infect. Dis. 1996, 23, 40–46. [Google Scholar] [CrossRef] [PubMed]
- Rohner, P.; Pepey, B.; Auckenthaler, R. Comparative evaluation of BACTEC aerobic Plus/F and Septi-Chek Release blood culture media. J. Clin. Microbiol. 1996, 34, 126–129. [Google Scholar] [CrossRef] [PubMed]
- Rohner, P.; Auckenthaler, R. Review on evaluations of currently available blood-culture systems. Clin. Microbiol. Infect. 1999, 5, 513–529. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Rimmer, K.; Cabot, M. Comparison of Bactec NR-660 and Signal systems. J. Clin. Pathol. 1988, 41, 676–678. [Google Scholar] [CrossRef]
- Hellinger, W.C.; Cawley, J.J.; Alvarez, S.; Hogan, S.F.; Harmsen, W.S.; Ilstrup, D.M.; Cockerill, F.R. Clinical comparison of the isolator and BacT/Alert aerobic blood culture systems. J. Clin. Microbiol. 1995, 33, 1787–1790. [Google Scholar] [CrossRef]
- Ombelet, S.; Ronat, J.-B.; Walsh, T.; Yansouni, C.P.; Cox, J.; Vlieghe, E.; Martiny, D.; Semret, M.; Vandenberg, O.; Jacobs, J.; et al. Clinical bacteriology in low-resource settings: Today’s solutions. Lancet Infect. Dis. 2018, 18, e248–e258. [Google Scholar] [CrossRef]
- Grand View Research. Blood Culture Tests: Market Analysis & Segment Forecast from 2014–2025; Grand View Research, Inc.: San Francisco, CA, USA, 2018. [Google Scholar]
- WHO. Global Action Plan on Antimicrobial Resistance. 2015, p. P078. Available online: https://www.who.int/publications/i/item/9789241509763 (accessed on 30 January 2022).
- WHO. World Health Organization Model List of Essential In Vitro Diagnostics First Edition. 2018. Available online: http://www.who.int/medical_devices/diagnostics/Selection_in-vitro_diagnostics%0A2 (accessed on 30 January 2022).
- Lim, S.H.; Mix, S.; Xu, Z.; Taba, B.; Budvytiene, I.; Berliner, A.N.; Queralto, N.; Churi, Y.S.; Huang, R.S.; Eiden, M.; et al. Colorimetric Sensor Array Allows Fast Detection and Simultaneous Identification of Sepsis-Causing Bacteria in Spiked Blood Culture. J. Clin. Microbiol. 2013, 52, 592–598. [Google Scholar] [CrossRef]
- Templier, V.; Livache, T.; Boisset, S.; Maurin, M.; Slimani, S.; Mathey, R.; Roupioz, Y. Biochips for Direct Detection and Identification of Bacteria in Blood Culture-Like Conditions. Sci. Rep. 2017, 7, 9457. [Google Scholar] [CrossRef]
- Drees, C.; Vautz, W.; Liedtke, S.; Rosin, C.; Althoff, K.; Lippmann, M.; Zimmermann, S.; Legler, T.J.; Yildiz, D.; Perl, T.; et al. GC-IMS headspace analyses allow early recognition of bacterial growth and rapid pathogen differentiation in standard blood cultures. Appl. Microbiol. Biotechnol. 2019, 103, 9091–9101. [Google Scholar] [CrossRef]
- Dailey, P.J.; Osborn, J.; Ashley, E.A.; Baron, E.J.; Dance, D.A.B.; Fusco, D.; Fanello, C.; Manabe, Y.C.; Mokomane, M.; Newton, P.N.; et al. Defining System Requirements for Simplified Blood Culture to Enable Widespread Use in Resource-Limited Settings. Diagnostics 2019, 9, 10. [Google Scholar] [CrossRef]
- Maia, M.R.G.; Marques, S.; Cabrita, A.R.; Wallace, R.J.; Thompson, G.; Fonseca, A.; Oliveira, H.M. Simple and Versatile Turbidimetric Monitoring of Bacterial Growth in Liquid Cultures Using a Customized 3D Printed Culture Tube Holder and a Miniaturized Spectrophotometer: Application to Facultative and Strictly Anaerobic Bacteria. Front. Microbiol. 2016, 7, 1381. [Google Scholar] [CrossRef] [PubMed]
- Ombelet, S.; Natale, A.; Ronat, J.-B.; Kesteman, T.; Vandenberg, O.; Jacobs, J.; Hardy, L. Biphasic versus monophasic manual blood culture bottles for low-resource settings: An in-vitro study. Lancet Microbe 2021, 3, e124–e132. [Google Scholar] [CrossRef]
- Sasidharan, K.; Martinez-Vernon, A.; Chen, J.; Fu, T.; Soyer, O. A low-cost DIY device for high resolution, continuous measurement of microbial growth dynamics. bioRxiv 2018, 407742. [Google Scholar] [CrossRef]
- Wikipedia. HSL and HSV. Available online: https://en.wikipedia.org/wiki/HSL_and_HSV#From_RGB (accessed on 30 January 2022).
- Sprouffske, K.; Wagner, A. Growthcurver: An R package for obtaining interpretable metrics from microbial growth curves. BMC Bioinform. 2016, 17, 172. [Google Scholar] [CrossRef] [PubMed]
- Arp, M.; Larsen, S.O. Comparison of detection speed and yield in agitated and non-agitated aerobic blood culture bottles. APMIS 1992, 100, 1061–1065. [Google Scholar] [CrossRef]
- Prag, J.; Nir, M.; Jensen, J.; Arpi, M. Should aerobic blood cultures be shaken intermittently or continuously? APMIS 1991, 99, 1078–1082. [Google Scholar] [CrossRef]
- Ombelet, S.; Kpossou, G.; Kotchare, C.; Agbobli, E.; Sogbo, F.; Massou, F.; Lagrou, K.; Barbé, B.; Affolabi, D.; Jacobs, J. Blood culture surveillance in a secondary care hospital in Benin: Epidemiology of bloodstream infection pathogens and antimicrobial resistance. BMC Infect. Dis. 2022, 22, 119. [Google Scholar] [CrossRef]
- Tack, B.; Phoba, M.-F.; Van Puyvelde, S.; Kalonji, L.M.; Hardy, L.; Barbé, B.; AB van der Sande, M.; Monsieurs, E.; Deborggraeve, S.; Lunguya, O.; et al. Salmonella Typhi From Blood Cultures in the Democratic Republic of the Congo: A 10-Year Surveillance. Clin. Infect. Dis. 2019, 68, S130–S137. [Google Scholar] [CrossRef]
- Stoesser, N.; Moore, C.E.; Pocock, J.M.; An, K.P.; Emary, K.; Carter, M.; Sona, S.; Poda, S.; Day, N.; Kumar, V.; et al. Pediatric Bloodstream Infections in Cambodia, 2007 to 2011. Pediatr. Infect. Dis. J. 2013, 32, e272–e276. [Google Scholar] [CrossRef]
- Vlieghe, E.R.; Phe, T.; De Smet, B.; Veng, H.C.; Kham, C.; Lim, K.; Koole, O.; Lynen, L.; Peetermans, W.E.; Jacobs, J.A. Bloodstream Infection among Adults in Phnom Penh, Cambodia: Key Pathogens and Resistance Patterns. PLoS ONE 2013, 8, e59775. [Google Scholar] [CrossRef]
- Reddy, E.A.; Shaw, A.V.; Crump, J.A. Community-acquired bloodstream infections in Africa: A systematic review and meta-analysis. Lancet Infect. Dis. 2010, 10, 417–432. [Google Scholar] [CrossRef]
- Kern, W.; Rieg, S. Burden of bacterial bloodstream infection—A brief update on epidemiology and significance of multi-drug-resistant pathogens. Clin. Microbiol. Infect. 2020, 26, 151–157. [Google Scholar] [CrossRef] [PubMed]

| N° | Strain | Mc Farland (McF) Suspension | Incubation Time |
|---|---|---|---|
| 1 | Haemophilus influenzae ATCC 49247 | 0.5 McF | 96 h |
| 2 | Escherichia coli ATCC 25922 | 0.5 McF | 20 h |
| 3 | Staphylococcus aureus ATCC 25923 | 0.5 McF | 36 h |
| 4 | Streptococcus pneumoniae ATCC 49619 | 0.5 McF | 36 h |
| 5 | Burkholderia cepacia ATCC 25416 | 0.75 McF | 96 h |
| 6 | Salmonella Typhimurium ATCC 14028 | 0.5 McF | 20 h |
| 7 | Pseudomonas aeruginosa ATCC 27853 | 0.5 McF | 24 h |
| 8 | Candida albicans ATCC 66027 | 0.5 McF | 96 h |
| 9 | Klebsiella pneumoniae ATCC 700603 | 0.5 McF | 20 h |
| 10 | Neisseria subflava MO00045/1 (clinical isolate) | 0.5 McF | 96 h |
| Strain | Run | Rep * | Growth Rate r | Mean ± SD r | Inflection Point t_mid | Mean ± SD t_mid | Doubling Time t_gen | Mean ± SD t_gen | Goodness- of-Fit Sigma | Mean ± SD Sigma | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Escherichia coli | 1 | 1 | 2.71 | 2.59 ± 0.24 | 2.27 ± 0.41 | 12.09 | 12.09 ± 0.20 | 12.09 ± 0.17 | 0.26 | 0.27 ± 0.03 | 0.31 ± 0.06 | 0.05 | 0.05 ± 0.01 | 0.05 ± 0.01 |
| 2 | 2.31 | 12.28 | 0.30 | 0.05 | ||||||||||
| 3 | 2.76 | 11.88 | 0.25 | 0.06 | ||||||||||
| 2 | 1 | 2.21 | 1.95 ± 0.23 | 11.97 | 12.10 ± 0.18 | 0.31 | 0.36 ± 0.04 | 0.05 | 0.05 ± 0.01 | |||||
| 2 | 1.78 | 12.30 | 0.39 | 0.04 | ||||||||||
| 3 | 1.85 | 12.02 | 0.37 | 0.05 | ||||||||||
| Streptococcus pneumoniae | 1 | 1 | 1.26 | 1.04 ± 0.21 | 0.92 ± 0.22 | 12.23 | 12.45 ± 0.34 | 12.50 ± 0.36 | 0.55 | 0.69 ± 0.14 | 0.79 ± 0.19 | 0.03 | 0.03 ± 0.00 | 0.03± 0.01 |
| 2 | 0.84 | 12.84 | 0.83 | 0.02 | ||||||||||
| 3 | 1.02 | 12.27 | 0.68 | 0.03 | ||||||||||
| 2 | 1 | 0.67 | 0.80 ± 0.19 | 12.93 | 12.54 ± 0.46 | 1.03 | 0.90 ± 0.19 | 0.04 | 0.04 ± 0.01 | |||||
| 2 | 0.70 | 12.66 | 0.99 | 0.04 | ||||||||||
| 3 | 1.01 | 12.01 | 0.68 | 0.03 | ||||||||||
| Salmonella Typhimurium | 1 | 1 | 1.98 | 1.75 ± 0.42 | 1.36 ± 0.52 | 11.93 | 12.00 ± 0.11 | 12.26 ± 0.29 | 0.35 | 0.41 ± 0.12 | 0.57 ± 0.20 | 0.04 | 0.03 ± 0.01 | 0.03 ± 0.01 |
| 2 | 2.01 | 12.13 | 0.34 | 0.04 | ||||||||||
| 3 | 1.27 | 11.94 | 0.55 | 0.02 | ||||||||||
| 2 | 1 | 1.11 | 0.96 ± 0.13 | 12.51 | 12.51 ± 0.03 | 0.62 | 0.73 ± 0.10 | 0.04 | 0.03 ± 0.01 | |||||
| 2 | 0.86 | 12.54 | 0.80 | 0.03 | ||||||||||
| 3 | 0.91 | 12.49 | 0.76 | 0.03 | ||||||||||
| Klebsiella pneumoniae | 1 | 1 | 2.14 | 2.08 ± 0.18 | 2.16 ± 0.17 | 12.19 | 11.85 ± 0.30 | 11.73 ± 0.24 | 0.32 | 0.34 ± 0.03 | 0.32 ± 0.03 | 0.07 | 0.06 ± 0.01 | 0.06 ± 0.01 |
| 2 | 2.22 | 11.64 | 0.31 | 0.07 | ||||||||||
| 3 | 1.88 | 11.72 | 0.37 | 0.05 | ||||||||||
| 2 | 1 | 2.37 | 2.24 ± 0.13 | 11.58 | 11.62 ± 0.11 | 0.29 | 0.31 ± 0.02 | 0.07 | 0.06 ± 0.01 | |||||
| 2 | 2.10 | 11.74 | 0.33 | 0.07 | ||||||||||
| 3 | 2.23 | 11.53 | 0.31 | 0.05 | ||||||||||
| Strain | TTD * Turbidity | TTD Colour Change | TTD Turbidimeter | TTD Reference System | TTD Turbidimeter Minus Reference System |
|---|---|---|---|---|---|
| Escherichia coli | 12.1 | 11.4 | 11.4 | 12.0 | −0.6 |
| Streptococcus pneumoniae | 12.5 | 17.4 | 12.5 | 11.6 | 0.9 |
| Salmonella Typhimurium | 12.3 | 10.9 | 10.9 | 13.2 | 2.3 |
| Klebsiella pneumoniae | 11.7 | 14.7 | 11.7 | 12.0 | −0.3 |
| Staphylococcus aureus | No detection | 21.9 | 21.9 | 13.8 | 8.2 |
| Burkholderia cepacia | No detection | 27.3 | 27.3 | 20.5 | 6.8 |
| Pseudomonas aeruginosa | No detection | 18.7 | 18.7 | 16.7 | 2.0 |
| Haemophilus influenzae | No detection | 39.1 | 39.1 | 14.6 | 24.5 |
| Neisseria subflava | No detection | 57.4 | 57.4 | 17.0 | 40.4 |
| Candida albicans | No detection | 45.4 | 45.4 | 32.0 | 13.4 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Barbé, B.; Corsmit, E.; Jans, J.; Kaur, K.; Baets, R.; Jacobs, J.; Hardy, L. Pilot Testing of the “Turbidimeter”, a Simple, Universal Reader Intended to Complement and Enhance Bacterial Growth Detection in Manual Blood Culture Systems in Low-Resource Settings. Diagnostics 2022, 12, 615. https://doi.org/10.3390/diagnostics12030615
Barbé B, Corsmit E, Jans J, Kaur K, Baets R, Jacobs J, Hardy L. Pilot Testing of the “Turbidimeter”, a Simple, Universal Reader Intended to Complement and Enhance Bacterial Growth Detection in Manual Blood Culture Systems in Low-Resource Settings. Diagnostics. 2022; 12(3):615. https://doi.org/10.3390/diagnostics12030615
Chicago/Turabian StyleBarbé, Barbara, Ellen Corsmit, Jasper Jans, Kamalpreet Kaur, Roel Baets, Jan Jacobs, and Liselotte Hardy. 2022. "Pilot Testing of the “Turbidimeter”, a Simple, Universal Reader Intended to Complement and Enhance Bacterial Growth Detection in Manual Blood Culture Systems in Low-Resource Settings" Diagnostics 12, no. 3: 615. https://doi.org/10.3390/diagnostics12030615
APA StyleBarbé, B., Corsmit, E., Jans, J., Kaur, K., Baets, R., Jacobs, J., & Hardy, L. (2022). Pilot Testing of the “Turbidimeter”, a Simple, Universal Reader Intended to Complement and Enhance Bacterial Growth Detection in Manual Blood Culture Systems in Low-Resource Settings. Diagnostics, 12(3), 615. https://doi.org/10.3390/diagnostics12030615








